Abstract
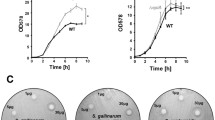
Persisters (P) are the cells surviving, but not reproducing in the presence of lethal doses of antibiotics (AB). These are the cells of a scarce (0.001–1.0%) subpopulation, emerging due to a phenotypic transition, which may be stochastic or induced by stress impacts. Growing in fresh medium, P reproduce the structure of the parent population, which is considered the cause of: (a) recurrent infections; (b) inefficiency of antibiotic treatment; and (c) development of antibiotic resistance, as was discussed in numerous articles and reviews. The goal of the present work was to investigate P formation by bacteria contaminating human skin and their properties, which have not been studied previously. Two approaches were used to measure the frequency of occurrence was determined for P in the populations of stationary-phase cultures of Staphylococcus capitis Р205-1 (UNIQEM, Russia), which is sensitive to polymyxin, Na sulfacyl, and co-trimoxazole, and Staphylococcus epidermidis 14990 АТСС, which is resistant to these antibiotics: (1) traditional, by application of lethal AB doses and (2) by using a lysis solution (LS), which killed the ordinary vegetative cells and selected the persister cells resistant to AB and LS (PAB and PLS, respectively). This is the first report on applicability of the method involving the lysis solution for determination of P abundance in strains with multidrug AB resistance (for the S. epidermidis strain). Unlike most bacterial strains retrieved from collections , the share of P in stationary-phase cultures of staphylococci inhabiting human skin was quite high: ~10% for both S. epidermidis PLS and S. capitis. Heterogeneity in resistance to the lysis solution was revealed in the PAB obtained by the standard procedure: PLS share in S. capitis was ~1% of PAB abundance. These PLS were resistant to AB cross-action. The differences in PLS abundance in the studied strains (10% for S. epidermidis and below 1.0% for S. capitis) correlated with their AB resistance. Long-term incubation (4 months) under standard conditions resulted in the cycle of staphylococci development ending by emergence of dormant forms (DF); their share for both strains was ~0.1% of the number of viable cells in the stationary-phase cultures. The DF of staphylococci had all characteristics of dormant cells: emerged in the cycle of culture development; preserved viability under nongrowth conditions; had no experimentally detectable metabolism (absence of respiration); had specific ultrastructural organization; and were heat-resistant (200‒600 times higher than the vegetative cells at 90°C for 15 min). The correlation between PLS abundance in the cultures of staphylococci and the share of heat-resistant DF confirms our earlier suggestion on persisters as DF percursors.





Similar content being viewed by others
REFERENCES
Balaban, N., Merrin, I., Chait, R., Kowalik, L., and Leibler, S., Bacterial persistence as a phenotypic switch, Science, 2004, vol. 305, pp. 1622‒1625.
Balaban, N.Q., Helaine, S., Lewis, K., Ackermann, M., Aldridge, B., Andersson, D.I., Brynildsen, M.P., Bumann, D., Camilli, A., Collins, J.J., Dehio, C., Fortune, S., Ghigo, J.M., Hardt, W.D., Harms, A., et al., Definitions and guidelines for research on antibiotic persistence, Nat. Rev. Microbiol., 2019, vol. 17, pp. 441‒448.
Bettenworth, V., Steinfeld, B., Duin, H., Petersen, K., S-treit, W.R., Bischofs, I., and Becker, A., Phenotypic heterogeneity in bacterial quorum sensing systems, J. Mol. B-iol., 2019, vol. 431, pp. 4530‒4546.
Biel, F.M., Allen, F.A., and Häse, C.C., Autolysis in Vibrio tubiashii and Vibrio coralliilyticus,Can. J. Microbiol., 2014, vol. 60, pp. 57‒63.
Bigger, J., Treatment of staphylococcal infections with penicillin by intermittent sterilization, The Lancet, 1944, vol. 244, pp. 497‒500.
Bukharin, O.V., Gintsburg, A.L., Romanova, Yu.N., and El’-Registan, G.I., Mekhanizmy vyzhivaniya bakterii (Mechanisms of Bacterial Survival), Moscow: Meditsina, 2005.
Canas-Duarte, S.J., Restrepo, S., and Pedraza, J.M., Novel protocol for persister cells isolation, PLoS One, 2014, vol. 9, no. 2. e88660.
Demkina, E.V., Loiko, N.G., Mulyukin, A.L., Nikolaev, Y.A., El’-Registan, G.I., Smirnova, T.A., Gaponov, A.M., Pisarev, V.M., and Tutel’yan, A.V., Effect of inherent immunity factors on development of antibiotic tolerance and survival of bacterial potulations under antibiotic attack, Microbiology (Moscow), 2015, vol. 84, pp. 764‒774.
El-Registan, G.I., Mulyukin, A.L., Nikolaev, Yu.A., Galchenko, V.F., Suzina, N.E., and Duda, V.I., Adaptogenic functions of extracellular autoregulators of microorganisms, Microbiology (Moscow), 2006, vol. 75, pp. 380‒389.
Gollan, B., Grabe, G., Michaux, C., and Helaine, S., Bacterial persisters and infection: past, present, and progressing, Annu. Rev. Microbiol., 2019, vol. 73, pp. 359‒385.
Grice, E.A., Kong, H.H., Conlan, S., Deming, C.B., Davis, J., Young, A.C., Bouffard, G.G., Blakesley, R.W., Murray, P.R., Green, E.D., Turner, M.L., and Segre, J.A., NISC Comparative Sequencing Program, Science, 2009, vol. 324, pp. 1190–1192.
Kaldalu, N., Joers, A., Ingelman, H., and Tenson, T., A general method for measuring persister levels in Escherichia coli cultures, in Bacterial Persistence. Methods and Protocols, Michiels, J. and Fauvart, M., Eds., Humana, 2016, pp. 29‒42.
Levin-Reisman, I., Ronin, I., Gefen, O., Braniss, I., Shoresh, N., and Balaban, N.Q., Antibiotic tolerance facilitates the evolution of resistance, Science, 2017, vol. 355, pp. 826‒830.
Lewis, K. and Shan, Y., Why tolerance invites resistance, Science, 2017, vol. 355, p. 796.
Lewis, K., Persister cells, Annu. Rev. Microbiol., 2010, vol. 64, pp. 357‒372.
Leygeber, M., Lindemann, D., Sachs, C.C., Kagano-vitch, E., Wiechert, W., Nöh, K., and Kohlheyer, D., Analyzing microbial population heterogeneity—expanding the toolbox of microfluidic single-cell cultivations, J. Mol. Bi-ol., 2019, vol. 431, pp. 4569‒4588.
Loiko, N.G., Kozlova, A.N., Nikolaev, Y.A., El’-Registan, G.I., Gaponov, A.M., and Tutel’yan, A.V., Effect of stress on emergence of antibiotic-tolerant Escherichia coli cells, Microbiology (Moscow), 2015, vol. 84, pp. 595‒609.
Maisonneuve, E. and Gerdes, K., Molecular mechanisms underlying bacterial persisters, Cell, 2014, vol. 157, pp. 539‒548.
Mulyukin, A.L., Kozlova, A.N., Sorokin, V.V., El’-Regis-tan, G.I., Suzina, N.E., Cherdyntseva, T.A., Kotova, I.B., Gaponov, A.M., and Tutel’yan, A.V., Surviving forms of antibiotic treated Pseudomonas aeruginosa,Microbiology (Moscow), 2015, vol. 84, pp. 751‒763.
Popp, P.F. and Mascher, T., Coordinated cell death in isogenic bacterial populations: sacrificing some for the benefit of many?, J. Mol. Biol., 2019, vol. 431, pp. 4656–4669.
Post-Conference Report “a World without Antibiotics. Conclusions from Uppsala Health Summit,” 2–3 June 2015. http://apps.who.int/medicinedocs/documents/ s22152en/s22152en.pdf.
Salina, E.G., Grigorov, A.S., Bychenko, O.S., Skvortsova, Y.V., Mamedov, I.Z., Azhikina, T.L., and Kaprelyants, A.S., Resuscitation of dormant “non-culturable” Mycobacterium tuberculosis is characterized by immediate transcriptional burst, Front. Cell Infect. Microbiol., 2019, vol. 9, article 272. https://doi.org/10.3389/fcimb.2019.00272
Sherwood, L., Willey, J., and Woolverton, C., NIH News, in Prescott’s Microbiology, 9th ed., New York: McGraw Hill, 2013, pp. 713–721.
Sudo, S.Z. and Dworkin, M., Comparative biology of procaryotic resting cells, Adv. Microbiol. Physiol., 1973, vol. 9, pp. 153‒224.
The Human Microbiome Project Consortium, Structure, function and diversity of the healthy human microbiome, Nature, 2012, vol. 486, no. 7402, pp. 207–214.
Thomas, P., Sekhar, A.C., Upreti, R., Mujawar, M.M., and Pasha, S.S., Optimization of single plate-serial dilution spotting (SP-SDS) with sample anchoring as an assured method for bacterial and yeast CFU enumeration and single colony isolation from diverse samples, Biotechnol. Rep. (Amst.), 2015, vol. 8, pp. 45–55.
Van den Bergh, B., Fauvart, M., and Michiels, J., Formation, physiology, ecology, evolution and clinical importance of bacterial persisters, FEMS Microbiol. Rev., 2017, vol. 41, pp. 219‒251.
Van den Bergh, B., Michiels, J.E., and Michiels, J., Experimental evolution of Escherichia coli persister levels using cyclic antibiotic treatments, in Bacterial Persistence. Methods and Protocols, Michiels, J. and Fauvart, M., Eds., Humana, 2016, pp. 131‒146.
Wilmaerts, D., Windels, E.M., Verstraeten, N., and Michiels, J., General mechanisms leading to persister formation and awakening, Trends Genet., 2019, vol. 35, pp. 401‒411.
Windels, E.M., Michiels, J.E., Van den Bergh, B., Fauvart, M., and Michiels, J., Antibiotics: combatting tolerance to stop resistance, mBio, 2019, vol. 10. e02095-19. https://doi.org/10.1128/mBio.02095-19
Xu, T., Wang, X.-Y., Cui, P., Zhang, Y.-M., Zhang, W.-H., and Zhang, Y., The Agr quorum sensing system represses persister formation through regulation of phenol soluble modulins in Staphylococcus aureus,Front. Microbiol., 2017, vol. 8, art. 2189. https://doi.org/10.3389/fmicb.2017.02189
Funding
This work was supported by the Russian Science Foundation (grant no. 19-74-10071). The work on species identification of Staphylococcus capitis strain Р205-1 was partially supported by the Ministry of Science and Higher Education of the Russian Federation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare that they have no conflict of interest. This article does not contain any studies involving animals or human participants performed by any of the authors.
Additional information
Translated by A. Oleskin
Rights and permissions
About this article
Cite this article
Nikolaev, Y.A., Pankratov, T.A., Gannesen, A.V. et al. Formation and Properties of Persister Cells of Staphylococcus capitis and Staphylococcus epidermidis, Bacteria Inhabiting Human Skin. Microbiology 89, 425–434 (2020). https://doi.org/10.1134/S0026261720040104
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0026261720040104