Abstract
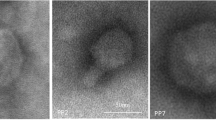
Phages of the phytopathogenic Pectobacteriaceae species causing black leg and soft rot of potato were investigated. These phages are promising as biocontrol agents to prevent the loss of seed and ware potato tubers. The present work characterizes a new podovirus PP16, infecting a broad range of Pectobacterium carotovorum strains. Based on its genomic composition, phage PP16 was assigned to a separate phylogenetic branch of the genus Phimunavirus, subfamily Autographivirinae. Bacteriophage PP16 efficiently inhibited development of bacterial infection both in vitro and in planta. The field experiment demonstrated a substantial increase of plant germination after the treatment of seed potato with phage PP16.







Similar content being viewed by others
REFERENCE
Ackermann H.-W. Basic phage electron microscopy, in Bacteriophages: Methods and Protocols, 2009, vol. 1, pp. 113–126. https://doi.org/10.1007/978-1-60327-164-6
Adeolu, M., Alnajar, S., Naushad, S., and Gupta, R.S., Genome-based phylogeny and taxonomy of the ‘Enterobacteriales’: proposal for Enterobacterales ord. nov. divided into the families Enterobacteriaceae, Erwiniaceae fam. nov., Pectobacteriaceae fam. nov., Yersiniaceae fam. nov., Hafniaceae fam. nov., Int. J. Syst. Evol. Microbiol., 2016, vol. 66, pp. 5575–5599. https://doi.org/10.1099/ijsem.0.001485
Altschul, S.F., Madden, T.L., Schäffer, A.A., Zhang, J., Zhang, Z., Miller, W., and Lipman, D.J., Gapped BLAST and PSI-BLAST: a new generation of protein database search programs, Nucl. Acids Res., 1997, vol. 25, pp. 3389–3402. https://doi.org/10.1093/nar/25.17.3389
Balogh, B., Jones, J.B., Iriarte, F.B., and Momol, M.T., Phage therapy for plant disease control, Curr. Pharm. Biotechnol., 2010, vol. 11, pp. 48–57. https://doi.org/10.2174/138920110790725302
Bankevich, A., Nurk, S., Antipov, D., Gurevich, A.A., Dvorkin, M., Kulikov, A.S., Lesin, V.M., Nikolenko, S.I., Pham, S., Prjibelski, A.D., Pyshkin, A.V., Sirotkin, A.V., Vyahhi, N., Tesler, G., Alekseyev, M.A., and Pevzner, P.A., SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing, J. Computat. Biol., 2012, vol. 19, pp. 455–477. https://doi.org/10.1089/cmb.2012.0021
Besemer, J., Lomsadze, A., and Borodovsky, M., GeneMarkS: a self-training method for prediction of gene starts in microbial genomes. Implications for finding sequence motifs in regulatory regions, Nucl. Acids Res., 2001, vol. 29, pp. 2607–2618. doi: 11410670
Buttimer, C., Lucid, A., Neve, H., Franz, C.M.A.P., O’Mahony, J., Turner, D., Lavigne, R., and Coffey, A., Pectobacterium atrosepticum phage VB_PatP_CB5: a member of the proposed genus “Phimunavirus”, Viruses, 2018, vol. 10. pii: E394. https://doi.org/10.3390/v10080394
Clokie, M.R.J. and Kropinski, A.M., Bacteriophages: Methods and Protocols, vol. 1: Isolation, Characterization, and Interactions, in Methods in Molecular Biology, Humana, 2009. https://doi.org/10.1007/978-1-60327-164-6
Czajkowski, R., Ozymko, Z., de Jager, V., Siwinska, J., Smolarska, A., Ossowicki, A., Narajczyk, M., and Lojkowska, E., Genomic, proteomic and morphological characterization of two novel broad host lytic bacteriophages ΦPD10.3 and ΦPD23.1 infecting pectinolytic Pectobacterium spp. and Dickeya spp., PLoS One, 2015, vol. 10, article e0119812. https://doi.org/10.1371/journal.pone.0119812
Dees, M.W., Lysøe, E., Rossmann, S., Perminow, J., and Brurberg, M.B., Pectobacterium polaris sp. nov., isolated from potato (Solanum tuberosum), Int. J. Syst. Evol. Microbiol., 2017, vol. 67, pp. 5222–5529. https://doi.org/10.1099/ijsem.0.002448
Eriksson, H., Maciejewska, B., Latka, A., Majkowska-Skrobek, G., Hellstrand, M., Melefors, Ö., Wang, J.-T., Kropinski, A.M., Drulis-Kawa, Z., and Nilsson, A.S., A suggested new bacteriophage genus, ‘Kp34likevirus’, within the Autographivirinae subfamily of Podoviridae, Viruses, 2015, vol. 7, pp. 1804–1822. https://doi.org/10.3390/v7041804
Finn, R.D., Clements, J., and Eddy, S.R., HMMER web server: interactive sequence similarity. searching, Nucl. Acids Res., 2011, vol. 39, suppl. 2. https://doi.org/10.1093/nar/gkr367
Gardan, L., Gouy, C., Christen, R., and Samson, R., Elevation of three subspecies of Pectobacterium carotovorum to species level: Pectobacterium atrosepticum sp. nov., Pectobacterium betavasculorum sp. nov. and Pectobacterium wasabiae sp. nov., Int. J. Syst. Evol. Microbiol., 2003, vol. 53, pp. 381–391. https://doi.org/10.1099/ijs.0.02423-0
Hirata, H., Kashihara, M., Horiike, T., Suzuki, T., Dohra, H., Netsu, O., and Tsuyumu, S., Genome sequence of Pectobacterium Carotovorum phage PPWS1, isolated from Japanese horseradish [Eutrema Japonicum (Miq.) Koidz] showing soft-rot symptoms, Genome Announc., 2016, vol. 4, no. 2. pii: e01625-15. https://doi.org/10.1128/genomeA.01625-15
Humphris, S.N., Cahill, G., Elphinstone, J.G., Kelly, R., Parkinson, N.M., Pritchard, L., Toth, I.K., and Saddler, G.S., Detection of the bacterial potato pathogens Pectobacterium and Dickeya spp. using conventional and real-time PCR, in Methods in Molecular Biology, Clifton, NJ, 2015, pp. 1–16. https://doi.org/10.1007/978-1-4939-2620-6_1
Lavigne, R., Sun, W.D., and Volckaert, V., PHIRE, a deterministic approach to reveal regulatory elements in bacteriophage genomes, Bioinformatics, 2004, vol. 20, pp. 629–635. https://doi.org/10.1093/bioinformatics/btg456
Lavigne, R., Seto, D., Mahadevan, P., Ackermann, H.-W., and Kropinski, A.M., Unifying classical and molecular taxonomic classification: analysis of the Podoviridae using BLASTP-based tools, Res. Microbiol., 2008, vol. 159, pp. 406–414. https://doi.org/10.1016/j.resmic.2008.03.005
Lim, J.A., Heu, S., Park, J., and Roh, E., Genomic characterization of bacteriophage VB_PcaP_PP2 infecting Pectobacterium carotovorum subsp. carotovorum, a new member of a proposed genus in the subfamily Autographivirinae, Arch. Virol., 2017, vol. 162, pp. 2441–2444. https://doi.org/10.1007/s00705-017-3349-6
Mansfield, J., Genin, S., Magori, S., Citovsky, V., Sriariyanum, M., Ronald, P., Dow, M., Verdier, V., Beer, S.V., Machado, M.A., Toth, I., Salmond, G., and Foster, G.D., Top 10 plant pathogenic bacteria in molecular plant pathology, Mol. Plant Pathol., 2012, vol. 13, pp. 614–629. https://doi.org/10.1111/j.1364-3703.2012.00804.x
Martin, B., Humbert, O., Camara, M., Guenzi, E., Walker, J., Mitchell, T., Andrew, P., Prudhomme, M., Alloing, G., Hakenbeck, R., Morrison, D.A., Boulnois, G.J., and Claverys, J.-P., A highly conserved repeated DNA element located in the chromosome of Streptococcus pneumoniae, Nucl. Acids Res., 1992, vol. 20, pp. 3479–3483. https://doi.org/10.1093/nar/20.13.3479
Meier-Kolthoff, J.P. and Göker, M., VICTOR: Genome-based phylogeny and classification of prokaryotic viruses, Bioinformatics, 2017, vol. 33, pp. 3396–3404. https://doi.org/10.1093/bioinformatics/btx440
Pérombelon, M.C.M., Potato diseases caused by soft rot erwinias: an overview of pathogenesis, Plant Pathol., 2002, vol. 51, pp. 1–12. https://doi.org/10.1046/j.0032-0862.2001
Prokhorov, N.S., Riccio, C., Zdorovenko, E.L., Shneider, M.M., Browning, C., Knirel, Y.A., Leiman, P.G., and Letarov, A.V., Function of bacteriophage G7C esterase tailspike in host cell adsorption, Mol. Microbiol., 2017, vol. 105, pp. 385–398. https://doi.org/10.1111/mmi.13710
Roucourt, B. and Lavigne, R., The role of interactions between phage and bacterial proteins within the infected cell: a diverse and puzzling interactome, Environ. Microbiol., 2009, vol. 11, pp. 2789–2805. https://doi.org/10.1111/j.1462-2920.2009.02029.x
Sambrook, J., Fritsch, E.F., and Maniatis, T., Molecular Cloning: A Laboratory Manual, 2nd ed., Cold Spring Harbor Laboratory Press, 1989.
Sarfraz, S., Riaz, K., Oulghazi, S., Cigna, J., Sahi, S.T., Khan, S.H., and Faure, D., Pectobacterium punjabense sp. nov., isolated from blackleg symptoms of potato plants in Pakistan, Int. J. Syst. Evol. Microbiol., 2018, vol. 68, pp. 3551–3556. https://doi.org/10.1099/ijsem.0.003029
Shirshikov, F.V., Korzhenkov, A.A., Miroshnikov, K.K., Kabanova, A.P., Barannik, A.P., Ignatov, A.N., and Miroshnikov, K.A., Draft genome sequences of new genomospecies “Candidatus Pectobacterium maceratum” strains, which cause soft rot in plants, Genome Announc., 2018, vol. 6, article e00260-18. https://doi.org/10.1128/genomeA.00260-18
Waleron, M., Misztak, A., Waleron, M., Franczuk, M., Wielgomas, B., and Waleron, K., Transfer of Pectobacterium carotovorum subsp. carotovorum strains isolated from potatoes grown at high altitudes to Pectobacterium peruviense sp. nov., Syst. Appl. Microbiol., 2018, vol. 41, pp. 85–93. https://doi.org/10.1016/J.SYAPM.2017.11.005
Yagodin, B.A., Zhukov, Yu.P., and Kobzarenko, V.I., Agrokhimiya (Agochemistry), Moscow: Kolos, 2002.
Young, R., Phage lysis: three steps, three choices, one outcome, J. Microbiol., 2014, vol. 52, pp. 243–258. https://doi.org/10.1007/s12275-014-4087-z
Funding
The research, except for the study of bacteriophage morphology, was supported by the grant of the Russian Science Foundation no. 16-16-00073.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors declare that they have no conflict of interest. This article does not contain any studies involving animals or human participants performed by any of the authors.
Additional information
Translated by E. Makeeva
Rights and permissions
About this article
Cite this article
Voronina, M.V., Bugaeva, E.N., Vasiliev, D.M. et al. Characterization of Pectobacterium carotovorum subsp. carotovorum Bacteriophage PP16 Prospective for Biocontrol of Potato Soft Rot. Microbiology 88, 451–460 (2019). https://doi.org/10.1134/S0026261719040118
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0026261719040118