Abstract
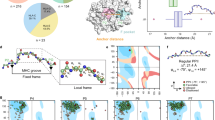
The coupling between peptides and MHC-II proteins in the human immune system is not well understood. This work presents an evidence-based hypothesis of a guiding intermolecular force present in every human MHC-II protein (HLA-II). Previously, we examined the spatial positions of the fully conserved residues in all HLA-II protein types. In each one, constant planar patterns were revealed. These molecular planes comprise of amino acid groups of the same chemical species (for example, Gly) distributed across the protein structure. Each amino acid plane has a unique direction and this directional element offers spatial selectivity. Constant within all planes, too, is the presence of an aromatic residue possessing electrons in movement, leading the authors to consider that the planes generate electromagnetic fields that could serve as an attractive force in a single direction. Selection and attraction between HLA-II molecules and antigen peptides would, therefore, be non-random, resulting in a coupling mechanism as effective and rapid as is clearly required in the immune response. On the basis of planar projections onto the HLA-II groove, modifications were made by substituting the key residues in the class II-associated invariant chain peptide—a peptide with a universal binding affinity—resulting in eight different modified peptides with affinities greater than that of the unmodified peptide. Accurate and reliable prediction of MHC class II-binding peptides may facilitate the design of universal vaccine-peptides with greatly enhanced binding affinities. The proposed mechanisms of selection, attraction and coupling between HLA-II and antigen peptides are explained further in the paper.
Similar content being viewed by others
References
Reinherz E.L., Tan K., Tang L., Kern P., Liu J.-h., Xiong Y., Hussey R.E., Smolyar A., Hare B., Zhang R. 1999. The crystal structure of a T cell receptor in complex with peptide and MHC class II. Science. 286, 1913–1921.
Chicz R.M., Urban R.G., Gorga J.C., Vignali D., Lane W.S., Strominger J.L. 1993. Specificity and promiscuity among naturally processed peptides bound to HLA-DR alleles. J. Exp. Med. 178, 27–47.
Jones E.Y., Fugger L., Strominger J.L., Siebold C. 2006. MHC class II proteins and disease: A structural perspective. Nat. Rev. Immunol. 6, 271–282.
Brown J.H., Jardetzky T.S., Gorga J.C., Stern L.J., Urban R.G., Strominger J.L., Wiley D.C. 1993. Threedimensional structure of the human class II histocompatibility antigen HLA-DR1. Nature. 364, 33–39.
Leckband D. 2000. Measuring the forces that control protein interactions. Annu. Rev. Bioph. Biom. 29, 1–26.
Beretta S., Chirico G., Baldini G. 2000. Short-range interactions of globular proteins at high ionic strengths. Macromolecules. 33, 8663–8670.
Bischof M., Del Giudice E. 2013. Communication and the emergence of collective behavior in living organisms: A quantum approach. Mol. Biol. Int. 2013, 987549. doi 10.1155/2013/987549
Melkikh A.V. 2013. Biological complexity, quantum coherent states and the problem of efficient transmission of information inside a cell. Biosystems. 111, 190–198.
Ćosić I., Pirogova E., Vojisavljević V., Fang Q. 2006. Electromagnetic properties of biomolecules. FME Trans. 34, 71–80.
Pirogova E., Cosic I., Vojisavljevic V. 2009. Biological effects of electromagnetic radiation. In: Biomedical Engineering. Ed. Barros de Mello C.A.InTechOpen, pp. 87–106. doi 10.5772/787810.5772/7878
Vojisavljevic V., Pirogova E., Cosic I. 2007. Influence of electromagnetic radiation on enzyme kinetics. Conf. Proc. IEEE Eng. Med. Biol. Soc. 2007, pp. 5021–5024.
Vojisavljevic V., Pirogova E., Cosic I. 2010. Review of studies on modulating enzyme activity by low intensity electromagnetic radiation. Conf. Proc. IEEE Eng. Med. Biol. Soc. 2010, pp. 835–838. doi 10.1109/IEMBS.2010.5626786
Cortés A., Coral J., Benítez Benítez R. 2013. Hallazgo de patrones para péptidos-vacuna con capacidad de acople universal en moléculas de HLA-II. Acta Bioquim. Clin. L. 47, 541–549.
Cortes A., Coral J. 2015. Campos electromagneticos planares permiten explicar el acople entre peptidos y moleculas de HLA-II. Acta Bioquim. Clin. L. 49, 221–228.
Wiesner M., Stepniak D., de Ru A.H., Moustakis A.K., Drijfhout J.W., Papadopoulos G.K., van Veelen P.A., Koning F. 2008. Dominance of an alternative CLIP sequence in the celiac disease associated HLA-DQ2 molecule. Immunogenetics. 60, 551–555.
Neefjes J., Jongsma M.L., Paul P., Bakke O. 2011. Towards a systems understanding of MHC class I and MHC class II antigen presentation. Nat. Rev. Immunol. 11, 823–836.
Bordner A., Mittelmann H. 2010. Prediction of the binding affinities of peptides to class II MHC using a regularized thermodynamic model. BMC Bioinform. 11, 41.
Bordner A., Mittelmann H. 2010. MultiRTA: A simple yet reliable method for predicting peptide binding affinities for multiple class II MHC allotypes. BMC Bioinform. 11, 482.
Painter C.A., Negroni M.P., Kellersberger K.A., Zavala-Ruiz Z., Evans J.E., Stern L.J. 2011. Conformational lability in the class II MHC 310 helix and adjacent extended strand dictate HLA-DM susceptibility and peptide exchange. Proc. Natl. Acad. Sci. U.S. A. 108, 19329–19334.
Dai S., Murphy G.A., Crawford F., Mack D.G., Falta M.T., Marrack P., Kappler J.W., Fontenot A.P. 2010. Crystal structure of HLA-DP2 and implications for chronic beryllium disease. Proc. Natl. Acad. Sci. U.S. A. 107, 7425–7430.
Hennecke J., Carfi A., Wiley D.C. 2000. Structure of a covalently stabilized complex of a human αβ T-cell receptor, influenza HA peptide and MHC class II molecule, HLA-DR1. EMBO J. 19, 5611–5624.
Yin Y., Li Y., Kerzic M.C., Martin R., Mariuzza R.A. 2011. Structure of a TCR with high affinity for self-antigen reveals basis for escape from negative selection. EMBO J. 30, 1137–1148.
Parry C.S., Gorski J., Stern L.J. 2007. Crystallographic structure of the human leukocyte antigen DRA, DRB3* 0101: Models of a directional alloimmune response and autoimmunity. J. Mol. Biol. 371, 435–446.
Kim C.-Y., Quarsten H., Bergseng E., Khosla C., Sollid L.M. 2004. Structural basis for HLA-DQ2- mediated presentation of gluten epitopes in celiac disease. Proc. Natl. Acad. Sci. U.S. A. 101, 4175–4179.
Tollefsen S., Hotta K., Chen X., Simonsen B., Swaminathan K., Mathews I.I., Sollid L.M., Kim C.-Y. 2012. Structural and functional studies of trans-encoded HLA-DQ2.3 (DQA1*03:01/DQB1*02:01) protein molecule. J. Biol. Chem. 287, 13611–13619.
Li Y., Huang Y., Lue J., Quandt J.A., Martin R., Mariuzza R.A. 2005. Structure of a human autoimmune TCR bound to a myelin basic protein self-peptide and a multiple sclerosis-associated MHC class II molecule. EMBO J. 24, 2968–2979.
Deng L., Langley R.J., Brown P.H., Xu G., Teng L., Wang Q., Gonzales M.I., Callender G.G., Nishimura M.I., Topalian S.L. 2007. Structural basis for the recognition of mutant self by a tumor-specific, MHC class II–restricted T cell receptor. Nat. Immunol. 8, 398–408.
Harkiolaki M., Holmes S.L., Svendsen P., Gregersen J.W., Jensen L.T., McMahon R., Friese M.A., Van Boxel G., Etzensperger R., Tzartos J.S. 2009. T cell-mediated autoimmune disease due to low-affinity crossreactivity to common microbial peptides. Immunity. 30, 348–357.
Sethi D.K., Schubert D.A., Anders A.-K., Heroux A., Bonsor D.A., Thomas C.P., Sundberg E.J., Pyrdol J., Wucherpfennig K.W. 2011. A highly tilted binding mode by a self-reactive T cell receptor results in altered engagement of peptide and MHC. J. Exp. Med. 208, 91–102.
Yin Y., Wang X.X., Mariuzza R.A. 2012. Crystal structure of a complete ternary complex of T-cell receptor, peptide–MHC, and CD4. Proc. Natl. Acad. Sci. U.S. A. 109, 5405–5410.
Siebold C., Hansen B.E., Wyer J.R., Harlos K., Esnouf R.E., Svejgaard A., Bell J.I., Strominger J.L., Jones E.Y., Fugger L. 2004. Crystal structure of HLADQ0602 that protects against type 1 diabetes and confers strong susceptibility to narcolepsy. Proc. Natl. Acad. Sci. U.S. A. 101, 1999–2004.
Broughton S.E., Petersen J., Theodossis A., Scally S.W., Loh K.L., Thompson A., van Bergen J., Kooy-Winkelaar Y., Henderson K.N., Beddoe T. 2012. Biased T cell receptor usage directed against human leukocyte antigen DQ8-restricted gliadin peptides is associated with celiac disease. Immunity. 37, 611–621.
Murthy V.L., Stern L.J. 1997. The class II MHC protein HLA-DR1 in complex with an endogenous peptide: Implications for the structural basis of the specificity of peptide binding. Structure. 5, 1385–1396.
Li Y., Depontieu F.R., Sidney J., Salay T.M., Engelhard V.H., Hunt D.F., Sette A., Topalian S.L., Mariuzza R.A. 2010. Structural basis for the presentation of tumor-associated MHC class II-restricted phosphopeptides to CD4+ T cells. J. Mol. Biol. 399, 596–603.
Yin L., Crawford F., Marrack P., Kappler J.W., Dai S. 2012. T-cell receptor (TCR) interaction with peptides that mimic nickel offers insight into nickel contact allergy. Proc. Natl. Acad. Sci. U.S. A. 109, 18517–18522.
Dessen A., Lawrence C.M., Cupo S., Zaller D.M., Wiley D.C. 1997. X-ray crystal structure of HLA-DR4 (DRA* 0101, DRB1* 0401) complexed with a peptide from human collagen II. Immunity. 7, 473–481.
Lang H.L., Jacobsen H., Ikemizu S., Andersson C., Harlos K., Madsen L., Hjorth P., Sondergaard L., Svejgaard A., Wucherpfennig K. 2002. A functional and structural basis for TCR cross-reactivity in multiple sclerosis. Nat. Immunol. 3, 940–943.
Wang M., Claesson M.H. 2014. Classification of human leukocyte antigen (HLA) supertypes. Methods Mol. Biol. 1184, 309–317. doi 10.1007/978-1-4939-1115-8_17
Kautsch S.J. 2006. The Nature of Flat Galaxies. Basel: Univ. of Basel Press.
Zhu Q., Vera C., Asaro R.J., Sche P., Sung L.A. 2007. A hybrid model for erythrocyte membrane: A single unit of protein network coupled with lipid bilayer. Biophys. J. 93, 386–400.
Author information
Authors and Affiliations
Corresponding author
Additional information
The article is published in the original.
Published in Russian in Molekulyarnaya Biologiya, 2017, Vol. 51, No. 3, pp. 524–533.
Rights and permissions
About this article
Cite this article
Cortés, A., Coral, J., McLachlan, C. et al. Planar molecular arrangements aid the design of MHC class II binding peptides. Mol Biol 51, 465–473 (2017). https://doi.org/10.1134/S002689331702008X
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S002689331702008X