Abstract
Background
Accumulating evidence indicates that long non-coding RNAs (lncRNAs) are involving in the tumorigenesis and metastasis of lung cancer. The aim of the study is to systematically characterize the lncRNA-associated competing endogenous RNA (ceRNA) network and identify key lncRNAs in the development of stage I lung adenocarcinoma (LUAD).
Methods
Totally, 1,955 DEmRNAs, 165 DEmiRNAs and 1,107 DElncRNAs were obtained in 10 paired normal and LUAD tissues. And a total of 8,912 paired lncRNA-miRNA-mRNA network was constructed. Using the Cancer Genome Atlas (TCGA) dataset, the module of ME turquoise was revealed to be most relevant to the progression of LUAD though Weighted Gene Co-expression Network Analysis (WGCNA).
Results
Of the lncRNAs identified, LINC00639, RP4-676L2.1 and FENDRR were in ceRNA network established by our RNA-sequencing dataset. Using univariate Cox regression analysis, FENDRR was a risk factor of progression free survival (PFS) of stage I LUAD patients (HRs = 1.69, 95%CI 1.07–2.68, P < .050). Subsequently, diffe rential expression of FENDRR in paired normal and LUAD tissues was detected significant by real-time quantitative (qRT-PCR) (P < 0.001).
Conclusions
This study, for the first time, deciphered the regulatory role of FENDRR/miR-6815-5p axis in the progression of early-stage LUAD, which is needed to be established in vitro and in vivo.
Similar content being viewed by others
Introduction
Lung cancer remains the prominent contributor of cancer-related mortality, with the worldwide 5-year survival rate of which is around 16.6% [1, 2]. Recently, with the wide application of low-dose computed tomography for early screening and the rapid development of target drugs for genetic mutations, the progress against lung cancer has achieved profound success. As of 2017, the mortality rate of lung cancer dropped from its peak by 51% among males and by 26% among females [2]. Nonetheless, the 5-year survival rate for lung cancer patients is still not well manifested. Additionally, the early-stage patients’ prognosis displays quite disparate from those of advanced-stage, with 5-year survival ranging from 85 to 6% [3], but its recurrence rate is still up to 90% after local resection or radical excision [4].
Currently, non-small cell lung cancer (NSCLC), accounting for approximately 85% of all lung cancer, has been endowed with several therapeutic options, including surgery, chemotherapy, radiation, target therapy and immunotherapy [5]. And surgery is the first choice of curative treatments for the medically operable. Owing to its readily entry into regional lymph nodes and apt to metastasize at an early stage, however, the recurrence rate accounts for approximately 27% to 38% for stage I NSCLC patients [6,7,8]. Therefore, intensive efforts have been directed to elucidate the molecular mechanism of premalignancy development and progression, and to identify potential molecular signatures for early diagnosis and interception.
Recently, long noncoding RNAs (lncRNAs) have attracted significant attentions in various cancers. LncRNA is a class of transcripts with length of more than 200 nucleotides that possesses limited or no protein-coding capacity, which is transcribed by RNA polymerase II, spliced, 5’capped, and polyadenylated [9]. It has been identified that lncRNAs were involved into diverse cellular, physiological and pathological process via a series of mechanism [10, 11], including serving as critical regulators of tumorigenesis and metastasis [12]. Furthermore, accumulating evidences revealed that lncRNA could disrupt miRNA-mediated degradation of target mRNAs by acting as “miRNA sponges” [13], indicating coding and noncoding RNAs could control one another through their ability to compete for miRNA binding locus, which termed as “ceRNA”. Under this hypothesis, a growing number of evidence revealed that ceRNA axis could contribute to tumorigenesis, progression and metastasis of cancer [14, 15]. For instance, researchers have demonstrated that LINC00336, an novel regulator of ferroptosis, could act as a ceRNA to affect tumor genesis and progression and mediate the expression of cystathionine-β-synthase (CBS) by sponging miR6852, which may serve as a potential therapeutic target of lung cancer [16]. Also, a novel lncRNA CCAT1 was reported to directly bind to miR-218 response elements and to promote gefitinib resistance of NSCLC by acting as a ceRNA of miR-218 to regulate HOXA1 expression [17]. However, most of previous studies on lncRNAs acting as ceRNAs were carried out on NSCLC patients with ignoring the heterogeneity of staging, and the clinical significance of tumor progression in early stage remains largely unknown.
Lung adenocarcinoma (LUAD) accounts for the most common subtype of NSCLC, which brings to light the necessity of distinguishing LUAD from other subtypes at an early stage to propose individual treatment. To date, it has been revealed that EGFR and KRAS mutations, and multiple other oncogenic and tumor suppressor genes were involved in the process of initiation and pathogenesis [18, 19]. Given tumor’s biological behaviors and molecular characteristic displaying heterogeneous at an early stage, some patients could experience a long-term survival, while others could not. To address this, we performed RNA-sequencing for stage I LUAD patients to identify differentially expressed lncRNAs (DElncRNAs) in paired normal and tumor tissues, following by ceRNAs network constructed. On this basis, lncRNA-related ceRNAs identified were validated by database from The Cancer Genome Atlas (TCGA) to establish clinical prognostic model. Furthermore, the hub gene related ceRNAs regulatory network of lncRNAs-miRNAs-mRNAs were established to broaden our knowledge of expression pattern.
Methods
Patients and clinical samples
A total of 10 paired LUAD and adjacent normal fresh tissues were obtained from Second Affiliated Hospital of Fujian Medical University between January, 2019 and May, 2019. The specimens were collected from LUAD patients with treatment-naïve who underwent primary surgical treatment. All of the specimens were immediately snap-frozen by liquid nitrogen after surgical resection, and stored at -80℃ until RNA extraction. The clinicopathological diagnosis were confirmed by two pathologists according to the guidelines of the World Health Organization (WHO, version 2015). All participants provided written informed consent, and the bioethical committees at The Second Affiliated Hospital of Fujian Medical University, China, gave written approval for the study (2020–206).
The other group resulted from the LUAD-related RNA-sequencing data in TCGA database, containing the expression data of mRNAs, miRNAs and lncRNAs. A total of 595 paired LUAD tissues RNA-seq data were retrieved, with their clinicopathological features. RNA-sequencing data and corresponding clinical information in LUAD were obtained from TCGA database as following criteria: 1) histologically diagnosed as LUAD; 2) data with complete clinical information; 3) diagnosed as stage I patients. Finally, a total of 260 paired LUAD tissues was included for further analysis. The present study meets the criteria of data usage and publishing of the National Cancer Institute of NIH, and no approval from the ethics committee was required. Progression-Free Survival (PFS) was calculated from the date when patients first received treatment until the date of progression or the last follow-up or death.
RNA extraction and sequencing
Total RNA of LUAD tissues and corresponding normal tissues was extraction from frozen tissues using RNeasy Mini Kit (Qiagen, Germany) following the manufacturer’s protocol. The RNA concentration was evaluated by the Qubit 4.0 (Thermo Fisher Scientific, Wilmington, DE, USA), and the RNA quality was evaluated by agarose gel eletrophoresis.
Then, ribosomal RNA was removed from total RNA to obtain the maximum residual ncRNA. After fragment of rRNA-depleted RNA, the cDNA library was constructed using the TruSeq RNA sample Prep Kit (Illumina, San Diego, CA, USA). LncRNA/mRNA sequencing libraries were prepared using VAHTS total RNA-seq Library Prep kit for Illumina (Vazyme NR603, China) according to the manufacturer’s protocol. The cDNA fragments with 150-bp paired-end reads were generated for RNA sequencing. Additionally, NEBNext® Multiplex Small RNA Library Prep Set for Illumina® (NEB) was used to prepare the miRNA library for samples. 12 libraries were pooled and sequenced in a single lane of Illumina HiSeq Xten sequencing platform. And Illumina’s TruSeq small RNA library preparation kit was used to prepare the miRNA library with 50-bp paired-end reads generated. After library construction, RNA sequencing for both lncRNA/mRNA and miRNA was performed using Illumina HiSeq Xten platform.
Identifications of DEmRNAs, DEmiRNA and DElncRNAs
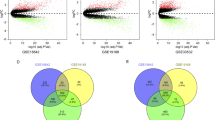
Mirdeep2 (v2.0.0.5) was used for new miRNA prediction, whose expression was calculated and standardized using CPM (counts per million read, CPM). LncRNAs were annotated by the following database, including PLEX (https://sourceforge.net/projects/plek/), CPAT (https://sourceforge.net/projects/rna-cpat/), CNCI (https://github.com/www-bioinfo-org/CNCI) and CPC2 (http://cpc2.cbi.pku.edu.cn/). The intersection was exhibited using Venn diagram. (Figure S 1).
DESeq2Rpackage(https://bioconductor.org/packages/release/bioc/html/DESeq2.html) in Bioconductor project was used to screen the differentially expressed mRNA (DEmRNAs), differentially expressed miRNA (DEmiRNAs) and differentially expressed lncRNA (DElncRNAs) between LUAD and normal tissues. |Log2(fold change)| (|log2FC|) ≥ 1 and statistical P value ≤ 0.05 were set as cut-off criteria. Finally, unsupervised hierarchical clustering was carried out for DE-RNAs, and the expression patterns of which in paired tissues were displayed in form of heatmap using the pheatmap R package (https://cran.r-project.org/web/packages/pheatmap/index.html).
Analysis of the DElncRNAs Enrichment pathway
Gene ontology (GO, http://www.bioconductor.org/packages/release/bioc/html/topGO.html) analysis was conducted to screen enrichment of target genes to determine the biological functions regulated by lncRNAs, including biological processes (BP), cellular component (CC), and molecular function (MF) annotations. Kyoto Encyclopedia of Genes and Genomes (KEGG, https://www.kegg.jp/kegg/kegg1.html) analysis was performed to annotate the signaling pathways mainly involved for target genes [20, 21]. The GO and pathway analysis with enriched gene count ≥ 2 and P value < 0.05 as the threshold for statistical significance.
Predication of miRNA regulation relationship
The predication of miRNA-gene analysis of DEmiRNA obtained was performed using the online website miRWalk 2.0 (http://zmf.umm.uni-heidelberg.de/pps/zmf/mirwalk2/). Furthermore, miRWalk, miRanda, miRDB, miRMap and TargetScan databases were used to predict the possible DEmiRNA-DEmRNA regulatory relationships. Using the StarBase (http://starbase.sysu.edu.cn/) database, the miRNA-lncRNA regulatory relationships by DEmiRNA were predict. Then the DEmRNA-DEmiRNA and DEmiRNA-DElncRNA regulatory regulation relationship were constructed based on shared miRNAs with which both lncRNAs and mRNAs interact, and were illustrated by Cytoscape software.
Construction of lncRNA-miRNA-mRNA ceRNA network
Under the hypothesis of miRNA sponge, the positive correlation expression of lncRNAs-mRNAs was focused on, therefore, and the positive correlation coexpression relationship between mRNAs and lncRNAs simultaneously regulated by miRNAs was obtained. ceRNA network were constructed based on shared miRNAs with which mRNAs and lncRNAs interact. For further analysis, we used a hypergeometric cumulative distribution function test to determine potential ceRNA pairs [22], with correlation coefficient > 0.5 and P value < 0.05 as a ceRNA triplet with statistical significance.
Weighted gene correlation network analysis (WGCNA)
As described previously [23, 24], gene co-expressed network analysis was performed on LUAD and adjacent normal tissues using R WGCNA package. The expression matrix was restricted to expressed lncRNAs, following by a Pearson correlation matrix and a weighted adjacency matrix generated. The module eigengene (ME) was calculated for each module, the values of which clinical traits associated with was calculated by Pearson’s correlation. In the present study, we set the optimal soft-thresholding power at 4 (scale-free R2 = 0.85), cut height at 0.25, and minimal modules sized to 30, to identify key modules. topological overlap matrix (TOM) was constructed and transformed. Then, the lncRNAs were grouped into co-expression modules using tree pruning of gene hierarchical clustering dendrograms by cutreeDynamic method, with correlated modules merged. The module significantly correlated with sample traits were used for hub genes selection.
LUAD-specific prognostic lncRNA signatures identification
Progression-free survival (PFS) time was calculated from the date when patients first received chemotherapy until the date of progression or the last follow-up or death. Using TCGA dataset, the associations between DElncRNAs in ceRNA network and PFS in stage I LUAD patients were evaluated using univariate Cox regression analysis. The samples were divided into high expression and low expression groups based on the median FPKM of lncRNAs. Kaplan–Meier (KM) survival analysis and Log-rank (LR) test were performed, and LR P value, hazard ratio (HR) with 95% confidence interval (CI) were computed. Survival and survminer R package were used for Cox regression analysis, and the statistical P < 0.05 was considered as significant.
RNA extraction and quantitative real-time RT-PCR assay
Total RNA was extracted using (Trizol®; Sigma) from 26 paired LUAD tissues, according to the manufacturer’s recommendations. After purification, the RNA was transcribed into cDNA using Reverse Transcription Kit (Takara, Tokyo, Japan). RT-PCR analysis was performed using the SYBR Green (Takara). The primers used for RT-PCR were: FENDRR primers were AGTCACAGCACCAGAAAGCCAAC (sense) and TGATGTTCTCCTTCTT GCCTCAGC (anti-sense); LINC00639 primers were GTGAGTGTTCAGACATGCCAGGAG (sense) and AGCCGAGTGGATTCAGCGAGAG (anti-sense); RP4-676L2.1 primers were TGTTTGAAGCCGTGAGACTGAGTG (sense) and CTCCTTTGCTGGCTC TTCCTCATC (anti-sense); GAPDH primers were CTCCTGCACCACCAACTGCTTAG (sense) and GACGCCTGCTTCACCACCTTC (anti-sense); miR-6815-5p primers were AGGTGGCGCCGGAGGA (sense) and AGTGCAGGGTCCGAGGTATT (anti-sense); U6 primers were CTCGCTTCGGCAGCACA (sense) and AACGCTTCACGAATTTGCGT (anti-sense). The cycle conditions were as follows: 95 °C 10 min, followed by 40 cycles of 95 °C for 15 s, 65 °C for 30 s, and 72 °C for 30 s. Relative quantification was calculated using the 2−ΔΔCT method.
Results
Identification of DEmRNAs, DEmiRNAs and DElncRNAs
The schematic diagram was constructed to display the methods of the present study (Fig. 1). Initially, to explore the role of lncRNAs in stage I LUAD, we performed RNA-seq analysis of total RNA in 10 paired tumor and adjacent normal tissues. According to the criteria of |log2FC|≥ 1 and P value ≤ 0.05 cut-off, a total of 1,955 DEmRNAs (1,372 up- and 583 down-regulated), 165 DEmiRNAs (111 up- and 54 down-regulated) and 1,107 DElncRNAs (775 up- and 332 down-regulated) were detected from dataset. The heatmaps of DEmRNAs, DEmiRNAs and DElncRNAs were presented in Fig. 2A-C, and the top 20 of which were displayed in Table S1, 2, 3, respectively.
To explore the significance of DElncRNAs that might involve in LUAD, we performed functional enrichment analysis of target mRNAs. It implied that the most significant BP, CC and MF by DElncRNAs enriched were embryo development (GO:0,009,790, P = 3.10E-09), presequence translocase-associated import motor (GO:0,001,405, P = 0.0002) and nucleic acid binding transcription factor activity (GO:0,001,071, P = 6.10E-07), respectively. Figure 2D displayed the top 20 GO terms enriched by DElncRNAs. Additionally, pathway enrichment analysis was performed to reveal potential biological function of DElncRNAs. It showed that the top 20 pathways were significantly enriched using Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis, with glycerophospholipid metabolism pathway enriching the most significant genes in the network (Fig. 2E and F).
ceRNA network construction
With the criteria of total score ≥ 150 and total energy ≤ -20, we used miRanda to predict the regulation relationship between miRNA and mRNA, as well as the relationship between miRNA and lncRNA (Fig. 3A and B). According to the regulation of DElncRNA-DEmiRNA and DEmRNA-DEmiRNA, lncRNAs and mRNAs regulated by the same miRNA were screened. Using the parameter of Pearson correlation (PPC) > 0.5 and P value < 0.05 as a threshold, a total of 8,912 interaction relationships of lncRNA-miRNA-mRNA with statistical significance were predicted in 10 paired LUAD tissues, and the top 20 ceRNA network were listed in Table S4.
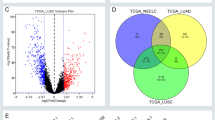
Analysis of expression profile and function enrichment. A-C The heatmaps of DEmRNAs, DEmiRNAs and DElncRNAs in the expression profile in 10 paired LUAD and adjacent normal tissue. D Top 20 GO terms enriched in DElncRNAs. E–F Top 20 pathways enriched in DElncRNAs using Kyoto Encyclopedia of Genes and Genomes (KEGG) analysis and displayed in bubble chart (included predominant metabolic pathway—map 00,564). BP: biological processes; CC: cellular component; MF: molecular function
Co-expression module identification and hub gene selection
To further explore the potential function of lncRNAs in the progression of LUAD, the transcriptomic data of 260 stage I LUAD patients and their corresponding clinical data in TCGA database were extracted to construct WGCNA network. In this study, we chose β = 4 (R2 = 0.85) as a soft threshold to construct a scale-free topology network (Fig. 4A). The TOM including all genes was depicted by topological overlapping heat map. Totally, 43 gene co-expression modules were identified by linkage hierarchical clustering according to TOM-based dissimilarity measure (1-TOM). Finally, we identified 7 modules relevant to clinical traits, as eigengene adjacency heatmap descripted (Fig. 4B). Of these, the turquoise module was most highly correlated with DElncRNAs (Fig. 4B and C). Interestingly, we found that the correlation coefficient between royalblue module and progressive disease reached to 0.94, indicating that the royalblue module is a gene set specifically associated with progression of disease. The royalblue module is also the most relevant module to distant metastasis (cor = 0.72), indicating the correlation of lncRNAs with prognosis (Fig. 4D). By setting the module membership (MM) to > 0.8 and the gene significance (GS) to > 0.4 as threshold, we selected a total of 81 hub lncRNAs from the modules (Table S 5), most of which were involved in the recurrence or metastasis of disease. The results indicated that the DElncRNAs identified were involved in the prognosis of stage I LUAD patients.
Construction of ceRNA network. A-B The regulation relationship between miRNA and mRNA predicted by miRanda, and the relationship between miRNA and lncRNA predicted by miRanda, with the criteria of total score ≥ 150 and total energy ≤ -20. The green node represents lncRNAs; the red node represents miRNAs; the blue node represents mRNAs
Subsequently, we found that 19 genes were associated with distant metastasis, 5 genes associated with locoregional recurrence and 11 genes associated with distant metastasis, and another 65 genes associated with new primary tumor. Of the hub genes identified, FENDRR, LINC00639 and RP4-676L2.1 were also predicted as ceRNAs in stage I LUAD constructed by our database. Furthermore, we constructed ceRNA network using the three lncRNAs by Cytoscape (Fig. 5A). Finally, ceRNA network containing 174 lncRNA-miRNA-mRNA regulatory relation for FENDRR, 937 lncRNA-miRNA-mRNA regulatory relation for LINC00639, and 54 regulatory relation for RP4-676L2.1. It showed that and LINC00639 were hub nodes that could target more miRNAs and mRNAs in the network. These findings indicated that DElncRNAs regulated mRNA expression via interaction with miRNAs.
WGCNA and identification of significant modules. A The soft-thresholding power in WGCNA. B The eigengene of each colored module were calculated to establish an adjacent matrix. C Cluster dendrogram obtained from lncRNAs data of stage I LUAD in TCGA dataset with average hierarchical linkage clustering. The color row underneath the dendrogram represents the module assigned by Dynamic Tree Cut. D Module-trait relationship heatmap. The row represents the modules, while the column represents the trait. The values in the box represents the correlation and P values
The relationship and expression between lncRNAs and miRNAs. A The ceRNA network constructed by FENDRR, LINC00639 and RP4-676L2.1. B-C The FENDRR and miR-6815-5p expression levels of stage I LUAD tissues and paired normal tissues were tested by RT-PCR (n = 26). D Prognostic significance of FENDRR expression on PFS for stage I LUAD patients by the median value as cutoff
Interestingly, one of subnetworks showed that lncRNA FENDRR only act as a sponge for hsa-miR-6815-5p to regulated mRNAs, including TNS1, PDLIM2, PPFIBP1, SCMH1, PLXDC1 and so on, which mainly involved in adherens junction, chromatin silence, blood vessel development (Figure S2A). Thus, we further explore the relationship between FENDRR and hsa-miR-6815-5p. In our analysis of RNA sequencing, the results showed that FENDRR and LINC00639 were significantly downregulated, while RP4-676L2.1 and hsa-miR-6815-5p was marly upregulated in stage I LUAD patients (P value < 0.05, Figure S 2B). Further, RT-PCR indicated that expression of FENDRR was significantly decreased and expression of hsa-miR-6815-5p was markedly increased in LUAD tissues compared with paired normal tissues (P value < 0.001) (Fig. 5B and C), while the expression of RP4-676L2.1 and LINC00639 were not significant (Figure S2C). Thus, the results revealed that down-regulation of FENDRR maybe involved in tumorigenesis via upregulating of hsa-miR-6815-5p.
To further explore the FENDRR identified, we evaluated its effects on recurrence by univariate Cox regression analysis in TCGA dataset. It showed that FENDRR was significantly associated with PFS (HRs = 1.69, 95%CI 1.07–2.68, P < 0.05), indicating that FENDRR could be regarded as a prognostic factor of tumor recurrence. Using median FPKM of FENDRR as cutoff value, we divided the patients into high-expression group and low-expression group. KM survival curve for the two groups indicated that PFS in the high FENDRR expression group displayed significantly longer than that in the low FENDRR expression group (Fig. 5D), while PFS for RP4-676L2.1 and LINC00639 between two groups were not significantly different (Figure S2D).
Discussion
Although it has achieved great improvement on clinical management for LUAD aided by the discovery of genetic mutations, there is still main obstacles on improving prognosis of patients resulting from its apt to migration and invasion. Therefore, the promise of utilization of lncRNAs in predicating clinical prognosis has attracted much attention in translational research. In this study, we identify three lncRNAs, FENDRR, LINC00639 and RP4-676L2.1, which might be related with prognosis of stage I LUAD patients. However, only FENDRR is predicted to be associated with PFS of patients, and validated to be down-regulated in LUAD tissues by RT-PCR assay. The primary results uncover the potential role of FENDRR in the progression of early-stage LUAD. Importantly, the study gives a novel hint of the mechanism by which FENDRR might involve in the progression of disease.
Previous studies have revealed that lncRNAs could exert their biological function via interaction with DNA, RNA and protein depending on their location within cells [25]. Although there is still a lot to learn about lncRNA, accumulating studies demonstrated its function on relieving target mRNAs degradation mediated by miRNA, playing a role as ceRNAs, which could act as vital regulators in various physiological and pathological process of tumor [26, 27]. To date, despite the plethora of reports, several lncRNAs have been identified to be involved in recurrence and metastasis of lung cancer [28,29,30]. In this study, we explored the most likely significant lncRNAs profile alternations of stage I LUAD, and revealed that FENDRR, LINC00639 and RP4-676L2.1 were predicted to be associated with progression of disease. However, only FENDRR were validated as predictive lncRNA of PFS by cox regression analysis, indicating its underlying function responsible for progression of LUAD.
LncRNA fetal-lethal non-coding developmental regulatory RNA (FENDRR), also named as FOXF1-AS1, is an intergenic lncRNA with consisting of seven exons. It is located at 3q13.31, 1,354 bp upstream of transcriptional start site, which is transcribed from a bidirectional promoter shared with the protein coding gene Foxf1 and Pitx2. Previous studies revealed that FENDRR could regulate cell migration, invasion, and lymphatic metastasis, demonstrating its inhibitory regulation in tumor progression [31]. It was reported that FENDRR was highly expressed in the lung, while was lowly expressed in the liver, colon and brain tissues, and the level of which was associated with prognosis of patients [32]. In breast cancer, the low level of FENDRR was associated with poor prognosis of patients, including a shorter survival and a shorter PFS [33]. Also, FENDRR was found to be related with survival of gastric cancer patients, the expression of which could be suppressed in gastric cancer associated fibroblasts by hypermethylation [34]. In agreement with previous studies [31], the level of FENDRR expression in tumor was also shown to be suppressed compared with paired normal tissue.
The mechanism of anti-malignant effects for FENDRR might involve in inhibiting cell migration, invasion and mediated stem-like properties by regulating epithelial mesenchymal transition (EMT) [35,36,37]. Previous studies reported that FENDRR could anchor PRC2 and/or TrxG/MLL complexes at its target promoters, increasing PRC2 occupancy and H2K27 trimethylation, which lead to the attenuation of target gene expression [38]. In gastric cancer, FENDRR was found to increase cell migration and invasion via up-regulation of FN1, MMP2 and MMP9 [34]. In vitro, FENDRR was revealed to decrease the IC50 for cisplatin in A549/DDP cells, and depressed chemotherapy resistance to cisplatin in NSCLC [37]. Hitherto, the contribution of FENDRR on progression of LUAD has not been well clarified. In our dataset, FENDRR could targeted miR-6815-5p in ceRNA network constructed, suggesting that it might exert function on regulation expression of target mRNAs by ‘sponging’ miR-6815-5p. To date, only one study was found that miR-6815-5p was significantly upregulated in exosomes of HPV16-infected cervical-vaginal fluid (CVF) based on microarray analysis, while its biological functions was not reported and unclear [39]. Hence, further studies on miR-6815-5p are warranted. To the best our knowledge, this is the first study where predicts the regulation of miR-6815-5p on targeted mRNAs in LUAD. Apart from the predication of lncRNAs on PFS, the main breakthrough of our study is centred around the regulation of miR-6815-5p on FENDRR, which needs to explore its molecular mechanisms to increase further confidence to this result.
Regarding RP4-676L2.1 and LINC00639, we speculated that they could be related with recurrence of disease by WGCNA using TCGA data set, suggesting their prognostic value in stage I LUAD. However, they were not significantly correlated with PFS of stage I LUAD patients by cox regression analysis. Surprisingly, RP4-676L2.1 and LINC00639 were also not successfully validated to be consistent difference of expression between tumor and normal lung tissue by qPCR. We speculated that it may lie with the bioinformatic analysis. Still, further studies are needed to validate using larger samples.
The discovery of lncRNAs contributes significantly to clinical prognosis in stage I LUAD patients. Its strength lies in not only testing using the sequencing dataset, but also validating using TCGA dataset by bioinformatics methods. Still, some limitations must be noted. Firstly, we speculated that FENDRR/miR-6815-5p axis may play an important role in the biological behavior of LUAD based on bioinformatics analysis and RT-PCR, but the specific mechanism still needs to be further verified such as the functional experiments of LUAD cells and RNA-RNA interaction verification experiments. Secondly, due to the low number of samples performed RNA-seq, the WGCNA was constructed using TCGA dataset, which might induce racial bias. This is not surprising, a substantial number of studies on lncRNAs have been validated using TCGA dataset. Thirdly, the lncRNAs identified from RNA-seq were inconsistent with the qPCR results, which may result from incorrect annotation in the bioinformatic analysis.
Conclusions
Taken together, this study contributed significantly to the wider knowledge of lncRNAs and ceRNAs involvement of progression of stage I LUAD. Out of the three lncRNAs validated, FENDRR is validated to involve in the recurrence of disease, which confirm previous findings. Based on the FENDRR-related ceRNA network constructed, it was revealed the regulation of miR-6815-5p on FENDRR for the first time, therefore open to further research to explore molecular mechanism. However, studies are still needed to establish the role of FENDRR/miR-6815-5p axis in the progression of early-stage LUAD.
Availability of data and materials
The datasets generated during the current study are not publicly available due to concerns regarding patient confidentiality and proprietary information but are available upon reasonable request from the corresponding author. we provided a reviewer link of unpublished BioProject. Use the following URL: https://dataview.ncbi.nlm.nih.gov/object/PRJNA732584?reviewer=jo5ph00rjrkr7u19thhft8eho.
Abbreviations
- lncRNAs:
-
Long non-coding RNAs
- NSCLC:
-
Non-small cell lung cancer
- LUAD:
-
Lung adenocarcinoma
- ceRNA:
-
Competing endogenous RNA
- PFS:
-
Progression free survival
- CBS:
-
Cystathionine-β-synthase
- RIG:
-
Retinoid-inducible protein
- TCGA:
-
The Cancer Genome Atlas
- DElncRNAs:
-
Identify differentially expressed lncRNAs
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- BP:
-
Biological processes
- CC:
-
Cellular component
- MF:
-
Molecular function
- WGCNA:
-
Weighted gene correlation network analysis
- KM:
-
Kaplan–Meier
- HR:
-
Hazard ratio
- FENDRR:
-
Fetal-lethal non-coding developmental regulatory RNA
References
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68(6):394–424. https://doi.org/10.3322/caac.21492.
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2020. CA Cancer J Clin. 2020;70(1):7–30. https://doi.org/10.3322/caac.21590.
Goldstraw P, Chansky K, Crowley J, Rami-Porta R, Asamura H, Eberhardt WE, Nicholson AG, Groome P, Mitchell A, Bolejack V, et al. The IASLC Lung Cancer Staging Project: Proposals for Revision of the TNM Stage Groupings in the Forthcoming (Eighth) Edition of the TNM Classification for Lung Cancer. J Thorac Oncol. 2016;11(1):39–51. https://doi.org/10.1016/j.jtho.2015.09.009.
Vansteenkiste J, De Ruysscher D, Eberhardt WE, Lim E, Senan S, Felip E, Peters S, ESMO Guidelines Working Group. Early and locally advanced non-small-cell lung cancer (NSCLC): ESMO Clinical Practice Guidelines for diagnosis, treatment and follow-up. Ann Oncol. 2013;24(Suppl 6):vi89–98. https://doi.org/10.1093/annonc/mdt241.
Herbst RS, Morgensztern D, Boshoff C. The biology and management of non-small cell lung cancer. Nature. 2018;553(7689):446–54. https://doi.org/10.1038/nature25183.
Hung JJ, Hsu WH, Hsieh CC, Huang BS, Huang MH, Liu JS, Wu YC. Post-recurrence survival in completely resected stage I non-small cell lung cancer with local recurrence. Thorax. 2009;64(3):192–6. https://doi.org/10.1136/thx.2007.094912.
Spiro SG, Tanner NT, Silvestri GA, Janes SM, Lim E, Vansteenkiste JF, Pirker R. Lung cancer: progress in diagnosis, staging and therapy. Respirology. 2010;15(1):44–50. https://doi.org/10.1111/j.1440-1843.2009.01674.x.
Guerrera F, Errico L, Evangelista A, Filosso P, Ruffini E, Lisi E, Bora G, Asteggiano E, Olivetti S, Lausi P, et al. Exploring Stage I non-small-cell lung cancer: development of a prognostic model predicting 5-year survival after surgical resection†. Eur J Cardiothorac Surg. 2015;47(6):1037–43. https://doi.org/10.1093/ejcts/ezu410.
Rinn J, Chang HJArob. Genome regulation by long noncoding RNAs. 2012;81:145–66. https://doi.org/10.1146/annurev-biochem-051410-092902.
Cabili M, Trapnell C, Goff L, Koziol M, Tazon-Vega B, Regev A, Rinn JJG. Development: Integrative annotation of human large intergenic noncoding RNAs reveals global properties and specific subclasses. 2011;25(18):1915–27. https://doi.org/10.1101/gad.17446611.
Geisler S, Coller JJNrMcb. RNA in unexpected places: long non-coding RNA functions in diverse cellular contexts. 2013;14(11):699–712. https://doi.org/10.1038/nrm3679.
Weidle U, Birzele F, Kollmorgen G, Rüger RJCg. Proteomics: Long Non-coding RNAs and their Role in Metastasis. Cancer Genomics Proteomics. 2017;14(3):143–60. https://doi.org/10.21873/cgp.20027.
Karreth F, Tay Y, Perna D, Ala U, Tan S, Rust A, DeNicola G, Webster K, Weiss D, Perez-Mancera P, et al. In vivo identification of tumor- suppressive PTEN ceRNAs in an oncogenic BRAF-induced mouse model of melanoma. 2011;147(2):382–95. https://doi.org/10.1016/j.cell.2011.09.032.
Cai J, Fang L, Huang Y, Li R, Xu X, Hu Z, Zhang L, Yang Y, Zhu X, Zhang H, et al. Simultaneous overactivation of Wnt/β-catenin and TGFβ signalling by miR-128–3p confers chemoresistance-associated metastasis in NSCLC. Nat Commun. 2017;8:15870. https://doi.org/10.1038/ncomms15870.
Yuan Y, Liao H, Pu Q, Ke X, Hu X, Ma Y, Luo X, Jiang Q, Gong Y, Wu M, et al. miR-410 induces both epithelial-mesenchymal transition and radioresistance through activation of the PI3K/mTOR pathway in non-small cell lung cancer. Signal Transduct Target Ther. 2020;5(1):85. https://doi.org/10.1038/s41392-020-0182-2.
Wang M, Mao C, Ouyang L, Liu Y, Lai W, Liu N, Shi Y, Chen L, Xiao D, Yu F, et al. Long noncoding RNA LINC00336 inhibits ferroptosis in lung cancer by functioning as a competing endogenous RNA. Cell Death Differ. 2019;26(11):2329–43. https://doi.org/10.1038/s41418-019-0304-y.
Jin X, Liu X, Zhang Z, Guan Y. lncRNA CCAT1 Acts as a MicroRNA-218 Sponge to Increase Gefitinib Resistance in NSCLC by Targeting HOXA1. Molecular therapy Nucleic acids. 2020;19:1266–75. https://doi.org/10.1016/j.omtn.2020.01.006.
Pasquali S, Chiswell K, Hall M, Thibault D, Romano J, Gaynor J, Shahian D, Jacobs M, Gaies M, O'Brien S, et al. Estimating Resource Utilization in Congenital Heart Surgery. 2020;110(3):962–68. https://doi.org/10.1016/j.athoracsur.2020.01.013.
Kadota K, Sima C, Arcila M, Hedvat C, Kris M, Jones D, Adusumilli P, Travis WJTAjosp. KRAS Mutation Is a Significant Prognostic Factor in Early-stage Lung Adenocarcinoma. 2016;40(12):1579–90. https://doi.org/10.1097/pas.0000000000000744.
Kanehisa M, Furumichi M, Tanabe M, Sato Y, Morishima K. KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Res. 2017;45:D353–61. https://doi.org/10.1093/nar/gkw1092.
Kanehisa M, Sato Y, Kawashima M, Furumichi M, Tanabe M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 2016;44:D457–62. https://doi.org/10.1093/nar/gkv1070.
Sumazin P, Yang X, Chiu HS, Chung WJ, Iyer A, Llobet-Navas D, Rajbhandari P, Bansal M, Guarnieri P, Silva J, Califano A. An extensive microRNA-mediated network of RNA-RNA interactions regulates established oncogenic pathways in glioblastoma. Cell. 2011;147(2):370–81. https://doi.org/10.1016/j.cell.2011.09.041.
Langfelder P, Horvath SJBb. WGCNA: an R package for weighted correlation network analysis. BMC Bioinformatics. 2008;9:559. https://doi.org/10.1186/1471-2105-9-559.
Langfelder P, Horvath S. Fast R Functions for Robust Correlations and Hierarchical Clustering. J Stat Softw. 2012;46(11):i11.
Chen C, He W, Huang J, Wang B, Li H, Cai Q, Su F, Bi J, Liu H, Zhang B, et al. LNMAT1 promotes lymphatic metastasis of bladder cancer via CCL2 dependent macrophage recruitment. Nat Commun. 2018;9(1):3826. https://doi.org/10.1038/s41467-018-06152-x.
Xu J, Meng Q, Li X, Yang H, Xu J, Gao N, Sun H, Wu S, Familiari G, Relucenti M, et al. Long Noncoding RNA MIR17HG Promotes Colorectal Cancer Progression via miR-17–5p. Cancer Res. 2019;79(19):4882–95. https://doi.org/10.1158/0008-5472.Can-18-3880.
Yu W, Ding J, He M, Chen Y, Wang R, Han Z, Xing E, Zhang C, Yeh SJO. Estrogen receptor β promotes the vasculogenic mimicry (VM) and cell invasion via altering the lncRNA-MALAT1/miR-145–5p/NEDD9 signals in lung cancer. Oncogene. 2019;38(8):1225–38. https://doi.org/10.1038/s41388-018-0463-1.
Yuan S, Xiang Y, Wang G, Zhou M, Meng G, Liu Q, Hu Z, Li C, Xie W, Wu N, et al. Hypoxia-sensitive LINC01436 is regulated by E2F6 and acts as an oncogene by targeting miR-30a-3p in non-small cell lung cancer. Mol Oncol. 2019;13(4):840–56. https://doi.org/10.1002/1878-0261.12437.
Wu D, Yang B, Chen J, Xiong H, Li Y, Pan Z, Cao Y, Chen J, Li T, Zhou S, et al. Upregulation of long non-coding RNA RAB1A-2 induces FGF1 expression worsening lung cancer prognosis. Cancer Lett. 2018;438:116–25. https://doi.org/10.1016/j.canlet.2018.09.016.
Li C, Wan L, Liu Z, Xu G, Wang S, Su Z, Zhang Y, Zhang C, Liu X, Lei Z, et al. Long non-coding RNA XIST promotes TGF-β-induced epithelial-mesenchymal transition by regulating miR-367/141-ZEB2 axis in non-small-cell lung cancer. Cancer Lett. 2018;418:185–95. https://doi.org/10.1016/j.canlet.2018.01.036.
Acha-Sagredo A, Uko B, Pantazi P, Bediaga NG, Moschandrea C, Rainbow L, Marcus MW, Davies MPA, Field JK, Liloglou T. Long non-coding RNA dysregulation is a frequent event in non-small cell lung carcinoma pathogenesis. Br J Cancer. 2020;122(7):1050–8. https://doi.org/10.1038/s41416-020-0742-9.
Sauvageau M, Goff LA, Lodato S, Bonev B, Groff AF, Gerhardinger C, Sanchez-Gomez DB, Hacisuleyman E, Li E, Spence M, Liapis SC, et al. Multiple knockout mouse models reveal lincRNAs are required for life and brain development. Elife. 2013;2:e01749. https://doi.org/10.7554/eLife.01749.
Li Y, Zhang W, Liu P, Xu Y, Tang L, Chen W, Guan X. Long non-coding RNA FENDRR inhibits cell proliferation and is associated with good prognosis in breast cancer. Onco Targets Ther. 2018;11:1403–12. https://doi.org/10.2147/ott.S149511.
Xu TP, Huang MD, Xia R, Liu XX, Sun M, Yin L, Chen WM, Han L, Zhang EB, Kong R, De W, Shu YQ. Decreased expression of the long non-coding RNA FENDRR is associated with poor prognosis in gastric cancer and FENDRR regulates gastric cancer cell metastasis by affecting fibronectin1 expression. J Hematol Oncol. 2014;7:63. https://doi.org/10.1186/s13045-014-0063-7.
Wei H, Nickoloff J, Chen W, Liu H, Lo W, Chang Y, Yang P, Wu C, Williams D, Gelovani J, et al. FOXF1 mediates mesenchymal stem cell fusion-induced reprogramming of lung cancer cells. Oncotarget. 2014;5(19):9514–29. https://doi.org/10.18632/oncotarget.2413.
Miao L, Huang Z, Zengli Z, Li H, Chen Q, Yao C, Cai H, Xiao Y, Xia H, Wang Y. Loss of long noncoding RNA FOXF1-AS1 regulates epithelial-mesenchymal transition, stemness and metastasis of non-small cell lung cancer cells. Oncotarget. 2016;7(42):68339–49. https://doi.org/10.18632/oncotarget.11630.
Kun-Peng Z, Chun-Lin Z, Xiao-Long M. Antisense lncRNA FOXF1-AS1 Promotes Migration and Invasion of Osteosarcoma Cells Through the FOXF1/MMP-2/-9 Pathway. Int J Biol Sci. 2017;13(9):1180–91. https://doi.org/10.7150/ijbs.21722.
Grote P, Wittler L, Hendrix D, Koch F, Währisch S, Beisaw A, Macura K, Bläss G, Kellis M, Werber M, Herrmann BG. The tissue-specific lncRNA Fendrr is an essential regulator of heart and body wall development in the mouse. Dev Cell. 2013;24(2):206–14. https://doi.org/10.1016/j.devcel.2012.12.012.
Wu Y, Wang X, Meng L, Li W, Li C, Li P, Xu S. Changes of miRNA Expression Profiles from Cervical-Vaginal Fluid-Derived Exosomes in Response to HPV16 Infection. Biomed Res Int. 2020;2020:7046894. https://doi.org/10.1155/2020/7046894.
Acknowledgements
Not applicable.
Funding
The work was supported by Fujian Provincial Health Fund for Young and Middle-aged People (2019-ZQNB-7) and Quanzhou major science and technology projects (2018-QDZX-9).
Author information
Authors and Affiliations
Contributions
XY, LGF, LYF, LXB and ZYM participated in the study design. XY, LGF and LYF performed the most of the RNA-seq analysis and drafted the paper. LYF and LXB performed the WGCNA. LH, GZF and XYX participated in the design of figures. LXB, CSH and YJS participated in the acquisition of LUAD tissues. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
The ethics committee of The Second Affiliated Hospital of Fujian Medical University approved this study and is in compliance with the Helsinki Declaration. Written informed consent was obtained from all individual participants included in the study (2020–206).
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1:
Table S1. Top 20 DEmRNAs identified between tumor and adjacent normal tissues in stage I LUAD patients. Table S2. Top 20 DEmiRNAs identified between tumor and adjacent normal tissues in stage I LUAD patients. Table S3. Top 20 DElncRNAs identified between tumor and adjacent normal tissues in stage I LUAD patients. Table S4. Top 20 ceRNA network constructed in 10 stage I LUAD patients. Table S5. Hub lncRNAs predicted by WGCNA using TCGA dataset.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Xu, Y., Lin, G., Liu, Y. et al. An integrated analysis of the competing endogenous RNA network associated of prognosis of stage I lung adenocarcinoma. BMC Cancer 22, 188 (2022). https://doi.org/10.1186/s12885-022-09290-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12885-022-09290-0