Abstract
Expansins and xyloglucan endotransglycosylases play an important role in the regulation of plant growth under optimal and stressful conditions. Transgenic tobacco plants overexpressing NtEXPA1 and NtEXPA5 expansin genes and NtEXGT xyloglucan endotransglycosylase of Nicotiana tabacum L. have been previously created by the authors. The aim of this work was the morphophysiological analysis of the roots of these transgenic tobacco plants under conditions of cadmium stress. Transgenic tobacco plants were characterized by increased root length compared to wild type plants, both under optimal conditions and when exposed to cadmium. The area of parenchyma cells of roots of transgenic tobacco plants overexpressing NtEXPA1 and NtEXPA5 expansin genes was greater than the wild type, while the cell sizes, on the contrary, were smaller in the case of the transgene NtEXGT. Overexpression of NtEXPA1, NtEXPA5, and NtEXGT genes contributed to an increase in the total antioxidant capacity and activity of ascorbate peroxidases and a decrease in the content of proline in the roots under the action of cadmium. In the shoots of plants transgenic for the expansin genes, a lower content of MDA was found both under optimal conditions and under the action of cadmium. Thus, it has been shown that NtEXPA1 and NtEXPA5 transgenes have a stimulating effect on the growth of tobacco roots under conditions of cadmium stress by enhancing cell expansion and a positive effect on the components of the antioxidant system. The NtEXGT gene is also involved in root growth under the action of cadmium, including through the effect on the antioxidant system.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
INTRODUCTION
The cell wall of a growing plant cell consists of several layers of cellulose microfibrils immersed in a matrix consisting of glycans and pectic substances. Expansins are nonenzymatic proteins that cause reversible destruction of hydrogen bonds between glycans and cellulose microfibrils [1, 2]. Xyloglucan endotransglycosylases (XTHs) are apoplastic enzymes that carry out cleavage and transglycosylation reactions of glycans [3]. The essence of the cell wall expansion process is the tandem and synergistic work of expansins, XTHs, and some other enzymes by modifying the polymer network represented by cellulose microfibrils and glycans.
Overexpression of expansins and XTHs can positively affect plant productivity and stress resistance not only due to stimulation of cell expansion but also through the effect on the antioxidant system. For example, transgenic plants with constitutive expression of the OsEXPA7 rice expansin gene were characterized by a reduced amount of ROS and an increase in total antioxidant activity compared to the wild type [4]. However, the interaction of expansins and XTHs with the antioxidant system remains a poorly understood area.
Increased expression of expansins and XTHs genes improves root growth due to stimulation of cell expansion [5–7]. Expansins and XTHs have also been shown to be involved in plant stress resistance to drought, cold, and salinity by supporting the growth of root cells under conditions of moisture deficiency [6, 8–10]. The content of expansins NtEXPA1 [11], NtEXPA5 [12], and xyloglucan endotransglycosylase NtEXGT [6] gene transcripts increases in response to salinity, drought, and low positive temperatures, and tobacco plants transgenic according to these genes had improved growth parameters under the action of the above stress factors [6, 7]. However, how the roots of these plants would grow under the influence of cadmium remained unknown.
Cadmium and its compounds have a serious negative effect on all physiological and biochemical processes in plants [13]. Very little is known about the involvement of expansins and XTHs in cadmium stress tolerance. For example, overexpression of the PtoEXPA12 poplar expansin gene in tobacco plants [14] and TaEXPA2 wheat expansin gene [15] positively influenced the formation of plant resistance to cadmium stress. In this regard, it seems relevant to conduct additional studies on the contribution of expansins and XTHs to the regulation and maintenance of cadmium resistance.
The aims of this study were morphometric and microscopic analysis and assessment of the state of the antioxidant system in the roots of transgenic tobacco plants with overexpression of NtEXPA1, NtEXPA5, and NtEXGT genes with cadmium stress.
MATERIALS AND METHODS
Obtaining Transgenic Plants
Transgenic tobacco plants Nicotiana tabacum L. cultivar Petit Havana of line SR1 with overexpression of NtEXPA1 [11] and NtEXPA5 [12] expansin and NtEXGT xyloglucan endotransglycosylase genes [6] were obtained by agrobacterium-mediated transformation of leaf disks; the protocols are described in detail earlier [6, 11, 12]. The selection of transgenic plants was carried out based on the results of PCR analysis for the presence of target genes of expansins and NtEXGT as well as the 35SCaMV promoter. Morphometric analysis was carried out on second-generation transgenic plants grown on Murashige-Skoog selective medium (MS) with the addition of hygromycin to eliminate nontransgenic seedlings. The content of transgene transcripts was determined in the shoots and roots of the transgenic plants by real-time quantitative RT-PCR. Lines of transgenic plants characterized by a high content of NtEXPA1, NtEXPA5, and NtEXGT transgene transcripts were selected for subsequent analysis. Three lines of transgenic tobacco plants with overexpression of NtEXPA1 and NtEXPA5 expansin and NtEXGT xyloglucan endotransglycosylase genes were selected.
Real-Time Quantitative RT-PCR
Total RNA from the roots and shoots of the studied tobacco plants was isolated using Trizol, the first strand of cDNA was built using an oligo(dT) primer and MMuLV-revertase (NEB, United States). Quantification of mRNA content of NtEXPA1, NtEXPA5, and NtEXGT genes was carried out by real-time PCR in the presence of SYBR Green I intercalating dye on a Rotor-Gene thermal cyclerTM 6000 (Corbett Research, Australia). As a reference gene, as in our previous studies [6, 7, 10, 11], we used the gene EF-1α N. tabacum, characterized by the most stable level of expression under changing environmental conditions. The amplification reaction was carried out in 0.2-mL test tubes (AXYGEN, Inc., United States) in a volume of 25 µL using the reaction mixture for real-time PCR (Synthol, cat. no. M-427, Russia). Primer sequences for RT-PCR of EF-1α, NtEXPA1, NtEXPA5, and NtEXGT genes are presented in additional materials (Table S1).
Estimation of Root Growth Parameters of Transgenic Tobacco Plants by the Action of Cadmium Acetate
Transgenic plants were germinated in Binder climate chambers (Germany) at a temperature of +25°С, illumination of approximately 140 µM/(m2 s), and a photoperiod of 16/8 h (light/dark) on MS nutrient medium. After 10 days of germination on a selective medium with hygromycin, seedlings with the same root sizes were transferred to vertically oriented Petri dishes with MS medium, and root growth (length change) was determined at the norm (control) and under the action of 100, 200, and 400 μM acetate cadmium after 10 days. The same concentrations of cadmium acetate have been used in our previous studies and have been shown to significantly inhibit tobacco root growth [16]. The length of the roots of transgenic plants was measured before the start of the experiment and after a 10-day period. Later, when processing the results of the experiment, only the growth of the roots of transgenic plants for 10 days was taken into account. The control line was N. tabacum cultivar Petit Havana line SR1 wild type tobacco plants. The sample consisted of 60 plants for each line. The results of the studies were presented in the form of histograms with the mean values of the sample. The bars denote the standard error of the mean. The significance of differences in all experiments was assessed using the Duncan test (P < 0.05) [17].
Fixation and Microscopic Analysis of Roots
Tobacco roots were fixed in 4% formalin in phosphate buffer (pH 7.2) for 4 h at room temperature. They were then transferred into 30% glycerol prepared with 2% dimethyl sulfoxide and kept for 30 min at room temperature. The roots were then transferred to the “clearing solution” to prepare for viewing preparations under a microscope. The composition of the “clearing solution” was as follows: 3.7 M KI and 12.5 mM Na2S2O3 in 100 mL of 2% dimethyl sulfoxide. Next, 35 mL of this solution was mixed with 65 mL of 100% glycerol. Tobacco roots were kept in the “clearing solution” for at least 1.5 h, after which temporary preparations were prepared in 50% glycerol [18]. The size of root cells was studied in the variants: optimal growth conditions and growth under the action of 200 μM CdAc. In both variants, the plants were grown for 10 days before root fixation. Each repetition included ten roots per experimental variant in each line under study (n = 10). Root parenchyma cells were measured in the central part of the elongation zone. The root growth zone was determined visually during microscopic analysis by location between other zones, as well as by the characteristic cell structure. The cell size and area were analyzed using a Biozero BZ-8100 fluorescent microscope (Japan). We analyzed 150 cells for each variation of the experiment for each line of the studied plants (n = 150).
Analysis of the Antioxidant System of Transgenic Tobacco Plants under Conditions of Cadmium Stress
The activity of ascorbate peroxidases (APOCs) was determined according to the method described by Verma and Dubey [19]. The method is based on the calculation of the decomposition rate of H2O2 enzyme ascorbate peroxidase to form H2Oh and dehydroascorbate. The content of malonic dialdehyde (MDA) was measured using thiobarbituric acid according to the method described by Taylor and Millar [20]. The method for determining the amount of proline was taken from the work of Bates et al. [21] with modifications by Khedr et al. [22]. The assessment of the total antioxidant capacity (TAC) was carried out on methanol (80%) extracts by the reduction of molybdenum (VI) to molybdenum (V) in an acidic medium [23].
RESULTS
Content of NtEXPA1, NtEXPA5, and NtEXGT Transcripts in Transgenic Tobacco Plants
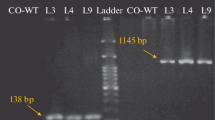
In the course of previous work on agrobacterium-mediated transformation, the following was obtained: 11 lines of transgenic plants with the NtEXPA1 expansin gene, 15 lines of transgenic plants with the NtEXPA5 expansin gene, and 12 lines of transgenic plants with the NtEXGT XTH gene. In all obtained plants, transgene expression was under the control of the constitutive 35SCaMV promoter. The transgenicity of all obtained lines was proven by PCR and RT-PCR analysis of the studied genes. The high content of transcripts of expansin genes and NtEXGT was found in shoots of all studied plants. A high content of gene transcripts in the roots was found in only a few obtained plants (additional material, Table S2). Further, we used lines of transgenic tobacco with an increased expression of the studied genes in the roots by a factor of three or more compared to the wild type in the experiments (Fig. 1, additional material, Table 2).
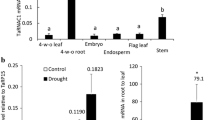
Content of (a) NtEXPA1 and (b) NtEXPA5 expansin gene transcripts and (c) NtEXGT xyloglucan endotransglycosylase in the roots of transgenic tobacco plants. Asterisks (*) denote significantly different results of the content of transcripts between wild type and transgenic plants after statistical analysis according to the Duncan test (P < 0.05).
In addition, the growth of roots over 10 days (Fig. 2) of these lines under optimal conditions was studied, which confirmed the relationship between changes in the content of transcripts and the growth of roots of transgenic plants with overexpression of NtEXPA1 (Fig. 2a) and NtEXPA5 (Fig. 2e) expansin genes. Under optimal conditions, the growth of the roots of these plants was, on average, 1.7 times greater than the growth of the roots of wild type tobacco. In transgenic plants with NtEXGT gene overexpression (Fig. 2i), the average growth of roots under optimal conditions was 1.9 times compared with the wild type.
Growth in 10 days of roots of transgenic tobacco plants with (a–d) NtEXPA1, (e–h) NtEXPA5, and (i–l) NtEXGT gene overexpression. (a, e, i) Optimal conditions, (b, f, j) 100 µM CdAc, (c, g, k) 200 µM CdAc, (d, h, l) 400 µM CdAc. WT: wild type. Asterisks (*) indicate significantly different results of root growth between wild type and transgenic plants after statistical analysis according to the Duncan test (P < 0.05).
Root Growth of Transgenic Tobacco Plants with NtEXPA1, NtEXPA5, and NtEXGT Overexpression when Exposed to Cadmium Acetate
The growth of roots during cultivation for 10 days on vertically oriented Petri dishes was analyzed in all lines of transgenic tobacco plants, which showed an increased relative content of NtEXPA1, NtEXPA5, and NtEXGT gene transcripts in the roots compared to the wild type. Under optimal conditions (+25°C), a significant increase in the length of the roots compared to the wild type was found in all lines of transgenic tobacco plants with overexpression of the NtEXPA1 gene (Fig. 2a), in lines 49 and 56 of transgenic tobacco plants with NtEXPA5 gene overexpression, and in the line of 17 transgenic tobacco plants with NtEXGT gene overexpression (Fig. 2i).
When plants were grown on MS medium with the addition of cadmium acetate at a concentration of 100 μM, increased rates of root growth compared to the wild type were detected in transgenic tobacco plants of line 49 with the NtEXPA5 gene (Fig. 2f) and lines 17 and 21 of transgenic tobacco plants with NtEXGT gene overexpression (Fig. 2j). Lines 5 and 13 of transgenic tobacco plants with NtEXPA1 gene overexpression (Fig. 2b) and line 22 of transgenic tobacco plants with the NtEXGT gene showed a decrease in root growth relative to the wild type when the content of cadmium acetate in the medium was 100 μM. Under the action of 200 μM cadmium acetate, all lines of the studied transgenic plants were characterized by faster root growth rates (Figs. 2c, 2g, 2k) compared to the wild type. Under the action of cadmium acetate at a concentration of 400 μM, almost all lines also showed a significantly greater increase in roots compared to the increase in wild type roots, with the exception of line 22 of transgenic tobacco plants with the NtEXGT gene (Figs. 2d, 2h, 2l).
Microscopic Analysis of the Roots of Transgenic Tobacco Plants Exposed to Cadmium Acetate
Parenchyma cells of wild type tobacco roots and transgenic plants overexpressing NtEXPA1, NtEXPA5, and NtEXGT genes under optimal conditions and under the influence of cadmium acetate at a concentration of 200 μM were analyzed (Fig. 3). Analysis and fixation of cells were carried out 10 days after planting on vertically oriented Petri dishes. The shape of the root cells was cylindrical and elongated under optimal conditions and under the influence of cadmium acetate. The cells acquired a cubic shape only in transgenic tobacco plants with overexpression of the NtEXPA1 expansin gene upon exposure to cadmium acetate 200 μM (Fig. 3d). At the same time, no visual changes in the thickness of the cell walls were observed.
We found that the cell area in transgenic tobacco plants with NtEXPA1 and NtEXPA5 gene overexpression was greater than the wild type on nutrient media with cadmium acetate at a concentration of 200 μM. In transgenic plants with NtEXGT gene overexpression, the cell area of the root parenchyma was less than that of the wild type, both under optimal conditions and under the action of cadmium acetate (Table 1).
Analysis of the Antioxidant System in Transgenic Tobacco Plants Exposed to Cadmium Acetate
Cadmium has a negative effect on plants through an increase in the level of oxidative stress and the formation of reactive oxygen species (ROS) in cells. ROS, in turn, disrupt the structure and components of the plant cell [24]. Since cadmium affects the entire plant, it was decided to analyze the antioxidant system in both roots and shoots of transgenic tobacco plants with overexpression of the studied genes. Plants for experiments were grown for 10 days on vertically oriented Petri dishes, then appropriate analyzes were carried out.
The total antioxidant capacity (TAC) of shoots exposed to cadmium at a concentration of 200 μM in transgenic tobacco plants overexpressing all the studied genes was lower than in the wild type (Fig. 4a). TAC in transgenic roots by NtEXPA5 and NtEXGT genes of plants exposed to cadmium acetate was higher compared to the wild type (Fig. 4b).
Analysis of the antioxidant system of wild type tobacco and transgenic plants with NtEXPA1, NtEXPA5, and NtEXGT gene overexpression under normal conditions and when exposed to cadmium acetate at a concentration of 200 μM 10 days after the start of the experiment on vertically oriented Petri dishes: (a) TAC in shoots; (b) TAC in roots; (c) APOC activity in shoots; (d) APOC activity in roots; (e) content of MDA in shoots; (f) content of MDA in roots; (g) proline content in shoots; (h) Proline content in roots. WT: wild type. Asterisks (*) denote significantly different results between wild-type and transgenic plants after statistical analysis according to the Duncan test (P < 0.05).
In the roots of most of the analyzed transgenic plants, an increased activity of APOC enzymes was found in comparison with the wild type, both under optimal conditions and when exposed to cadmium acetate at a concentration of 200 µM (Figs. 4c, 4d).
In transgenic plants with overexpression of expansin genes, the content of MDA in shoots, both under optimal conditions and under the action of cadmium acetate, was lower than in the wild type (Fig. 4e). However, the content of MDA was higher than in the wild type in NtEXGT transgenic gene plants in the roots and shoots, both under normal and under cadmium stress (Figs. 4e, 4f).
The content of proline in the shoots of all transgenic plants under the action of cadmium acetate (Fig. 4g) was higher than in the wild type. In the roots, the amount of proline under the action of cadmium increased both in the wild type and in all plant lines; however, that in NtEXPA5 and NtEXGT transgenic plants was to a lesser extent than in the wild type (Fig. 4h).
DISCUSSION
An important role of expansins and XTHs is to ensure the growth of roots and shoots under the action of such abiotic stress factors as drought, hypothermia, and salinity [2]. Our data prove the participation of expansins and XTHs in the growth of roots and shoots also under the action of cadmium. Similar data were obtained by Ren et al. [15]: tobacco plants overexpressing the wheat expansin gene TaEXPA2, had a higher rate of germination, root elongation, and accumulated more biomass compared to wild type plants after treatment with CdCl2. For XTHs genes, such results have not been shown before our studies, although data are available on the activation of XTHs gene expression under the action of cadmium [25]. Our results in transgenic tobacco plants prove that overexpression of both expansins and XTHs genes increases plant resistance to cadmium acetate.
The transgenic tobacco plants, overexpressing NtEXPA1 and NtEXPA5 genes, had an increased root length compared to the wild type, both under optimal conditions and when exposed to cadmium at a concentration of 200 μM due to an increase in the size of parenchymal cells in the elongation zone by expansion the roots. Lee et al. [26] also found a correlation between the overexpression of the GmEXP1 expansin gene at the tips of soybean roots with cell expansion. Overexpression of an OsEXPA8 rice root-specific gene improved root growth by stimulating cell expansion [27]. It was shown that the OsEXPA10 rice expansin gene participates in the elongation of rice root cells under the action of aluminum [28]. Thus, the overexpression of expansin genes positively affects the growth of root cells by elongation under the action of cadmium. Roots of 35S:NtEXGT transgenic plants also grew faster than in the wild type under both normal and cadmium stress conditions. However, in this case, there was no correlation between root growth and cell size. Conversely, NtEXGT gene overexpression possibly stimulated cell division (Table 1).
35S::NtEXPA5 and 35S::NtEXGT plants were characterized by increased TAC in the roots when exposed to cadmium compared to the wild type (Fig. 4b). In the roots of the transgenic plants we analyzed, a higher ascorbate peroxidase activity was also observed than in the wild type (Fig. 4d). Ren et al. [15] also found that transgenic plants overexpressing the wheat expansin gene TaEXPA2 have increased resistance to CdCl2 by increasing the activity of the components of the antioxidant system. It should be noted that such changes in TAC and peroxidase activity were not found in shoots of transgenic plants. Apparently, this is due to a lower stress load in the organs of the shoot compared to the roots.
In the shoots of plants transgenic for the expansin genes, a lower content of MDA was found compared with the wild type under cadmium stress (Fig. 4e). This may indicate a lower stress load experienced by transgenic plants, possibly due to the protective effect of expansins [14]. Protective effect of the NtEXGT gene product is apparently realized through other mechanisms since even more MDA accumulated in the roots of transgenic plants 35S:NtEXGT under cadmium stress than in the wild type. Such a high accumulation of MDA can be explained in this case by the higher growth rates of the roots of transgenic plants compared to the wild type.
The amount of free proline in plants increases many times in response to the impact of heavy metals, which indicates the sensitivity of plants to this stressor [29]. However, in roots transgenic for genes NtEXPA5 and NtEXGT plants, we observed a lower content of proline than in the wild type (Fig. 4h). These data may also indicate the presence of a protective effect of expansins and XTHs in cadmium stress [15].
Changes in the activity of the antioxidant system during overexpression of expansin genes have been described previously [15, 30], while a very intriguing question still remains unanswered: what is the specific mechanism of the effect of these cell wall proteins, which are known mainly only as regulators of cell expansion, on the antioxidant system? It is well known that some ROS, for example, the hydroxyl radical, are positive regulators of cell growth, directly participating in the breakdown of cell wall polysaccharides, i.e., they act synergistically with expansins and XTHs [31]. Based on this, it can be suggested that a compensatory decrease in the content of ROS occurs in the roots of transgenic plants with overexpression of the expansins and XTHs genes. Confirmation of this hypothesis requires further research.
Thus, expansins have a stimulating effect on root growth under conditions of cadmium stress by enhancing cell expansion and positive changes in the antioxidant system. Xyloglucan endotransglycosylases also have a positive effect on root growth under the action of cadmium, which is expressed by the same changes in the antioxidant system: an increase in peroxidase activity and overall antioxidant capacity. NtEXPA1, NtEXPA5, and NtEXGT genes can be proposed as targets for the creation of transgenic plants with increased resistance to cadmium.
Change history
08 May 2023
An Erratum to this paper has been published: https://doi.org/10.1134/S1021443723330014
REFERENCES
Sharova, E.I., Expansins: Proteins involved in cell wall softening during plant growth and morphogenesis, Russ. J. Plant Physiol., 2007, vol. 54 (6), p. 805. https://doi.org/10.1134/S1021443707060015
Cosgrove, D.J., Plant expansins: diversity and interactions with plant cell walls, Curr. Opin. Plant Biol., 2015, vol. 25, p. 162. https://doi.org/10.1016/j.pbi.2015.05.014
Van Sandt, V.S., Suslov, D., Verbelen, J.P., and Vissenberg, K., Xyloglucan endotransglucosylase activity loosens a plant cell wall, Ann. Bot., 2007, p. 1467. https://doi.org/10.1093/aob/mcm248
Jadamba, C., Kang, K., Paek, N.C., Lee, S.I., and Yoo, S.C., Overexpression of rice expansin 7 (Osexpa 7) confers enhanced tolerance to salt stress in rice, Int. J. Mol. Sci., 2020, vol. 21, p. 454. https://doi.org/10.3390/ijms21020454
Lin, C., Choi, H.S, and Cho, H.T., Root hair-specific EXPANSIN A7 is required for root hair elongation in Arabidopsis, Mol. Cells., 2011, vol. 31, p. 393. https://doi.org/10.1007/s10059-011-0046-2
Kuluev, B.R., Mikhaylova, E.V., Berezhneva, Z.A., Nikonorov, Y.M., Postrigan, B.N., Kudoyarova, G.R., and Chemeris, A.V., Expression profiles and hormonal regulation of tobacco NtEXGT gene and its involvement in abiotic stress response, Plant Physiol. Biochem., 2017, vol. 111, p. 203. https://doi.org/10.1016/j.plaphy.2016.12.005
Kuluev, B.R., Berezhneva, Z.A., Mikhaylova, E.V., and Chemeris, A.V., Growth of transgenic tobacco plants with changed expression of genes encoding expansins under the action of stress factors, Russ. J. Plant Physiol., 2018, vol. 65, p. 211. https://doi.org/10.1134/S1021443718020036
Zhao, M.R., Li, F., Fang, Y., Gao, Q., and Wang, W., Expansin-regulated cell elongation is involved in the drought tolerance in wheat, Protoplasma, 2011, vol. 248, p. 313. https://doi.org/10.1007/s00709-010-0172-2
Xu, Q., Xu, X., Shi, Y., Xu, J., and Huang, B., Transgenic tobacco plants overexpressing a grass PpEXP1 gene exhibit enhanced tolerance to heat stress, PLoS One, 2014, vol. 9, p. e100792. https://doi.org/10.1371/journal.pone.0100792
Kuluev, B.R., Avalbaev, A.M., Mikhaylova, E.V., Nikonorov, Y.M., Berezhneva, Z.A., and Chemeris, A.V., Expression profiles and hormonal regulation of tobacco expansin genes and their involvement in abiotic stress response, J. Plant Physiol., 2016, vol. 206, p. 1. https://doi.org/10.1016/j.jplph.2016.09.001
Kuluev, B.R., Knyazev, A.V., Nikonorov, Yu.M., and Chemeris A.V., The role of the NtEXPA1 and NtEXPA4 genes in the regulation of cell elongation during the growth of tobacco leaves, Genetics, 2014, vol. 50, p. 560. https://doi.org/10.1134/S1022795414010074
Kuluev, B.R., Safiullina, M.G., Knyazev, A.V., and Chemeris, A.V., Effect of ectopic NtEXPA5 gene expression on cell size and organ growth in transgenic tobacco plants, Ontogenez, 2013, vol. 44, p. 34. https://doi.org/10.1134/S1062360413010049
Titov, A.F., Kaznina, N.M., and Talanova, V.V., Heavy metals and plants, Petrozavodsk: Karelian Scientific Center of the RAS, 2014, 194 p.
Zhang, H., Ding, Y., Zhi, J., Li, X., Liu, H., and Xu J., Over-expression of the poplar expansin gene PtoEXPA12 in tobacco plants enhanced cadmium accumulation, Int. J. Biol. Macromol., 2018, vol. 116, p. 676. https://doi.org/10.1016/j.ijbiomac.2018.05.053
Ren, Y., Chen, Y., An, J., Zhao, Z., Zhang, G., Wang, Y., and Wang, W., Wheat expansin gene TaEXPA2 is involved in conferring plant tolerance to Cd toxicity, Plant Sci., 2018, vol. 270, p. 245. https://doi.org/10.1016/j.plantsci.2018.02.022
Berezhneva, Z.A., Kashafutdinova, A.R., and Kuluev, B.R., Growth of roots of transgenic Nicotiana tabacum L. plants with constitutive expression of the rapeseed glutathione synthetase BnGSH gene under the action of stress factors, Bull. Plant Protect, 2017, vol. 3(93), p. 55.
Duncan, D.B., Multiple range and multiple F-test, Biometrics, 1955, vol. 11, p. 1. https://doi.org/10.2307/3001478
Filin, A.N. and Ivanov, V.B., Influence of 2,4-D on cell proliferation and elongation in the roots of Arabidopsis thaliana, Plant Physiol., 2016, vol. 63, p. 174. https://doi.org/10.7868/S0015330316010061
Verma, S., Dubey, R.S., Lead toxicity induces lipid peroxidation and alert the activities of antioxidant enzymes in grooving rice plants, Plant Sci., 2003, vol. 64, p. 645. https://doi.org/10.1016/S0168-9452(03)00022-0
Taylor, N.L. and Millar, A.H., Oxidative stress and plant mitochondria, Met. Mol. Biol., 2007, vol. 372, p. 389. https://doi.org/10.1007/978-1-59745-365-3_28
Bates, L.S., Waldren, R.P., and Teare, I.D., Rapid determination of free proline for water-stress studies, Plant Soil., 1973, vol. 39, p. 205. https://doi.org/10.1007/BF00018060
Khedr, A.H.A., Abbas, M.A., Abdel, W.A.A., Quick, W.P., and Abogadallah, G.M., Proline induces the expression of salt-stress-responsive proteins and may improve the adaptation of Pancratium maritimum L. to salt-stress, J. Exp. Bot., 2003, vol. 54, p. 2553. https://doi.org/10.1093/jxb/erg277
Boestfleisch, C., Wagenseil, N.B., Buhmann, A.K., Seal, C.E., Wade, E.M., Muscolo, A., and Papenbrock, J., Manipulating the antioxidant capacity of halophytes to increase their cultural and economic value through saline cultivation, AoB Plants, 2014, vol. 13, p. 6. https://doi.org/10.1093/aobpla/plu046
Sazanova, K.A., Bashmakov, D.I., and Lukatkin, A.S., Generation of superoxide anion-radical in plant leaves under chronic exposure to heavy metals, Tr. Karel. Res. Cent. of RAS, Experimental biology, 2012, vol. 2, p. 119.
Liu, Y., Yu, X., Feng, Y., Zhang, C., Wang, C., Zeng, J., Huang, Z., Kang, H., Fan, X., Sha, L., Zhang, H., Zhou, Y., Gao, S., and Chen, Q., Physiological and transcriptome response to cadmium in cosmos (Cosmos bipinnatus Cav.) seedlings, Sci. Rep., 2017, vol. 7, p. 14691. https://doi.org/10.1038/s41598-017-14407-8
Lee, D.K., Ahn, J.H., Song, S.K., Choi, Y.D., and Lee, J.S., Expression of an expansin gene is correlated with root elongation in soybean, Plant Physiol., 2003, vol. 131, p. 985. https://doi.org/10.1104/pp.009902
Ma, N., Wang, Y., Qiu, S., Kang, Z., Che, S., Wang, G., and Huang, J., Overexpression of OsEXPA8, a root-specific gene, improves rice growth and root system architecture by facilitating cell extension, PLoS One, 2013, vol. 8, e75997. https://doi.org/10.1371/journal.pone.0075997
Che, J., Yamaji, N., Shen, R.F., and Ma, J.F., An Al-inducible expansin gene, OsEXPA10 is involved in root cell elongation of rice, Plant J., 2016, vol. 88, p. 132. https://doi.org/10.1111/tpj.13237
Kuznetsov, V.V. and Shevyakova, N.I., Proline under stress: biological role, metabolism, regulation, Plant Physiol., 1999, vol. 46, p. 321.
Kuluev, B.R., Musin, H.G., and Yakupova, A.B., The NtEXPA5 expansin gene increases the stress resistance of tobacco hairy roots through its effect on the antioxidant system, Ecological genetics, 2021, vol. 19 (1), p. 5.
Liszkay, A., Zalm, E., and Schopfer, P., Production of reactive oxygen intermediates (O2 –, H2O2, and OH) by maize roots and their role in wall loosening and elongation growth, Plant Physiol., 2004, vol. 136, p. 3114.
Funding
The work was carried out within the framework of State Assignment no. AAAA-A19-119021190011-0 with the support of a grant from the president of the Russian Federation, MD-2304.2020.4.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
This article does not contain any studies involving humans and people as subjects. The authors declare that they have no conflicts of interest.
Additional information
Abbreviations: APOC, ascorbate peroxidase; OAC, total antioxidant capacity; CdAc, cadmium acetate; XTH/XTHs, xyloglucan endotransglycosylase/s.
The original online version of this article was revised: Due to a retrospective Open Access order.
Supplementary Information
Rights and permissions
Open Access. This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Berezhneva, Z.A., Musin, K.G. & Kuluev, B.R. Root Growth of Transgenic Tobacco Plants with Overexpression of Expansin and Xyloglucan endotransglycosylase Genes under Cadmium Stress. Russ J Plant Physiol 69, 96 (2022). https://doi.org/10.1134/S102144372205003X
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1134/S102144372205003X