Abstract
SARS-CoV-2, the cause of the COVID-19 pandemic, has introduced a challenging era characterized by the persistent emergence of subvariants. Even after the World Health Organization announced the end of the pandemic, the virus continues to evolve, posing significant challenges to public health responses. This comprehensive review examines the multifaceted impacts of these subvariants, emphasizing their significance across diverse dimensions. SARS-CoV-2 has genetic variability, especially at the spike protein region, which has given rise to Variants of Concern, including Beta, Delta, Gamma, Alpha, and the highly mutable Omicron, which differently exhibit varying levels of immune evasion, disease severity, and transmissibility. Subvariants within the Omicron lineage, including BA.1, BA.2, BA.3, and others, further complicate the landscape with distinct genetic signatures and varying infectivity levels. The impacts extend to diagnostic techniques, treatment strategies, and vaccine effectiveness, underscoring the need for a comprehensive public health response emphasizing preventive measures, genomic surveillance, and vaccination campaigns. Sustaining these interventions is critical, necessitating long-term strategies considering socio-political factors, community involvement, continuous adaptation of healthcare approaches, robust monitoring, and sustainable public health interventions to effectively combat the virus's ever-changing landscape.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
The COVID-19 pandemic caused by SARS-CoV-2 has resulted in a significant global health crisis, leading to 770,875,433 total cases as well as 6,959,316 mortalities by September 27, 2023, according to the World Health Organization (WHO) [1, 2]. SARS-CoV-2 variants vary in terms of disease severity, clinical course, and transmissibility. They could additionally have effects on disease diagnosis, treatment susceptibility, vaccine-mediated protection against severe sickness, and lasting medical impacts following infection with SARS-CoV-2, commonly referred to as long-term consequences [2]. Similar to many other RNA viruses, COVID-19 experiences fast evolution, an observable and quantitative process that happens in a matter of months or years [3]. Factors contributing to this evolutionary process include transmission of the virus, immunological profiles and human movement, epidemiological factors, and all linkages that characterize RNA viruses [4]. Other variables impacting the virus’s evolution include ecological changes in humidity, air pollution, and climate change, all of which increase temperature and cause thermal denaturation of viral proteins and genomes [5].
The rate at which mutations appear and propagate throughout populations determines how quickly viruses evolve. Natural selection favours advantageous mutations, such as the D614G mutation that boosts transmissibility. As viruses reproduce, change, and evolve within people with complexity, they also successfully adapt for person-to-person transmission on a different scale [6]. As viral lineages evolve into potentially separate strains, higher-level dynamics like lineage competition and extinction come into play in addition to these population-level processes [4]. These dynamics play a role in the trajectory and evolution of the virus, providing insights into the emergence and persistence of certain variants [1]. Lineage competition refers to the process by which variants enter into competition for dominance within a population [2]. This competition occurs because of different levels of transmissibility and pathogenicity, influenced by factors like host immune responses, transmission dynamics, and environmental conditions. Mutations can give certain variants advantages, allowing them to outcompete others [1]. On the other hand, less fit variants may face extinction due to selective pressures when a lineage of the virus loses its ability to replicate and transmit effectively and subsequently disappears from the population [3]. Extinction events can occur at both individual variant and broader lineage levels. Variant-level random phenomena and difficulties can lead to the loss of certain genetic variants, while entire viral lineages may become extinct due to factors like widespread immunity, changes in host susceptibility, or the emergence of more successful competitors [1, 7].
The evolution of the virus has led to over 2500 fatalities and roughly 1.5 million new COVID-19 cases worldwide between July 10, 2023, and August 6, 2023, despite the worldwide public health crisis having been declared over by the WHO in May 2023 [5]. This shows that the disease and its variants continue to strike, thus adding to the global effect of the pandemic with its evolution. The variants are currently divided into three distinct groups: variants of concern (VOC), variants of interest (VOI), and variants under monitoring (VUM) [6]. This classification was based on criteria that include genetic changes within the viral genome, which include the potential to increase virus transmission and its virulence, cumulative cases, and the need for risk assessment [6]. Certain mutations may confer a selective advantage, enhancing the virus’s ability to spread efficiently from host to host. These genetic alterations can influence key viral properties such as infectivity, replication kinetics, and immune evasion mechanisms. Additionally, the potential to increase virus transmission is intricately linked to the molecular interactions between the virus and its host. Viral proteins involved in receptor binding, cell entry, and immune evasion are central players in determining the virus's transmissibility [8, 9]. This demonstrates the need to explore the variants and subvariants that have been identified to provide the evidence necessary to support health policy-makers in improving population health [10]. This article aims to comprehensively examine the multifaceted implications and challenges posed by the ongoing infection caused by SARS-CoV-2 subvariants in the pandemic and post-pandemic eras.
2 Methodology
We conducted a comprehensive internet-based literature search on Scopus, PubMed, and Google Scholar to identify relevant literature. A snowballing method was utilized to identify additional relevant articles to ensure that all relevant literature was captured, thereby enhancing the comprehensiveness of the review. Types of research included original research, perspectives, commentaries, correspondence, systematic literature reviews and meta-analyses, literature reviews, and grey literature addressing the evolution and impact of SARS-CoV-2 variants during and in post-COVID-19 public health emergencies. The time since the WHO proclaimed COVID-19 to be no longer a pandemic was the definition of the post-pandemic period. To generate search strings, Boolean operators ‘‘OR’’ and ‘‘AND’’ were used to combine keywords such as ‘‘COVID-19’’, ‘‘SARS-CoV-2’’, ‘‘Variants’’, ‘‘Subvariants’’, ‘‘Public health emergency’’, and ‘‘Viral evolution’’. A bibliometric search of the included papers was also carried out to find more pertinent research for the review. We included articles based on their relevance and quality in line with our objectives, published between 2019 and 2024. Using a rigorous qualitative synthesis technique, the two first authors (EM and OJO) critically evaluated and narratively summarized the findings of the included papers under the relevant subheadings, given the limited nature of the review.
3 Results
3.1 The evolution amid the COVID-19 global emergency
RNA viruses are generally prone to developing variations as a result of mutations in specific proteins, leading to Variants of Concern (VOCs) [11]. Since the COVID-19 pandemic began, other mutant variants have been detected that are produced through genetic evolution. During the SARS-CoV-2 outbreaks, the WHO has designated the majority of mutants as "variants of concern, which include omicron, beta, delta, alpha, and gamma [12]. Omicron stands out from the other variants due to its 30 hallmark mutations, including 23 major ones [10, 13]. Beta (B.1.351) was discovered in the last month of 2020 in South Africa [14], Gamma (P.1) at the start of January 2021 in Brazil [1], Delta (B.1.617.2) in India around December 2020 [14], as well as Omicron (B.1.1.529) in November 2021 around South Africa, all before Beta (B.1.351) was discovered there [15]..
The Alpha mutant (B.1.1.7 ancestry), originally discovered in the UK in late December 2020, was initially identified as GRY (previously GR/501Y.V1) utilizing the complete genome sequencing of SARS-CoV-2-positive clinical specimens [16, 17]. To find this variant, a commercial test was used to detect S-gene target failure, SGTF, in polymerase chain reaction specimens [18]. The Alpha variant's viral genome has 17 mutations, including eight at the region of the spike (S) protein (Δ144 deletion, D1118H, N501Y, P681H, Δ69-70 deletion, T716I, S982A, A570D) [16, 17, 19]. Notably, the N501Y mutation increases the spike-associated protein's attraction for ACE 2 receptors, allowing viral integration and infiltration inside the cells of the host. Whereas it did not show initial significant changes in hospitalization or mortality risk when compared to other variants, later research demonstrated an increase in disease severity [19]. In comparison to previously circulating strains, research indicated a death hazard ratio of 1.64 for B.1.1.7-infected patients, supported by other studies that also found an elevated risk of death linked to this variant, highlighting its potential health hazard [20].
The beta variants (B.1.351 lineage), found around October 2020 in South Africa, constituted part of the subsequent outbreak of COVID-19 cases and carried some spike protein mutations. These include nine alterations, three of which (N501Y, E484K, and K417N) belong to the hormone receptor binding domain (RBD), increasing spike protein's binding for ACE sensors [21]. At the end of January 2021, this variant was detected in the United States, where it resulted in a greater likelihood of transmission and was less vulnerable to being neutralized by convalescent sera, monoclonal antibodies, and post-vaccination sera [22]. The Gamma mutant (P.1 lineage) is a B.1.1.28 variant with ten spike gene mutations, three of which (E484K, L18F, and K417N) have been identified in the RBD, identical to the B.1.351 variant. It was discovered in the United States in January 2021, following its initial discovery in Brazil in December 2020 [23]. The Delta B.1.617.2 variant, which first became known around December 2020 in India, was linked to the devastating second round of the pandemic that hit India in April 2021 and was subsequently identified in March 2021 in the US, with ten alterations made to its spike protein [14].
The Kappa variant, which belongs to the B.1.617 sublineage [24] and is referred to as B.1.617.1, was discovered in India for the first time in December 2020. It was identified as VUI-21APR-01, a variation under investigation, by Public Health England. This variation comes with some unique alterations, including P681R, E484Q, and L452R [25]. The Omicron variation has had extensive spike protein mutations, just as other viral genome parts have undergone mutation. Those other parts include the envelope, nucleocapsid protein, matrix, receptor-binding domain, and non-structural proteins. It was therefore unsurprising that the Omicron mutation was swiftly designated as VOC since its initial detection on November 23rd, 2021, in South Africa, owing to the fast rise in COVID-19 cumulative cases [25]. Omicron’s intricacy and diversity have been highlighted by the identification of greater cases of subvariants, such as BA.5, BA.4, BA.3, BA.2, and BA.1 [26]. BA.2, BA.1.1, and BA.1. being the most common as of February 2022 [27]. These subvariants also have an important number of mutations. For instance, BA.2 had an additional 28 mutations, with four mutations in the spike protein, while BA.1 (21 K) comprises 60 mutations, with 32 in the region of the spike protein [27]. Because BA.2 (21L)'s genome lacked the distinctive STGF pattern, it was initially mistaken for the delta variants in PCR tests, earning it the moniker "Stealth Omicron''.
Although BA.2 has been called ‘‘stealth,’’ this subvariant is thought to be more contagious than BA.1 [28] with its sub-ancestry being referred to as BA.1 (B.1.1.529.1), while others are referred to as BA.3 (B.1.1.529.3) and BA.2 (B.1.1.529.2), among others. Additionally, BA.2 subvariants like BA.2.12.1 and BA.2.12 arose and outlived BA.2 by a wide margin [29]. In January 2022, the recombinant of BA.1 and BA.2, producing the XE subvariant, appeared in the UK, where it has been contagious rapidly [30]. Around January 2022, in South Africa, BA.5 and BA.4, comparable to BA.2, but keeping STGF (69-70del) from BA.1, triggered new COVID-19 waves around the world beginning in April 2022 [31]. These successfully resist the immune system by spike protein changes, such as L452R, F486V, and Q493, thus being designated VOC by the European CDC and UK Health Security Agency after causing new COVID-19 waves globally [32].
Relevant to the Omicron variant, a new circulating variant referred to as BA.2.75, first reported in May 2022 in India, was also identified elsewhere afterward. Although this subvariant is thought to be more contagious than other Omicron subvariants, it has not yet been designated as a VOI or VUM because of scant data and is still under monitoring by the ECDC [33]. Several omicron variants have been documented. BA.5, also known as BF.7, has recently come to light due to its greater transmissibility, quicker incubation period, and higher risk of reinfection [34]. Another new variety, XBB.1.5, has attracted interest because of its potential for widespread transmission, immune evasion, and greater transmissibility. It carries several spike protein mutations, including the uncommon F486P alteration [29]. XBB.1.16, a combination of two BA.2 offspring and a relative of XBB.1.5, was originally discovered on January 9th, 2023 [35]. Due to its genetic resemblance to XBB.1.5 and the addition of two amino acid changes (K478R and E180V) at the region of the spike protein compared to their parent, XBB.1.1, it was first classified as a VUM on March 22nd, 2023, but was later elevated to a VOI on April 17th, 2023 [36].
Although WHO ended the COVID-19 global health emergency as of May 2023, the virus continues to mutate, with new subvariants on the rise [37, 38]. As the WHO was closely watching XBB.1.16, changes and decreasing herd immunity that allow the pathogen to evade remaining immunity among the people were identified as the cause of the spike in COVID-19 infections, which has supplanted other variations in circulation in India and is now spreading in the United States. For instance, as of April 17th, 2023, 33 different nations had reported 3648 sequences of the Omicron XBB.1.16 variation, and there have been 60,300 continuing COVID-19 cases registered just in India [35, 36]. At the beginning of August 2023, the Omicron subvariant XBB.1.9.2 was found to be connected to the variant EG.5, known as "Eris," which has been affecting nations including the US, the UK, and China. Although the variant had a higher prevalence and traits related to immune evasion and growth advantage, the WHO initially found that EG.5 poses a low public health risk globally until August 9th, 2023, when it was upgraded to VOI from VUM [39, 40], having documented a novel mutation in EG.5’s spike protein in a genomic sample analysis.
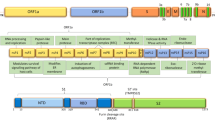
Interestingly, it has been argued whether EG.5 would continue to be considered a VOI [41]. Likewise, in the same month of August 2023, following the spike in the incidence of COVID-19 in the US and the UK, BA.2.86, an ancestral line descended from BA.2, first discovered in 2022 in Nigeria, was reported in the UK, US, South Africa, Denmark, and Israel by August 23, 2023 [42]. Its numerous genomic variations that distinguish it from its ancestor, BA.2, and from other XBB-derived SARS-CoV-2 mutants are currently widespread, leading the WHO to designate it as a VUM [43] (Fig. 1).

Source: Emerging SARS-CoV-2 variants of concern and potential intervention approaches. 2021 [44]
Image showing major mutations of SARS-CoV-2 variants of concerns.
3.2 C0VID-19 variants in post-COVID-19 public health emergency
The state of COVID-19 PHEIC, which was initially declared on January 30, 2020, by the WHO, was ended by the WHO Director-General on May 5, 2023 [43]. However, the declaration distinguished COVID-19 as a global emergency and a pandemic. While the former is over, the WHO diplomat highlighted the relevance of the latter as the SARS-CoV-2 variants keep mutating, demonstrating the significance of transitioning and integrating the COVID-19 pandemic into a nationwide infectious disease management program. The occurrence of variant mutations raises significant concerns in the post-COVID-19 pandemic global emergency era, considering their historical surge that caused a multitudinous number of infections and deaths [43, 45].
In addition to their fast global spread, these variants have potential immune evasion properties, thereby becoming a major issue in the post-pandemic era [45]. For instance, the Omicron variant, which has been the most contagious variant of any other, has not stopped evolving. Whereas no variant was declared VOC as of December 1, 2023, the ECDC listed four VOI, including XBB.1.5-like + F456L, BA.2.75, XBB.1.5-like, and BA.2.86 [46].
Having been first documented on February 17, 2023, EG.5, also known as Eris, a descendant offspring of XBB.1.9.2, has F456L amino acid variations at the spike protein, thus having immune-evasion capability and concerning effects on individuals. It caused just around 68.3% of COVID-19 cumulative cases in China [10]. Globally, this subvariant has spread in around 66 countries and caused just over 50% of cumulative cases during the summer of 2023, thus fulfilling the criteria of being declared a VOI by the WHO on August 9, 2023 [47]. Interestingly, data from 71 countries illustrated that COVID-19 cases caused by EG.5 steadily increased from 26.3% in mid-August to 33.10% on September 10, 2023 [48]. Equally important, using evaluation risk criteria that include genetic features, immune growth rate, and escape characteristics, the recent WHO report assessed the variant as low risk predicted; however, EG.5 could significantly spread, resulting in rising cases and hospitalizations [48]. The mutations in EG.5 are concerning, and their descendant lineages are significant in the post-pandemic period. This concern lies in the fact that its subvariants accounted for around 51.8% of all VOI and VUM with the F456L mutation, thus being the most common subvariants [48]. It is then unsurprising that one of its subvariants-HV.1 was dubbed the highly contagious subvariant as of December 2023. Its rapid spread has caused nearly 33% of cases in the USA, thus being the most predominant and overtaking EG.5 in the country [49]. Similarly, COVID-19 cases, caused by the HV.1 subvariant have risen from 2.1% to 34.4% in Canada as of October 29, 2023, thus being the most prevalent subvariant [50].
Another variant of interest as of December 1, 2023, was BA.2.86 [46]. Following the occurrence of cases, WHO designated it a VUM on August 17, 2023, but classified it a VOI later. According to the GISAID data from 46 countries, the BA.2.86 variant has steadily increased, with its global prevalence having risen from 1.8% on October 8, 2023, to 8.9% as of November 5, 2023. Interestingly, the variant has significantly increased in countries that include the United Kingdom (3.6% to 14.2%), France (3.1% to 13.8%), and Sweden (5.5% to 12.0%) between October 30 and November 5, 2023 [51]. The rapid increase of new infections highlights the reason it has been categorized as VOI and shows the need for continuous monitoring and evaluation, including assessment of vaccine and treatment efficacy against evolving variants [52]. The WHO designated the JN.1 subvariant of COVID-19 as a VOI owing to its fast spread. JN.1, first discovered in the US in September 2023, and its subvariants, JN.1.7, JN.1.13.1, and JN.1.18, have spread to 41 nations, accounting for more than 25% of the global sequences that GISAID has examined [53, 54]. The prevalence of JN.1 in the US, where it is responsible for more than 60% of COVID-19 cases, and the UK, Iceland, and Portugal, prompts worries about what might happen to nations with less robust healthcare systems. Compared to its predecessors, such as BA.2.86, JN.1 is more transmissible, and its illness severity is not appreciably different from other variants [55]. Nonetheless, JN.1 seemed less transmissible than earlier strains, indicating a possible trade-off between transmissibility and immune evasion. FLiRT variants, descendants of JN.1, have caused a surge in cases during the winter. KP.2, which has emerged as a prominent variant, made up 25% of newly sequenced cases. While KP.1.1 and other FLiRT variants were not as widespread in the U.S, they represented 7.5% of illnesses across the country in the last term of 2023, leading to further investigation of their transmission rates, possible severity of sickness, and vaccination resistance [56, 57]. In addition to the already mentioned omicron subvariants from lineage XBB.1.5-like + F456L and BA.2.86, the other two that include XBB.1.5-like and BA.2.75 have been reportedly grouped among VOI as of December 1, 2023. Taking into account its first letter-XBB.1.5-like, the X means that the subvariant is from more or just two sublineages, BA.2.75 and BA.2.10.1 specifically [58]. Its extreme hACE2-binding affinity, strengthened by significant advantageous growth, gives it the potential for immune evasion mutations, thus spreading considerably and explaining its prevalence in certain communities [59]. Omicron subvariant BA.2.75, which was first identified and was more prominent in India in 2022 than in any other country [60], is still prevalent in the last term of 2023. Its immune resistance and evasion are due to the receptor binding affinity and spike protean frequency of several mutations, such as N460K, F157L, I210V, W152R, G257S, D339H, G446S, and Q493 [61] (Fig. 2).

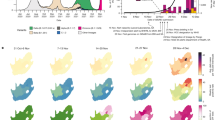
Source: Evolutionary trajectory of SARS-CoV-2 and emerging variants, 2021 [62]
Mutations found on Human coronaviruses.
Red dots in the genomes represent certain amino acid residues that have undergone intense positive selection, leading to the emergence of a particular mutation that is now prevalent in the area from which it originated. Genomic areas with significant polymorphisms within a CoV species are indicated by purple bars, while red bars indicate deletions that have been chosen. The blue shaded area represents beta-CoV Lineage B (Sarbecovirus), the yellow shaded area represents beta-CoV Lineage C (Merbecovirus), the red shaded area represents beta-CoV Lineage A (Embecovirus), and the green shaded area represents alpha-CoVs. The top remark indicates the genome length in kilobases (kb) [62]
3.3 SARS-CoV-2 variations and their impact on public health
The impact of those subvariants is multiple. Notably, the evolution of the virus represents a significant concern for the emergence of COVID-19 herd immunity while the virus is still developing [63]. While herd immunity can potentially be produced through natural infection and vaccination, viral alterations may impair the efficacy of existing immunity [64]. As a result, it is necessary to regularly assess and adjust vaccination strategies to address emerging variants and maintain effective protection against the evolving virus [65]. The communities must watch the virus closely, take all necessary precautions, and restore adequate protective behaviour to prevent the possible quick spread of the virus. Institutions must work together to adopt a test and vaccine to tackle the virus.
In addition, those variants have significantly impacted SARS-CoV-2 diagnostics, necessitating testing method adaptation [64]. Viral variants that can alter illness severity, symptoms, and tissue tropism, as well as host-specific characteristics including age and immunization status, can impact the virus's effects on individuals [66]. On a therapeutic level, certain variants, like the Omicron variant, require less drastic measures, whereas others, like the Delta variant, may lead to hospitalization and affect the efficacy of antiviral medications and monoclonal antibody therapy [67, 68], thus leading to resistance. These mutations, particularly at the RBD regions and the N-terminal mutations of the spike protein, can reduce the efficacy of treatment approaches [69, 70]. For instance, the D614G mutation improved the virus's infectivity by improving its binding to ACE2 receptors. However, studies show inconsistencies in the virus's immune escape mechanism, with mutations like E484K rendering treatments like Bamlanivimab ineffective [70, 71]. Other mutations, such as S477G, led to immune escape from monoclonal antibodies. Omicron variants, with over 30 mutations, contribute to immune evasion and increased transmissibility. Variants with unique spike protein mutations, such as BA.5 and BA.4, may enhance transmissibility and immune evasion [29, 52]. The SARS-CoV-2 variations’ tendency to evolve has had an impact on vaccination efficiency, resulting in varied degrees of protection against various forms [69, 70]. While vaccinations have typically continued to protect against serious illness, their effectiveness has gradually declined over time due to recently discovered variations. This is the case for people who received a single dose of the vaccination but were challenged by variants, such as delta [72]. The Omicron version has demonstrated decreased vaccination efficacy, particularly after booster doses, due to several variations in the spike protein. In addition, the efficacy of vaccines has been further questioned by subvariants like BA.4 and BA.5 [73]. As a result, variant-adapted vaccinations have been developed to improve defence against these shifting strains. In addition, the use of seasonal updates, like influenza vaccines, has been debated as the long-term plan for COVID-19 vaccines [29, 73]. SARS-CoV-2 transmission occurs through the air, droplets, aerosols, or contact with contaminated surfaces [74]. The virus attaches to host cells through the complex relationship between the ACE2 receptor and the spike protein, with recent studies suggesting an ACE2-independent mode of infection. Antibodies targeting SARS-CoV-2 target the antigenic alterations in the protein's structure that are associated with viral immune escape [75, 76]. SARS-CoV-2 variations include Omicron, Gamma, Beta, Alpha, and Delta, with multiple variations centered at the spike protein area [77]. These antigenic changes enhance transmission and facilitate immune escape by increasing spike protein affinity for the human ACE2 receptor [78].
SARS-CoV-2 could impair the CNS through ACE2 receptors in humans and mice, potentially leading to brain infection. Neuroinvasion can occur through transsynaptic transfer, vascular endothelial cell infection, or leukocyte migration [75]. The virus is capable of impacting the cardiovascular system, resulting in direct cytotoxicity and proinflammatory cytokines that cause myocarditis and cardiac arrhythmias. COVID-19 is also associated with acute coronary syndrome, potentially linked to the virus's thrombogenicity and inflammatory cytokine release [79]. SARS-CoV-2's impact on various physiological systems is multifactorial and not fully understood. Pathogenesis in the gastrointestinal tract involves viral cytotoxicity, inflammation, vascular abnormalities, and gut dysbiosis [80]. The hepatobiliary system's response to the virus is unclear, with hypotheses including immune-mediated inflammatory responses, viral replication, drug-induced liver injury, and hypoxic damage [81]. The renal system's involvement in COVID-19-related kidney disorders is multifactorial, potentially caused by viral cytotoxicity, RAAS disturbances, cytokine-induced hyperinflammatory states, and prothrombotic conditions [1].
The elderly, immunocompromised people and people with pre-existing medical issues are among the vulnerable groups that are more likely to experience severe disease and death from COVID-19. The management of the pandemic presents major hurdles for poor and developing countries, as they have inadequate resources and infrastructure for healthcare [82]. This can result in overburdened healthcare systems, a scarcity of medical supplies, and challenges in distributing vaccines. The pandemic has made already-existing disparities worse, disproportionately harming underprivileged populations who have little access to social services and healthcare. Strict lockdown procedures and social distance guidelines have also interfered with day-to-day activities, resulting in financial difficulties, joblessness, and mental health problems [83]. To stop the virus from spreading, governments all over the world have enacted emergency laws and regulations, which have an influence on people's liberties and social norms. These laws and regulations include mandatory mask use, travel limitations, and quarantine requirements. The risk of COVID-19 transmission is increased in vulnerable settings, including prisons, refugee camps, and highly populated urban areas, because of overcrowding, inadequate sanitation, and restricted access to healthcare [84]. In order to mitigate disparities and foster resilience in the face of future challenges, comprehensive strategies are needed to address the multifaceted impacts of SARS-CoV-2 on vulnerable populations and settings. These strategies include equitable vaccine distribution, strengthened healthcare systems, socioeconomic support measures, and community-based interventions [82,83,84].
4 Public health response and future directions
As we move into the post-Global Emergence Period of COVID-19, it is essential to focus on implementing comprehensive vaccination programs. This includes ensuring that vaccines are readily accessible to communities, addressing any vaccine hesitancy through education and awareness campaigns, and prioritizing equitable distribution to underserved populations [85]. Relevant to that, several vaccines have been developed and deployed as part of this comprehensive plan, with 322 candidates undergoing clinical studies, 17 gaining Emergency Use Authorization (EUA), and six having full WHO approval [86]. These vaccines cover a variety of technological platforms, such as mRNA-based vaccines like BNT162b2 and mRNA-1273, inactivated virus vaccines like Covaxin, CoronaVac, and Sinopharm, and adenovirus vector vaccines like those produced by Oxford-AstraZeneca and Johnson & Johnson [86, 87]. The rapid mutation rate of COVID-19 has led to significant challenges in vaccine development and deployment, requiring a balance between expediency and adherence to established research ethics. The urgency to curb the virus's spread has led to unprecedented collaboration and resource mobilization within the scientific community [88]. However, concerns have been raised about the potential compromise of ethical standards, particularly in terms of safety, efficacy, and equitable access to vaccines. The ethical implications of the vaccine race include issues such as informed consent, transparency in clinical trial data, and prioritizing vulnerable populations in vaccine distribution efforts [88]. It was reported, however, that there was not a clear strategy to achieve vaccination among the vulnerable population [89]. Fostered translational research is necessary to stay ahead of the new variants, monitor and understand the virus's mutations, develop new treatment options, and advance technology for rapid testing and surveillance [90]. This further implies the need to invest in and strengthen public health infrastructure to better prepare for any future health emergencies. This should include increasing funding for public health agencies in low-resource settings to expand testing and contact tracing capabilities and improve health governance and health service delivery [91].
The management of SARS-CoV-2 variants necessitates a multifaceted approach. The vaccines' efficacy varies, with 90% of real-world effectiveness influenced by unique patient characteristics. Breakthrough infections, even in highly vaccinated populations, highlight the need for continued vigilance. Factors influencing these infections include exposure levels, antibody affinity, and avidity [92]. Understanding the long-term effects of breakthrough infections is crucial for assessing vaccine efficacy against emerging variants. Further research is needed to define protection correlates and enhance vaccine responses. Addressing vaccine hesitancy through tailored messaging and transparent communication is essential. Combating misinformation and engaging local communities in vaccine dialogue is crucial for widespread vaccination coverage [92, 93]. Sustaining efforts to promote behavioral and hygiene practices, such as handwashing, mask-wearing, and social distancing, remains important in preventing the spread of COVID-19. Public health messaging should emphasize the ongoing need for these practices while adapting to the changing dynamics of the post-pandemic period [94]. Strategies for promoting local policy implementation should include regular emergency communication, cooperation among actors, and fostering openness and information exchange [95]. For instance, the SARS-CoV-2 Interagency Group (SIG) has been tracking the changes in COVID-19 variations in real-time in partnership with institutions like the Centers for Disease Control and Prevention (CDC), WHO, the National Institutes of Health (NIH), the Food and Drug Administration (FDA), the Biomedical Advanced Research and Development Authority (BARDA), and international health agencies. Similarly, the Department of Biotechnology and the Indian Council of Medical Research are cooperating with the INSACOG in India to manage the situation [46]. These international and regional counterparts use whole genome sequencing (WGS) as a fundamental technique to examine the most common strains, evaluate the most important mutations, and develop management and prevention plans [94]. A crucial step towards identifying and containing SARS-CoV-2 on a national and international scale is the global implementation of integrated genomic surveillance systems. The prediction and mitigation of potential pandemic strains are possible with real-time genetic data [46, 94] and wastewater-based surveillance [96]. The current global health approach to combating SARS-CoV-2 is a complex strategy that should combine preventative and curative measures [87].
5 Conclusion
WHO declared the end of the COVID-19 global health emergency. However, SARS-CoV-2 subvariants continue to arise in the post-pandemic age of COVID-19. Public health interventions face major difficulties as a result of these subvariants, particularly the highly changeable Omicron lineage and its numerous subvariants. These differences affect diagnosis, treatment plans, and vaccine effectiveness, highlighting the necessity of constant monitoring and strengthening of the healthcare system and practices. The focus of the public health approach has been prevention, non-pharmaceutical therapies, genetic surveillance, and vaccinations, among the core elements of the strategy. In the continuous battle to eradicate SARS-CoV-2 and its altering subvariants, the sustainability of public health interventions and a continued emphasis on monitoring infection rates and variant-specific trends remain paramount.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Cascella M, Rajnik M, Aleem A et al. Features, Evaluation, and Treatment of Coronavirus (COVID-19). StatPearls. Treasure Isl StatPearls Publ. 2023; https://www.ncbi.nlm.nih.gov/books/NBK554776/
Mizrahi B, Sudry T, Flaks-Manov N, Yehezkelli Y, Kalkstein N, Akiva P, et al. Long covid outcomes at one year after mild SARS-CoV-2 infection: nationwide cohort study. BMJ. 2023. https://doi.org/10.1136/bmj-2022-072529.
Markov PV, Ghafari M, Beer M, Lythgoe K, Simmonds P, Stilianakis NI, et al. The evolution of SARS-CoV-2. Nat Rev Microbiol. 2023;21(6):361–79.
Volz E, Hill V, McCrone JT, Price A, Jorgensen D, O’Toole Á, et al. Evaluating the effects of SARS-CoV-2 spike mutation D614G on transmissibility and pathogenicity. Cell. 2021;184(1):64-75.e11.
Zhan J, Liu QS, Sun Z, Zhou Q, Hu L, Qu G, et al. Environmental impacts on the transmission and evolution of COVID-19 combing the knowledge of pathogenic respiratory coronaviruses. Environ Pollut. 2020;267:115621.
Chakraborty C, Chatterjee S, Bhattacharya M, Chopra H, Bhattacharya P, Islam MA, et al. The D614G mutation helps to increase the transmissibility and reduce the virulence of SARS-CoV-2 variants through natural selection. Int J Surg. 2023;109(2):171–4. https://doi.org/10.1097/JS9.0000000000000155.
Chen J, Gu C, Ruan Z, Tang M. Competition of SARS-CoV-2 variants on the pandemic transmission dynamics. Chaos Solitons Fractals. 2023. https://doi.org/10.1016/j.chaos.2023.113193.
Ramesh S, Govindarajulu M, Parise RS, Neel L, Shankar T, Patel S, Lowery P, Smith F, Dhanasekaran M, Moore T. Emerging SARS-CoV-2 variants: a review of its mutations, its implications and vaccine efficacy. Vaccines (Basel). 2021;9(10):1195. https://doi.org/10.3390/vaccines9101195.PMID:34696303;PMCID:PMC8537675.
Mengist HM, Kombe Kombe AJ, Mekonnen D, Abebaw A, Getachew M, Jin T. Mutations of SARS-CoV-2 spike protein: implications on immune evasion and vaccine-induced immunity. Semin Immunol. 2021. https://doi.org/10.1016/j.smim.2021.101533.
Manirambona E. Health policy and systems research in Sub-Saharan Africa during the COVID-19 Pandemic. Annals of Public Health. 2022. https://doi.org/10.55085/aph.2022.609.
Hirabara SM, Serdan TDA, Gorjao R, Masi LN, Pithon-Curi TC, Covas DT, et al. SARS-COV-2 variants: differences and potential of immune evasion. Front Cell Infect Microbiol. 2022. https://doi.org/10.3389/fcimb.2021.781429/full.
World Health Organization. Tracking SARS-CoV-2 variants. 2023. https://www.who.int/activities/tracking-SARS-CoV-2-variants
Sundararaj Stanleyraj J, Sethuraman N, Gupta R, Thiruvoth S, Gupta M, Ryo A. Treating COVID-19: are we missing out the window of opportunity? J Antimicrob Chemother. 2021;76(2):283–5.
Raman R, Patel KJ, Ranjan K. COVID-19: unmasking emerging SARS-CoV-2 variants, vaccines and therapeutic strategies. Biomolecules. 2021;11(7):993.
Chenchula S, Karunakaran P, Sharma S, Chavan M. Current evidence on efficacy of COVID-19 booster dose vaccination against the Omicron variant: a systematic review. J Med Virol. 2022;94(7):2969–76. https://doi.org/10.1002/jmv.27697.
Galloway SE, Paul P, MacCannell DR, Johansson MA, Brooks JT, MacNeil A, et al. Emergence of SARS-CoV-2 B.1.1.7 Lineage—United States, December 29, 2020–January 12, 2021. MMWR Morb Mortal Wkly Rep. 2021;70(3):95–9.
Volz E, Mishra S, Chand M, Barrett JC, Johnson R, Geidelberg L, et al. Assessing transmissibility of SARS-CoV-2 lineage B.1.1.7 in England. Nature. 2021;593(7858):266–9.
Davies NG, Abbott S, Barnard RC, Jarvis CI, Kucharski AJ, Munday JD, et al. Estimated transmissibility and impact of SARS-CoV-2 lineage B.1.1.7 in England. Science. 2021. https://doi.org/10.1126/science.abg3055.
Davies NG, Barnard RC, Jarvis CI, Russell TW, Semple MG, Jit M, et al. Association of tiered restrictions and a second lockdown with COVID-19 deaths and hospital admissions in England: a modelling study. Lancet Infect Dis. 2021;4:482–92.
Grint DJ, Wing K, Williamson E, McDonald HI, Bhaskaran K, Evans D, et al. Case fatality risk of the SARS-CoV-2 variant of concern B.1.1.7 in England, 16 November to 5 February. Eurosurveillance. 2021. https://doi.org/10.2807/1560-7917.ES.2021.26.11.2100256.
Tegally H, Wilkinson E, Giovanetti M, Iranzadeh A, Fonseca V, Giandhari J, et al. Detection of a SARS-CoV-2 variant of concern in South Africa. Nature. 2021;592(7854):438–43.
Mwenda M, Saasa N, Sinyange N, Busby G, Chipimo PJ, Hendry J, et al. Detection of B1351 SARS-CoV-2 variant strain—Zambia. MMWR Morb Mortal Wkly Rep. 2021;70(8):280–2.
Faria NR, Mellan TA, Whittaker C, Claro IM, Candido D da S, Mishra S, et al. Genomics and epidemiology of a novel SARS-CoV-2 lineage in Manaus, Brazil. medRxiv Prepr Serv Heal Sci. 2021.
Saville JW, Mannar D, Zhu X, Srivastava SS, Berezuk AM, Demers JP, et al. Structural and biochemical rationale for enhanced spike protein fitness in delta and kappa SARS-CoV-2 variants. Nat Commun. 2022;13(1):742.
Callaway E. Heavily mutated Omicron variant puts scientists on alert. Nature. 2021;600(7887):21–21.
Gu H, Krishnan P, Ng DYM, Chang LD, Liu GYZ, Cheng SSM, et al. Probable transmission of SARS-CoV-2 omicron variant in quarantine hotel, Hong Kong China. Emerg Infect Dis. 2022;28(2):460–2.
Arora P, Zhang L, Rocha C, Sidarovich A, Kempf A, Schulz S, et al. Comparable neutralisation evasion of SARS-CoV-2 omicron subvariants BA.1, BA.2, and BA.3. Lancet Infect Dis. 2022;22(6):766–7.
del Rio C, Malani PN. COVID-19 in 2022—the beginning of the end or the end of the beginning? JAMA. 2022;327(24):2389.
Gupta P, Gupta V, Singh CM, Singhal L. Emergence of COVID-19 variants: an update. Cureus. 2023. https://www.cureus.com/articles/130199-emergence-of-covid-19-variants-an-update
Mohapatra RK, Kandi V, Tuli HS, Chakraborty C, Dhama K. The recombinant variants of SARS-CoV-2: concerns continues amid COVID-19 pandemic. J Med Virol. 2022;94(8):3506–8. https://doi.org/10.1002/jmv.27780.
Maxmen A. Are new Omicron subvariants a threat Heres how scientists are keeping watch. Nature. 2022;604(7907):605–6.
Tegally H, Moir M, Everatt J, Giovanetti M, Scheepers C, Wilkinson E, et al. Emergence of SARS-CoV-2 omicron lineages BA.4 and BA.5 in South Africa. Nat Med. 2022;28(9):1785–90.
Callaway E. Coronavirus variant XBB15 rises in the United States—is it a global threat. Nature. 2023;613(7943):222–3.
NDTV. Covid-19 Variant BF.7 Detected In India: Symptoms, Precautions, Transmission Rate And More. https://www.ndtv.com/health/omicron-sub-variant-bf-7-detected-in-india-symptoms-precautions-transmission-rate-and-more-3628167?utm_medium=email&utm_source=transaction
TIME. What to Know About the New XBB.1.16 COVID-19 Variant. https://time.com/6268467/omicron-subvariant-xbb116-covid-19/
John OO, Olabode ON, Lucero-Prisno DE, Adebimpe OT, Singh A. XBB116 Omicron subvariant rise to a variant of interest: Implications for global alertness and preparedness. J Taibah Univ Med Sci. 2023;18(6):1285–7.
Kupferschmidt K, Wadman M. End of COVID-19 emergencies sparks debate. Science. 2023. https://doi.org/10.1126/science.adi6511.
Lenharo M. WHO declares end to COVID-19’s emergency phase. Nature. 2023. https://doi.org/10.1038/d41586-023-01559-z.
Rahimi F, Darvishi M, Bezmin Abadi AT. The end’—or is it? Emergence of SARS-CoV-2 EG.5 and BA.2.86 subvariants. Future Virol. 2024. https://doi.org/10.2217/fvl-2023-0150.
The Guardian. WHO declares ‘Eris’ Covid strain a variant of interest as cases rise globally. https://www.theguardian.com/world/2023/aug/09/who-declares-eris-covid-strain-a-variant-of-interest-as-uk-cases-rise
Dyer O. Covid-19: Infections climb globally as EG5 variant gains ground. BMJ. 2023. https://doi.org/10.1136/bmj.p1900.
Nigeria Centre for Disease Prevention and Control. Official Statement on the new COVID-19 Subvariants. https://ncdc.gov.ng/news/493/official-statement-on-the-new-covid-19-subvariants#:~:text=The recently discovered%2Freported BA,Africa%2C and the United States.
Okesanya OJ, Manirambona E, Buban JMA, Olabode ON. Coronavirus disease, emergency is over but the pandemic is not: implications for a new global order. Int J Surg Glob Heal. 2019. https://doi.org/10.1097/GH9.0000000000000207.
Khateeb J, Li Y, Zhang H. Emerging SARS-CoV-2 variants of concern and potential intervention approaches. Crit Care. 2021;25:244. https://doi.org/10.1186/s13054-021-03662-x.
Ao D, He X, Hong W, Wei X. The rapid rise of SARS-CoV-2 Omicron subvariants with immune evasion properties: XBB1.5 and BQ1.1 subvariants. MedComm. 2023. https://doi.org/10.1002/mco2.239.
European Centre for Disease Prevention and Control. SARS-CoV-2 variants of concern as of 5 Jan 2024. https://www.ecdc.europa.eu/en/covid-19/variants-concern
Euronews.net. “It’s not gone. It’s changing. It’s killing”: The COVID variants the WHO is watching closely. 2023. https://www.euronews.com/next/2023/11/24/its-not-gone-its-changing-its-killing-the-covid-variants-who-is-watching-closely
WHO. SARS-CoV-2 variant risk evaluation framework, 30 Aug 2023. https://www.who.int/publications/i/item/WHO-2019-nCoV-Variants-Risk-assessment-2023.1
TODAY. COVID variant HV.1 is still spreading. These are its most common symptoms. https://www.today.com/health/coronavirus/new-covid-variant-hv1-symptoms-rcna124198
Global News. ‘It’s here’: COVID subvariant HV.1 growing in Canada. What to know so far. https://globalnews.ca/news/10088387/covid-omicron-subvariant-hv-1-canada/
Health and Lifestyle. New COVID-19 Variant Expands Internationally. https://learningenglish.voanews.com/a/new-covid-19-variant-expands-internationally/7247990.html
Alam MS. Heliyon Insight into SARS-CoV-2 Omicron variant immune escape possibility and variant independent potential therapeutic opportunities. Heliyon. 2023;9(2):e13285. https://doi.org/10.1016/j.heliyon.2023.e13285.
Idris I, Adesola RO. Emergence and spread of JN1 COVID-19 variant. Bull Natl Res Cent. 2024;48(1):27. https://doi.org/10.1186/s42269-024-01183-5.
Quarleri J, Delpino MV, Galvan V. Anticipating the future of the COVID-19 pandemic: insights into the emergence of SARS-CoV-2 variant JN.1 and its projected impact on older adults. GeroScience. 2024;46(3):2879–83. https://doi.org/10.1007/s11357-024-01066-7.
Satapathy P, Kumar P, Mehta V, Suresh V, Khare A, Rustagi S, et al. Global spread of COVID-19’s JN1 variant: implications and public health responses. New Microbes New Infect. 2024;57:101225.
European Centre for disease prevention and control. SARS-CoV-2 variants of concern as of 26 April 2024. https://www.ecdc.europa.eu/en/covid-19/variants-concern
TIME. What to Know About the ‘FLiRT’ Variants of COVID-19. https://time.com/6972143/COVID-19-flirt-variants-kp-2/
Xbb T, Mahase E. Covid-19 : What do we know about XBB . 1 . 5 and should we be worried ? 2023;(January):1–2.
News Medical Life Science. Study shows XBB.1.5 exhibits a substantially higher hACE2-binding affinity compared to BQ.1.1 and XBB/XBB.1 [Internet]. https://www.news-medical.net/news/20230106/Study-shows-XBB15-exhibits-a-substantially-higher-hACE2-binding-affinity-compared-to-BQ11-and-XBBXBB1.aspx
Chakraborty C, Bhattacharya M, Dhama K. Cases of BA.2.75 and recent BA.2.75.2 subvariant of Omicron are increasing in India: Is it alarming at the global level? Ann Med Surg. 2022. https://doi.org/10.1016/j.amsu.2022.104963.
Chavda VP, Balar P, Vaghela D, Solanki HK, Vaishnav A, Hala V, et al. Omicron variant of SARS-CoV-2: an indian perspective of vaccination and management. Vaccines. 2023;11:1–22.
Singh J, Pandit P, McArthur AG, et al. Evolutionary trajectory of SARS-CoV-2 and emerging variants. Virol J. 2021;18:166. https://doi.org/10.1186/s12985-021-01633-w.
Korber B, Fischer WM, Gnanakaran S, Yoon H, Theiler J, Abfalterer W, et al. Tracking changes in SARS-CoV-2 spike: evidence that D614G Increases Infectivity of the COVID-19 Virus. Cell. 2020;182(4):812-827.e19.
Giurgea LT, Morens DM. Great expectations of COVID-19 herd immunity. MBio. 2022;13(1):e0349521.
Bazargan M, Elahi R, Esmaeilzadeh A. OMICRON : Virology, immunopathogenesis, and laboratory diagnosis. J Gene Med. 2022;27:1–15.
Bruijns B, Folkertsma L, Tiggelaar R. FDA authorized molecular point-of-care SARS-CoV-2 tests: a critical review on principles, systems and clinical performances. Biosens Bioelectron X. 2022;11:100158.
Du X, Tang H, Gao L, Wu Z, Meng F, Yan R, et al. Omicron adopts a different strategy from delta and other variants to adapt to host. Signal Transduct Target Ther. 2022;7(1):45.
Mahilkar S, Agrawal S, Chaudhary S, Parikh S, Sonkar SC, Verma DK, et al. SARS-CoV-2 variants: impact on biological and clinical outcome. Front Med. 2022. https://doi.org/10.3389/fmed.2022.995960/full.
Hachmann NP, Miller J, Collier A, et al. Neutralization escape by SARS-CoV-2 Omicron Subvariants BA.2.12.1, BA.4, and BA.5. N Engl J Med. 2022;387(1):86–8. https://doi.org/10.1056/NEJMc2206576.
McKeigue PM, McAllister DA, Hutchinson SJ, Robertson C, Stockton D, Colhoun HM. Vaccine efficacy against severe COVID-19 in relation to delta variant (B.1.617.2) and time since second dose in patients in Scotland (REACT-SCOT): a case-control study. Lancet Respir Med. 2022;10(6):566–72.
Yao L, Zhu KL, Jiang XL, Wang XJ, Zhan BD, Gao HX, et al. Omicron subvariants escape antibodies elicited by vaccination and BA.2.2 infection. Lancet Infect Dis. 2022;8:1116–7.
Andrews N, Stowe J, Kirsebom F, Toffa S, Rickeard T, Gallagher E, et al. Covid-19 vaccine effectiveness against the omicron (b.1.1.529) variant. N Engl J Med. 2022;386(16):1532–46. https://doi.org/10.1056/NEJMoa2119451.
Qu P, Evans JP, Faraone JN, Zheng YM, Carlin C, Anghelina M, et al. Enhanced neutralization resistance of SARS-CoV-2 Omicron subvariants BQ.1, BQ.1.1, BA.4.6, BF.7, and BA.2.75.2. Cell Host Microbe. 2023;31(1):9–17.
Zhou P, Yang XL, Wang XG, Hu B, Zhang L, Zhang W, et al. Addendum: A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature. 2020;588(7836):E6–E6.
Puray-Chavez M, LaPak KM, Schrank TP, Elliott JL, Bhatt DP, Agajanian MJ, et al. Systematic analysis of SARS-CoV-2 infection of an ACE2-negative human airway cell. Cell Rep. 2021;36(2):109364.
Yan K, Dumenil T, Tang B, Le TT, Bishop CR, Suhrbier A, et al. Evolution of ACE2-independent SARS-CoV-2 infection and mouse adaption after passage in cells expressing human and mouse ACE2. Virus Evol. 2022. https://doi.org/10.1093/ve/veac063/6648308.
Harvey WT, Carabelli AM, Jackson B, Gupta RK, Thomson EC, Harrison EM, et al. SARS-CoV-2 variants, spike mutations and immune escape. Nat Rev Microbiol. 2021;19(7):409–24.
Gravagnuolo AM, Faqih L, Cronshaw C, Wynn J, Klapper P, Wigglesworth M. High throughput diagnostics and dynamic risk assessment of SARS-CoV-2 variants of concern. EBioMedicine. 2021;70:103540.
Huang C, Wang Y, Li X, Ren L, Zhao J, Hu Y, et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan China. Lancet. 2020;395(10223):497–506.
Patel KP, Patel PA, Vunnam RR, Hewlett AT, Jain R, Jing R, et al. Gastrointestinal, hepatobiliary, and pancreatic manifestations of COVID-19. J Clin Virol. 2020;128:104386.
Gabarre P, Dumas G, Dupont T, Darmon M, Azoulay E, Zafrani L. Acute kidney injury in critically ill patients with COVID-19. Intensive Care Med. 2020;46(7):1339–48. https://doi.org/10.1007/s00134-020-06153-9.
Liu E, Dean CA, Elder KT. Editorial: The impact of COVID-19 on vulnerable populations. Front Public Health. 2023;8(11):1267723. https://doi.org/10.3389/fpubh.2023.1267723.PMID:37744477;PMCID:PMC10515203.
Wünsch K, Anastasiou OE, Alt M, Brochhagen L, Cherneha M, Thümmler L, van Baal L, Madel RJ, Lindemann M, Taube C, Witzke O, Rohn H, Krawczyk A, Jansen S. COVID-19 in elderly, immunocompromised or diabetic patients-from immune monitoring to clinical management in the hospital. Viruses. 2022;14(4):746. https://doi.org/10.3390/v14040746.PMID:35458476;PMCID:PMC9024512.
Bwire G, Ario AR, Eyu P, et al. The COVID-19 pandemic in the African continent. BMC Med. 2022;20:167. https://doi.org/10.1186/s12916-022-02367-4.
Uwizeyimana T, Manirambona E, Saidu Musa S, Uwiringiyimana E, Bazimya D, Mathewos K. Achieving COVID-19 herd immunity in Rwanda Africa. Public Health Chall. 2023;2:e75. https://doi.org/10.1002/puh2.75.
Dubey A, Choudhary S, Kumar P, Tomar S. Emerging SARS-CoV-2 variants: genetic variability and clinical implications. Curr Microbiol. 2022;79(1):20. https://doi.org/10.1007/s00284-021-02724-1.
Tregoning JS, Flight KE, Higham SL, Wang Z, Pierce BF. Progress of the COVID-19 vaccine effort: viruses, vaccines and variants versus efficacy, effectiveness and escape. Nat Rev Immunol. 2021;21(10):626–36.
Excler J, Saville M, Privor-Dumm L, et al. Factors, enablers and challenges for COVID-19 vaccine development. BMJ Glob Health. 2023;8: e011879.
Manirambona E, Hague O, Trajano LF, Killen A, Wilkins L, Nkeshimana M, Lucero-Prisno Iii DE. COVID-19 vaccines: ensuring social justice and health equity among refugees in Africa. Ann Glob Health. 2021;87(1):106. https://doi.org/10.5334/aogh.3415.
Manirambona E, Okesanya OJ, Shomuyiwa DO, Musa SS, Lucero-Prisno DE. Mitigating the threat of disease to global health security. New Microbes New Infect. 2024;57:101223.
Manirambona E, Musa SS, Irakoze S, Uwizeyimana T, Gyeltshen D, Bizoza D, et al. The impact of COVID-19 on international development aid and health systems strengthening in low-income countries. Ann Med Surg. 2022. https://doi.org/10.1016/j.amsu.2022.104772.
Haque A, Pant AB. Mitigating Covid-19 in the face of emerging virus variants, breakthrough infections and vaccine hesitancy. J Autoimmun. 2022;127:102792. https://doi.org/10.1016/j.jaut.2021.102792.
Rabaan AA, Al-Ahmed SH, Albayat H, Alwarthan S, Alhajri M, Najim MA, AlShehail BM, Al-Adsani W, Alghadeer A, Abduljabbar WA, Alotaibi N, Alsalman J, Gorab AH, Almaghrabi RS, Zaidan AA, Aldossary S, Alissa M, Alburaiky LM, Alsalim FM, Thakur N, Verma G, Dhawan M. Variants of SARS-CoV-2: influences on the vaccines’ effectiveness and possible strategies to overcome their consequences. Medicina. 2023;59(3):507. https://doi.org/10.3390/medicina59030507.
Alsaqqa HH. Sustaining the public health intervention strategies in confronting the Covid-19 pandemic. J Public health Res. 2022;11(2):227990362211024. https://doi.org/10.1177/22799036221102493.
European Centre for Disease Preventyion and Control. Public health control measures for COVID-19. https://www.ecdc.europa.eu/en/infectious-disease-topics/z-disease-list/covid-19/facts/public-health-control-measures-covid-19
Manirambona E, Lucero-Prisno DE, Shomuyiwa DO, Denkyira SA, Okesanya OJ, Haruna UA, et al. Harnessing wastewater based surveillance WBS in Africa a historic turning point towards strengthening the pandemic control. Discov Water. 2024;4(1):9.
Acknowledgements
Not applicable.
Funding
We declare that no funding was received for this work.
Author information
Authors and Affiliations
Contributions
EM and OJO conceptualized and wrote the original draft-revised the manuscript. NOO, TAO, and DELP III reviewed, edited, and approved the final draft of the manuscript. All authors contributed equally to the writing of this paper and have read and approved the final draft of this article for publication.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Approval from the ethical committee was not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Manirambona, E., Okesanya, O.J., Olaleke, N.O. et al. Evolution and implications of SARS-CoV-2 variants in the post-pandemic era. Discov Public Health 21, 16 (2024). https://doi.org/10.1186/s12982-024-00140-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12982-024-00140-x