Abstract
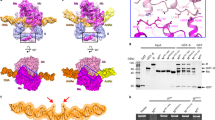
DNA mimicking ArdA anti-restriction proteins specifically inhibit restriction (endonuclease) activity of the type I restriction-modification (RM) system. An ArdA monomer is comprised of three α-β domains (the N-domain, Central domain, and C-domain), each with a different fold. Here we describe an alignment of the amino acid (a.a.) sequences of the ArdA with a conserved 20-a.a. motif in the N domain. The N domains of ArdA proteins of the Gram-positive bacteria Arthrobacter sp. and Bifidobacterium longum, and the Gram-negative bacteria Pseudomonas plecoglossicida are capable of inhibiting the repressive activity of the H-NS global silencer protein in Escherichia coli cells. The presence of the H-NS inhibiting N domain in the ArdA structure enables horizontal gene transfer by mobile elements, including conjugative plasmids and transposons. Specifically, it aids in overcoming intercellular restriction barriers, allowing faster adaption to the genome context of the recipient bacterium.





Similar content being viewed by others
REFERENCES
Belogurov A.A., Yussifov T.N., Kotova V.Yu., Zavilgelsky G.B. 1985. The novel gene(s) (ard) of plasmid pKM101: Alleviation of EcoKI restriction. Mol. Gen. Genet. 198, 509–513.
Delver E.P., Kotova V.Yu., Zavilgelsky G.B., Belo-gurov A.A. 1991. Nucleotide sequence of the gene (ard) encoding the antirestriction protein of plasmid ColIb-P9. J. Bacteriol. 173 (18), 5887–5892.
Chilley P.M., Wilkins B.M. 1995. Distribution of the ardA family of antirestriction genes on conjugative plasmids. Microbiology. 141, 2157–2164.
Serfiotis-Mitsa D., Roberts G.A., Cooper L.P., White J.H., Nutley M., Cooper A., Blakely G.W., Dryden D.T.F. 2008. The Orf18 gene product from conjugative transposon Tn916 is an ArdA antirestriction protein that inhibits type I DNA restriction–modification systems. J. Mol. Biol. 383, 970–981.
McMahon S.A., Roberts G.A., Johnson K.A., Cooper L.P., Liu H., White J.H., Carter L.G., Sanghvi B., Oke M., Walkinshaw M.D., Blakely G.W., Naismith J.H., Dryden D.T.F. 2009. Extensive DNA mimicry by the ArdA anti-restriction protein and its role in the spread of antibiotic resistance. Nucleic Acids Res. 37, 4887–4897.
Belogurov A.A., Delver E.P. 1995. A motif conserved among the type I restriction-modification enzymes and antirestriction proteins: A possible basis for mechanism of action of plasmid-encoded anti-restriction functions. Nucleic Acids Res. 23, 785–787.
Shen Y., Volrath S.L., Weatherly S.C., Elich T.D., Tong L. 2004. A mechanism for the potent inhibition of eukaryotic acetyl-coenzyme A carboxylase by soraphen A, a macrocyclic polyketide natural product. Mol. Cell. 16, 881–891.
Nekrasov S.V., Agafonova O.V., Belogurova N.G., Delver E.P., Belogurov A.A. 2007. Plasmid-encoded antirestriction protein ArdA can discriminate between type I methyltransferase and complete restriction-modification system. J. Mol. Biol. 365, 281–297.
Zavilgelsky G.B., Kotova V.Yu., Rastorguev S.M. 2008. Comparative analysis of antirestriction activities of ArdA (ColIb-P9) and Ocr (T7) proteins. Biochemistry (Moscow). 73 (8), 906–911.
Walkinshaw M.D., Tylor P., Sturrock S.S., Atanasiu C., Berge T., Henderson R.M., Edwardson J.M., Dryden D.T.F. 2002. Structure of Ocr from bacteriophage T7, a protein that mimics B-form of DNA. Mol. Cell. 9, 187–194.
Melkina O.E., Goryanin I.I., Zavilgelsky G.B. 2016. The DNA-mimic antirestriction proteins ArdA ColIb-P9, Arn T4, and Ocr T7 as activators of H-NS-dependent gene transcription. Microbiol. Res. 192, 283–291.
Robin S., Togashi D., Ryder A.G., Wall J.G. 2009. Trigger factor from psychrophilic bacterium Psychrobacter frigidicola is a monomeric chaperone. J. Bacteriol. 191, 1162–1169.
Ulitzur S., Matin A., Fraley C., Meighen E. 1997. H‑NS protein represses transcription of the lux systems of Vibrio fischeri and other luminous bacteria cloned into Escherichia coli. Curr. Microbiol. 35, 336–342.
Ulitzur S. 1998. H-NS controls the transcription of three promoters of Vibrio fischeri lux cloned in Escherichia coli. J. Biolumin. Chemilumin. 13, 185–188.
Van Dyk T., Rosson R.A. 1998. Photorhabdus luminescens luxCDABE promoter probe vectors. In: Methods in Molecular Biology. Ed. LaRossa R.A. Totowa, NY: Humana Press, 102, 85–95.
Sambrook J., Fritsch E.F., Maniatis T. 1989. Molecular Cloning: A Laboratory Manual, 2nd ed. New York: Cold Spring Harbor Lab. Press.
Liu Q., Richardson C.C. 1993. Gene 5.5 protein of bacteriophage T7 inhibits the nucleoid protein H-NS of Escherichia coli. Proc. Natl. Acad. Sci. U. S. A. 90, 1761–1765.
Ali S.S., Beckett E., Bac S.J., Nawarre W.W. 2011. The 5.5 protein of phage T7 inhibits H-NS through interactions with the central oligomerization domain. J. Bacteriol. 193, 4881–4892.
Redgwell R.T., Michniewski S., Harrison D.C., Millard A. 2020. GenBank CAA6800530. University of Warwick, Coventry, UK.
Zhu B., Lee S.-J., Tan M., Wang E.-D., Richardson C.C. 2012. Gene 5.5 of bacteriophage T7 in complex with Escherichia coli nucleoid protein H-NS and transfer RNA masks transfer RNA priming in T7 replication. Proc. Natl. Acad. Sci. U. S. A. 109, 8050–8059.
Dorman C.J. 2007. H-NS, the genome sentinel. Nat. Rev. Microbiol. 5, 157–161.
Dillon S.C., Dorman C.J. 2010. Bacterial nucleoid-associated proteins, nucleoid structure and gene expression. Nat. Rev. Microbiol. 8, 185–195.
ACKNOWLEDGMENTS
The authors are grateful to the staff of the NRC Kurchatov Institute, Sergei Rastorguev and Aleksei Kozhenkov, for their assistance in genome-wide sequencing of bacteria and the search for ardA nucleotide gene sequences.
Funding
This work was supported by the Russian Foundation for Basic Research (project no. 19-04-00495).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare they have no conflict of interest. The study contains no research using humans or animals as objects of study.
Additional information
Abbreviations: a.a., amino acid residue (with number).
Rights and permissions
About this article
Cite this article
Melkina, O.E., Zavilgelsky, G.B. N-Domain of ArdA Antirestriction Proteins Inhibits the Repression Activity of the Histone-Like H-NS Protein. Mol Biol 55, 424–431 (2021). https://doi.org/10.1134/S0026893321020266
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0026893321020266