Abstract—
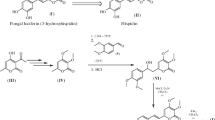
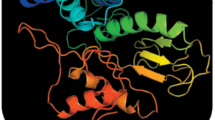
Bioluminescence is a phenomenon of light emission resulting from oxidation of a substrate, luciferin, catalyzed by the enzyme luciferase. The fungus Neonothopanus nambi is the first eukaryotic organism with a fully deciphered bioluminescent system: the structure of luciferin was established, the luciferase gene was described, and intermediates and enzymes involved in the luciferin biosynthesis pathway were identified. One of the crucial reactions in this pathway is the formation of luciferin by hydroxylation of hispidin catalyzed by hispidin-3-hydroxylase (nnH3H). To fully understand the mechanism of action and substrate specificity of the enzyme, it is necessary to carry out structural studies of the molecule. To do that, it is necessary to develop a protocol for obtaining a highly purified and functionally active nnH3H in the appropriate quantities. We describe a robust approach to produce a soluble and enzymatically active nnH3H fused with SUMO and coexpressed with GroEL/ES chaperonin at low temperature in Escherichia coli. The yield of recombinant nnH3H achieved was 20 mg per 100 mL of bacterial culture. Additionally, we show for the first time that FAD is a cofactor of fungal hispidin-3-hydroxylase.





Similar content being viewed by others
REFERENCES
Kaskova, Z.M., Tsarkova, A.S., and Yampolsky, I.V., Chem. Soc. Rev., 2016, vol. 45, pp. 6048–6077. https://doi.org/10.1039/C6CS00296J
Oba, Y., Stevani, C.V., Oliveira, A.G., Tsarkova, A.S., Chepurnykh, T.V., and Yampolsky, I.V., Photochem. Photobiol., 2017, vol. 93, pp. 405–415. https://doi.org/10.1111/php.12704
Kotlobay, A.A., Sarkisyan, K.S., Mokrushina, Y.A., Marcet-Houben, M., Serebrovskaya, E.O., Markina, N.M., Somermeyer, L.G., Gorokhovatsky, A.Y., Vvedensky, A., Purtov, K.V., Petushkov, V.N., Rodionova, N.S., Chepurnyh, T.V., Fakhranurova, L.I., Guglya, E.B., Ziganshin, R., Tsarkova, A.S., Kaskova, Z.M., Shender, V., Abakumov, M., Abakumova, T.O., Povolotskaya, I.S., Eroshkin, F.M., Zaraisky, A.G., Mishin, A.S., Dolgov, S.V., Mitiouchkina, T.Y., Kopantzev, E.P., Waldenmaier, H.E., Oliveira, A.G., Oba, Y., Barsova, E., Bogdanova, E.A., Gabaldon, T., Stevani, C.V., Lukyanov, S., Smirnov, I.V., Gitelson, J.I., Kondrashov, F.A., and Yampolsky, I.V., Proc. Natl. Acad. Sci. U. S. A., 2018, vol. 115, pp. x12728–12732. https://doi.org/10.1073/pnas.1803615115
Medvedev, K.E., Kinch, L.N., Schaeffer, R.D., and Grishin, N.V., PLoS Comput. Biol., 2019, vol. 15. e1007569. https://doi.org/10.1371/journal.pcbi.1007569
Rosano, G.L. and Ceccarelli, E.A., Front. Microbiol., 2014, vol. 5, p. 172. https://doi.org/10.3389/fmicb.2014.00172
McCoy, J. and Lavallie, E., Curr. Protoc. Mol. Biol., 2001, chapter 16, unit 16.8. https://doi.org/10.1002/0471142727.mb1608s28
Zhang, W., Lu, J., Zhang, S., Liu, L., Pang, X., and Lv, J., Microb. Cell Fact., 2018, vol. 17, p. 50. https://doi.org/10.1186/s12934-018-0894-y
Butt, T.R., Edavettal, S.C., Hall, J.P., and Mattern, M.R., Protein Expr. Purif., 2005, vol. 43, pp. 1–9. https://doi.org/10.1016/j.pep.2005.03.016
Kobayashi, H., Yoshida, T., and Inouye, M., Appl. Environ. Microbiol., 2009, vol. 75, pp. 5356–5362. https://doi.org/10.1128/AEM.00691-09
Lamppa, J.W., Tanyos, S.A., and Griswold, K.E., J. Biotechnol., 2013, vol. 164, pp. 1–8. https://doi.org/10.1016/j.jbiotec.2012.11.007
Mossessova, E. and Lima, C.D., Mol. Cell, 2000, vol. 5, pp. 865–876. https://doi.org/10.1016/s1097-2765(00)80326-3
Loomans, E.E., Roelen, A.J., Van Damme, H.S., Bloemers, H.P., Gribnau, T.C., and Schielen, W.J., J. Immunol. Methods, 1995, vol. 184, pp. 207–217. https://doi.org/10.1016/0022-1759(95)00089-s
Sambrook, J., Fritsch, E.F., and Maniatis, T., Molecular Cloning. A Laboratory Manual, Cold Spring Harbor Laboratory Press, 2012, 4th ed.
Kurien, B.T. and Scofield, R.H., BioTechniques, 1995, vol. 18, pp. 1023–1026.
Laemmli, U.K., Nature, 1970, vol. 227, pp. 680–685. https://doi.org/10.1038/227680a0
Zeder-Lutz, G., Cherouati, N., Reinhart, C., Pattus, F., and Wagner, R., Protein Expr. Purif., 2006, vol. 50, pp. 118–127. https://doi.org/10.1016/j.pep.2006.05.017
Goffin, P., Dewerchin, M., De Rop, P., Blais, N., and Dehottay, P., Biotechnol. J., 2017, vol. 12. https://doi.org/10.1002/biot.201700168
Bubyrev, A.I., Tsarkova, A.S., and Kaskova, Z.M., Russ. J. Bioorg. Chem., 2019, vol. 45, pp. 183–185. https://doi.org/10.1134/S106816201902002X
Funding
The research was carried out with the financial support of the Russian Foundation for Basic Research (grant No. 18-34-20134) and the grant of the President of the Russian Federation for state support of the leading scientific schools of the Russian Federation No. NSH-2605.2020.4.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
COMPLIANCE WITH ETHICAL STANDARDS
The article does not contain a description of research carried out by any of the authors of this work, involving humans or using animals as objects.
Conflict of Interests
I.V. Yampolsky is the founder of Planta LLC. Planta has applied for patents related to the use of components of the fungal bioluminescent system. A.S. Gerasimov, S.O. Rogozhkin, E.S. Shakhova, T.V. Chepurnykh, A. Yu. Gorokhovatsky, N.M. Myshkina and A.V. Balakireva declare no conflict of interest.
Additional information
Abbreviations: nnH3H, hispidin-3-hydroxylase N. nambi; Kd, dissociation constant.
Rights and permissions
About this article
Cite this article
Gerasimov, A.S., Rogozhkin, S.O., Shakhova, E.S. et al. Recombinant Production of Hispidin-3-Hydroxylase: the Key Enzyme in Fungal Luciferin Biosynthesis. Russ J Bioorg Chem 47, 1066–1076 (2021). https://doi.org/10.1134/S1068162021040099
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1068162021040099