Abstract
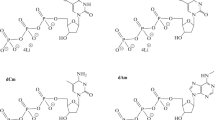
The effects of modified deoxyuridine triphosphates (mod-dUTPs) with different substituents at the C5 position of the pyrimidine cycle on the kinetics of PCR with Taq and Vent (exo-) DNA polymerases are studied. Substituents in mod-dUTP include carboxamide group and groups that are part of the side chains of alanine, valine, leucine, phenylalanine, tryptophan, or tyrosine. For each mod-dUTP, the yields of the target product are measured with the full substitution of dTTP. A fragment of bacterial DNA with a certain nucleotide sequence and a synthetic combinatorial DNA library of random nucleotide sequences are used as templates for amplification. For each mod-dUTP–template–polymerase combination, the correlation between the amplification efficiencies and yields of the target product are investigated. PCR product accumulation curves are influenced by both the template used and the presence of a modified substrate. The catalytic activity of Taq polymerase is higher when mod-dUTPs with short aliphatic substituents are used and decreases when the derivatives with long aliphatic, phenyl, and indole substituents are utilized. Vent (exo-) polymerase is less sensitive to the chemical structure of mod-dUTP. The dynamic measuring of DNA accumulation may be useful for optimizing the temperature–time PCR profiles individually for each of the mod-dUTP. The derivatives may be used in combination with Vent (exo-) polymerase to obtain modified DNA sequences for the method of selection of modified aptamers (mod-SELEX).





Similar content being viewed by others
REFERENCES
Zhuo Z., Yu Y., Wang M., Li J., Zhang Z., Liu J., Wu X., Lu A., Zhang G., Zhang B. 2017. Recent advances in SELEX technology and aptamer applications in biomedicine. Int. J. Mol. Sci. 18, e2142.
Lapa S.A., Chudinov A.V., Timofeev E.N. 2016. The toolbox for modified aptamers. Mol. Biotechnol. 58, 79–92.
Gold L., Ayers D., Bertino J., Bock C., Bock A., Brody E.N., Carter J., Dalby A.B., Eaton B.E., Fitzwater T., Flather D., Forbes A., Foreman T., Fowler C., Gawande B., et al. 2010. Aptamer-based multiplexed proteomic technology for biomarker discovery. PLoS One. 5, e15004.
Gold L. 2015. SELEX: How it happened and where it will go. J. Mol. Evol. 81, 140–143.
Latham J.A., Johnson R., Toole J.J. 1994. The application of a modified nucleotide in aptamer selection: Novel thrombin aptamers containing 5-(1-pentynyl)-2'-deoxyuridine. Nucleic Acids Res. 22, 2817–2822.
Davies D.R., Gelinas A.D., ZhangC., Rohloff J.C., Carter J.D., O’Connell D., Waugh S.M., Wolk S.K., Mayfield W.S., Burgin A.B., Edwards T.E., Stewart L.J., Gold L., Janjic N., Jarvis T.C. 2012. Unique motifs and hydrophobic interactions shape the binding of modified DNA ligands to protein targets. Proc. Natl. Acad. Sci. U. S. A. 109, 19971–19976.
Kopylov A.M., Spiridonova V.A. 2000. Combinatorial chemistry of nucleic acids: SELEX. Mol. Biol. (Moscow). 34 (6), 940–955.
Kuwahara M., Hanawa K,Ohsawa K, Kitagata R, Ozaki H, Sawai H. 2006. Direct PCR amplification of various modified DNAs having amino acids: convenient preparation of DNA libraries with high-potential activities for in vitro selection. Bioorg. Med. Chem. 14, 2518–2526.
Masud M.M., Kuwahara M., Ozaki H., Sawai H. 2004. Sialyllactose-binding modified DNA aptamer bearing additional functionality by SELEX. Bioorg. Med. Chem. 12, 1111–1120.
Lapa S.A., Romashova K.S., Spitsyn M.A., Shershov V.E., Kuznetsova V.E., Guseinov T.O., Zasedateleva O.A., Radko S.P., Timofeev E.N., Lisitsa A.V., Chudinov A.V. 2018. Preparation of modified combinatorial DNA libraries via emulsion PCR with subsequent strand separation. Mol. Biol. (Moscow). 52 (6), 854–864.
Vaught J.D., Bock C., Carter J., Fitzwater T., Otis M., Schneider D., Rolando J., Vaugh S., Wilcox S.K., Eaton B.E. 2010. Expanding the chemistry of DNA for in vitro selection. J. Am. Chem. Soc. 132, 4141–4151.
Tolle F., Wilke J., Wengel J., Mayer G. 2014. By-product formation in repetitive PCR amplification of DNA libraries during SELEX. PLoS One. 9, e114693.
Chudinov A.V., Kiseleva Ya.Yu., Kuznetsova V.E., Shershov V.E., Spitsyn M.A., Guseinov T.O., Lapa S.A., Timofeev E.N., Archakov A.I., Lisitsa A.V., Radko S.P., Zasedatelev A.S. 2017. Structural and functional analysis of biopolymers and their complexes: Enzymatic synthesis of high-modified DNA. Mol. Biol. (Moscow). 51, 534–544.
Mikhailovich V., Lapa S., Gryadunov D., Sobolev A., Strizhkov B., Chernyh N., Skotnikova O., Irtuganova O., Moroz A., Litvinov V., Vladimirskii M., Perelman M., Chernousova L., Erokhin V., Zasedatelev A., Mirzabekov A. 2001. Identification of rifampin-resistant Mycobacterium tuberculosis strains by hybridization, PCR, and ligase detection reaction on oligonucleotide microchips. J. Clin. Microbiol. 39, 2531–2540.
Gryadunov D., Mikhailovich V., Lapa S., Roudinskii N., Donnikov M., Pan’kov S., Markova O., Zasedatelev A., Mirzabekov A., Kuz’min A., Chernousova L., Skotnikova O., Moroz A. 2005. Evaluation of hybridisation on oligonucleotide microarrays for analysis of drug-resistant Mycobacterium tuberculosis. Clin. Microbiol. Infect. 11, 531–539.
Ramakers C., Ruijter J.M., Deprez R.H., Moorman A.F. 2003. Assumption-free analysis of quantitative real-time polymerase chain reaction (PCR) data. Neurosci. Lett. 339, 62–66.
Peirson S.N., Butler J.N., Foster R.G. 2003. Experimental validation of novel and conventional approaches to quantitative real-time PCR data analysis. Nucleic Acids Res. 31, e73.
Liu W., Saint D.A. 2002. A new quantitative method of real time reverse transcription polymerase chain reaction assay based on simulation of polymerase chain reaction kinetics. Anal. Biochem. 302, 52–59.
Ahmed M., Kim D.R. 2018. PCR: An R package for quality assessment, analysis and testing of qPCR data. Peer J. 6, e4473.
Lapa S.A., Volkova O.S., Spitsyn M.A., Shershov V.E., Kuznetsova V.E., Guseinov T.O., Zasedatelev A.S., Chudinov A.V. 2019. Amplification efficiency and substrate properties of fluorescent-labeled deoxyuridine triphosphates in PCR with DNA polymerases devoid of 3′-5′ exonuclease activity. Russ. J. Bioorg. Chem. 45 (in press).
Ioannou A.K., Alexiadou D.K., Kouidou S.A., Voulgaropoulos A.N., Girousi S.T. 2010. Electroanalytical study of SYBR Green I and ethidium bromide intercalation in methylated and unmethylated amplicons. Anal. Chim. Acta. 657, 163–168.
Mao F., Leung W.Y., Xin X. 2007. Characterization of EvaGreen and the implication of its physicochemical properties for qPCR applications. BMC Biotechnol. 7, 76.
Khan S.A., Sung K., Nawaz M.S.(2011. Detection of aacA-aphD, qacEδ1, marA, floR, and tetA genes from multidrug-resistant bacteria: Comparative analysis of real-time multiplex PCR assays using EvaGreen® and SYBR® Green I dyes. Mol. Cell. Probes. 25, 78–86.
Author information
Authors and Affiliations
Corresponding author
Additional information
Translated by N. Onishchenko
Rights and permissions
About this article
Cite this article
Lapa, S.A., Pavlov, A.S., Kuznetsova, V.E. et al. Enzymatic Preparation of Modified DNA: Study of the Kinetics by Real-Time PCR. Mol Biol 53, 460–469 (2019). https://doi.org/10.1134/S0026893319030099
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0026893319030099