Abstract
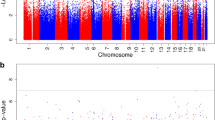
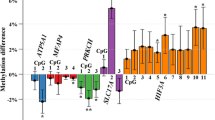
Repeat pregnancies with different perinatal outcomes minimize underlying maternal genetic diversity and provide unique opportunities to investigate nongenetic risk factors and epigenetic mechanisms of pregnancy complications. We investigated gestational diabetes mellitus (GDM)-related differential DNA methylation in early pregnancy peripheral blood samples collected from women who had a change in GDM status in repeat pregnancies. Six study participants were randomly selected from among women who had 2 consecutive pregnancies, only 1 of which was complicated by GDM (case pregnancy) and the other was not (control pregnancy). Epigenome-wide DNA methylation was profiled using Illumina HumanMethylation 27 BeadChips. Differential Identification using Mixture Ensemble and false discovery rate (<10%) cutoffs were used to identify differentially methylated targets between the 2 pregnancies of each participant. Overall, 27 target sites, 17 hypomethylated (fold change [FC] range: 0.77-0.99) and 10 hypermethylated (FC range: 1.01-1.09), were differentially methylated between GDM and control pregnancies among 5 or more study participants. Novel genes were related to identified hypomethylated (such as NDUFC1, HAPLN3, HHLA3, and RHOG) or hypermethylated sites (such as SEP11, ZAR1, and DDR). Genes related to identified sites participated in cell morphology, cellular assembly, cellular organization, cellular compromise, and cell cycle. Our findings support early pregnancy peripheral blood DNA methylation differences in repeat pregnancies with change in GDM status. Similar, larger, and repeat pregnancy studies can enhance biomarker discovery and mechanistic studies of GDM.
Similar content being viewed by others
References
Robertson K. DNA methylation and human disease. Nat Rev Genet. 2005;6(8):597–610.
Kirchner H, Osier ME, Krook A, Zierath JR. Epigenetic flexibility in metabolic regulation: disease cause and prevention? Trends Cell Biol. 2013;23(5):203–209.
Toperoff G, Aran D, Kark JD, et al. Genome-wide survey reveals predisposing diabetes type 2-related DNA methylation variations in human peripheral blood. Hum Mol Genet. 2012;21(2):371–383.
American Diabetes Association. Standards of medical care in diabetes-2014. 2014;37(suppl 1):S14–S80.
Di Cianni G, Ghio A, Resi V, Volpe L. Gestational diabetes mellitus: an opportunity to prevent type 2 diabetes and cardiovascular disease in young women. Womens Health (Lond Engl). 2010;6(1): 97–105.
Enquobahrie DA, Williams MA, Qiu C, Luthy DA. Early pregnancy lipid concentrations and the risk of gestational diabetes mellitus. Diabetes Res Clin Pract. 2005;70(2): 134–142.
Chatzi L, Plana E, Pappas A, et al. The metabolic syndrome in early pregnancy and risk of gestational diabetes mellitus. Diabetes Metab. 2009;35(6):490–494.
Cheng J, Tang L, Hong Q, et al. Investigation into the promoter DNA methylation of three genes (CAMK1D, CRY2 and CALM2) in the peripheral blood of patients with type 2 diabetes. Exp Ther Med. 2014;8(2):579–584.
Ross SA, Dwyer J, Umar A, et al. Introduction: diet, epigenetic events and cancer prevention. Nutr Rev. 2008;66(suppl 1):S1–S6.
American Diabetes Association. Gestational diabetes mellitus. Diabetes Care. 2004;27(suppl 1):S88–S90.
Sim S, Huang YW, Yan PS, Huang TH, Lin S. Preprocessing differential methylation hybridization microarray data. BioData Min. 2011;4:13.
Taslim C, Huang T, Lin S. DIME: R-package for identifying differential ChIP-seq based on an ensemble of mixture models. Bioinformatics. 2011;27(11):1569–1570.
Khalili A, Huang T, Lin S. A robust unified approach to analyzing methylation and gene expression data. Comput Stat Data Anal. 2009;53(5):1701–1710.
Maier S, Olek A. Diabetes: a candidate disease for efficient DNA methylation profiling. J Nutr. 2002;132(8 suppl):2440S–2443S.
Yokomori N, Tawata M, Onaya T. DNA demethylation during the differentiation of 3T3-L1 cells affects the expression of the mouse GLUT4 gene. Diabetes. 1999;48(4):685–690.
Poirier LA, Brown AT, Fink LM, et al. Blood S-adenosylmethionine concentrations and lymphocyte methylenetetrahydrofolate reductase activity in diabetes mellitus and diabetic nephropathy. Metabolism. 2001;50(9):1014–1018.
El Hajj N, Pliushch G, Schneider E, et al. Metabolic programming of MEST DNA methylation by intrauterine exposure to gestational diabetes mellitus. Diabetes. 2013;62(4): 1320–1328.
Bouchard L, Hivert MF, Guay SP, St-Pierre J, Perron P, Brisson D. Placental adiponectin gene DNA methylation levels are associated with mothers’ blood glucose concentration. Diabetes. 2012;61(5):1272–1280.
Bouchard L, Thibault S, Guay SP, et al. Leptin gene epigenetic adaptation to impaired glucose metabolism during pregnancy. Diabetes Care. 2010;33(11):2436–2441.
Zhao J, Goldberg J, Bremner JD, Vaccarino V. Global DNA methylation is associated with insulin resistance: a monozygotic twin study. Diabetes. 2012;61(2):542–546.
Ton C, Hwang DM, Dempsey AA, Liew CC. Identification and primary structure of five human NADH-ubiquinone oxidoreductase subunits. Biochem Biophys Res Commun. 1997;241(2): 589–594.
Rak M, Rustin P. Supernumerary subunits NDUFA3, NDUFA5 and NDUFA12 are required for the formation of the extramembrane arm of human mitochondrial complex I. FEBS Lett. 2014; 588(9): 1832–1838.
Patti ME, Corvera S. The role of mitochondria in the pathogenesis of type 2 diabetes. Endocr Rev. 2010;31(3):364–395.
Qiu C, Enquobahrie DA, Frederick IO, et al. Early pregnancy urinary biomarkers of fatty acid and carbohydrate metabolism in pregnancies complicated by gestational diabetes. Diabetes Res Clin Pract. 2014;104(3):393–400.
Peterson EA, Petty EM. Conquering the complex world of human septins: implications for health and disease. Clin Genet. 2010; 77(6):511–524.
Hall PA, Russell SE. The pathobiology of the septin gene family. J Pathol. 2004;204(4):489–505.
Roeseler S, Sandrock K, Bartsch I, Zieger B. Septins, a novel group of GTP-binding proteins: relevance in hemostasis, neuropathology and oncogenesis. Klin Padiatr. 2009;221(3):150–155.
Laird PW. The power and the promise of DNA methylation markers. Nat Rev Cancer. 2003;3(4):253–266.
Gong T, Hartmann N, Kohane IS, et al. Optimal deconvolution of transcriptional profiling data using quadratic programming with application to complex clinical blood samples. PLoS One. 2011; 6(11):e27156.
Shen-Orr SS, Tibshirani R, Khatri P, et al. Cell type-specific gene expression differences in complex tissues. Nat Methods. 2010; 7(4):287–289.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Enquobahrie, D.A., Moore, A., Muhie, S. et al. Early Pregnancy Maternal Blood DNA Methylation in Repeat Pregnancies and Change in Gestational Diabetes Mellitus Status—A Pilot Study. Reprod. Sci. 22, 904–910 (2015). https://doi.org/10.1177/1933719115570903
Published:
Issue Date:
DOI: https://doi.org/10.1177/1933719115570903