Abstract
Background
The basic helix-loop-helix (bHLH) transcription factors play important roles in many processes in plant growth, metabolism and responses to abiotic stresses. Although, the sequence of Chinese white pear genome (cv. ‘Dangshansuli’) has already been reported, there is still a lack of clarity regarding the bHLH family genes and their evolutionary history.
Results
In this work, a genome-wide identification of the bHLH genes in Chinese white pear was performed, and we characterized the functional roles of these PbrbHLH genes in response to abiotic stresses. Based on the phylogenetic analysis and structural characteristics, 197 identified bHLH genes could be well classified into 21 groups. Expansion of PbrbHLH gene family was mainly driven by WGD and dispersed duplication with the purifying selection from the recent WGD. The functional annotation enrichment showed that the majority of PbrbHLHs were enriched in the GO terms and KEGG pathways involved in responds to stress conditions as TFs. Transcriptomic profiles and qRT-PCR revealed that PbrbHLH7, PbrbHLH8, PbrbHLH128, PbrbHLH160, PbrbHLH161 and PbrbHLH195 were significantly up-regulated under cold and drought treatments. In addition, PbrbHLH195-silenced pear seedlings display significant reduced cold tolerance, exhibiting reduced chlorophyll content, as well as increased electrolyte leakage and concentrations of malondialdehyde and H2O2.
Conclusion
For the first time, a comprehensive analysis identified the bHLH genes in Chinese white pear and demonstrated that PbrbHLH195 is involved in the production of ROS in response to cold stress, suggesting that members of the PbrbHLH family play an essential role in the stress tolerance of pear.
Similar content being viewed by others
Background
Transcription factors (TFs) are protein molecules with special structure and function of regulating gene expression, which plays many crucial roles in plant growth and development [1]. The basic helix-loop-helix (bHLH) transcription factor family is the second largest family in plants. The members of this family are designated by a highly conserved domain called the bHLH which are able to bind and form DNA dimers [2]. The conserved bHLH domain consists of about 60 amino acids and has two functional segments, the basic region and the HLH region. The N-terminal basic region, which contains 13–17 major basic amino acids, serves as the DNA binding domain to identify and specifically bind to DNA motifs in the promoter of the target gene [3,4,5,6]. The HLH region is located at the C-terminus of the bHLH domain, which consists of two parental α-helixes, mainly composed of hydrophobic residues, connected by a relatively dispersed (length and primary sequence) loop region. The HLH domain promotes protein-protein interactions and allows the formation of homo-dimer or hetero-dimer complexes [7]. bHLH transcription factors are involved in many process about plant growth and metabolism, such as stomata development [8], light signal transduction [9, 10], flowering regulation [11], anthocyanin and secondary metabolism [12,13,14]. There have been reported that bHLH genes are mainly involved in abiotic stress in plants, such as the responses to drought, low temperature, salt, ABA and mechanical damage [15, 16]. For example, AtbHLH006, AtbHLH17, AtbHLH32, AtbHLH92, AtbHLH122, AtbHLH128 and AtbHLH130 are directly or indirectly involved in ABA signaling pathway to improve drought resistance in Arabidopsis [17]. The over-expression of bHLH TF MYC-type ICE1, ICE2 and CBF enhanced the tolerance of Arabidopsis to low temperature stress [18]. In wheat, TabHLH1 is able to regulate ABA-mediated stress tolerance pathway to improve plant adaptability to drought and salt stresses [19]. The TabHLH39 gene is involved in regulating gene expression levels in stress responses, thereby increasing salt tolerance in over-expressing plants [20]. In rice, OsbHLH148 and OsbHLH006 (RERJ1) respond to drought stress through the jasmonic acid signaling pathway [21, 22]. The bHLH transcription factor RsICE1 can improve the cold tolerance of transgenic rice [23]. The expression of the PebHLH35 gene in populus increased during drought and ABA induction, and PebHLH35 had an active regulatory effect under drought stress, which mentioned plant tolerance [24]. Similarly, it was shown that VabHLH1 and VvbHLH1 are positive regulators of response to low temperature stress in Chinese wild Vitis amurensis and Vitis vinifera cv. Cabernet Sauvignon, and able to confer enhanced low temperature tolerance to transgenic plants by regulating the expression level of cold regulated (COR) genes [25].
To date, based on the rapid development of genome sequencing, a number of plant bHLH TF genes have been identified and characterized in many species. Although, there are 162, 167, 155, 124 and 188 bHLHs have been identified in Arabidopsis, rice, bean, potato and apple, respectively [26], there has been no report about the bHLH family in pear. Pear is an important cash crop and widely distributed in the world. However, pears were suffered from abiotic stresses such as drought, low temperature, and salt during the growth and development process, which not only restricts the cultivation area, but also affects their growth, development and yield. Therefore, investigating of pear bHLH transcription factors are necessary to elucidate the biological processes underlying pear stress responses.
In this study, we identified 197 pear bHLH (PbrbHLH) genes from the Chinese white pear genomic sequence and carried out phylogenetic analysis to determine the relationships among these genes. Analysis results of protein motifs and intron/exon structures support the classification of the bHLH family. At the same time, we identified duplication events that likely contributed to the expansion of the bHLH family. In addition, RNA-Seq data showed that the expression patterns of PbrbHLHs differed in response to drought and cold stresses. The data from this study will increase our understanding of PbrbHLH functions associated with stress responses. Meanwhile, our systematic analysis provided a foundation for further mechanisms of cold-tolerance and drought-tolerance for bHLH genes in pear, especially for aiming to identify candidate genes that may be involved in the cold- and drought-tolerance of pears.
Results
Identification, classification and function annotation of bHLH genes in Chinese white pear
To identify the PbrbHLH genes, we performed local HMM-search with the HMM file (PF00010) against Chinese white pear genome, with default parameters. 200 putative PbrbHLH protein sequences were identified. SMART and NCBI Batch CD-Search tools were used to confirm the existence of the conserved bHLH domain, and redundant sequences were removed. We finally obtained 197 sequences in pear bHLH family. According to the order of gene ID, these genes were named from PbrbHLH1 to PbrbHLH197 (Table 1 and Table S1). 168 PbrbHLH genes are randomly distributed on all 17 chromosomes ranging from 1 to 25 per chromosome, and the others were localized to 25 unanchored scaffolds (Table 1). Chromosome 15 has the most PbrbHLHs (25 genes), followed by chr 5 (21 genes) and chr 14 (15 genes).
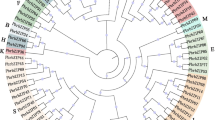
The exact number of subgroup classifications for plant bHLH proteins is unknown, but is thought to be 15–32 [7, 8, 27]. To classify these genes and investigate their evolutionary relationships, phylogenetic tree was built applying the NJ method (Fig. 1, Fig. 2a and Fig. S1). The unrooted tree revealed that PbrbHLH gene family could be separated into 21 clades with the subfamily names A to U, which is the same number as those found in tomato [28] and Phaseolus vulgaris [29]. Unlike other clades, clade P and Q contained a single bHLH protein, respectively, meaning that PbrbHLH32 and PbrbHLH184 are unique. Furthermore, the NJ-tree built with these two PbrbHLHs and 167 AtbHLHs protein sequences indicated that the correlation between PbrbHLH132 and PbrbHLH184 and other bHLH proteins were relatively low (Fig. S1), which is consistent with the un-rooted tree. Except clade P and Q, the gene numbers of each clade varied wildly from 3 (clade L and M) to 22 (clade U). The results of gene structure analysis also showed that the PbrbHLH gene family have a broad range of exon numbers as well the gene structural diversity (Fig. 2c), such as the fact that there is no characteristic distribution pattern of exons and UTRs within most of certain subfamilies. However, the distribution patterns of exons were relatively conserved in clade D, F, G, H, J, K and U, and genes in these clades have a high similarity in exons number, exon pattern and the length of each exon, such as PbrbHLH73, PbrbHLH74, PbrbHLH75, PbrbHLH76 and PbrbHLH180 in clade F and PbrbHLH47 to PbrbHLH77 in clade H.
Un-rooted phylogenetic tree of PbrbHLH proteins.MEGA 7 was used to construct the phylogenetic tree based on the protein sequences. iTOL was used to annotate and review the phylogenic tree. The proteins were clustered into 21 groups. Different background colors indicate the different group of the PbrbHLH proteins
Schematics of the gene structure and conserved motifs in the PbrbHLH family. a Subgroup classification. Phylogenetic tree was generated using 197 PbrbHLH genes with MEGA7. The subgroup names were labeled accordingly. b Conserved motif analysis. Twenty distinct motifs were identified with MEME suite and each motif was represented with different color. c. Gene structural analysis
The characteristics of the PbrbHLH family and their coding genes are shown in Table 1 and Table S1. The protein molecular weights of bHLHs were from 10.38 to 274.01 kD. Protein isoelectric points (pI) ranged from 4.24 to 10.62, and 120 of them were lower than 7 (Table 1). The grand average of hydropathy (GRAVY) for all bHLH proteins in pear was positive, suggesting that all PbrbHLHs were likely soluble proteins which are consistent with their potential functions as TFs.
The annotation information from GO and KEGG databases were able to depict potential function of these genes. To predict the functions of identified PbrbHLH genes, the functional enrichment analysis of PbrbHLHs and a blastp analysis against the protein sequences of reported AtbHLH genes were all performed. As shown in Fig. S2a and Table S1, PbrbHLHs were mainly enriched in the terms of binding, cell part, cellular process, metabolic process and some regulation function, and all of these functions and processes are closely related to TFs. In addition, the KEGG enrichment result showed that these genes were largely enriched in circadian rhythm, MAPK signaling and plant hormone signal transduction pathways (Fig. S2b), all of which are the main mechanisms by which bHLH family TFs regulate the expression of downstream genes. Furthermore, the blastp result also showed that the crucial TF of these pathway were detected, including AtbHLH004 (the orthologous ofPbrbHLH62, PbrbHLH80, PbrbHLH102, PbrbHLH155, PbrbHLH156 and PbrbHLH162) and AtbHLH003 (the orthologous of PbrbHLH38, PbrbHLH39, PbrbHLH112 and PbrbHLH113), thepositive and negative regulator of jasmonate responses, respectively; AtbHLH008 (the orthologous of PbrbHLH24, PbrbHLH28, PbrbHLH115 and PbrbHLH135), a negative regulator of phyB signalling; and AtbHLH098 (the orthologous of PbrbHLH47, PbrbHLH61, PbrbHLH81 and PbrbHLH142), a substrate of kinases MPK3 and MPK6.
Synteny analysis of PbrbHLHs
The gene duplication events, such as WGD/segmental duplication, tandem duplication and transposition events, are the main causes for gene family expansion and affect the evolution of protein-coding gene families [30]. By using the MCScanX package, we detected the duplication events of bHLH gene family, and each of genes was assigned to one of five different duplication types: singleton, WGD/segmental, tandem, proximal or dispersed. Five duplication types were all detected driving the expansion of the PbrbHLH genes (Table 2 and Table S2). The results showed that 58.9% of bHLH genes in Chinese white pear were duplicated and retained from WGD/segmental events, and almost one quarter (23.9%) of PbrbHLHs was belonged to dispersed type.
To explore the evolutionary process behind the PbrbHLH genes, we performed intragenomic synteny analysis to identify conservation chromosome blocks within Chinese white pear. The landscape of ortholog PbrbHLH genes pairs were shown in Fig. 3 and their chromosomal distribution was random. The evolutionary date of WGD/segmental duplication events could be estimated by the Ks value (synonymous substitutions per site) [31]. As the previous reports, based on Ks values, the genome of pear had undergone two genome-wide duplication events: the ancient WGD (Ks ~ 1.5–1.8) that took place ~ 140 MYA [32] and the recent WGD (Ks ~ 0.15–0.3) occurred at 30–45 MYA [33] in pear. Therefore, we used Ks values to estimate the evolutionary date of the gene duplication events among the PbrbHLH gene family. The Ks values implied that most PbrbHLH genes were duplicated from a date around the recent WGD event, and some of others were originated from the ancient WGD (Table 3). The selection intensity and direction could be represented by Ka/Ks ratio, Ka/Ks value of one indicates neutral evolution, positive selection was indicated by a Ka/Ks value greater than one, and purifying selection was indicated by a Ka/Ks value less than one [34]. The Ka/Ks ratios of almost all homologous PbrbHLH genes were less than one (except the gene pair PbrbHLH110-PbrbHLH152), which implying that PbrbHLHs mainly evolved under purifying selection (Table 3).
Conserved motif analysis of PbrbHLH gene family
The types and composition of inner motifs mainly determine the protein function, and the evolutionary relationships among these PbrbHLH proteins were also determined by analyzing their conserved motifs. To further identify motif constructions of the PbrbHLH proteins, the online MEME program was used in this study to detect motif patterns. As showed in Fig. 2b, 20 conserved motifs with low E-values were recognized. The details of motif patterns were shown in Table S3. These composition patterns were nearly consistent with the phylogenetic analysis results, which were similar within the same group, but varying greatly between groups. Among PbrbHLHs, although pattern [#1,2] was detected in all members as the conserved motif pattern for bHLH TF in Chinese white pear, some of the other motifs were present only in certain groups, including motif #8 in group B, I and K; motif #10 in group F and O; motif #11 in group N; motif #9 in group S and motif #19 in T. However, some unique motifs patterns also only could be detected in specific subfamilies. Such as the pattern [#13,12,10,1,2,6,3,14] in clade F, the pattern [#15,7,5,18,1,2,6,3] in clade K and the pattern [#1,2,6,3,20] in clade J. We found that many subfamilies had relatively certain motif composition and there were significant differences among each other. However, there were some groups that have more than one pattern, and no conserved pattern was detected in some other clades, indicating that PbrbHLHs in these groups were not conservative in the evolutionary process, and the division among groups might have occurred in an early period.
Expression profile and patterns of PbrbHLH genes in response to drought and cold stresses
Previous transcriptome data revealed the expression patterns of candidate PbrbHLH genes in response to drought stress and cold stress, respectively (Fig. 4) [35, 36]. Overall, the results indicated that although the background expression of some PbrbHLH genes was rarely detected, that of others was significantly different among these investigated time points. Several differentially expressed genes showed up-regulation trend under both two stress conditions, to varying degrees, such as PbrbHLH119 and PbrbHLH120 in clade E, PbrbHLH7 and PbrbHLH160 in clade G and PbrbHLH128 to PbrbHLH80 from clade K. This suggested that these genes may be involved in some close-related pathways in response to drought and cold stresses. Interestingly, compared to the expression of these genes in cold treatment, the peak expression of them under drought condition was showed at a relatively late time point. In contrast, some other PbrbHLHs showed different (or even opposite) expression patterns, indicating that their responses might vary according to the different stress conditions.
To further verify the functions of these identified PbrbHLHs, eight differentially expressed PbrbHLH genes (PbrbHLH119 from clade E, PbrbHLH7,PbrbHLH8 and PbrbHLH160 from clade G, PbrbHLH80, PbrbHLH128, PbrbHLH161 and PbrbHLH195 from clade K) were selected to examine the expression in response to drought and cold stresses, respectively (Fig. 5). Comparing with the expression at 0 hpt (hours post treatment), except PbrbHLH8 and PbrbHLH80 in drought treatment as well PbrbHLH7 and PbrbHLH119 under cold stress (data not shown), expressions for all other genes were significantly altered in the early stage of drought or cold treatment. Their responses tended to be more rapid under drought conditions, usually changing within the first 12 h. Under cold stress, expression of PbrbHLH8, PbrbHLH128, PbrbHLH160 and PbrbHLH161 initially showed down-regulating trend before being up-regulated as well as the expression of PbrbHLH7 and PbrbHLH195 under drought stress. The opposite trends between cold and drought stresses were noted for PbrbHLH128 and PbrbHLH160. Under drought stress, both were up-regulated at first and then down-regulated, whereas, under cold stress, their expression initially decreased before increasing. These results indicated that PbrbHLH genes were indeed involved in the responses to drought and cold stresses, and the pathways they taken part in under these stresses condition seemed to be different.
Expression analysis of PbrbHLHs undercold and drought stresses. a Relative expression of PbrbHLH8, PbrbHLH80, PbrbHLH128, PbrbHLH160, PbrbHLH161 and PbrbHLH195 with cold treatment. b Relative expression of PbrbHLH7, PbrbHLH119, PbrbHLH128, PbrbHLH160, PbrbHLH161 and PbrbHLH195 with drought treatment. The pear Actin was used as internal reference for the normalization. The statistical analyses were performed using student’s t-test (* p < 0.05)
Silencing of PbrbHLH195 reduced cold tolerance of P. betulaefolia
To understand whether PbrbHLHs is required for cold tolerance in pear, the VIGS system was employed to silence PbrbHLH195, which is significantly up-regulated under cold condition, in P. betulaefolia. The transcript abundance of PbrbHLH195 in the positive plants was substantially reduced by 50–90%, compared with that of WT (Fig. 6j, k). The positive silent plants (p-TRV1, p-TRV2 and p-TRV3) and WT plants were morphologically indistinguishable under normal growing conditions (Fig. 6a, d). However, upon exposure to 0 °C for 8 days, the silent plants displayed more severe damage in comparison with WT (Fig. 6a). The electrolyte leakage (EL) and malondialdehyde (MDA) concentrations in silent plants were significantly higher than those in WT under cold stress (Fig. 6b, c). Meanwhile, when they were subjected to cold treatment, Chl fluorescence in silent plants were prominently repressed, accompanied by significantly lower Fv/Fm ratio and Chl content, in comparison with WT (Fig. 6d-g). In addition, compared with silent plants, WT had lower H2O2 content (Fig. 6h, i). In situ accumulation of H2O2 was histochemically stained with DAB. In the presence of low temperature, the staining became darker, but silent plants staining was deeper and stronger than that of WT (Fig. 6h, i), which was further confirmed by quantitative measurement (Fig. 6i), which means that silencing plants accumulate more reactive oxygen species than WT. These results indicated that silencing of PbrbHLH195 promotes cold susceptibility in P. betulaefolia.
Cold tolerance assay of PbrbHLH195-silenced Pyrus betulaefolia plants. a-c Phenotype of 1-month-old PbrbHLH195-silenced plants before and after cold treatment for 8 days (a). Electrolyte leakage (EL) (b) and malondialdehyde (MDA) concentrations (c) after cold treatment. d-g Chlorophyll fluorescence imaging of silenced plants and WT plants(d), Fv/Fm ratios (e), Chl content (f) and phenotype (g) of WT and pTRV-PbrbHLH195 silencing plants (pTRV-1, pTRV-2 and pTRV-3) at the end of the chilling stress. h-i In situ accumulation of H2O2 of WT and silencing plants, as revealed by histochemical staining with 3, 3-diaminobenzidine (DAB) (h) after cold treatment. Quantitative measurement of H2O2 (i) levels after cold treatment. The expression of PbrbHLH195 was detected by RT-PCR (j) and qRT-PCR (k) at 8d after cold treatment
Discussion
After the release of the Chinese white pear genome sequencing data, there were many TF genes at the whole-genome level have been identified and characterized, including NAC-TF, BAM-TF and WRKY-TF et.al [22, 37, 38].. bHLH transcription factors are involved in many pathways in plant growth and metabolism [12]. However, no such detailed studies have been done with the bHLH family, and only a few examinations have been made of PbrbHLHs in pear. Here, we identified 197 PbrbHLH genes in Chinese white pear. Results of the phylogenetic analysis, gene structure and protein conserved motif analysis enable us to classify these PbrbHLH proteins into 21 groups, which is the same number reported in tomato and apple [28, 39], even though those organisms have fewer SlbHLHs (159) and MdbHLHs (188) than the members of PbrbHLHs in pear. On the basis of phylogenetic analysis, the un-rooted tree showed that PbrbHLHs were well separated into 19 clades with the wildly varied gene numbers from 3 (clade L and M) to 22 (clade U) and two one-gene clade P and Q. The gene and protein structure analysis showed that PbrbHLH family also has a broad diversity in intron/exon organizations as well the protein motif patterns. Although, the distribution pattern of exons and UTRs in clade D, F, G, H, J, K and U were relatively conserved, there was a broad range of exon numbers and structural diversity in many other clades, which is similar to the results of protein motif pattern analysis. By using online MEME software, 20 conserved protein motifs were detected among PbrbHLHs with low E values, and pattern [#1,2] were existed in all bHLHs which was regarded as the characteristic pattern for PbrbHLH TF. Meanwhile, some other motifs were present only in certain groups, including the motif #8 in group B, I and K and motif #10 in group F and O. Furthermore, three unique motif patterns only could be detected in specific subfamilies, respectively, such as the pattern [#13,12,10,1,2,6,3,14] in clade F, the pattern [#15,7,5,18,1,2,6,3] in clade K and the pattern [#1,2,6,3,20] in clade J. These results suggested that the PbrbHLH gene family may play diverse roles in the adaptive evolution to environmental stresses, and the division among groups might have occurred in an early period.
Gene duplication analysis revealed that the main driving force for the expansion of PbrbHLH family was WGD/segmental events, which is same as the case in apple. For instance, by applying MCScanX, 58.9% of bHLH genes in Chinese white pear were categorized into WGD/segmental type. Although pear was undergone the recent WGD events, almost one quarter of bHLH genes were duplicated from dispersed events. This may be due to the high ratio of self-incompatibility and the domestication process of pear. These results showed that WGD/segmental and dispersed gene duplications play critical roles in the expansion of the bHLH gene family in Chinese white pear. Furthermore, Ks values analysis implied that almost all WGD type PbrbHLH genes were duplicated from a date around the recent WGD event, and the Ka/Ks ratios indicated that PbrbHLHs evolved mainly under purifying selection and they seem to be necessary for adaptation to the current environment in their evolutionary history.
The function enrichment analysis showed that PbrbHLH genes were mainly enriched in the functions and processes closely related to TF, and the pathways they classified in were the main mechanisms by which bHLH family TFs regulate the expression of downstream genes, such as circadian rhythm, MAPK signaling and plant hormone signal transduction pathways. For example, OsbHLH148 and OsbHLH006 (REJ1) can improve drought stress by jasmonic acid signaling pathway in rice. Under salt and drought stress, in grapes VvbHLH1 confers a dominant effect on salinity and drought tolerance thought increasing the accumulation of flavonoids and ABA signaling in transgenic Arabidopsis thaliana. In addition, bHLH protein is also involved in plant stress resistance. Arabidopsis AtbHLH112 gene improves drought tolerance by increasing osmotic substances, eliminating ROS content and reducing water diversion. The results indicated that PbrbHLHs might play roles as other bHLHs.
By analyzing previous transcriptome data, we revealed the expression patterns of PbrbHLHs under cold and drought stress conditions. The results showed that, except some genes, the expression of most PbrbHLHs was significantly altered. For example, under both two stresses, PbrbHLH genes including PbrbHLH7, PbrbHLH119, PbrbHLH120, PbrbHLH160 and PbrbHLH128 to PbrbHLH80 in clade K had an up-regulation trend, which suggested that these genes might play similar roles in some close-related pathways in response to drought and cold stresses. Comparing with cold treatment, the peak expression of them under drought condition was showed at a relatively late time point, indicating that the responses of PbrbHLHs varied according to the treatment applied. To verify whether PbrbHLHs were involved in the response to cold or drought stresses, we performed stress treatments and qRT-PCR analysis. The results showed that the expression of all tested genes was significantly altered in the early stage of drought or cold treatments, however, the responses of same gene between two treatmentes could be diverse. For instance, under cold treatment, expression of PbrbHLH7, PbrbHLH8, PbrbHLH161, PbrbHLH128, PbrbHLH160 and PbrbHLH195 showed down-regulating trend at first before being up-regulated, whereas, under drought stress, PbrbHLH128 and PbrbHLH160 were up-regulated at first and then down-regulated. Furthermore, as a high up-regulated gene induced in both cold and drought stress conditions, the interference of PbrbHLH195 in transcription level significantly reduced the cold tolerance of the RNAi pear seedlings. These results indicate that PbrbHLH genes were involved in the responses to drought and cold stresses in pear, and the pathways they involved in seemed to be different under various stress conditions.
Our works in this study highlight the importance of bHLH TF in the cold and drought tolerance of pear. This is the first study to identify the PbrbHLH genes and examine their expression patterns in pear. QRT-PCR analysis showed that PbrbHLH is involved in stress tolerance pathways and functional analysis showed that PbrbHLH195 plays an important role in pear abiotic stress tolerance. However, further investigation will be required to understand the roles of PbrbHLHs in the stress response pathways, and the characterization of key (even the marker) bHLH TFs under each stress condition was also crucial to the revealing of the functional mechanisms of bHLH in pear.
Conclusions
In this study, we identified 197 PbrbHLH genes from Chinese white pear and carried out phylogenetic analysis to determine the relationships among these genes. Based on the results of protein motifs and intron/exon characteristics and phylogenetic analysis, PbrbHLH family was classified into 21 groups. According to the analysis of collinearity, WGD and dispersed duplication might have a role in the evolution of the PbrbHLH family. In addition, RNA-seq data, qRT-PCR and VIGS results revealed that PbrbHLH genes might have important roles in response to abiotic stresses, and the expression patterns of them differed in response to drought and cold stresses. The underlined collected data from this study provided a foundation for advanced studies to evaluate the mechanisms of cold-tolerance and drought-tolerance for bHLH genes in pear.
Methods
Plant materials and bacterial strains
The pear seedlings were grown in the greenhouse under a 16 h/8 h light/dark photoperiod, 75% relative humidity and 25 °C. A. tumefaciens GV3101 was grown in LB media supplemented with kanamycin and Rif at 28 °C in an orbital shaker at 200 rpm and harvested during the log phase of growth for infiltration.
Identification and functional annotation of bHLH gene family in Chinese white pear
To identify the bHLH genes in Chinese white pear, we performed multiple database-based searches. We downloaded all needed sequences and annotation file of Chinese white pear from Pear Centre of Nanjing Agricultural University (http://peargenome.njau.edu.cn/) and the seed file of bHLH conserved domain (PF00010) was downloaded from Pfam (http://pfam.sanger.ac.uk/). HMMER (Hidden Markov Model, HMM) software was used to detect conserved Pfam domain with default parameters E-value < 0.05 [40]. Then we checked the predicted bHLH transcription factors by using the NCBI Batch CD-Search tools (Batch CD-Search: https://www.ncbi.nlm.nih.gov/Structure/bwrpsb/bwrpsb.cgi) based on CDD v3.18 and SMART v6.0 databases to verify the existence of bHLH domain (Table S1). The proteins with E-values greater than 1e− 6 or without a bHLH domain were removed. The relevant gene ID of PbrbHLH genes were shown in Table 1. The annotation information for Chinese white pear was extracted from the GFF file, and the result was visualized by a R script. The BLASTP was performed against 167 reported AtbHLH protein sequences [5], and the protein sequences were downloaded from TAIR (The Arabidopsis Information Resource, https://www.arabidopsis.org/).
Structure and conserved motif analysis of the PbrbHLH genes
The Gene Structure Display Server (GSDS 2.0) (http://gsds.cbi.pku.edu.cn/) was used to analyze the structures of the bHLH genes by aligning the cDNA sequences with their corresponding genomic DNA sequences [41]. Conserved motif analysis of bHLH proteins was performed by online Multiple Expectation Maximization for Motif Elicitation (MEME) (http://meme.nbcr.net/meme/cgibin/meme.cgi) with default parameters, and maximum number parameter of motifs were set as 20 [42]..
Phylogenetic analysis
The phylogenetic tree was built with Neighbor-Joining (NJ) method and a bootstrap of 1000 in MEGA7.0 (http://www.megasoftware.net/) [43]. The p-distance was used and the optional parameters for pairwise deletion were considered.
Chromosomal localization and synteny analysis
The chromosomal localization information was extracted from the GFF file. The same procedure used in the PGDD (http://chibba.agtec.uga.edu/duplication/) was performed to analyze the synteny among the PbrbHLHs. Primarily, the local all-vs-all BLASTP searches among identified PbrbHLH genes were conducted (E < 1e− 10). Afterward, MCScanX was employed for the determination of syntenic gene pairs with the BLASTP result and gene location information used as input files [44]. The downstream analysis tool (duplicate_gene_classifier) in the MCScanX package was employed for the identification of tandem, proximal dispersed, and segmental/whole-genome duplications (WGD) of PbrbHLH family genes. The results were visualized using circos-0.69 software [45]. The Ka and Ks values were analyzed via KaKs_Calculator 2.0 [46]. For the estimation of the date of segmental duplication events, the succeeding pairs of homologous genes within 100 Kb on all sides of the PbrbHLH genes, considered for the mean Ks calculation.
Expression analysis of PbrbHLH genes under drought and cold stress conditions
Published transcriptomic data (FPKM values) characterizing the total RNA of drought treatment samples (D0, D1, D3 and D6 indicating the samples harvested at 0 hpt (hour post treatment), 1 hpt, 3 hpt and 6 hpt under drought stress) were downloaded from Li et al. (2016) [35]; cold treatment samples (C0, C5, C12 and C24 indicating the samples harvested at 0 hpt, 5 hpt, 12 hpt and 24 hpt under cold stress) were downloaded from Yang and Huang (2018) [36]. We determined the expression patterns of PbrbHLH family genes under drought and cold stress conditions. The differentially expressed genes were identified with the threshold |log2FC| > 1. TBtools v1.068 was used to visualize the results [47].
For the qRT-PCR analysis, 9-week-old pear seedlings were treated with drought and cold, respectively. The leaves were cryopreserved with liquid nitrogen at 0 hpt, 1 hpt, 3 hpt, 6 hpt, 12 hpt and 24 hpt after drought stress treatment as well the leaves with cold treatment at 0 hpt, 1 hpt, 3 hpt,9 hpt, 12 hpt and 24 hpt. Total RNA extraction and the synthesis of cDNA were according to the instructions of RNA kit (Tiangen, Beijing, China) and PrimeScript RT reagent Kit (Trans Gen). Specialized primers of the constitutive TUB and eight tested PbrbHLH genes were designed via NCBI online tool Primer-BLAST (https://www.ncbi.nlm.nih.gov/tools/primer-blast/index.cgi?LINK_LOC=BlastHome) with the Specificity Parameters Organism option set as Pyrus bretschneideri (taxid:225117) (Table S4). The qRT-PCR assays were conducted with three technical copies. QRT-PCR reactions (20 μl per hole) were performed as previously reported [48]. The expression was evaluated for each sample via the 2−ΔΔCt method, and Duncan’s multiple range test was conducted. A p-value of less than 0.05 was the considerable variation and the differentially expressed genes were identified with |log2FC| > 1.
Generation of silenced plants
Virus-induced gene silencing (VIGS)-mediated suppression of PbrbHLH195 was performed according to previous methods [49]. A 182 bp fragment of PbrbHLH195 open reading frame (ORF) was inserted into EcoR I and BamH I sites of tobacco rattle virus-based vector 2 (TRV2) to generate the pTRV2-PbrbHLH195 construct. The vectors pTRV1, pTRV2 and pTRV2-PbrbHLH195 were transformed into A. tumefaciens strain GV3101 by heat shock. The bacterial cells (OD600 = 1.0) containing pTRV1 were mixed with pTRV2-PbrbHLH195 or pTRV2 in a 1: 1 volume ratio in 2-(Nmorpholino) ethanesulfonic acid (MES) buffer (10 mM MgCl2, 200 mM acetosyringone, and 10 mM MES, pH 5.6) and kept slowly shaking in dark for 4 h at room temperature. The bacterial mixtures were injected into the leaves of seedlings and rinsed with water, grown in soil pots and transferred to a controlled growth chamber. Two weeks later, un-injected leaves were collected from each plant and subjected to genomic PCR and qRT-PCR analyses, and the VIGS plants exhibiting similar magnitude of PbrbHLH195 suppression were used for further analyses.
Physiological analysis
EL was measured according to [50]. MDA and H2O2 were detected according to the instructions using the corresponding detection kit (Nanjing Jiancheng Bioengineering Institute, Nanjing, China). Chl fluorescence was measured by Imaging PAM CHL fluorometer, Fv/FM ratio was calculated by imaging Winge software (Walz, Germany). Chl was extracted and analyzed according to [51].
Availability of data and materials
All needed genome sequences and genome annotation files of Chinese white pear were obtained from the Nanjing Agricultural University pear genome project website (http://peargenome.njau.edu.cn). All data generated or analysed during this study are included in this published article and its supplementary information files.
Abbreviations
- bHLH:
-
Basic helix-loop-helix
- TF:
-
Transcription factor
- COR gene:
-
Cold regulated gene
- pI:
-
Protein isoelectric points
- GRAVY:
-
Grand average of hydropathy
- Ks:
-
Synonymous substitutions per site
- hpt:
-
Hours post treatment
- EL:
-
Electrolyte leakage
- MDA:
-
Malondialdehyde
- HMM:
-
Hidden Markov Model
- GSDS:
-
Gene Structure Display Server
- MEME:
-
Multiple Expectation Maximization for Motif Elicitation
- NJ:
-
Neighbor-Joining
- RPKM:
-
Fragments per kilobase of exon model per million mapped reads
- WGD:
-
Whole-genome duplications
- VIGS:
-
Virus-induced gene silencing
- ORF:
-
Open reading frame
- MES:
-
2-(Nmorpholino) ethanesulfonic acid
References
Niu X, Guan Y, Chen S, Li H. Genome-wide analysis of basic helix-loop-helix (bHLH) transcription factors in Brachypodium distachyon. BMC Genomics. 2017;18(1):1–20. https://doi.org/10.1186/s12864-017-4044-4.
Ledent V, Vervoort M. The basic helix-loop-helix protein family: comparative genomics and phylogenetic analysis. Genome Res. 2001;11(5):754–70. https://doi.org/10.1101/gr.177001.
Massari ME, Murre C. Helix-loop-helix proteins: regulators of transcription in Eucaryotic organisms. Mol Cell Biol. 2000;20(2):429–40. https://doi.org/10.1128/mcb.20.2.429-440.2000.
Toledo-Ortiz G, Huq E, Quail PH. The Arabidopsis basic/helix-loop-helix transcription factor family. Plant Cell. 2003;15(8):1749–70. https://doi.org/10.1105/tpc.013839.
Li X, Duan X, Jiang H, et al. Genome-wide analysis of basic/helix-loop-helix transcription factor family in rice and Arabidopsis. Plant Physiol. 2006;141(4):1167–84. https://doi.org/10.1104/pp.106.080580.
Huang XS, Wang W, Zhang Q, Liu JH. A basic helix-loop-helix transcription factor, PtrbHLH, of Poncirus trifoliata confers cold tolerance and modulates peroxidase-mediated scavenging of hydrogen peroxide. Plant Physiol. 2013;162(2):1178–94. https://doi.org/10.1104/pp.112.210740.
Buck MJ, Atchley WR. Phylogenetic analysis of plant basic helix-loop-helix proteins. J Mol Evol. 2003;56(6):742–50. https://doi.org/10.1007/s00239-002-2449-3.
Pires N, Dolan L. Origin and diversification of basic-helix-loop-helix proteins in plants. Mol Biol Evol. 2010;27(4):862–74. https://doi.org/10.1093/molbev/msp288.
Roig-Villanova I, Bou-Torrent J, Galstyan A, et al. Interaction of shade avoidance and auxin responses: a role for two novel atypical bHLH proteins. EMBO J. 2007;26(22):4756–67. https://doi.org/10.1038/sj.emboj.7601890.
Leivar P, Monte E, Al-Sady B, et al. The Arabidopsis phytochrome-interacting factor PIF7, together with PIF3 and PIF4, regulates responses to prolonged red light by modulating phyB levels. Plant Cell. 2008;20(2):337–52. https://doi.org/10.1105/tpc.107.052142.
Ito S, Song YH, Josephson-Day AR, et al. FLOWERING BHLH transcriptional activators control expression of the photoperiodic flowering regulator CONSTANS in Arabidopsis. Proc Natl Acad Sci U S A. 2012;109(9):3582–7. https://doi.org/10.1073/pnas.1118876109.
Nesi N, Debeaujon I, Jond C, Pelletier G, Caboche M, Lepiniec L. The TT8 gene encodes a basic helix-loop-helix domain protein required for expression of DFR and BAN genes in Arabidopsis siliques. Plant Cell. 2000;12(10):1863–78. https://doi.org/10.1105/tpc.12.10.1863.
Ohno S, Hosokawa M, Hoshino A, et al. A bHLH transcription factor, DvIVS, is involved in regulation of anthocyanin synthesis in dahlia (Dahlia variabilis). J Exp Bot. 2011;62(14):5105–16. https://doi.org/10.1093/jxb/err216.
Xie XB, Li S, Zhang RF, et al. The bHLH transcription factor MdbHLH3 promotes anthocyanin accumulation and fruit colouration in response to low temperature in apples. Plant Cell Environ. 2012;35(11):1884–97. https://doi.org/10.1111/j.1365-3040.2012.02523.x.
Chinnusamy V, Ohta M, Kanrar S, Lee B-h, Hong X, Agarwal M, Zhu J-K. ICE1: a regulator of cold-induced transcriptome and freezing tolerance in Arabidopsis. Genes Dev. 2003;17:1043–54. https://doi.org/10.1101/gad.1077503.CRT.
Kiribuchi K, Sugimori M, Takeda M, et al. RERJ1, a jasmonic acid-responsive gene from rice, encodes a basic helix-loop-helix protein. Biochem Biophys Res Commun. 2004;325(3):857–63. https://doi.org/10.1016/j.bbrc.2004.10.126.
Le Hir R, Castelain M, Chakraborti D, Moritz T, Dinant S, Bellini C. AtbHLH68 transcription factor contributes to the regulation of ABA homeostasis and drought stress tolerance in Arabidopsis thaliana. Physiol Plant. 2017;160(3):312–27. https://doi.org/10.1111/ppl.12549.
Fursova OV, Pogorelko GV, Tarasov VA. Identification of ICE2, a gene involved in cold acclimation which determines freezing tolerance in Arabidopsis thaliana. Gene. 2009;429(1–2):98–103. https://doi.org/10.1016/j.gene.2008.10.016.
Yang T, Yao S, Hao L, Zhao Y, Lu W, Xiao K. Wheat bHLH-type transcription factor gene TabHLH1 is crucial in mediating osmotic stresses tolerance through modulating largely the ABA-associated pathway. Plant Cell Rep. 2016;35(11):2309–23. https://doi.org/10.1007/s00299-016-2036-5.
Zhai Y, Zhang L, Xia C, et al. The wheat transcription factor, TabHLH39, improves tolerance to multiple abiotic stressors in transgenic plants. Biochem Biophys Res Commun. 2016;473(4):1321–7. https://doi.org/10.1016/j.bbrc.2016.04.071.
Seo JS, Joo J, Kim MJ, et al. OsbHLH148, a basic helix-loop-helix protein, interacts with OsJAZ proteins in a jasmonate signaling pathway leading to drought tolerance in rice. Plant J. 2011;65(6):907–21. https://doi.org/10.1111/j.1365-313X.2010.04477.x.
Zhao L, Gong X, Gao J, et al. Transcriptomic and evolutionary analyses of white pear (Pyrus bretschneideri) β-amylase genes reveals their importance for cold and drought stress responses. Gene. 2019;689:102–13. https://doi.org/10.1016/j.gene.2018.11.092.
Man L, Xiang D, Wang L, Zhang W, Wang X, Qi G. Stress-responsive gene RsICE1 from Raphanus sativus increases cold tolerance in rice. Protoplasma. 2017;254(2):945–56. https://doi.org/10.1007/s00709-016-1004-9.
Dong Y, Wang C, Han X, et al. A novel bHLH transcription factor PebHLH35 from Populus euphratica confers drought tolerance through regulating stomatal development, photosynthesis and growth in Arabidopsis. Biochem Biophys Res Commun. 2014;450(1):453–8. https://doi.org/10.1016/j.bbrc.2014.05.139.
Wang Y, Jiang CJ, Li YY, Wei CL, Deng WW. CsICE1 and CsCBF1: two transcription factors involved in cold responses in Camellia sinensis. Plant Cell Rep. 2012;31(1):27–34. https://doi.org/10.1007/s00299-011-1136-5.
Wang L, Xiang L, Hong J, Xie Z, Li B. Genome-wide analysis of bHLH transcription factor family reveals their involvement in biotic and abiotic stress responses in wheat (Triticum aestivum L.). 3. Biotech. 2019;9(6):1–12. https://doi.org/10.1007/s13205-019-1742-4.
Carretero-Paulet L, Galstyan A, Roig-Villanova I, Martínez-García JF, Bilbao-Castro JR, Robertson DL. Genome-wide classification and evolutionary analysis of the bHLH family of transcription factors in Arabidopsis, poplar, rice, moss, and algae. Plant Physiol. 2010;153(3):1398–412. https://doi.org/10.1104/pp.110.153593.
Sun H, Fan HJ, Ling HQ. Genome-wide identification and characterization of the gene family in tomato. BMC Genom. 2015;16(1). https://doi.org/10.1186/s12864-014-1209-2.
Kavas M, Baloğlu MC, Atabay ES, Ziplar UT, Daşgan HY, Ünver T. Genome-wide characterization and expression analysis of common bean bHLH transcription factors in response to excess salt concentration. Mol Gen Genomics. 2016;291(1):129–43. https://doi.org/10.1007/s00438-015-1095-6.
Maher C, Stein L, Ware D. Evolution of Arabidopsis microRNA families through duplication events. Genome Res. 2006;16(4):510–9. https://doi.org/10.1101/gr.4680506.
Qiao X, Li M, Li L, Yin H, Wu J, Zhang S. Genome-wide identification and comparative analysis of the heat shock transcription factor family in Chinese white pear (Pyrus bretschneideri) and five other Rosaceae species. BMC Plant Biol. 2015;15(1):1–16. https://doi.org/10.1186/s12870-014-0401-5.
Fawcett JA, Maere S, Van De Peer Y. Plants with double genomes might have had a better chance to survive the cretaceous-tertiary extinction event. Proc Natl Acad Sci U S A. 2009;106(14):5737–42. https://doi.org/10.1073/pnas.0900906106.
Wu J, Wang Z, Shi Z, et al. The genome of the pear (Pyrus bretschneideri Rehd.). Genome Res. 2013;23(2):396–408. https://doi.org/10.1101/gr.144311.112.
Yang Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol Biol Evol. 2007;24(8):1586–91. https://doi.org/10.1093/molbev/msm088.
Li KQ, Xu XY, Huang XS. Identification of differentially expressed genes related to dehydration resistance in a highly drought-tolerant pear, Pyrus betulaefolia, as through RNA-Seq. PLoS One. 2016;11(2):1–21. https://doi.org/10.1371/journal.pone.0149352.
Yang T, Huang XS. Deep sequencing-based characterization of transcriptome of Pyrus ussuriensis in response to cold stress. Gene. 2018;661:109–18. https://doi.org/10.1016/j.gene.2018.03.067.
Gong X, Zhao L, Song X, et al. Genome-wide analyses and expression patterns under abiotic stress of NAC transcription factors in white pear (Pyrus bretschneideri). BMC Plant Biol. 2019;19(1):1–18. https://doi.org/10.1186/s12870-019-1760-8.
Liu Y, Yang T, Lin Z, et al. A WRKY transcription factor PbrWRKY53 from Pyrus betulaefolia is involved in drought tolerance and AsA accumulation. Plant Biotechnol J. 2019;17(9):1770–87. https://doi.org/10.1111/pbi.13099.
Mao K, Dong Q, Li C, Liu C, Ma F. Genome wide identification and characterization of apple bHLH transcription factors and expression analysis in response to drought and salt stress. Front Plant Sci. 2017;8(April). https://doi.org/10.3389/fpls.2017.00480.
Eddy SR. Accelerated profile HMM searches. PLoS Comput Biol. 2011;7(10). https://doi.org/10.1371/journal.pcbi.1002195.
Hu B, Jin J, Guo AY, Zhang H, Luo J, Gao G. GSDS 2.0: an upgraded gene feature visualization server. Bioinformatics. 2015;31:1296–7. https://doi.org/10.1093/bioinformatics/btu817.
Bailey TL, Williams N, Misleh C, Li WW. MEME: Discovering and analyzing DNA and protein sequence motifs. Nucleic Acids Res. 2006;34(WEB. SERV. ISS):369–73. https://doi.org/10.1093/nar/gkl198.
Kumar S, Stecher G, Tamura K. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets brief communication. Mol Biol Evol. 2016;33(7):1870–4. https://doi.org/10.1093/molbev/msw054.
Tang H, Wang X, Bowers JE, Ming R, Alam M, Paterson AH. Unraveling ancient hexaploidy through multiply-aligned angiosperm gene maps. Genome Res. 2008;706:1944–54. https://doi.org/10.1101/gr.080978.108.1944.
Connors J, Krzywinski M, Schein J, et al. Circos : an information aesthetic for comparative genomics. Genome Res. 2009;19(604):1639–45. https://doi.org/10.1101/gr.092759.109.19.
Wang D, Zhang Y, Zhang Z, Zhu J, Yu J. KaKs_Calculator 2.0: a toolkit incorporating gamma-series methods and sliding window strategies. Genom Proteom Bioinf. 2010;8(1):77–80. https://doi.org/10.1016/S1672-0229(10)60008-3.
Chen C, Chen H, Zhang Y, et al. TBtools - an integrative toolkit developed for interactive analyses of big biological data. Mol Plant. 2020:1–9. https://doi.org/10.1016/j.molp.2020.06.009.
Chen Q, Li Q, Qiao X, Yin H, Zhang S. Genome-wide identification of lysin motif containing protein family genes in eight rosaceae species , and expression analysis in response to pathogenic fungus Botryosphaeria dothidea in Chinese white pear. BMC Genomics. 2020;21(612):1–20.
Jiang X, Zhang C, Lü P, et al. RhNAC3, a stress-associated NAC transcription factor, has a role in dehydration tolerance through regulating osmotic stress-related genes in rose petals. Plant Biotechnol J. 2014;12(1):38–48. https://doi.org/10.1111/pbi.12114.
Dahro B, Wang F, Peng T, Liu JH. PtrA/NINV, an alkaline/neutral invertase gene of Poncirus trifoliata, confers enhanced tolerance to multiple abiotic stresses by modulating ROS levels and maintaining photosynthetic efficiency. BMC Plant Biol. 2016;16(1):1–18. https://doi.org/10.1186/s12870-016-0761-0.
Liu JH, Moriguchi T. Changes in free polyamine titers and expression of polyamine biosynthetic genes during growth of peach in vitro callus. Plant Cell Rep. 2007;26(2):125–31. https://doi.org/10.1007/s00299-006-0223-5.
Acknowledgements
This work has been supported by the National Key Research and Development Program of China (2019YFD1000102), the National Science Foundation of China (31872070; 32072538), the Jiangsu Agriculture Science and Technology Innovation Fund (CX(18)3065), the Excellent Youth Natural Science Foundation of Jiangsu Province (SBK2017030026), the Fundamental Research Funds for the Central Universities of Nanjing Agricultural University (KYZ201607), the SRT project of the Nanjing Agriculture University (202011YX05), and the Undergraduate Training Program for Innovation and Entrepreneurship (S20190040).
Author information
Authors and Affiliations
Contributions
HZD, YQD and WJH designed and carried out the experiments and QMC carried out all bioinformatics analysis and wrote the manuscript. XSH and SLZ directed and revised the manuscript. All authors read, reviewed and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1 Table S1.
Detailed characteristics of PbrbHLHs.Table S2. Duplication type of PbrbHLH genes in Chinese white pear. Table S3. Sequence informations of 20 detected motifs in MEME analysis. Table S4. The primers of PbrbHLHs for qRT-PCR and vector construction.
Additional file 2 Fig. S1.
Phylogenetic tree of 167 AtbHLHs and the two unique PbrbHLH proteins. MEGA 7 was used to construct the phylogenetic tree based on the protein sequences. iTOL was used to annotate and review the phylogenic tree. Fig. S2. Functional annotation enrichment analysis. (a) GO (Gene ontology) term enrichment analysis of PbrbHLH proteins. (b) KEGG enrichment analysis of PbrbHLH proteins.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Dong, H., Chen, Q., Dai, Y. et al. Genome-wide identification of PbrbHLH family genes, and expression analysis in response to drought and cold stresses in pear (Pyrus bretschneideri). BMC Plant Biol 21, 86 (2021). https://doi.org/10.1186/s12870-021-02862-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12870-021-02862-5