Abstract
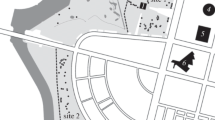
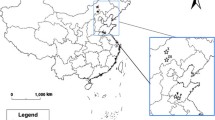
Over the past half-century, the common hamster (Cricetus cricetus), along with range-wide decline of natural populations, has actively populated the cities. The study of the genetic structure of urban populations of common hamster may shed light on features of the habitation of this species in urban landscapes. This article is focused on the genetic structure of common hamster populations in Simferopol (Crimea), one of the largest known urban populations of this species. On the basis of the analysis of nucleotide sequences of the cytochrome b gene and mtDNA control region, and the allelic composition of ten microsatellite loci of nDNA, we revealed that, despite the fact that some individuals can move throughout the city at considerable distances, the entire population of the city is represented by separate demes confined to different areas. These demes are characterized by a high degree of the genetic isolation and reduced genetic diversity compared to that found for the city as a whole.
Similar content being viewed by others
References
Haye, M.J.J., Neumann, K., and Koelewijn, H.P., Strong decline of gene diversity in local population of the highly endangered common hamster (Cricetus cricetus) in the western part of its European range, Conserv. Genet., 2012, vol. 13, pp. 311–322. doi 10.1007/s10592-011-0278-x
Neumann, K., Jansman, H., Kayser, A., et al., Multiple bottlenecks in threatened western European populations of the common hamster Cricetus cricetus (L.), Conserv. Genet., 2004, vol. 5, pp. 181–193. doi 10.1023/B:COGE.0000014055.95035cd
Neumann, K., Michaux, J., Maak, S., et al., Genetic spatial structure of European common hamsters (Cricetus cricetus)—a result of repeated range expansion and demographic bottlenecks, Mol. Ecol., 2005, vol. 14, pp. 1473–1483. doi 10.1111/j.1365-294X.2005. 02519x
Nechay, G., Status of Hamster: Cricetus cricetus, Cricetulus migratorius, Mesocricetus newtoni and Other Hamster Species in Europe, in Nature and Environment Series, Strasbourg: Council of Europe, 2000.
Korbut, Z., Rusin, M.Y., and Banaszek, A., The distribution of the common hamster (Cricetus cricetus) in western Ukraine, Zool. Poloniae, 2013, vol. 58, nos. 3–4, pp. 99–112. doi 10.2478/zoop-2013-0008
Franceschini-Zink, C. and Millesi, E., Population development and life expectancy in common hamsters, Biosyst. Ecol. Ser., 2008, vol. 25, pp. 45–59.
Schmelzer, E. and Millesi, E., Surface activity patterns in a population of European hamsters (Cricetus cricetus) in an urban environment, in The Common Hamster in Europe: Ecology, Management, Genetics, Conservation, Reintroduction (Proc. Meet. Int. Hamster Workgroup), Nechay, G., Ed., 2008, pp. 19–22.
Banaszek, A. and Ziomek, J., cricetus L.) population in the city of Lublin, Ann. Univ. Mariae Curie-Sklodowska, Sect. C, 2010, vol. 65, no. 1, pp. 59–66. doi 10.2478/v10067-011-0005-5
Vohralík, V., New records of Cricetus cricetus in the Czech Republic (Rodentia: Cricetidae), Lynx (Praha), 2011, vol. 42, pp. 189–196.
Anády, A., New site of the European hamster (Cricetus cricetus) in the urban environment of Ko ice city (Slovakia), Zool. Ecol., 2013, vol. 23, no. 1, pp. 61–65. doi 10.1080/21658005.2013.769772
Karaseva, E.V., Telitsyna, A.Yu., and Samoilov, B.L., Mlekopitayushchie Moskvy v proshlom i nastoyashchem (Mammals of Moscow in the Past and Present), Moscow: Nauka, 1999.
Sidorov, G.N., Kassal, B.Yu., Goncharova, O.V., et al., Teriofauna Omskoi oblasti (promyslovye gryzuny) (Theriofauna of the Omsk Oblast (Fur-Trapping Rodents)), Omsk: Nauka, 2011.
Feoktistova, N.Yu., Surov, A.V., Tovpinetz, N.N., et al., The common hamster as a synurbist: a history of settlement in European cities, Zool. Poloniae, 2013, vol. 58, nos. 3–4, pp. 113–126. doi 10.2478/zoop-2013-0009
Tovpinets, N.N., Evstaf’ev, I.L., and Karaseva, E.V., Inclination to synanthropy of the common hamster (Cricetus cricetus) based on observations in Crimea, in Fauna v antropogennom landshafte (Fauna in Anthropogenic Landscape) (Proc. Teriol. Shkoly, issue 8), Lugansk, 2006, pp. 136–145.
Luniak, M., Synurbization—adaptation of animal wildlife to urban development Proceedings 4th International Urban Wildlife Symposium, Shaw, W.W., Harris, K.L., and Druff, L., Eds., Tucson: Univ. Arizona, 2004, pp. 50–55.
Kajdacsi, B., Costa, F., Hyseni, C., et al., Urban population genetics of slum dwelling rats (Rattus norvegicus) in Salvador, Brazil, Mol. Ecol., 2013, vol. 22, pp. 5056–5070. doi 10.1111/mec.12455
Chiappero, M.B., Panzetta-Dutari, G.M., Gomez, D., et al., Contrasting genetic structure of urban and rural populations of the wild rodent Calomys musculinus (Cricetidae, Sigmodontinae), Mammal. Biol., 2011, vol. 76, pp. 41–50. doi 10.1016/jmambio.2010.02.003
Munshi-South, J. and Nagy, C., Urban park characteristics, genetic variation, and historical demography of white-footed mouse (Peromyscus leucopus) populations in New York City, Peer J., 2014, vol. 2, pp. 310–315. doi 10.7717/peerj.310
Kocher, T.D., Thomas, W.K., Meyer, A., et al., Dynamics of mitochondrial DNA evolution in animals: amplification and sequencing with conserved primers, Proc. Natl. Acad. Sci. U.S.A., 1989, vol. 86, pp. 6196–6200. doi 10.1073/pnas.86.16.6196
Xie, J. and Zhang, Z., Mitochondrial DNA phylogeography of populations of Cricetulus triton in the North China Plain, J. Mammal., 2005, vol. 84, pp. 833–840. doi 10.1644/09-mamm-a-098r 1.1
Montgelard, C., Bentz, S., Tirard, C., et al., Molecular systematics of Sciurognathi (Rodentia): the mitochondrial cytochrome b and 12S rRNA genes support the Anomaluroidea (Pedetidae and Anomaluridae), Mol. Phylogenet. Evol., 2002, vol. 22, no. 2, pp. 220–233. doi 10.1006/mpev.2001.1056
Hall, T.A., BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT, Nucleic Acids Symp. Ser., 1999, vol. 41, pp. 95–98.
Excoffier, L., Laval, G., and Schneider, S., Arlequin ver. 3.0: an integrated software package for population genetics data analysis, Evol. Bioinform. Online, 2005, vol. 1, pp. 47–50. PMCID: PMC2658868
Rousset, F., Genepop’007: a complete re-implementation of the Genepop software for Windows and Linux, Mol. Ecol. Notes, 2008, vol. 8, pp. 103–106. doi 10.1111/j.1558-5646.2012.01606x
Bandelt, H.-J., Forster, P., Röhl, A., Median-joining networks for inferring intraspecific phylogenies, Mol. Biol. Evol., 1999, vol. 16, pp. 37–48. http://www. fluxus-engineeringcom. doi 10.1093/oxfordjournalsmolbeva026036
Pritchard, J.K., Stephens, M., and Donnelly, P.J., Inference of population structure using multilocus genotype data, Genetics, 2000, vol. 155, pp. 945–959.
Documentation for structure software: version 2.3. http://pritchardlabstanfordedu/structure_software/release_versions/v2.3.4/structure_docpdf
Author information
Authors and Affiliations
Corresponding author
Additional information
Original Russian Text © N.Yu. Feoktistova, I.G. Meschersky, A.V. Surov, P.L. Bogomolov, N.N. Tovpinetz, N.S. Poplavskaya, 2016, published in Genetika, 2016, Vol. 52, No. 2, pp. 221–230.
Rights and permissions
About this article
Cite this article
Feoktistova, N.Y., Meschersky, I.G., Surov, A.V. et al. Genetic structure of urban population of the common hamster (Cricetus cricetus). Russ J Genet 52, 194–203 (2016). https://doi.org/10.1134/S1022795416020046
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1022795416020046