Abstract
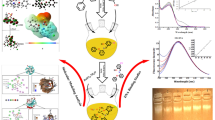
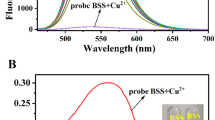
A new mononuclear copper complex [Cu(dpq)(L–Met)Cl] ⋅ H2O has been designed and synthesized using the phenanthroline derivative dipyridoquinoxaline (dpq). This complex has been characterized by element analysis and X-ray single crystal diffraction. The crystal structure of the complex belongs to a monoclinic system with space group C2/c. Binding of the complex to DNA has been investigated by absorption spectroscopy, fluorescence spectroscopy, and agarose gel electrophoresis. Spectrophotometric studies have showed that the mode of binding is intercalation with a binding constant Kb of 1.25 × 103 L mol–1. Additionally, the complex could quench an ethidium bromide-DNA complex. The apparent binding constant value for the complex is 1.89 × 10 5 L mol–1, which is slightly lower than the classical binding constant of 107 L mol–1. In addition, the quenching mechanism of the complex has been not static or dynamic. Agarose gel electrophoresis has shown that the complex cleaved circular pBR322 plasmid DNA to both nicked and linear forms; at higher concentrations and longer reaction times, the cutting results have been improved. The cleavage mechanism between the complex and plasmid DNA may involve singlet oxygen and hydroxyl radical as reactive oxygen species.






Similar content being viewed by others
REFERENCES
S. M. Zhang, H. Y. Zhang, Q. P. Qin, et al., J. Inorg. Biochem. 193, 52 (2019). https://doi.org/10.1016/j.jinorgbio.2019.01.009
S. A. Patil, C. T. Prabhakara, B. M. Halasangi, et al., Spectrochim. Acta A 137, 641 (2015). https://doi.org/10.1016/j.saa.2014.08.028
Q. P. Huang, S. N. Zhang, S. H. Zhang, et al., Molecules 22, 1813, 1 (2017). https://doi.org/10.3390/molecules22111813
N. Vamsikrishna, M. P. Kumar, G. Ramesh, et al., J. Chem. Sci. 129, 609 (2017). https://doi.org/10.1007/s12039-017-1273-7
L. F. Tan, J. L. Shen, X. J. Chen, et al., DNA Cell Biol. 28, 461 (2009). https://doi.org/10.1089/dna.2009.0889
Q. L. Zhang, J. H. Liu, X. Z. Ren, et al., Chin. J. Inorg. Chem. 19, 645 (2003). https://doi.org/10.3321/j.issn:1001-4861.2003.06.019
J. Kalyanmoy, D. Somnath, P. Horst, et al., Inorg. Chim. Acta 487, 128 (2019). https://doi.org/10.1016/j.ica.2018.12.007
X. L. Liang, X. Q. Zou, L. F. TAN, et al., J. Inorg. Biochem. 104, 1259 (2010). https://doi.org/10.1016/j.jinorgbio.2010.08.006
U. P. Sidhali, F. Joseph, B. Arnab, et al., J. Bio. Inorg. Chem. 23, 1331 (2018). https://doi.org/10.1007/s00775-018-1620-2
X. L. Liang, L. F. Tan, W. G. Zhu, DNA Cell Biol. 30, 61 (2011). https://doi.org/10.1089/dna.2010.1079
R. E. H. M. B. Osório, A. Neves, T. P. Camargo, et al., Inorg. Chim. Acta 435, 153 (2015). https://doi.org/10.1016/j.ica.2015.06.023
T. P. Camargo, R. A. Peralta, R. Moreira, et al., Inorg. Chem. Commun. 37, 34 (2013). https://doi.org/10.1016/j.inoche.2013.09.039
R. Bushtit, Z. Trávnícěk, J. Vancǒ, J. Inorg. Biochem. 116, 163 (2012). https://doi.org/10.1016/j.jinorgbio.2012.07.009
R. Novotná, R. Herchel, Z. Trávnícěk, Polyhedron 34, 56 (2012). https://doi.org/10.1016/j.poly.2011.12.016
B. P. Mateus, A. F. Liniquer, D. S. Josiéli, et al., Inorg. Chim. Acta 469, 561 (2018). https://doi.org/10.1016/j.ica.2017.09.063
S. Tabassum, S. Amir, F. Arjmand, et al., Eur. J. Med. Chem. 60, 216 (2013). https://doi.org/10.1016/j.ejmech.2012.08.019
Z. F. Chen, Y. X. Wu, Z. Z. Zhu, et al., Biometals 32, 227 (2019). https://doi.org/10.1007/s10534-019-00172-w
S. Zahra, C. Hossein, A. M. Amir, et al., J. Photochem. Photobiol. B: Biol. 162, 34 (2016). https://doi.org/10.1016/j.jphotobiol.2016.06.022
C. Madhuri, T. Deepak, C. Sulekh, J. Mol. Struct. 1179, 431 (2019). https://doi.org/10.1016/j.molstruc.2018.11.027
L. P. Lu, M. L. Zhu, P. Yang, J. Inorg. Biochem. 95, 31 (2003). https://doi.org/10.1016/S0162-0134(03)00049-7
S. Dhar, D. Senapati, P. K. Das, et al., J. Am. Chem. Soc. 125, 12 118 (2003). https://doi.org/10.1021/ja036681q
S. Ramakrishnan and M. Palaniandavar, J. Chem. Sci. 117, 179 (2005). https://doi.org/10.1007/bf03356114
R. Venugopal, K. Ramasamy, P. Mallayan, et al., Inorg. Chem. 46, 8208 (2007). https://doi.org/10.1021/ic700755p
X. F. Li, X. Q. Feng, H. Chang, et al., J. Mater. Sci. Eng. 35, 129 (2017).https://doi.org/10.14136/j.cnki.issn:1673-2812.2017.01.026
J. Q. Tao, W. Z. Shi, X. Zhuang, et al., Chin. J. Inorg. Chem. 18, 255 (2002). https://doi.org/10.3321/j.issn:1001-4861.2002.03.006
G. M. Sheldrick, Correction Software (Univ. of Göttingen, Göttingen, 1996).
G. M. Sheldrick, SHELXS 97: Program for the Solution of Crystal Structures (Univ. of Göttingen, Göttingen, 1997).
G. M. Sheldrick, SHELXL 97, Program for the Refinement of Crystal Structures, (Univ. of Göttingen, Göttingen, 1997).
J. Qian, W. Gu, H. Liu, et al., Dalton. Trans. 10, 1060 (2007). https://doi.org/10.1039/B615148E
C. Tu, Y. Shao, N. Gan, et al., Inorg. Chem. 43, 4761 (2004). https://doi.org/10.1021/ic049731g
S. A. Tysoe, A. D. Baker, and T. C. Strekas, J. Phys. Chem. 97, 1707 (1993). https://doi.org/10.1021/j100110a038
S. M. Yue, X. R. Yue, and Y. N. Chen, Synth. Met. 200, 1 (2015). https://doi.org/10.1016/j.synthmet.2014.12.022
S. Sharma, P. K. Sharma, N. Kumar, et al., Biomed. Pharmacother.65(4), 244 (2011). https://doi.org/10.1016/j.biopha.2011.04.005
A. Wolfe, G. H. Shimer, and T. Meehan, Biochem. 26, 6392 (1987). https://doi.org/10.1021/bi00394a013
J. R. Lakowicz and G. Webber, Biochem. 12, 4161 (1973). https://doi.org/10.1021/bi00745a020
M. Cory, D. D. Mckee, J. Kagan, et al., J. Am. Chem. Soc. 107, 2528 (1985). https://doi.org/10.1021/ja00294a054
M. Balón, M. A. Muñoz, C. Carmona, et al., Biophys. Chem. 80, 41 (1999). https://doi.org/10.1016/S0301-4622(99)00059-9
R. T. Huang, Z. D. Li, Q. Zhao, et al., Chem. Res. App. 28, 19 (2016). https://doi.org/10.3969/j.issn.1004-1656.2016.01.004
Y. Y. Kou, M. L. Li, and X. H. Ren, Spectrochim. Acta A 205, 435 (2018). https://doi.org/10.1016/j/saa.2018.07.050
Y. Y. Kou and X. H. Ren, Chin. J. Inorg. Chem. 33(8), 1429 (2017). https://doi.org/10.11862/CJIC.2017.157
Funding
This work was supported by the General Plan of Science and Technology Plan of Beijing Education Commission (no. KM201610016006) and the Fundamental Research Funds for Beijing University of Civil Engineering and Architecture (no. X18131).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
No potential conflict of interest was reported by the authors.
Rights and permissions
About this article
Cite this article
Ying-Ying Kou, Zhao, Q. & Wang, XR. Synthesis, Structure, and Chemical Nuclease Activity of DNA with [Cu(dpq)(L–Met)Cl] · H2O. Russ. J. Inorg. Chem. 64, 1762–1768 (2019). https://doi.org/10.1134/S0036023619140134
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0036023619140134