Abstract
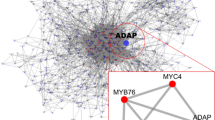
The methylerythritol phosphate (MEP) pathway for the production of isoprenoids is recently discovered. The current study aimed to identify MEP pathway disorder-related molecular mechanisms and potential genes in Arabidopsis thaliana. Microarray data (GSE61675) obtained from ceh1 mutant plants and corresponding parental lines were retrieved from Gene Expression Omnibus (GEO) database and were applied for differentially expressed genes (DEGs) screening. Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis of DEGs were performed. Protein-protein interaction (PPI) network was then constructed and displayed by Cytoscape software. Total 762 DEGs including 620 up-regulated and 142 down-regulated genes were screened. In addition, a great many of DEGs were mainly involved in biosynthesis and metabolism-related pathways, such as stilbenoid, diarylheptanoid, and gingerol biosynthesis, and biosynthesis of terpenoids and steroids. Moreover, a PPI network contained 90 down-regulated genes and 497 up-regulated genes were obtained. Up-regulated DEGs including glutaredoxin (GRX480, cytochrome BC1 synthase (BCS1, syntaxin of plants 121 (SYP121) and A. thaliana MAP kinase 11 (ATMPK11) with higher degree in this network were hub nodes. Pathways including stilbenoid, diarylheptanoid, and gingerol biosynthesis obtained in our study were consistent with previous studies. Importantly, GRX480, BCS1 and ATMPK11 could have close interactions with the MEP pathway and may play important roles in the biosynthesis of isoprenoids.
Similar content being viewed by others
Abbreviations
- DAVID:
-
database for annotation, visualization, and integrated discovery
- DEGs:
-
differentially expressed genes
- GEO:
-
Gene Expression Omnibus
- HPL:
-
hydroperoxide lyase
- IspH/HDR:
-
hydroxymethylbutenyl diphosphate reductase
- JA:
-
jasmonic acid
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- MAP kinases:
-
mitogen-activated protein kinases
- MEcPP:
-
methylerythritol cyclodiphosphate
- MEP:
-
methylerythritol phosphate
- MVA:
-
mevalonic acid
- NCBI:
-
National Center for Biotechnology Information
- PPI:
-
protein-protein interaction
- STRING:
-
Search Tool for the Retrieval of Interacting Genes
- SA:
-
salicylic acid
References
Rodríguez-Concepción, M., Campos, N., Ferrer, A., and Boronat, A., Biosynthesis of isoprenoid precursors in Arabidopsis, in Isoprenoid Synthesis in Plants and Microorganisms, Bach, T.J. and Rohmer, M., Eds., New York: Springer-Verlag, 2013, pp. 439–456.
Bloch, K., The biological synthesis of cholesterol, Science, 1965, vol. 150, pp. 19–28.
Rohmer, M., The discovery of a mevalonate-independent pathway for isoprenoid biosynthesis in bacteria, algae and higher plants, Nat. Prod. Rep., 1999, vol. 16, pp. 565–574.
Janthawornpong, K., Krasutsky, S., Chaignon, P., Rohmer, M., Poulter, C.D., and Seemann, M., Inhibition of IspH, a [4Fe–4S]2+ enzyme involved in the biosynthesis of isoprenoids via the methylerythritol phosphate pathway, J. Am. Chem. Soc., 2013, vol. 135, pp. 1816–1822.
Rodríguez-Concepción, M., The MEP pathway: a new target for the development of herbicides, antibiotics and antimalarial drugs, Curr. Pharm. Des., 2004, vol. 10, pp. 2391–2400.
Steinbacher, S., Kaiser, J., Eisenreich, W., Huber, R., Bacher, A., and Rohdich, F., Structural basis of fosmidomycin action revealed by the complex with 2-C-methyl-D-erythritol 4-phosphate synthase (IspC). Implications for the catalytic mechanism and anti-malaria drug development, J. Biol. Chem., 2003, vol. 278, pp. 18401–18407.
Wiesner, J., Reichenberg, A., Hintz, M., Ortmann, R., Schlitzer, M., van Calenbergh, S., Borrmann, S., Lell, B., Kremsner, P.G., and Hutchinson, D., Fosmidomycin as an antimalarial agent, Isoprenoid Synthesis in Plants and Microorganisms, Bach, T.J. and Rohmer, M., Eds., New York: Springer-Verlag, 2013, pp. 119–137.
Vigani, G., Zocchi, G., Bashir, K., Philippar, K., and Briat, J.-F., Signals from chloroplasts and mitochondria for iron homeostasis regulation, Trends Plant Sci., 2013, vol. 18, pp. 305–311.
Xiao, Y., Savchenko, T., Baidoo, E.E., Chehab, W.E., Hayden, D.M., Tolstikov, V., Corwin, J.A., Kliebenstein, D.J., Keasling, J.D., and Dehesh, K., Retrograde signaling by the plastidial metabolite MEcPP regulates expression of nuclear stress-response genes, Cell, 2012, vol. 149, pp. 1525–1535.
Smyth, G., Limma: Linear Models for Microarray Data, Bioinformatics and Computational Biology Solutions Using R and Bioconductor, Genleman, R., Carey, V.J., Huber, W., Irizarry, R.A., and Dudoit, S., Eds., New York: Springer-Verlag, 2005, pp. 397–420.
Huang, D.W., Sherman, B.T., and Lempicki, R.A., Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources, Nat. Protoc., 2008, vol. 4, pp. 44–57.
Szklarczyk, D., Franceschini, A., Kuhn, M., Simonovic, M., Roth, A., Minguez, P., Doerks, T., Stark, M., Muller, J., and Bork, P., The STRING database in 2011: functional interaction networks of proteins, globally integrated and scored, Nucleic Acids Res., 2011, vol. 39: D561–D568.
Shannon, P., Markiel, A., Ozier, O., Baliga, N.S., Wang, J.T., Ramage, D., Amin, N., Schwikowski, B., and Ideker, T., Cytoscape: a software environment for integrated models of biomolecular interaction networks, Genome Res., 2003, vol. 13, pp. 2498–2504.
Paseshnichenko, V.A., Regulation of terpenoid biosynthesis in plants and its relation to the biosynthesis of phenolic-compounds, Russ. J. Plant Physiol., 1995, vol. 42, pp. 699–714.
Van der Werf, M.J. and Boot, A.M., Metabolism of carveol and dihydrocarveol in Rhodococcus erythropolis DCL14, Microbiology, 2000, vol. 146, pp. 1129–1141.
Drašar, P., Isoprenoids, Chem. Int., 2015, vol. 37, p. 29.
Rodríguez-Concepción, M., Plant isoprenoids: a general overview, Plant Isoprenoids, vol. 1153, Ser. Methods in Molecular Biology, Rodríguez-Concepción, M., Ed., New York: Springer-Verlag, 2014, pp. 1–5.
Dar, T.A., Moinuddin, N.H., Idrees, M., and Ali, A., Recent trends in jasmonate signaling pathway, in Plant Signaling: Understanding the Molecular Crosstalk, Hakeem, K.R., Rehman, R.U., and Tahir, I., Eds., New Delhi, India: Springer-Verlag, 2013, pp. 277–290.
Woodson, J.D. and Chory, J., Organelle signaling: how stressed chloroplasts communicate with the nucleus, Curr. Biol., 2012, vol. 22, pp. R690–R692.
Moore, J.W., Foundation technologies in synthetic biology: tools for use in understanding plant immunity, PhD Thesis, Edinburgh: University of Edinburgh, 2012.
Koornneef, A. and Pieterse, C.M., Cross talk in defense signaling, Plant Physiol., 2008, vol. 146, pp. 839–844.
Wagener, N. and Neupert, W., Bcs1, a AAA protein of the mitochondria with a role in the biogenesis of the respiratory chain, J. Struct. Biol., 2012, vol. 179, pp. 121–125.
Ostojic, J., Panozzo, C., Lasserre, J.-P., Nouet, C., Courtin, F., Blancard, C., Di Rago, J.-P., and Dujardin, G., The energetic state of mitochondria modulates complex III biogenesis through the ATP-dependent activity of Bcs1, Cell Metab., 2013, vol. 18, pp. 567–577.
Eckardt, N.A., A plastidial pathway for protein isoprenylation in tobacco cells, Plant Cell, 2009, vol. 21, p. 13.
Hirt, H., Multiple roles of MAP kinases in plant signal transduction, Trends Plant Sci., 1997, vol. 2, pp. 11–15.
Author information
Authors and Affiliations
Corresponding author
Additional information
The article is published in the original.
Rights and permissions
About this article
Cite this article
Lang, C., Xi, J. Bioinformatics identification of the methylerythritol phosphate pathway associated genes in Arabidopsis thaliana with ceh1 mutant. Russ J Plant Physiol 63, 293–299 (2016). https://doi.org/10.1134/S1021443716020096
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1021443716020096