Abstract
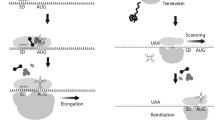
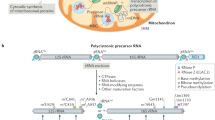
Mitochondria are obligate organelles of most eukaryotic cells that perform many different functions important for cellular homeostasis. The main role of mitochondria is supplying cells with energy in a form of ATP, which is synthesized in a chain of oxidative phosphorylation reactions on the organelle inner membrane. It is commonly believed now that mitochondria have the endosymbiotic origin. In the course of evolution, they have lost most of their genetic material as a result of genome reduction and gene transfer to the nucleus. The majority of mitochondrial proteins are synthesized in the cytosol and then imported to the mitochondria. However, almost all known mitochondria still contain genomes that are maintained and expressed. The processes of protein biosynthesis in the mitochondria — mitochondrial translation — substantially differs from the analogous processes in bacteria and the cytosol of eukaryotic cells. Mitochondrial translation is characterized by a high degree of specialization and specific regulatory mechanisms. In this review, we analyze available information on the common principles of mitochondrial translation with emphasis on the molecular mechanisms of translation initiation in the mitochondria of yeast and mammalian cells.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Abbreviations
- fMet-tRNA:
-
formylmethionine-tRNA
- mtIF2 and mtIF3:
-
translation initiation factors 2 and 3, respectively
- 5′-UTR:
-
5′-untranslated region of mRNA
References
Spinelli, J. B., and Haigis, M. C. (2018) The multifaceted contributions of mitochondria to cellular metabolism, Nat. Cell Biol., 20, 745–754.
Gray, M. W. (2012) Mitochondrial evolution, Cold Spring Harb. Perspect. Biol., 4, a011403; doi: https://doi.org/10.1101/cshperspect.a011403
Gray, M. W. (2015) Mosaic nature of the mitochondrial proteome: implications for the origin and evolution of mitochondria, Proc. Natl. Acad. Sci. USA, 112, 10133–10138; doi: https://doi.org/10.1073/pnas.1421379112.
Feagin, J. E., Harrell, M. I., Lee, J. C., Coe, K. J., Sands, B. H., Cannone, J. J., Tami, G., Schnare, M. N., and Gutell, R. R. (2012) The fragmented mitochondrial ribosomal RNAs of Plasmodium falciparum, PLoS One, 7, e38320; doi: https://doi.org/10.1371/journal.pone.0038320
Sloan, D. B., Alverson, A. J., Chuckalovcak, J. P., Wu, M., McCauley, D. E., Palmer, J. D., and Taylor, D. R. (2012) Rapid evolution of enormous, multichromosomal genomes in flowering plant mitochondria with exceptionally high mutation rates, PLoS Biol., 10, e1001241; doi: https://doi.org/10.1371/journal.pbio.1001241
Freel, K. C., Friedrich, A., and Schacherer, J. (2015) Mitochondrial genome evolution in yeasts: an all-encom-passing view, FEMS Yeast Res., 15, fov023; doi: https://doi.org/10.1093/femsyr/fov023
Taanman, J. W. (1999) The mitochondrial genome: structure, transcription, translation and replication, Biochim. Biophys. Acta, 1410, 103–123.
Burger, G., Gray, M. W., Forget, L., and Lang, B. F. (2013) Strikingly bacteria-like and gene-rich mitochondrial genomes throughout jakobid protists, Genome Biol. Evol., 5, 418–438; doi: https://doi.org/10.1093/gbe/evt008.
Finkelstein, A. V., Razin, S. V., and Spirin, A. S. (2018) Intersubunit mobility of the ribosome, Mol. Biol. (Moscow), 52, 921–934.
Eme, L., Spang, A., Lombard, J., Stairs, C. W., and Ettema, T. J. G. (2017) Archaea and the origin of eukaryotes, Nat. Rev. Microbiol., 15, 711–723.
Amunts, A., Brown, A., Toots, J., Scheres, S. H. W., and Ramakrishnan, V. (2015) The structure of the human mitochondrial ribosome, Science, 348, 95–98; doi: https://doi.org/10.1126/science.aaa1193.
Greber, B. J., Bieri, P., Leibundgut, M., Leitner, A., Aebersold, R., Boehringer, D., and Ban, N. (2015) The complete structure of the 55S mammalian mitochondrial ribosome, Science, 348, 303–308; doi: https://doi.org/10.1126/science.aaa3872.
Englmeier, R., Pfeffer, S., and Forster, F. (2017) Structure of the human mitochondrial ribosome studied in situ by cryoelectron tomography, Structure, 25, 1574–1581.e2; doi: https://doi.org/10.1016/j.str.2017.07.011
Bieri, P., Greber, B. J., and Ban, N. (2018) High-resolution structures of mitochondrial ribosomes and their functional implications, Curr. Opin. Struct. Biol., 49, 44–53.
Ramrath, D. J. F., Niemann, M., Leibundgut, M., Bieri, P., Prange, C., Horn, E. K., Leitner, A., Boehringer, D., Schneider, A., and Ban, N. (2018) Evolutionary shift toward protein-based architecture in trypanosomal mitochondrial ribosomes, Science, 362; doi: https://doi.org/10.1126/science.aau7735
Desai, N., Brown, A., Amunts, A., and Ramakrishnan, V. (2017) The structure of the yeast mitochondrial ribosome, Science, 355, 528–531; doi: https://doi.org/10.1126/science.aal2415.
Waltz, F., Nguyen, T. T., Arrive, M., Bochler, A., Chicher, J., Hammann, P., Kuhn, L., Quadrado, M., Mireau, H., Hashem, Y., and Giege, P. (2019) Small is big in Arabidopsis mitochondrial ribosome, Nat. Plants, 5, 106–117; doi: https://doi.org/10.1038/s41477-018-0339-y.
Waltz, F., Soufari, H., Bochler, A., Giege, P., and Hashem, Y. (2019) Cryo-EM structure of the RNA-rich plant mitochondrial ribosome, bioRxiv; doi: https://doi.org/10.1101/777342
Waltz, F., and Giege, P. (2019) Striking diversity of mitochondria-specific translation processes across eukaryotes, Trends Biochem. Sci., 45, 149–162; doi: https://doi.org/10.1016/j.tibs.2019.10.004.
Ott, M., Amunts, A., and Brown, A. (2016) Organization and regulation of mitochondrial protein synthesis, Annu. Rev. Biochem., 85, 77–101; doi: https://doi.org/10.1146/annurev-biochem-060815-014334.
Roll-Mecak, A., Shin, B. S., Dever, T. E., and Burley, S. K. (2001) Engaging the ribosome: universal IFs of translation, Trends Biochem. Sci., 26, 705–709.
Atkinson, G. C., Kuzmenko, A., Kamenski, P., Vysokikh, M. Y., Lakunina, V., Tankov, S., Smirnova, E., Soosaar, A., Tenson, T., and Hauryliuk, V. (2012) Evolutionary and genetic analyses of mitochondrial translation initiation factors identify the missing mitochondrial IF3 in S. cerevisiae, Nucleic Acids Res., 40, 6122–6134; doi: https://doi.org/10.1093/nar/gks272
Julian, P., Milon, P., Agirrezabala, X., Lasso, G., Gil, D., Rodnina, M. V., and Valle, M. (2011) The cryo-EM structure of a complete 30S translation initiation complex from Escherichia coli, PLoS Biol., 9, e1001095; doi: https://doi.org/10.1371/journal.pbio.1001095
Myasnikov, A. G., Simonetti, A., Marzi, S., and Klaholz, B. P. (2009) Structure-function insights into prokaryotic and eukaryotic translation initiation, Curr. Opin. Struct. Biol., 19, 300–309.
Suzuki, T., Nagao, A., and Suzuki, T. (2011) Human mitochondrial tRNAs: biogenesis, function, structural aspects, and diseases, Annu. Rev. Genet., 45, 299–329; doi: https://doi.org/10.1146/annurev-genet-110410-132531.
Tan, T. H. P., Bochud-Allemann, N., Horn, E. K., and Schneider, A. (2002) Eukaryotic-type elongator tRNAMet of Trypanosoma brucei becomes formylated after import into mitochondria, Proc. Natl. Acad. Sci. USA, 99, 1152–1157; doi: https://doi.org/10.1073/pnas.022522299.
Gaur, R., Grasso, D., Datta, P. P., Krishna, P. D., Das, G., Spencer, A., Agrawal, R. K., Spremulli, L., and Varshney, U. (2008) A single mammalian mitochondrial translation initiation factor functionally replaces two bacterial factors, Mol. Cell, 29, 180–190; doi: https://doi.org/10.1016/j.molcel.2007.11.021.
Yassin, A. S., Haque, M. E., Datta, P. P., Elmore, K., Banavali, N. K., Spremulli, L. L., and Agrawal, R. K. (2011) Insertion domain within mammalian mitochondrial translation initiation factor 2 serves the role of eubacterial initiation factor 1, Proc. Natl. Acad. Sci. USA, 108, 3918–3923; doi: https://doi.org/10.1073/pnas.1017425108.
Kummer, E., Leibundgut, M., Rackham, O., Lee, R. G., Boehringer, D., Filipovska, A., and Ban, N. (2018) Unique features of mammalian mitochondrial translation initiation revealed by cryo-EM, Nature, 560, 263–267.
Vambutas, A., Ackerman, S. H., and Tzagoloff, A. (1991) Mitochondrial translational initiation and elongation factors in Saccharomyces cerevisiae, Eur. J. Biochem., 201, 643–652; doi: https://doi.org/10.1111/j.1432-1033.1991.tb16325.x
Garofalo, C., Trinko, R., Kramer, G., Appling, D. R., and Hardesty, B. (2003) Purification and characterization of yeast mitochondrial initiation factor 2, Arch. Biochem. Biophys., 413, 243–252; doi: https://doi.org/10.1016/s0003-9861(03)00119-x.
Peske, F., Rodnina, M. V., and Wintermeyer, W. (2005) Sequence of steps in ribosome recycling as defined by kinetic analysis, Mol. Cell, 18, 403–412; doi: https://doi.org/10.1016/j.molcel.2005.04.009.
Elvekrog, M. M., and Gonzalez, R. L. (2013) Conformational selection of translation initiation factor 3 signals proper substrate selection, Nat. Struct. Mol. Biol., 20, 628–633; doi: https://doi.org/10.1038/nsmb.2554.
Valasek, L. S. (2012) “Ribozoomin” — translation initiation from the perspective of the ribosome-bound eukaryotic initiation factors (eIFs), Curr. Protein Pept. Sci., 13, 305–330.
Biou, V., Shu, F., and Ramakrishnan, V. (1995) X-ray crystallography shows that translational initiation factor IF3 consists of two compact alpha/beta domains linked by an alpha-helix, EMBO J., 14, 4056–4064; doi: https://doi.org/10.1002/J.1460-2075.1995.tb00077.x
Petrelli, D., La Teana, A., Garofalo, C., Spurio, R., Pon, C. L., and Gualerzi, C. O. (2001) Translation initiation factor IF3: two domains, five functions, one mechanism? EMBO J., 20, 4560–4569; doi: https://doi.org/10.1093/emboj/20.16.4560
Koc, E. C., and Spremulli, L. L. (2002) Identification of mammalian mitochondrial translational initiation factor 3 and examination of its role in initiation complex formation with natural mRNAs, J. Biol. Chem., 277, 35541–35549; doi: https://doi.org/10.1074/jbc.M202498200.
Haque, M. E., and Spremulli, L. L. (2008) Roles of the N- and C-terminal domains of mammalian mitochondrial initiation factor 3 in protein biosynthesis, J. Mol. Biol., 384, 929–940; doi: https://doi.org/10.1016/j.jmb.2008.09.077.
Bhargava, K., and Spremulli, L. L. (2005) Role of the N- and C-terminal extensions on the activity of mammalian mitochondrial translational initiation factor 3, Nucleic Acids Res., 33, 7011–7018; doi: https://doi.org/10.1093/nar/gki1007.
Christian, B. E., and Spremulli, L. L. (2009) Evidence for an active role of IF3 mt in the initiation of translation in mammalian mitochondria, Biochemistry, 48, 3269–3278; doi: https://doi.org/10.1021/bi8023493.
Koripella, R. K., Sharma, M. R., Haque, M. E., Risteff, P., Spremulli, L. L., and Agrawal, R. K. (2019) Structure of human mitochondrial translation initiation factor 3 bound to the small ribosomal subunit, iScience, 12, 76–86; doi: https://doi.org/10.1016/j.isci.2018.12.030.
Chicherin, I. V., Zinina, V. V., Levitskiy, S. A., Serebryakova, M. V., and Kamenski, P. A. (2019) Aim23p interacts with the yeast mitochondrial ribosomal small subunit, Biochemistry (Moscow), 84, 40–46; doi: https://doi.org/10.1134/S000629791901005X.
Levitskii, S., Derbikova, K., Baleva, M. V., Kuzmenko, A., Golovin, A. V., Chicherin, I., Krasheninnikov, I. A., and Kamenski, P. (2018) 60S dynamic state of bacterial ribosome is fixed by yeast mitochondrial initiation factor 3, PeerJ., 6, e5620; doi: https://doi.org/10.7717/peerj.5620
Derbikova, K., Kuzmenko, A., Levitskii, S., Klimontova, M., Chicherin, I., Baleva, M. V., Krasheninnikov, I. A., and Kamenski, P. (2018) Biological and evolutionary significance of terminal extensions of mitochondrial translation initiation factor 3, Int. J. Mol. Sci., 19; doi: https://doi.org/10.3390/ijms19123861
Kuzmenko, A., Derbikova, K., Salvatori, R., Tankov, S., Atkinson, G. C., Tenson, T., Ott, M., Kamenski, P., and Hauryliuk, V. (2016) Aim-less translation: loss of Saccharomyces cerevisiae mitochondrial translation initiation factor mIF3/Aim23 leads to unbalanced protein synthesis, Sci. Rep., 6, 18749; doi: https://doi.org/10.1038/srep18749.
Herrmann, J. M., Woellhaf, M. W., and Bonnefoy, N. (2013) Control of protein synthesis in yeast mitochondria: the concept of translational activators, Biochim. Biophys. Acta, 1833, 286–294.
Derbikova, K. S., Levitsky, S. A., Chicherin, I. V., Vinogradova, E. N., and Kamenski, P. A. (2018) Activation of yeast mitochondrial translation: who is in charge? Biochemistry (Moscow), 83, 87–97.
Montoya, J., Ojala, D., and Attardi, G. (1981) Distinctive features of the 5′-terminal sequences of the human mitochondrial mRNAs, Nature, 290, 465–470; doi: https://doi.org/10.1038/290465a0.
Weraarpachai, W., Antonicka, H., Sasarman, F., Seeger, J., Schrank, B., Kolesar, J. E., Lochmuller, H., Chevrette, M., Kaufman, B. A., Horvath, R., and Shoubridge, E. A. (2009) Mutation in TACO1, encoding a translational activator of COX I, results in cytochrome c oxidase deficiency and late-onset Leigh syndrome, Nat. Genet., 41, 833–837; doi: https://doi.org/10.1038/ng.390.
Green-Willms, N. S., Butler, C. A., Dunstan, H. M., and Fox, T. D. (2001) Pet111p, an inner membrane-bound translational activator that limits expression of the Saccharomyces cerevisiae mitochondrial gene COX2, J. Biol. Chem., 276, 6392–6397; doi: https://doi.org/10.1074/jbc.M009856200
Rudler, D. L., Hughes, L. A., Perks, K. L., Richman, T. R., Kuznetsova, I., Ermer, J. A., Abudulai, L. N., Shearwood, A. J., Viola, H. M., Hool, L. C., Siira, S. J., Rackham, O., and Filipovska, A. (2019) Fidelity of translation initiation is required for coordinated respiratory complex assembly, Sci. Adv., 5, eaay2118; doi: https://doi.org/10.1126/sciadv.aay2118
Author information
Authors and Affiliations
Corresponding author
Additional information
Russian Text © The Author(s), 2020, published in Biokhimiya, 2020, Vol. 85, No. 3, pp. 299–306.
Funding
This work was supported by the Russian Science Foundation (project 17-14-01005; testing hypotheses on the evolution of mitochondrial translation apparatus) and Russian Foundation for Basic Research (project 19-14-50206).
Conflict of interest
The authors declare no conflict of interest in financial or any other sphere.
Ethical approval
This article does not contain description of studies with human participants or animals performed by any of the authors.
Rights and permissions
Open access. This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Levitskii, S.A., Baleva, M.V., Chicherin, I.V. et al. Protein Biosynthesis in Mitochondria: Past Simple, Present Perfect, Future Indefinite. Biochemistry Moscow 85, 257–263 (2020). https://doi.org/10.1134/S0006297920030013
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S0006297920030013