Abstract
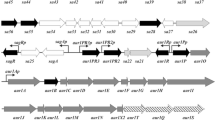
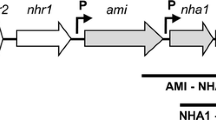
A new aliphatic amidase gene (ami), having a less than 77% level of similarity with the nearest homologs, was identified in the Rhodococcus erythropolis TA37 strain, which is able to hydrolyze a wide range of amides. The amidase gene was cloned within a 3.7 kb chromosomal locus, which also contains putative acetyl-CoA ligase and ABC-type transporter genes. The structure of this locus in the R. erythropolis TA37 strain differs from the structure of loci in other Rhodococcus strains. The amidase gene is expressed in Escherichia coli cells. It was demonstrated that amidase (generated in the recombinant strain) efficiently hydrolyzes acetamide (aliphatic amide) and does not use 4′-nitroacetanilide (N-substituted amide) as a substrate. Insertional inactivation of the amidase gene in the R. erythropolis TA37 strain results in a considerable decrease (by at least 6–7 times) in basal amidase activity, indicating functional amidase activity in the R. erythropolis TA37 strain.
Similar content being viewed by others
References
Debabov, V.G. and Yanenko, A.S., Biocatalytic hydrolyse of nitriles, Obz. Zh. Khim., 2011, vol. 1, no. 4, pp. 376–394.
Bork, P. and Koonin, E.V., A new family of carbon-nitrogen hydrolases, Protein Sci., 1994, vol. 3, no. 8, pp. 1344–1346.
Fournand, D. and Arnaud, A., Aliphatic and enantioselective amidases: from hydrolysis to acyl transfer activity, J. Appl. Microbiol., 2001, vol. 91, no. 3, pp. 381–393.
Lavrov, K.V. and Yanenko, A.S., Cloning of new acylamidase gene from Rhodococcus erythropolis and its expression in Escherichia coli, Russ. J. Genet., 2013, vol. 49, no. 10, pp. 1078–1081.
Ryabchenko, L.E., Polyakova, I.N., and Yanenko, A.S., Mobilizable plasmid vectors capable of conjugative transfer from Escherichia coli to Rhodococcus and their application to construction of Rhodococcus strains, Biotekhnol., 2005, no. 5, pp. 6–13.
Birnboim, H.C. and Doly, J., A rapid alkaline extraction procedure for screening recombinant plasmid DNA, Nucleic Acids Res., 1979, vol. 7, no. 6, pp. 1513–1523.
Ryabchenko, L.E., Podchernyaev, D.A., Kotlova, E.K., and Yanenko, A.S., Cloning the amidase gene from Rhodococcus rhodochrous M8 and its expression in Escherichia coli, Russ. J. Genet., 2006, vol. 42, no. 8, pp. 886–892.
Xu, M., Yu, H.M., Tan, T.W., et al., Molecular cloning of an amidase gene from Nocardia sp. and its expression in Escherichia coli, Sheng Wu Gong Cheng Xue Bao, 2006, vol. 22, no. 4, pp. 682–685.
Soubrier, F., Levy-Schil, S., Mayaux, J.F., et al., Cloning and primary structure of the wide-spectrum amidase from Brevibacterium sp. R312: high homology to the amiE product from Pseudomonas aeruginosa, Gene, 1992, vol. 116, no. 1, pp. 99–104.
Sharma, M., Sharma, N.N., and Bhalla, T.C., Purification studies on a thermo-active amidase of Geobacillus pallidus BTP-5x MTCC 9225 isolated from thermal springs of Tatapani, Appl. Biochem. Biotechnol., 2013, vol. 169, no. 1, pp. 1–14.
Solovyev, V. and Salamov, A., Automatic annotation of microbial genomes and metagenomic sequences, in Metagenomics and Its Applications in Agriculture, Biomedicine and Environmental Studies, Li, R.W., Ed., New York: Nova Science, 2011, pp. 61–78.
Bigey, F., Chebrou, H., Fournand, D., and Arnaud, A., Transcriptional analysis of the nitrile-degrading operon from Rhodococcus sp. ACV2 and high level production of recombinant amidase with an Escherichia coli-T7 expression system, J. Appl. Microbiol., 1999, vol. 86, no. 5, pp. 752–760.
Lavrov, K.V., Larikova, G.A., and Yanenko, A.S., Novel biocatalytic process of N-substituted acrylamide synthesis, Appl. Biochem. Microbiol., 2013, vol. 49, no. 8, pp. 1–4.
Author information
Authors and Affiliations
Corresponding author
Additional information
Original Russian Text © K.V. Lavrov, I.Yu. Karpova, A.S. Epremyan, A.S. Yanenko, 2014, published in Genetika, 2014, Vol. 50, No. 10, pp. 1145–1153.
Rights and permissions
About this article
Cite this article
Lavrov, K.V., Karpova, I.Y., Epremyan, A.S. et al. Cloning and analysis of a new aliphatic amidase gene from Rhodococcus erythropolis TA37. Russ J Genet 50, 1009–1016 (2014). https://doi.org/10.1134/S1022795414100056
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1022795414100056