Abstract
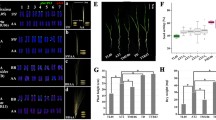
Polyploidization plays an important role in plant evolution, and it may trigger changes in genome structure and gene expression. The purpose of this study was to detect the extent of gene expression alterations in natural Brassica allotetraploids. Sequence-related amplified polymorphism (SRAP)-cDNA display was carried out on six species (13 accessions), three allotetraploids and their three ancestral diploids. Total of 1496 transcripts were screened, finding that 92 (6.1%) transcripts were subjected to silencing in allopolyploids, while 19 (1.3%) transcripts were activated. The results indicated that a significant fraction of duplicated genes have shown expression alterations in the natural allotetraploids compared with their diploid parents.
Similar content being viewed by others
References
Adams K.L., Wendel J.F. 2005a. Polyploidy and genome evolution in plants. Curr. Opin. Plant Biol. 8, 135–141.
Adams K.L., Wendel J.F. 2005b. Novel patterns of gene expression in polyploid plants. Trends Genet. 21, 539–543.
Blanc G., Hokamp K., Wolfe K.H. 2003. A recent polyploidy superimposed on older large-scale duplications in the Arabidopsis genome. Genome Res. 13, 137–144.
Rieseberg L.H., Willis J.H. 2007. Plant speciation. Science. 317, 910–914.
Chapman M.A., Burke J.M. 2007. Genetic divergence and hybrid speciation. Evolution. 61, 1773–1780.
Pires J.C., Zhao J., Schranz M.E., Leon E.J., Quijada P.A., Lukens L., Osborn T.C. 2004. Flowering time divergence and genomic rearrangements in resynthesized polyploids (Brassica). Biol. J. Linn. Soc. 82, 675–688.
Wang J., Tian L., Madlung A., Lee H.S., Chen M., Lee J.J., Watson B., Kagochi T., Comai L., Chen Z.J. 2004. Stochastic and epigenetic changes of gene expression in Arabidopsis polyploids. Genetics. 167, 1961–1973.
Luciano G.M., Juan-Pablo A.O., Juliana S., Francisco E., Camilo L.Q., Silvina C.P. 2005. A comprehensive analysis of gene expression alterations in a newly synthesized Paspalum notatum autotetraploid. Plant Sci. 169, 211–220.
Madlung A., Masuelli R., Watson B., Reynolds S., Davison J., Comai L. 2002. Remodeling of DNA methylation and phenotypic and transcriptional changes in synthetic Arabidopsis allotetraploids. Plant Physiol. 129, 733–746.
Pontes O., Neves N., Silva M., Lewis M.S., Madlung A., Comai L., Viegas W., Pikaard C.S. 2004. Chromosomal locus rearrangements are a rapid response to formation of the allotetraploid Arabidopsis suecica genome. Proc. Natl. Acad. Sci. USA. 101, 18240–18245.
Levy A.A., Feldman M. 2004. Genetic and epigenetic reprogramming of the wheat genome upon allopolyploidization. Biol. J. Linn. Soc. 82, 607–613.
Song K., Lu P., Tang K., Osborn T.C. 1995. Rapid genome change in synthetic polyploids of Brassica and its implications for polyploid evolution. Proc. Natl. Acad. Sci. USA. 92, 7719–7723.
Pikaard C.S. 2000. Nucleolar dominance: Uniparental gene silencing on a multi-megabase scale in genetic hybrids. Plant Mol. Biol. 43, 163–177.
Lee H.S., Chen Z.J. 2001. Protein-coding genes are epigenetically regulated in Arabidopsis polyploids. Proc. Natl. Acad. Sci. USA. 98, 6753–6758.
Kashkush K., Feldman M., Levy A.A. 2002. Gene loss, silencing, and activation in a newly synthesized wheat allotetraploid. Genetics. 160, 1651–1659.
Adams K.L., Cronn R., Percifield R., Wendel J.F. 2003. Genes duplicated by polyploidy show unequal contributions to the transcriptome and organ-specific reciprocal silencing. Proc. Natl. Acad. Sci. USA. 100, 4649–4654.
Adams K.L, Percifield R., Wendel J.F. 2004. Organ-specific silencing of duplicated genes in a newly synthesized cotton allotetraploid. Genetics. 168, 2217–2226.
Soltis D.E., Soltis P.S., Pires J.C., Kovarik A., Tate J.A., Mavrodiev E. 2004. Recent and recurrent polyploidy in Tragopogon (Asteraceae): Cytogenetic, genomic and genetic comparisons. Biol. J. Linn. Soc. 82, 485–501.
U N. 1935. Genome analysis in Brassica with special reference to the experimental formation of B. napus and peculiar mode of fertilization. Jap. J. Bot. 7, 389–452.
Zeng C.L., Wu X.M., Wang J.B. 2006. Seed coat development and its evolutionary implications in diploid and amphidiploid Brassica species. Acta Biol. Cracov. Bot. 48, 15–25.
Chen Z.J., Pikaard C.S. 1997a. Epigenetic silencing of RNA polymerase I transcription: Arole for DNA methylation and histone modification in nucleolar dominance. Gene Dev. 11, 2124–2136.
Chen Z.J., Pikaard C.S. 1997b. Transcriptional analysis of nucleolar dominance in polyploid plants: Biased expression/silencing of progenitor rRNA genes is developmentally regulated in Brassica. Proc. Natl. Acad. Sci. USA. 94, 3442–3447.
Li G., Quiros C.F. 2001. Sequence-related amplified polymorphism (SRAP), a new marker system based on a simple PCR reaction: Its application to mapping and gene tagging in Brassica. Theor. Appl. Genet. 103, 455–461.
Comai L., Tyagi A.P., Winter K., Holmes-Davis R., Reynolds S.H., Stevens Y., Byers B. 2000. Phenotypic instability and rapid gene silencing in newly formed Arabidopsis allotetraploids. Plant Cell. 12, 1551–1567.
Galitski T., Saldanha A.J., Styles C.A., Lander E.S., Fink G.R. 1999. Ploidy regulation of gene expression. Science. 285, 251–254.
Wang J., Lee J.J., Tian L., Lee H.S., Chen M., Rao S., Wei E.N., Doerge R.W., Comai L., Chen Z.J. 2005. Methods for genome-wide analysis of gene expression changes in polyploids. Meth. Enzymol. 395, 570–596.
Li G., Gao M., Yang B., Quiros F. 2004. Gene for gene alignment between the Brassica and Arabidopsis genomes by direct transcriptome mapping. Theor. Appl. Genet. 107, 168–180.
Wendel J.F. 2000. Genome evolution in polyploids. Plant Mol. Biol. 42, 225–249.
He P., Friebe B., Gill B., Zhou J.M. 2003. Allopolyploidy alters gene expression in the highly stable hexaploid wheat. Plant Mol. Biol. 52, 401–414.
Liu B., Brubaker C.L., Mergeai G., Cronn R.C., Wendel J.F. 2001. Polyploid formation in cotton is not accompanied by rapid genomic changes. Genome. 44, 321–330.
Author information
Authors and Affiliations
Corresponding author
Additional information
Original Russian Text © Qin Gui, Jialu Wang, Yanhao Xu, Jianbo Wang, 2009, published in Molekulyarnaya Biologiya, 2009, Vol. 43, No. 1, pp. 3–9.
The article is published in the original.
Rights and permissions
About this article
Cite this article
Gui, Q., Wang, J., Xu, Y. et al. Expression changes of duplicated genes in allotetraploids of Brassica detected by SRAP-cDNA technique. Mol Biol 43, 1–7 (2009). https://doi.org/10.1134/S0026893309010014
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0026893309010014