Abstract
Background
Endophytic fungi of medicinal plants, as special microorganisms, are important sources of antibacterial compounds. However, the diversity and antibacterial activity of endophytic fungi from Pinellia Tenore have not been systematically studied.
Results
A total of 77 fungi were isolated from roots, stems, leaves, and tubers of Pinellia ternata and P. pedatisecta. All fungi were belonged to five classes and twenty-five different genera. Biological activities tests indicated that 21 extracts of endophytic fungi exhibited antibacterial activities against at least one of the tested bacteria, and 22 fermentation broth of endophytic fungi showed strong phytotoxic activity against Echinochloa crusgalli with the inhibition rate of 100%. Furthermore, four compounds, including alternariol monomethyl ether (1), alternariol (2), dehydroaltenusin (3) and altertoxin II (4), and three compounds, including terreic acid (5), terremutin (6), citrinin (7), were isolated from Alternaria angustiovoidea PT09 of P. ternata and Aspergillus floccosus PP39 of P. pedatisecta, respectively. Compound 5 exhibited strong antibacterial activities against Escherichia coli, Micrococcus tetragenus, Staphylococcus aureus, and Pseudomonas syringae pv. actinidiae with the inhibition zone diameter (IZD) of 36.0, 31.0, 33.7, 40.2 mm and minimum inhibitory concentration (MIC) values of 1.56, 3.13, 1.56, 1.56 μg/mL respectively, which were better than or equal to those of positive gentamicin sulfate. The metabolite 7 also exhibited strong antibacterial activity against P. syringae pv. actinidiae with the IZD of 26.0 mm and MIC value of 6.25 μg/mL. In addition, the compound 7 had potent phytotoxic activity against E. crusgalli with the inhibition rate of 73.4% at the concentration of 100 μg/mL.
Conclusions
Hence, this study showed that endophytic fungi of P. ternata and P. pedatisecta held promise for the development of new antibiotic and herbicide resources.
Similar content being viewed by others
Background
There are five medicinal species belonging to Pinellia Tenore in China, including Pinellia cordata, P. integrifolia, P. pedatisecta, P. peltata and P. ternata [1]. Their tubers possess important medicinal values, such as antitussive and expectorant [2], antiemetic [3], antitumor [4], anti-fertility and terminate early pregnancy effects [5]. Pinellia Tenore also contains diverse phytochemicals, for instance, protein, lectins, alkaloids, sterols, flavonoids, and tannins [2,3,4,5]. Among them, alkaloids and lectins have been identified as the main active ingredients of Pinellia Tenore [6, 7]. P. pedatisecta and P. ternata are the main medicinal plants of the Pinellia Tenore and have similar bioactivities [6].
Endophytic fungi are widespread in the tissues of almost all plants [8], such as roots, stems and leaves [9,10,11]. Furthermore, endophytic fungi protect plants from pathogenic microorganisms and weeds by producing bioactive secondary metabolites [12, 13]. Secondary metabolites produced by endophytic fungi also have antibacterial, antifungal, insecticidal, herbicidal, cytotoxic, antioxidant and anticancer effects [14, 15]. For example, solanioic acid produced by Rhizoctonia solani isolated from Cyperus rotundus showed significant activity against Gram-positive bacteria (Bacillus subtilis, Staphylococcus aureus, and MRSA) [16]. Those bioactive secondary metabolites have significant potential applications in the pharmaceutical industry [14, 17].
Secondary metabolites of endophytic fungi of P. ternata have been shown to have outstanding antibacterial activity [18,19,20]. For instance, aspergillone A from endophytic Aspergillus cristatus of P. ternata exhibited remarkable antibacterial effects [20]. However, as far as we know, the diversity and antibacterial activity of its endophytic fungi have not been systematically studied. Besides, the lack of research focused on the endophytic fungi of P. pedatisecta. The main objective of this study is to identify the diversity of endophytic fungi from P. ternata and P. pedatisecta and to obtain endophytic fungal resources with antibacterial and herbicidal activity. This study further provides a basis for obtaining biotrophic fungi or mining bioactive natural products.
Results
Isolation and identification of endophytic fungi
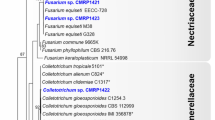
A total of 77 strains (Table 1) of endophytic fungi were isolated from P. ternata and P. pedatisecta for the first time. There were 53 and 24 strains of fungi that were isolated from all tissues of P. ternata and P. pedatisecta, respectively. The ITS1-ITS4 region of 77 strains was sequenced and compared with available GenBank reference sequences. The obtained 5.8S rDNA sequences were uploaded to NCBI under the accession numbers ON677855-ON677931. The results of sequence analysis showed that 77 fungi were attached to the phyla Ascomycota and Basidiomycota (Fig. 1). The 68 strains were grouped into four classes [Dothideomycetes (14.3%), Eurotiomycetes (32.5%), Saccharomycetes (2.6%) and Sordariomycetes (39.0%)] within the phylum Ascomycota. Nine other strains (11.7%) were distributed in the Agaricomycetes within the phylum Basidiomycota. The fungi of Sordariomycetes were the dominant species of cultivable fungi in phylogenetic diversity from P. ternata and P. pedatisecta. The largest number (Sordariomycetes, 30) of isolates was distributed in 5 orders, including the Glomerellales, Hypocreales, Pleurotheciales, Sordariales and Xylariales.
There were 10, 17, 16 and 34 strains, which belonged to 7, 9, 7 and 13 genera, from roots, stems, leaves, and tubers, respectively. Among them, Alternaria angustiovoidea, Macrophomina phaseolina and Schizophyllum commune were isolated from their roots only. Similarly, Clonostachys rosea, Lecanicillium dimorphum, Nigrospora pyreformis, Pseudoechria longicollis, Trichoderma atroviride and Zygosporium masonii were isolated from tubers only. Therefore, the results of phylogenetic diversity showed that the endophytic fungi were different in different plant tissues.
Diversity analyses of endophytic fungi from P. ternata and P. pedatisecta
The diversity analyses of endophytic fungi genera isolated from P. ternata and P. pedatisecta were summarized and shown in Table 2. The Species richness (S) and Margalef index (D') values showed positive correlation with endophytic fungal genera [21]. The higher the Shannon–Wiener index (H') and the Simpson diversity index (Ds), the more diverse the microbial community [22]. Therefore, the endophytic fungi of tubers from P. ternata showed high species richness and diversity, with the values of S (12), D' (3.3011), H' (2.2299), Ds (0.8673), and PIE (0.8995). However, the endophytic fungi of the leaf from P. pedatisecta had high species richness and diversity with the values of S (6), D' (2.4045), H' (1.7329), Ds (0.8125), and PIE (0.9286). The Pielou indexes (J) could reflect the level of uniformity in the distribution of the number of individuals of the species in the community [23]. In this study, the endophytic fungi of stem from P. ternata and tuber from P. pedatisecta showed high Pielou indexes (J) with values of 0.9732 and 0.9697, respectively, which indicated a convergence in the number of individuals of each species.
Antibacterial activities of the crude extracts of fungi
The filter paper disk method was used to evaluate the antibacterial activity of 77 fungal extracts from P. ternata and P. pedatisecta. The results showed that 21 extracts (27.3%) showed antibacterial activities against at least one pathogenic bacterium (Supplementary Table S1). Five of them (PT02, PP35, PP37, PP39, and PT58) had inhibitory activities against all four pathogens (Escherichia coli, Micrococcus tetragenus, Staphylococcus aureus, and Pseudomonas syringae pv. actinidiae). Among them, PP39 exhibited strong antibacterial activity against S. aureus with an IZD of 20.0 mm, which was equivalent to that of positive gentamicin sulfate with an IZD of 21.7 mm. PP39 also showed potent inhibition activities against M. tetragenus, E. coli and P. syringae pv. actinidiae with the IZD of 14.2, 15.2, and 14.0 mm, which were weaker than those of positive gentamicin sulfate with the IZD of 25.7, 26.7, and 24.3 mm, respectively. Besides, the strain PT83 showed strong inhibition activity against P. syringae pv. actinidiae with an IZD of 20.0 mm, which was comparable to that of positive gentamicin sulfate. In addition, the crude extracts of strains PT56 and PT82 also exhibited potent antibacterial activities against P. syringae pv. actinidiae with the IZD of 15.2 mm and 15.7 mm, respectively.
Phytotoxic assay
As the results shown in Table 3, 52 fermentation broth of endophytic fungi exhibited significant phytotoxic activity against the radicle of Echinochloa crusgalli with the inhibition rate of more than 50%. Among them, 22 strains showed strong phytotoxic activity against E. crusgalli with the inhibition rate of 100%. Interestingly, these fungal strains were mainly assigned to three genera (Aspergillus, Fusarium, and Talaromyces) and most of them (20 strains) were isolated from P. ternata. Besides, 16 strains showed outstanding phytotoxic activity against E. crusgalli with the inhibition rate of 80%-99%. Moreover, nine strains showed potent phytotoxic activity against E. crusgalli with the inhibition rate of 60%-79%. In addition, 25 strains showed relatively weak phytotoxic activity against E. crusgalli with the inhibition rate of 10%-60%.
Identification of the secondary metabolites isolated from PT09 and PP39
The fermentation broth PP39 from P. pedatisecta had the best antibacterial and phytotoxic activities among all tested endophytic fungi. PT09 from P. ternata also had strong phytotoxic activity against E. crusgalli with the inhibition rate of 91.1% and moderate antibacterial activity against S. aureus and P. syringae pv. actinidiae. Therefore, both PT09 and PP39 were selected as further research objects of active secondary metabolites.
Four monomer compounds (Fig. 2) were isolated from the liquid fermentation product of strain PT09. They were further identified as alternariol monomethyl ether (1), alternariol (2), dehydroaltenusin (3) and altertoxin II (4) by spectroscopic analyses, including HR–ESI–MS, NMR, and compared with data described in previous literature.
Alternariol monomethyl ether (1) (Figures S1-S2): colorless crystal; HR-ESI–MS: m/z: 271.0599 [M—H]−, calculated for C15H11O5 271.0621; 1H NMR (600 MHz, DMSO-d6) δ: 11.81 (s, OH-7), 10.35 (br s, OH-3), 7.20 (s, H-10), 6.72 (s, H-4), 6.63 (s, H-2), 6.60 (s, H-8), 3.90 (s, 3H, OCH3-9), 2.72 (s, 3H, H-11). Due to the chemical shifts and relative molecular masses in agreement with reported in the literature [24], the structure of the compound was determined to be alternariol monomethyl ether.
Alternariol (2) (Figures S3-S4) [25]: white crystal; HR-ESI–MS: m/z: 257.0453 [M—H]−, calculated for C14H9O5 257.0464; 1H NMR (600 MHz, Acetone-d6) δ: 11.93 (s, OH-3), 9.69 (br s, OH-5), 9.21 (br s, OH-4’), 7.34 (d, J = 2.4 Hz, H-6), 6.78 (d, J = 2.7 Hz, H-5’), 6.69 (d, J = 2.7 Hz, H-3’), 6.44 (d, J = 2.4 Hz, H-4), 2.76 (s, 3H, CH3-6’).
Dehydroaltenusin (3) (Figures S5-S7) [26]: yellow green solid; HR-ESI–MS: m/z: 287.0558 [M—H]−, calculated for C15H11O6 287.0570; 1H NMR (600 MHz, CDCl3) δ: 11.29 (s, OH-7), 6.73 (d, J = 2.3 Hz, H-10), 6.69 (s, H-1), 6.63 (d, J = 2.3 Hz, H-8), 6.41 (s, H-3), 6.28 (s, H-4), 3.91 (s, 3H, OCH3-9), 1.73 (s, 3H, CH3-4a); 13C NMR (150 MHz, CDCl3) δ: 180.9 (C-2), 167.5 (C-6), 166.5 (C-9), 164.9 (C-7), 153.3 (C-10b), 146.3 (C-3), 135.2 (C-10a), 121.0 (C-1), 116.3 (C-4), 104.5 (C-10), 103.9 (C-8), 100.0 (C-6a), 79.3 (C-4a), 56.1 (OCH3-9), 29.6 (CH3-4a).
Altertoxin II (4) (Figures S8-S10) [27]: white crystal; HR-ESI–MS: m/z: 351.0849 [M + H]+, calculated for C20H15O6 351.0854; 1H NMR (600 MHz, CDCl3) δ: 12.71 (s, OH-18), 12.12 (s, OH-13), 7.91 (d, J = 8.8 Hz, H-20), 7.86 (d, J = 8.8 Hz, H-11), 7.11 (d, J = 8.8 Hz, H-19), 7.06 (d, J = 8.7 Hz, H-12), 4.23 (d, J = 3.7 Hz, H-10), 3.71 (d, J = 3.5 Hz, H-9), 3.54 (s, H-6), 3.30 – 3.21 (m, H-15), 2.89 (m, H-14),2.83 (m, H-15), 2.41 (td, J = 13.6, 4.0 Hz, H-14); 13C NMR (150 MHz, CDCl3) δ: 204.3 (C-16), 196.8 (C-8), 163.5 (C-18), 162.9 (C-13), 139.0 (C-5), 133.7 (C-2), 133.0 (C-20), 132.7 (C-11), 124.1 (C-3), 122.6 (C-4), 120.0 (C-19), 118.2 (C-12), 114.8 (C-17), 113.7 (C-7), 68.5 (C-1), 55.9 (C-10), 52.9 (C-9), 45.3 (C-6), 33.4 (C-14), 32.3 (C-15).
Three monomeric compounds (Fig. 2) were isolated from the liquid fermentation product of strain PP39. The compounds were further analyzed and identified as terreic acid (5), terremutin (6), citrinin (7) by spectroscopic analysis, including HR–ESI–MS, NMR, and compared with data described in previous literature.
Terreic acid (5) (Figures S11-S13) [28]: white crystal; HR-ESI–MS: m/z: 153.0193 [M—H]−, calculated for C7H5O4 153.0202; 1H NMR (600 MHz, CDCl3) δ: 1.93 (s, 3H, 3-Me), 3.87 (d, J = 3.7 Hz, H-6), 3.90 (d, J = 3.5 Hz, H-5), 6.83 (s, 2-OH); 13C-NMR (150 MHz, CDCl3) δ: 190.78 (C-4), 187.66 (C-1), 152.03 (C-2), 120.57 (C-3), 53.97 (C-5), 51.74 (C-6), 8.88 (3-Me).
Terremutin (6) (Figures S14-S16) [29]: white crystal; HR-ESI–MS: m/z: 157.0495 [M + H]+, calculated for C7H9O4 157.0487; 1H NMR (600 MHz, Acetone-d6) δ: 1.65 (s, 3H, H-7), 3.35 (s, H-6), 3.64 (s, H-5), 4.60 (s, H-4); 13C-NMR (150 MHz, Acetone-d6) δ: 109.0 (C-2), 66.3 (C-4), 55.3 (C-6), 52.3 (C-5), 7.5 (C-7).
Citrinin (7) (Figures S17-S19) [30]: yellow crystal; HR-ESI–MS: m/z: 251.0913 [M + H]+, calculated for C13H15O5 251.0905; 1H NMR (600 MHz, CDCl3) δ: 1.23 (d, J = 7.3 Hz, 3H, H-10), 1.34 (d, J = 6.8 Hz, 3H, H-9), 2.02 (s, 3H, H-11), 2.98 (q, J = 7.3 Hz, H-4), 4.77 (q, J = 6.5 Hz, H-3), 8.23 (s, H-1); 13C-NMR (150 MHz, CDCl3) δ: 184.0 (C-6), 177.4 (C-8), 174.7 (C-12), 162.7 (C-1), 139.1 (C-4a), 123.3 (C-5), 107.7 (C-8a), 100.6 (C-7), 81.8 (C-3), 34.8 (C-4), 18.6 (C-9), 18.4 (C-10), 9.6 (C-11).
Antibacterial activities of secondary metabolites isolated from PT09 and PP39
The antibacterial activities of seven compounds isolated from strains PT09 and PP39 were tested as shown in Table 4. The results showed that compound 5 exhibited strong antibacterial activities against E. coli, M. tetragenus, S. aureus, and P. syringae pv. actinidiae with the IZD of 36.0, 31.0, 33.7, 40.2 mm and MIC values of 1.56, 3.13, 1.56, 1.56 μg/mL, which were better than or equal to those of positive gentamicin sulfate with the IZD of 26.7, 25.7, 21.7, 26.0 mm and MIC values of 3.13, 3.13, 1.56, 6.25 μg/mL, respectively. The metabolite 7 also exhibited strong antibacterial activity against P. syringae pv. actinidiae with the IZD of 26.0 mm and MIC value of 6.25 μg/mL, and moderate activity against S. aureus with the IZD of 10.0 mm and MIC value of 100 μg/mL. Moreover, compound 4 showed potent or moderate antibacterial activity against P. syringae pv. actinidiae and S. aureus with the IZD of 16.2, 13.3 mm, and MIC values of 25, 100 μg/mL, respectively. In addition, compounds 2 and 3 showed moderate antibacterial activities against P. syringae pv. actinidiae and S. aureus with the IZD of 15.3 mm and 14.2 mm, respectively. However, they were found to have MIC values of more than 100 μg/mL on both pathogenic bacteria. Besides, compounds 1 and 6 were found to have no effect on four tested pathogenic bacteria.
Phytotoxic assay of secondary metabolites isolated from PT09 and PP39
Metabolites 1–7 were assayed for their ability to inhibit radicle growth of E. crusgalli and Abutilon theophrasti using a Petri dish bioassay (Table 5). The results showed that metabolite 7 had potent phytotoxic activity against E. crusgalli and A. theophrasti with the inhibition rate of 73.4% and 41.7%, respectively, which was weaker than those of the positive 2,4-D with an inhibition rate of 100% at a concentration of 100 μg/mL. Compound 5 had moderate phytotoxic activity against E. crusgalli and A. theophrasti with the inhibition rates of 38.4% and 38.0%, respectively. However, metabolites 1–4 and 6 showed weak inhibitory activity in this bioassay with the inhibition rate of less than 31%.
Discussion
Pinellia Tenore is a well-known medicinal plant in China, and its tubers have high medicinal value. In this study, the diversity of endophytic fungi of P. ternata and P. pedatisecta was studied. A total of 77 fungi were isolated and identified by culture-dependent method and molecular biology sequencing for the first time. All fungi were distributed in 25 genera. Among them, 53 and 24 fungal strains were isolated from P. ternata and P. pedatisecta, respectively. The dominant genus from P. ternata and P. pedatisecta was Fusarium, which was also the dominant endophytic fungi from a medicinal plant Ligusticum chuanxiong Hort [31]. Howerver, endophytic fungal community composition of Panax bipinnatifidus var. bipinnatifidus was found to be significantly different from that of P. ternata and P. pedatisecta, with the dominant genus of the Plectosphaerella, Paraphoma, and Fusarium [32]. Aspergillus sp. [17,18,19, 33], Chaetomium globosum [34], and Stachybotrys chartarum [35] were reported to have been isolated from P. ternata. Among them, C. globosum and S. chartarum were different from the fungi isolated in this study. Therefore, many endophytic fungi of P. ternata might remain undiscovered.
Antibiotics are a class of compounds used to treat microbial infectious diseases, and they are widely used to treat human and animal diseases, as well as in agricultural production [36]. However, the overuse of antibiotics has led to a serious problem of antibiotic resistance, and the development of antibiotics is imminent [37]. Natural products are an important source of new antibiotics [38]. In this study, 77 strains of endophytic fungi from P. ternata and P. pedatisecta were evaluated for their antibacterial activity. The results showed that 21 strains had antibacterial effects. Furthermore, we investigated the secondary metabolites of PT09 and PP39 with good bioactive activities, which resulted in the isolation of alternariol monomethyl ether (1), alternariol (2), dehydroaltenusin (3), altertoxin II (4), terreic acid (5), terremutin (6), citrinin (7). Among them, metabolite 5 from A. floccosus PP39 showed strong inhibitory activity against all four tested bacteria in this study. The result was similar to it’s antibacterial activity against E. coli, P. aeruginosa and Klebsiella pneumoniae [39]. Further research into the antibacterial activity of metabolite 5 might be expected to develop novel antibiotics. Although metabolites 5 and 6 had similar structures, we found that metabolite 6 showed no obvious antibacterial activity. Compared with compound 5, hydroxy at position 1 was oxidized to a carbonyl in compound 6, suggesting that the 1-hydroxy was essential for the antibacterial activities [40]. Compound 7 exhibited strong antibacterial activity against P. syringae pv. actinidiae with the IZD of 26.0 mm and MIC value of 6.25 μg/mL. Furthermore, compound 7 has also been reported that it presented strong antibacterial activity against methicillin-resistant S. aureus and rifampicin-resistant S. aureus with the MIC values of 3.90 and 0.97 μg/mL, respectively [41]. However, there was evidence that compound 7 showed nephrotoxic, hepatotoxic, and carcinogenic activity [42, 43]. Therefore, it might not be suitable for developing antibiotic.
Herbicide resistance has become one of the most important issues in global crop production. New herbicides are needed to control weeds that are resistant to existing herbicides [44]. We investigated the herbicidal activity of fermentation broth of 77 endophytic fungi. The results revealed that 72 strains (93.5%) of endophytic fungi had herbicidal activity against E. crusgalli. It was reported that the secondary metabolites of 28 endophytic fungi from 21 plants also had great herbicidal effects [45]. Therefore, endophytic fungi might be a potential source of herbicidal resources [45].
The fermentation broth of both PT09 and PP39 had strong phytotoxic activity against E. crusgalli with the inhibition rate of more than 91%. However, the metabolites 1–7 from them had moderate or weak phytotoxic activities against E. crusgalli. Therefore, it was necessary to further explore the bioactive metabolites from strains PT09 and PP39 that inhibit E. crusgalli growth.
Conclusions
In this study, 77 fungi were isolated and identified from roots, stems, leaves, and tubers of P. ternata and P. pedatisecta. Sequences analysis showed that all fungi were attached to the phyla Ascomycota and Basidiomycota, 68 strains of which were grouped into four classes. The most common endophytic fungi were Fusarium and Aspergillus. Antibacterial activities tests indicated that 21 endophytic fungi extracts exhibited antibacterial activities against at least one of the tested bacteria. Metabolite 5 from the A. floccosus PP39 exhibited outstanding antibacterial activities against E. coli, M. tetragenus, S. aureus, and P. syringae pv. actinidiae with the IZD of 36.0, 31.0, 33.7, 40.2 mm and MIC values of 1.56, 3.13, 1.56, 1.56 μg/mL respectively, which were better than or equal to positive gentamicin sulfate. The metabolite 7 also exhibited strong antibacterial activity against P. syringae pv. actinidiae with an IZD of 26.0 mm and MIC value of 6.25 μg/mL. Phytotoxic effects of 77 fungi on the radicle growth of E. crusgalli were investigated under laboratory conditions, and 22 fungi showed strong phytotoxic activity with the inhibition rate of 100%. In addition, the metabolite 7 had potent phytotoxic activity against E. crusgalli with the inhibition rate of 73.4% at the concentration of 100 μg/mL. In conclusion, this study showed that the endophytic fungi of P. ternata and P. pedatisecta held promise for the development of new antibiotic and herbicide resources.
Methods
Sample collection and microbial isolations
The healthy individuals of P. ternata and P. pedatisecta were collected from Hezhang County (27.13°N, 104.72°E, Bijie city, China) in March 2021. Five P. pedatisecta and P. ternata samples were selected and collected. These samples were immediately placed in autoclaved bags, labelled, and shipped to the laboratory in ice boxes. Then, they were stored at 4℃ and processed within 3 days.
All samples were washed thoroughly with sterile water, and the roots, tubers, stems, and leaves systems were separated from the individuals. The different plant tissues were cut into small pieces and soaked in 75% ethanol for 2 min, then in 5% sodium hypochlorite for 2–5 min, and followed by 75% ethanol for 1 min, respectively [46]. All samples were washed with sterile water for 2–3 times and then put on sterile filter paper to eliminate water. For control, the final sterile water rinse was spread on the plate and observed after incubation. The surface-sterilized explant fragments were homogenized separately in 1 mL of sterilized water. Then, the homogenates were serially diluted to 10–1 through 10–4 dilution, and 100 µL from each dilution was spread onto isolation media (Czapek–Dox Medium: NaNO3 3 g, K2HPO4 1 g, MgSO4 0.5 g, KCI 0.5 g, FeSO4 0.01 g, sugar 30 g, agar 15 ~ 20 g, distilled water 1000 mL, natural pH; Potato Dextrose Agar (PDA) Medium: potato 200 g, glucose 20 g, agar 15 ~ 20 g, distilled water 1000 mL, natural pH; MEA Medium: Malt extract 30 g, soybean peptone 3 g, agar 15 ~ 20 g, distilled water 1000 mL, natural pH). All isolation media were added with 50 μg/mL ampicillin and streptomycin. A single fungal colony from the isolation medium was inoculated into new PDA medium. The purified fungal strains were stored on PDA slope at 4℃.
DNA Extraction and PCR Amplification
DNA sequencing was performed according to the previous methods with some modified [47]. Fungi were transferred into ME medium (20 g sucrose, 20 g malt extract, 1 g peptone, then added to 1 L with distilled water) and cultured at 28 ± 0.5℃ in Constant Temperature Incubator Shakers for 3 days. About 100 mg of fungal mycelium was used for gene DNA extraction. According to the manufacturer’s instructions, the DNeasy Plant Minikit (Qiagen, Germany) was utilized. The internal transcriptional spacer (ITS) of fungal ribosomal DNA was amplified using the forward primer ITS1 (5'-TCCGTAGGTGAACCTGCGG-3') and the reverse primer ITS4 (5'-TCCTCCGCTTATTGATATGC-3') [48]. The PCR products were sent to TSINGKE Biological Technology Corporation (Shandong, China) for purification and bi-directionally sequencing. Then, the obtained 5.8S rDNA sequences were uploaded to the National Center for Biotechnology Information (NCBI) database.
Identification and phylogenetic analysis of the endophytic fungi
As previously reported, the affiliations of all resultant sequences returned from TSINGKE Biological Technology Corporation were identified by valid data in BLAST from NCBI database [47]. Sequence alignment and Neighbor-joining Phylogenetic Analysis were carried out using MEGA software version 7.0. Bootstrap analysis of tree construction built on 1000 replicates of sequence intensities to estimate neighbor-joining information [49].
Diversity analyses of endophytic fungi from P. ternata and P. pedatisecta
The diversity of endophytic fungi was evaluated according to previously described methods [22]. The species richness was evaluated by the species richness index (S) and Margalef index (D'). The species diversity was assessed by the Shannon–Wiener index (H'), Simpson’s diversity index (Ds), and Simpson’s dominant index (λ). The probability of interspecific encounters and species evenness was assessed by the probability of interspecific encounter (PIE) index and Pielou’s evenness index (J), respectively. All indexes were calculated by equation:
where Ni is the number of isolates belonging to the ith genus, Nt is the total number of endophytic fungal isolates’ in each tissue, H is H' of each tissue, and S is the number of total genera in each tissue.
Preparation of extracts of fermentation broth of endophytic fungi
Each fungus was transferred to PDA medium and incubated at 28 ± 0.5℃ for 3–4 days. Then, fresh mycelia of each fungus were transferred to a 250 mL conical flask containing 150 mL of ME liquid medium and incubated in a constant temperature culture shaker rotating at 180 rpm for 7 days at 28 ± 0.5℃. The culture was passed through four layers of cotton gauze to obtain the fermentation broth, then the fermentation broth was extracted three times with ethyl acetate (EtOAc, 1:1, v/v). The crude fungal extracts were obtained by concentrating the ethyl acetate phase in vacuo.
Isolation of compounds from PT09 and PP39
A total of 16 L of fermentation broth of PT09 was filtered and extracted three times with EtOAc (3 × 16 L) at room temperature [34]. The black-brown crude extract (2 g) was obtained by vacuum evaporation of the EtOAc phase. The crude fungal extract was separated by column chromatography (CC) using silica gel (SiO2: 200–300 mesh) and eluted with a stepwise gradient of CH2Cl2/MeOH (100:0–100:32, v/v) to provide seven primary fractions (Fr1 to Fr7). White needle-like crystals (compound 1, 3 mg) were obtained from Fr1 by successive recrystallization with CH2Cl2/MeOH. Fr2 (CH2Cl2/MeOH, 100:1) was further separated on a silica gel chromatography column to obtain compounds 2 (2.5 mg) and 3 (2.5 mg). Compound 4 (1 mg) was isolated from Fr3 on a silica gel chromatography column with CH2Cl2/MeOH = 100:2.
The yellow–brown mixture of PP39 (5 g) was obtained using the above method. Compound 5 (24.5 mg) was isolated and purified from subfractions Fr1-5, which were obtained by silica gel column separation in component Fr1 (CH2Cl2/MeOH, 100:0). Compounds 6 (26.5 mg) and 7 (3.5 mg) were isolated and purified from component Fr3 (CH2Cl2/MeOH, 100:2) on a silica gel chromatography column.
Structural elucidation of metabolites
The structures of all compounds were initially analyzed by 1H/13C Nuclear Magnetic Resonance (NMR) spectroscopy and High-Resolution Mass Spectrometry (HR-ESI–MS). 1H/13C NMR data were acquired using an Agilent DD2 600 Hz spectrometer (Agilent, USA), and chemical shifts were reported as parts per million (δ) by referring to tetramethylsilane (TMS) as an internal standard. HR-ESI–MS spectral data were collected on a TripeTOF 4600 mass analyzer (Bruker, USA).
Antibacterial activity
The antibacterial activities of 77 fungal crude extracts and metabolites were assessed using the filter paper disk method [50]. Four tested bacteria (E. coli, M. tetragenus, S. aureus, and P. syringae pv. actinidiae) were used, and three of which (E. coli, M. tetragenus, and S. aureus) were cultured on trypticase soybean blood agar (TSBA) medium at 37℃. P. syringae pv. actinidiae was cultured on LB medium at 28℃. All tested crude extracts and metabolites were dissolved separately in acetone to obtain a concentration of 6 mg/mL. The gentamicin sulfate was used as positive control. All the tested crude extracts, metabolites and controls needed to be filtered by 0.22 μm sterile filter membrane. Next, sterile filter paper discs (6 mm in diameter) were added to 5 μL of the tested samples and then placed on the pre-prepared medium. Three replicates were established for each test. The petri dishes were incubated in a constant-temperature incubator for 24–36 h. The diameter of the inhibitory circle (in mm) was measured using the crossover method to assess the antibacterial activity.
The antibacterial activity of the minimum inhibitory concentration (MIC) of 96 well plates was determined by the continuous dilution method [20]. Compounds 1–7 were dissolved in dimethyl sulfoxide (DMSO) to prepare 10 mg/ml. The gentamicin sulfate was used as positive control.
Phytotoxic assay
According to the methods described in previous literature [51], the phytotoxic activity of endophytic fungi was evaluated on radicle growth of E. crusgalli. The fungi were fermented in 150 mL of ME liquid medium at 28 ± 0.5℃ for 7 days to obtain the fermentation broth, which was filtered to remove mycelia. The E. crusgalli seeds were surface disinfected by soaking them in 5% sodium hypochlorite for 20 min. Then, the seeds were washed several times with deionized water. The seeds were cultured in an illuminating incubator at 28℃ until germination. Then, 30 pregerminated seeds were placed in 9 cm diameter Petri dishes on filter paper disks imbibed with 5.0 mL fungal fermentation broth. Radicle length was measured after 2–3 days, and distilled water was used as the negative control.
All compounds were evaluated for phytotoxic activity against E. crusgalli and A. theophrasti using Petri dish bioassay [52, 53]. E. crusgalli seeds were germinated using the method described above. The seeds of A. theophrasti were soaked in water at 60℃ for 30 min and transferred to a 40 mmol/L CaCl2 solution for 12 h. Then, the seeds were surface disinfected according to the above method and transferred to a 28℃ illuminating incubator until germination. The compounds were dissolved in acetone and diluted to 100 μg/mL with 0.1% aqueous Tween-80. The bioassay of the phytotoxic activity for compounds was the same as that of fermentation broth. 2,4-Dichlorophenoxyacetic acid (2,4-D) was used as the positive control.
Statistical analysis
Statistical differences were analyzed by one-way ANOVA with post-hoc LSD test and t-test. Values were considered significantly different when P-value was less than 0.05.
Availability of data and materials
The datasets generated and/or analysed during the current study are available in the NCBI repository, ON677855-ON677931.
References
Cheng Y, Chen C, Xue M, Chen L. Relationships of five Pinellia species (Araceae) inferred from AFLP data. Acta Bot Yunnanica. 2006;28(6):559–64.
Tao J, Hou Y, Ma X, Liu D, Tong Y, Zhou H, Gao J, Bai G. An integrated global chemomics and system biology approach to analyze the mechanisms of the traditional Chinese medicinal preparation Eriobotrya japonica – Fritillaria usuriensis dropping pills for pulmonary diseases. BMC Complement Altern Med. 2016;16:4.
Kurata K, Tai T, Yang Y, Kinoshita K, Koyama K, Takahashi K, Watanabe K, Nunoura Y. Quantitative analysis of anti-emetic principle in the tubers of Pinellia ternata by enzyme immunoassay. Planta Med. 1998;64:645–8.
Du J, Ding J, Mu ZQ, Guan SH, Cheng CR, Liu X, Guo DA. Three new alkaloids isolated from the stem tuber of Pinellia pedatisecta. Chin J Nat Med. 2018;16:139–42.
Han L, Chen C, Wang B, Wang ZZ. The complete chloroplast genome sequence of medicinal plant Pinellia ternata. Mitochondrial DNA Part A. 2016;27:2921–2.
Ji X, Huang B, Wang G, Zhang C. The ethnobotanical, phytochemical and pharmacological profile of the genus Pinellia. Fitoterapia. 2014;93:1–17.
Yao JH, Zhao XY, Liao ZH, Lin J, Chen ZH, Chen F, Song J, Sun XF, Tang KX. Cloning and molecular characterization of a novel lectin gene from Pinellia ternata. Cell Res. 2003;13:301–8.
Zhong F, Fan X, Ji W, Hai Z, Hu N, Li X, Liu G, Yu C, Chen Y, Lian B, et al. Soil fungal community composition and diversity of culturable endophytic fungi from plant roots in the reclaimed area of the eastern coast of China. J Fungi. 2022;8:124.
Fan M, Chen X, Luo X, Zhang H, Liu Y, Zhang Y, Wu J, Zhao C, Zhao P. Diversity of endophytic fungi from the leaves of Vaccinium dunalianum. Lett Appl Microbiol. 2020;71:479–89.
Zheng H, Qiao M, Lv Y, Du X, Zhang KQ, Yu Z. New species of trichoderma isolated as endophytes and saprobes from Southwest China. J Fungi. 2021;7:467.
Zhu JN, Yu YJ, Dai MD, Zeng YL, Lu XJ, Wang L, Liu XH, Su ZZ, Lin FC. A new species in Pseudophialophora from wild rice and beneficial potential. Front Microbiol. 2022;13:845104.
Jana SK, Islam MM, Mandal S. Endophytic microbiota of rice and their collective impact on host fitness. Curr Microbiol. 2022;79:37.
Macías-Rubalcava ML, Garrido-Santos MY. Phytotoxic compounds from endophytic fungi. Appl Microbiol Biotechnol. 2022;106:931–50.
Gupta S, Chaturvedi P, Kulkarni MG, Van Staden J. A critical review on exploiting the pharmaceutical potential of plant endophytic fungi. Biotechnol Adv. 2020;39:107462.
Xie J, Wei JG, Wang KW, Luo J, Wu YJ, Luo JT, Yang XH, Yang XB. Three phytotoxins produced by Neopestalotiopsis clavispora, the causal agent of ring spot on Kadsura coccinea. Microbiol Res. 2020;238:126531.
Ratnaweera PB, Williams DE, Patrick BO, de Silva ED, Andersen RJ. Solanioic acid, an antibacterial degraded steroid produced in culture by the fungus Rhizoctonia solani isolated from tubers of the medicinal plant Cyperus rotundus. Org Lett. 2015;17:2074–7.
Martinez-Klimova E, Rodríguez-Peña K, Sánchez S. Endophytes as sources of antibiotics. Biochem Pharmacol. 2017;134:1–17.
Gombodorj S, Yang MH, Shang ZC, Liu RH, Li TX, Yin GP, Kong LY. New phenalenone derivatives from Pinellia ternata tubers derived Aspergillus sp. Fitoterapia. 2017;120:72–8.
Gu L, Sun FJ, Li CP, Cui LT, Yang MH, Kong LY. Ardeemins and citrinin dimer derivatives from Aspergillus terreus harbored in Pinellia ternate. Phytochem Lett. 2021;42:77–81.
Wang ML, Chen R, Sun FJ, Cao PR, Chen XR, Yang MH. Three alkaloids and one polyketide from Aspergillus cristatus harbored in Pinellia ternate tubers. Tetrahedron Lett. 2021;68:152914.
Jiang S, Wang W, Xue X, Cao C, Zhang Y. Fungal diversity in major oil-shale mines in China. J Environ Sci. 2016;41:81–9.
Li P, Wu Z, Liu T, Wang Y. Biodiversity, Phylogeny, and antifungal functions of endophytic fungi associated with Zanthoxylum bungeanum. Int J Mol Sci. 2016;17:1541.
Zhang J, Zhang K, Jiang YE, Li YJ, Fan JT, Yu WM, Chen ZZ. Diversity and structure of demersal fish community over the northern slope in the South China Sea. Front Mar Sci. 2022;9:809636.
Mikula H, Skrinjar P, Sohr B, Ellmer D, Hametner C, Fröhlich J. Total synthesis of masked Alternaria mycotoxins—sulfates and glucosides of alternariol (AOH) and alternariol-9-methyl ether (AME). Tetrahedron. 2013;69:10322–30.
Soukup ST, Kohn BN, Pfeiffer E, Geisen R, Metzler M, Bunzel M, Kulling SE. Sulfoglucosides as novel modified forms of the mycotoxins alternariol and alternariol monomethyl ether. J Agric Food Chem. 2016;64:8892–901.
Nakanishi S, Toki S, Saitoh Y, Tsukuda E, Kawahara K, Ando K, Matsuda Y. Isolation of myosin light chain kinase inhibitors from microorganisms: dehydroaltenusin, altenusin, atrovenetinone, and cyclooctasulfur. Biosci Biotechnol Biochem. 1995;59:1333–5.
Wu WB, Yue GC, Huang QL, Sun LL, Zhang W. A new compound from an endophytic fungus Alternaria tenuissima. J Asian Nat Prod Res. 2014;16:777–82.
Cazar ME, Schmeda-Hirschmann G, Astudillo L. Antimicrobial butyrolactone I derivatives from the ecuadorian soil fungus Aspergillus terreus Thorn. var terreus. World J Microbiol Biotechnol. 2005;21:1067–75.
Guo CJ, Sun WW, Bruno KS, Wang CCC. Molecular genetic characterization of terreic acid pathway in Aspergillus terreus. Org Lett. 2014;16:5250–3.
Do Rosário Marinho AM, Rodrigues-Fo E. Dicitrinol, a citrinin dimer, produced by Penicillium janthinellum. Helv Chimi Acta. 2011;94:835–41.
Tang Z, Qin Y, Chen W, Zhao Z, Lin W, Xiao Y, Chen H, Liu Y, Chen H, Bu T, et al. Diversity, chemical constituents, and biological activities of endophytic fungi isolated from Ligusticum chuanxiong Hort. Front Microbiol. 2021;12:771000.
An C, Ma S, Shi X, Liu C, Ding H, Xue W. Diversity and ginsenoside biotransformation potential of cultivable endophytic fungi associated with Panax bipinnatifidus var. bipinnatifidus in Qinling mountains, China. Front Pharmacol. 2022;13:762862.
Xin XQ, Chen Y, Zhang H, Li Y, Yang MH, Kong LY. Cytotoxic seco-cytochalasins from an endophytic Aspergillus sp. harbored in Pinellia ternata tubers. Fitoterapia. 2019;132:53–9.
An L, Li C-P, Zhang H, Wang M-L, Kong L-Y, Yang M-H. Four cytochalasin alkaloids produced by Chaetomium globosum. Tetrahedron Lett. 2020;61(19):151838.
Zhang H, Yang MH, Zhuo FF, Gao N, Cheng XB, Wang XB, et al. Seven new cytotoxic phenylspirodrimane derivatives from the endophytic fungus Stachybotrys chartarum. RSC Adv. 2019;9(7):3520–31.
Manzetti S, Ghisi R. The environmental release and fate of antibiotics. Mar Pollut Bull. 2014;79:7–15.
Qiao M, Ying GG, Singer AC, Zhu YG. Review of antibiotic resistance in China and its environment. Environ Int. 2018;110:160–72.
Newman DJ, Cragg GM. Natural products as sources of new drugs over the nearly four decades from 01/1981 to 09/2019. J Nat Prod. 2020;83:770–803.
Sharma R, Lambu MR, Jamwal U, Rani C, Chib R, Wazir P, et al. Escherichia coli N-acetylglucosamine-1-phosphate-uridyltransferase/glucosamine-1-phosphate-acetyltransferase (GlmU) inhibitory activity of terreic acid isolated from Aspergillus terreus. J Biomol Screen. 2016;21(4):342–53.
Yao G, Chen X, Zheng H, Liao D, Yu Z, Wang Z, et al. Genomic and chemical investigation of bioactive secondary metabolites from a marine-derived fungus Penicillium steckii P2648. Front Microbiol. 2021;12:600991.
Subramani R, Kumar R, Prasad P, Aalbersberg W. Cytotoxic and antibacterial substances against multi-drug resistant pathogens from marine sponge symbiont: Citrinin, a secondary metabolite of Penicillium sp. Asian Pac J Trop Biomed. 2013;3(4):291–6.
Liang B, Du X-J, Li P, Sun C-C, Wang S. Investigation of citrinin and pigment biosynthesis mechanisms in Monascus purpureus by transcriptomic analysis. Front Microbiol. 2018;9:1374.
Wu J, Yang C, Yang M, Liang Z, Wu Y, Kong X, et al. The role of ER stress and ATP/AMPK in oxidative stress meditated hepatotoxicity induced by citrinin. Ecotoxicol Environ Saf. 2022;237:11353.
Qu RY, He B, Yang JF, Lin HY, Yang WC, Wu QY, Li QX, Yang GF. Where are the new herbicides? Pest Manag Sci. 2021;77:2620–5.
Macías-Rubalcava ML, Garrido-Santos MY. Phytotoxic compounds from endophytic fungi. Appl Microbiol Biotechnol. 2022;106(3):931–50.
Zhao LX, Xu LH, Jiang CL. Methods for the study of endophytic microorganisms from traditional Chinese medicine plants. Methods Enzymol. 2012;517:3–21.
Xu X, Shao M, Yin C, Mao Z, Shi J, Yu X, Wang Y, Sun F, Zhang Y. Diversity, bacterial symbionts, and antimicrobial potential of termite-associated fungi. Front Microbiol. 2020;11:300.
AlAgha R, Chew KL, Tay WJ, Lum L, Munson E. Answer to photo quiz: phaeohyphomycosis caused by Exophiala oligosperma. J Clin Microbiol. 2021;59:e02195-e2120.
Kumar S, Stecher G, Tamura K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol. 2016;33:1870–4.
Li S, Shao MW, Lu YH, Kong LC, Jiang DH, Zhang YL. Phytotoxic and antibacterial metabolites from Fusarium proliferatum ZS07 isolated from the gut of long-horned grasshoppers. J Agric Food Chem. 2014;62:8997–9001.
Lu YH, Jin LP, Kong LC, Zhang YL. Phytotoxic, antifungal and immunosuppressive metabolites from Aspergillus terreus QT122 isolated from the gut of dragonfly. Curr Microbiol. 2017;74:84–9.
Zhang YL, Kong LC, Jiang DH, Yin CP, Cai QM, Chen Q, Zheng JY. Phytotoxic and antifungal metabolites from Curvularia sp. FH01 isolated from the gut of Atractomorpha sinensis. Bioresource Technology. 2011;102:3575–7.
Sun XL, Ji ZM, Wei SP, Ji ZQ. Design, synthesis, and herbicidal activity of N-benzyl-5-cyclopropyl-isoxazole-4-carboxamides. J Agric Food Chem. 2020;68:15107–14.
Acknowledgements
K.K. thanks Anhui Agricultural University for the award of a fee-less scholarship. State Key Laboratory of Functions and Applications of Medicinal Plants, Guizhou Medical University; School of Life Sciences, Anhui Agricultural University; and Biotechnology center of Anhui Agriculture University are thanked by K.K. for the facilities provided for the research work.
Funding
This work was financially supported by the State Key Laboratory of Functions and Applications of Medicinal Plants, Guizhou Medical University (Grant number FAMP202008K); Guizhou Science and Technology Platform Talents (QKHRCPT [2019]5106); the National Natural Science Foundation of China (NSFC) (32011540382) and Anhui Province Natural Science Funds for Distinguished Young Scholars (2108085J18).
Author information
Authors and Affiliations
Contributions
YL-Z and WD-P designed the research and supervised the study. K-K, ZD-H and SP-S performed the experiments and prepared the figures and/or tables. All authors contributed to the article and approved the submitted version.
Corresponding authors
Ethics declarations
Ethics approval and consent to participate
Experimental research and field studies on plants including the collection of plant material are comply with relevant guidelines and regulation. The individuals of Pinellia ternata and Pinellia pedatisecta were collected from Hezhang County in March 2021. The plant materials collected for the study were cultivars, which were from the State Key Laboratory of Functions and Applications of Medicinal Plants, Guizhou Medical University. We have permission to collect the plants for the study.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no conflict of interest.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/. The Creative Commons Public Domain Dedication waiver (http://creativecommons.org/publicdomain/zero/1.0/) applies to the data made available in this article, unless otherwise stated in a credit line to the data.
About this article
Cite this article
Kong, K., Huang, Z., Shi, S. et al. Diversity, antibacterial and phytotoxic activities of culturable endophytic fungi from Pinellia pedatisecta and Pinellia ternata. BMC Microbiol 23, 30 (2023). https://doi.org/10.1186/s12866-022-02741-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s12866-022-02741-5