Abstract
ADP-ribosyltransferases (ARTs) modify proteins with single units or polymers of ADP-ribose to regulate DNA repair. However, the substrates for these enzymes are ill-defined. For example, although histones are modified by ARTs, the sites on these proteins ADP-ribosylated following DNA damage and the ARTs that catalyse these events are unknown. This, in part, is due to the lack of a eukaryotic model that contains ARTs, in addition to histone genes that can be manipulated to assess ADP-ribosylation events in vivo. Here we exploit the model Dictyostelium to identify site-specific histone ADP-ribosylation events in vivo and define the ARTs that mediate these modifications. Dictyostelium histones are modified in response to DNA double strand breaks (DSBs) in vivo by the ARTs Adprt1a and Adprt2. Adprt1a is a mono-ART that modifies H2BE18 in vitro, although disruption of this site allows ADP-ribosylation at H2BE19. Although redundancy between H2BE18 and H2BE19 ADP-ribosylation is also apparent following DSBs in vivo, by generating a strain with mutations at E18/E19 in the h2b locus we demonstrate these are the principal sites modified by Adprt1a/Adprt2. This identifies DNA damage induced histone mono-ADP-ribosylation sites by specific ARTs in vivo, providing a unique platform to assess how histone ADP-ribosylation regulates DNA repair.
Similar content being viewed by others
Introduction
ADP-ribosyltransferases (ARTs) are primary sensors of DNA damage that catalyse the addition of ADP-ribose onto target proteins. 17 genes containing predicted ART catalytic domains have been identified in humans1 and the majority of these add single ADP-ribose moieties onto target proteins by a process known as mono-ADP-ribosylation (MARylation). However, several ARTs, including PARP1, PARP2, PARP5a and PARP5b catalyse ADP-ribose polymers that contain both linear and branched glycosidic linkages2,3. Whilst ADP-ribosylation has been implicated in a number of cellular processes including cell growth and differentiation, transcriptional regulation and programmed cell death3, its best defined role is in regulating DNA repair, with specific reference to DNA strand breaks. PARP1 and PARP2 are primary sensors of DNA damage that become activated on binding to DNA single strand breaks (SSBs) and modify a variety of substrates, including themselves, to facilitate accumulation of SSB repair factors at the break site4,5,6,7,8,9,10,11,12. ARTs have also been implicated in regulating DNA double strand break (DSB) repair by homologous recombination (HR) and non-homologous end-joining (NHEJ). PARP1 is required for HR-mediated restart of damaged and/or stalled replication forks13,14,15. It also promotes alternative-NHEJ (alt-NHEJ), an end-joining pathway activated in the absence of core NHEJ factors16,17,18,19. Whilst PARP1 has been implicated in regulating core NHEJ20,21, PARP3 promotes this repair pathway by facilitating accumulation of APLF and Ku at damage sites22,23,24,25,26.
Whilst the role of ARTs in DNA repair is well established, the identity of proteins ADP-ribosylated in response to DNA damage is less clear. Recent advances in mass spectrometry have begun to define the ADP-ribosylome27,28,29,30,31,32. However, the specific sites on these proteins modified in vivo and the ARTs that catalyse these events are largely unknown. This situation is exemplified by histones33. A fraction of all core histones isolated from cell nuclei are ADP-ribosylated34,35,36,37,38 and H2B/H3 are the principal histones modified in response DNA alkylating agents39,40. However, whether a similar pattern of histone ADP-ribosylation occurs in response to other varieties of DNA damage is unclear. Moreover, although the sites on histones modified by PARP1 have been established in vitro41, which sites on histones are ADP-ribosylated in response to DNA damage in vivo and the ARTs that catalyse these events are unknown. This, in part, is due to the lack of an appropriate eukaryotic experimental platform in which ARTs and histone genes can be manipulated to characterise ADP-ribosylation sites identified by biochemical or MS analysis. For example, high copy number arrays of histone genes in vertebrates makes it difficult to genetically manipulate these loci to assess how specific histone ADP-ribosylation events regulate a variety of cellular processes. This issue is compounded by the absence of ARTs in invertebrate models where histones genes are amenable to genetic manipulation, precluding an analysis of these pathways in these experimental systems.
Given these considerations, there is a call for a genetically tractable eukaryotic model that exhibits conservation of ARTs and key vertebrate DNA repair pathway components, in addition to single copy histone genes amenable to gene disruption and replacement strategies. All of these criteria are uniquely met in the eukaryotic amoeba Dictyostelium. Importantly, this organism has diverged less from humans than other invertebrates42. This is particularly striking with regards to DNA repair and several key human DNA repair pathway components are conserved in this organism that are absent in other models commonly used to study these pathways43,44,45,46,47,48,49,50,51. Importantly, the Dictyostelium genome contains 15 proteins with predicted ART catalytic domains47 and similar to vertebrates, two (Adprt1b and Adprt2) confer tolerance of cells to SSBs43,52. Moreover, in parallel with studies in mammalian cells22,24,26, we identified a further ART (Adprt1a) that whilst dispensable for SSBR, is required to promote NHEJ by facilitating accumulation of Ku at DSBs through a PAR interaction domain (PID) located in Ku7043,47.
Dictyostelium is ideally suited to assess how post-translational modification of histone proteins regulates a variety of cellular processes. Although, Dictyostelium possess the standard core histones, they exhibit a greater diversity of histone variants than budding yeast whilst retaining a simpler complement than metazoa, facilitating their analysis by both biochemical and genetic approaches53,54,55. The major post-translational modifications on histone N-terminal tails, including phosphorylation, acetylation and methylation, in addition the enzymes that catalyse these reactions, are also conserved in Dictyostelium53,55,56,57,58. Critically, in contrast to vertebrates, Dictyostelium contains single copies of most histone genes53,55, making their genetic manipulation relatively straightforward. Here we exploit the ability to manipulate histone genes in Dictyostelium, in addition to the conservation of ARTs in this organism, to assess DNA damage-induced site-specific ADP-ribosylation events on histones in vivo and define the ARTs that mediate these modifications. These data demonstrate, for the first time, mapping and confirmation of DNA damage induced histone ADP-ribosylation sites by a specific ARTs in vivo and will provide a unique experimental platform to assess the role of histone ADP-ribosylation in DNA repair and other cellular process.
Results
ADP-ribosylation of Dictyostelium histone proteins in response to DNA DSBs
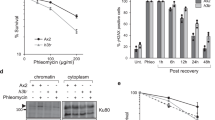
Previously, we identified that nuclear ADP-ribosylation is required for effective NHEJ in Dictyostelium and that this is dependent on the ARTs Adprt1a and Adprt243. To address which chromatin-associated proteins are targeted by these enzymes following DSBs, chromatin fractions were prepared from Dictyostelium Ax2 cells following DSB induction by phleomycin and ADP-ribosylation assessed by western blotting using an antibody that recognises PARylated proteins. PARylation of chromatin-associated proteins is induced in a time-dependent manner in Ax2 cells exposed to phleomycin (Fig. 1A) and several of these proteins migrate with molecular weights similar to histones (15–25 kDa). Our previous work illustrated that nuclear ADP-ribosylation is dependent on both Adprt1a and Adprt243. Therefore, we considered whether these ARTs are required for DSB-induced ADP-ribosylation of chromatin-associated proteins within the 15–25 kDa range. Similar to nuclear ADP-ribosylation43, disruption of Adprt1a or Adprt2 alone reduced ADPr of these proteins (data not shown) whilst these events were almost completely lost in the adprt1a−adprt2− strain (Fig. 1B).
(A) Ax2 cells were left untreated or exposed to 300 μg/ml phleomycin for the indicated times. Chromatin fractions were prepared and western blotting performed using the indicated antibodies. (B) Ax2 or adprt1a−adprt2− cells were left untreated or exposed to 300 μg/ml phleomycin for 60 minutes as indicated. Chromatin fractions were prepared and western blotting performed using the indicated antibodies. (C) Ax2 cells or the adprt1a−adprt2− strain were exposed to phleomycin for the times indicated. Histone-enriched acid extracts were prepared and western blotting performed with antibodies as indicated.
Next, we considered whether the PARylated proteins of between 15–25 kDa present in chromatin fractions following DSB-induction represent histone proteins. Consistent with this hypothesis, several PARylated polypeptides with molecular weights of between 15–25 kDa co-purify with histones during an optimised acid-extraction protocol developed to enrich Dictyostelium histones from vegetative cells (Fig. 1C)55. Similar to chromatin fractions, these proteins are absent in acid extracts prepared from adprt1a−adprt2− cells, indicating these modifications are dependent on Adprt1a and Adprt2. Taken together, these data indicate that several chromatin-associated proteins are PARylated in response to DSBs in an Adprt1a/Adprt2-dependent manner and that a subset of these proteins likely represents histones.
Dictyostelium H2B is ADP-ribosylated in response to DNA DSBs in vivo
Similar to Dictyostelium Adprt1a, human PARP3 promotes DSB repair by facilitating accumulation of NHEJ factors at damage sites22,23,24,25,26. All core histones are modified by PARP3 in vitro, although in the context of reconstituted nucleosomes H2B is the major histone modified by this ART59. Whether PARP3 is responsible for DNA damage-induced H2B ADP-ribosylation in vivo is unknown. Given Adprt1a is the functional equivalent of PARP343,52, we considered whether H2B is ADP-ribosylated in response to DNA DSBs in vivo. H2Bv3 is the principal H2B variant expressed in vegetative Dictyostelium53,55. To test whether this histone is ADP-ribosylated in response to DNA DSBs in vivo, we expressed H2Bv3 with 3x tandem C-terminal Flag tags in Ax2 cells and monitored its ADP-ribosylation status following DNA DSBs. Inclusion of the epitope tags on H2Bv3 increases the molecular weight of the protein to distinguish it from endogenous H2Bv3 and other modified histones during SDS-PAGE analysis. Similar to previous observations using a PAR-specific antibody (Fig. 1C), several polypeptides of between 15–25 kDa are detected by a reagent that recognises both MARylated and PARylated proteins specifically in extracts prepared from cells following exposure to phleomycin (Fig. 2A). The majority of these modifications are independent of H2Bv3-Flag expression, most likely reflecting ADP-ribosylation of endogenous histone proteins (Fig. 2A). Strikingly, a further DSB-induced ADP-ribosylated protein is present only in cells expressing H2Bv3-Flag. Moreover, a protein of the predicted molecular weight of H2Bv3-Flag is detected specifically in Flag-immunoprecipitates from extracts prepared after exposure of H2Bv3-Flag expressing cells to phleomycin (Fig. 2B), confirming ADP-ribosylation of H2Bv3-Flag in response to DSBs. Whilst DSB-induced ADP-ribosylation of H2BV3-Flag is detectable using the reagent that recognises both MARylation and PARylation events, it is not recognised by a reagent that recognises only PARylated proteins (Fig. 2C). From these data we conclude that H2Bv3 is ADP-ribosylated in response to DNA DSBs in vivo primarily by MARylation.
(A) Ax2 cells expressing H2Bv3-Flag, or control cells containing empty vector were left untreated or exposed to 300 μg/ml of phleomycin for 1 hour. Histone-enriched acid extracts were prepared and western blotting performed with the indicated antibodies. ADP-ribosylated H2Bv3-Flag is indicated with an arrow. (B) Ax2 cells expressing H2Bv3-Flag, or control cells containing empty vector, were treated as in (A). Proteins were released from chromatin and immunoprecipitated with Flag antibodies. Western blotting was performed with antibodies as indicated. ADP-ribosylated H2Bv3-Flag is indicated with an arrow. (C) Ax2 cells expressing H2Bv3-Flag, or control cells containing empty vector were treated as in (A). Western blotting was performed using a reagent that recognises both poly- and mono-ADP-ribosylation (left), or a reagent that recognises only poly-ADP-ribosylated proteins (right). ADP-ribosylated H2Bv3-Flag is indicated with an arrow.
Adprt1a is a mono-ART that modifies the N-terminus of H2Bv3 in vitro
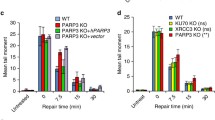
Although advances in mass spectrometry have begun to identify ADP-ribosylated proteins27,28,29,30,31,32, mapping site specific ADP-ribosylation events in vivo remains challenging. Given Adprt1a is the principle ART required for DSB-induced ADP-ribosylation43, we first considered the catalytic properties of Adprt1a in vitro and whether it is capable of modifying H2Bv3. It is well established that human DNA damage responsive ARTs are capable of auto-ADP-ribosylation in vitro and that this activity is activated by a variety of DNA strand breaks60. Similarly, we observe an ADP-ribosylated species that co-migrates with recombinant Adprt1a but not a catalytic-dead version of the protein (Adprt1acd) only in reactions that contain sheared salmon sperm DNA (Fig. 3A), indicating Adprt1a is activated by DNA to undergo auto-ADP-ribosylation in vitro. Given sheared DNA contains different DNA structures, most notably a variety of DNA single and double strand breaks, we next considered whether Adprt1a is activated by a specific variety of DNA strand break. Similar to human PARP361,62, Adprt1a is effectively activated by DNA DSBs and SSBs and enzyme activity increases in proportion to the number of breaks present in reactions (Fig. 3B). We also established whether Adprt1a auto-ribosylation represents MARylation or PARylation events. PARG removes Poly-ADP-ribose polymers from proteins63,64, whilst MacroD1 removes mono-ADP-ribose65,66,67. Incubation of auto-ADP-ribosylated Adprt1a with PARG does not significantly impact of the levels of auto-ribosylated Adprt1a, whilst inclusion of MacroD1 reduces levels of ADP-ribosylated Adprt1a (Fig. 3C). From these data we concluded that Adprt1a is a mono-ART that is activated by DNA strand breaks in vitro.
(A) Wild-type or catalytic dead (Adprt1acd) His-tagged Adprt1a was expressed and purified from bacteria (left) and employed in ADP-ribosylation assays using 32P-labeled NAD+ in the absence or presence of sheared salmon sperm DNA (sssDNA). Following SDS-PAGE, ADP-ribosylated proteins were detected by auto-radiography. (B) ADP-ribosylation assays were performed using His-Adprt1a as in (A) with the exception that pUC19 DNA either uncut, or cut with enzymes to produce the indicated numbers of breaks, was employed in assays instead of sssDNA. (C) His-Adprt1a (right) and human PARP1 (left) were auto-ribosylated by incubation with biotin-NAD+ prior to incubation with increasing concentrations of MacroD1 or PARG as indicated. ADP-ribosylated proteins were detected by Western blotting with Streptavidin conjugated HRP. (D) His-H2Bv3 or His-H2Bv3 lacking the N-terminal 25 amino acids (H2Bv3Δ25) were expressed and purified from bacteria (upper panel). ADP-ribosylation reactions were performed with Adprt1a in the presence of sssDNA as indicated. ADP-ribosylated proteins were detected as in (C). (E) Sequence alignment of the N-terminal 24 amino acids of Dictyostelium H2Bv3 with the equivalent regions of H2B from a variety of species. Identical amino acids are highlighted and potential ADP-ribosylation sites in the Dictyostelium protein indicated (*). (F) Wild-type His-H2Bv3, or His-H2Bv3 mutated at E18, E19, or both E18 and E19 were expressed and purified from bacteria (upper panel). Proteins were employed in ADP-ribosylation reactions as in (D).
Next, we established whether H2Bv3 is modified by Adprt1a in vitro and at which sites. Given human PARP1 is able to modify the N-terminus of all core histones in vitro41, we first tested whether full length H2Bv3, or H2Bv3 lacking the N-terminal 25 amino acids of the protein, are modified by Adprt1a (Fig. 3D). Full-length H2Bv3 is effectively ADP-ribosylated by Adprt1a in vitro and this is dependent on DNA and Adprt1a catalytic activity (Appendix Figure S1). Deletion of the N-terminal 25 amino acids of H2Bv3 almost completely ablates this modification (Fig. 3D), indicating the major ADP-ribose acceptor sites reside in this region of the protein. Although ARTs modify proteins at aspartate, glutamate, arginine, lysine, cysteine and asparagine amino acids1, the N-terminal 25 amino acids of Dictyostelium H2Bv3 contain only 5xLys and 2xGlu amino acids that are potential ADP-ribose acceptor sites (Fig. 3E). Given MacroD1 removes ADP-ribose moieties from glutamates65,66,67, and that this protein reverses Adprt1a catalysed ADP-ribosylation events (Fig. 3C), we hypothesised that E18 and/or E19 are modified by this enzyme. Therefore, we expressed and purified H2Bv3 from bacteria in which E18 and E19 have been mutated to Ala either alone or in combination, and assessed their ability to undergo Adprt1a-mediated ADP-ribosylation in vitro. Mutation of E18 of H2Bv3 has only a slight impact on Adprt1a-mediated ADP-ribosylation, whilst mutation of E19 increases the ability of Adprt1a to modify H2Bv3 (Fig. 3F). However, mutation of both E18 and E19 in combination dramatically reduces Adprt1a-mediated ADP-ribosylation of H2Bv3. These data demonstrate that whilst E18 of H2Bv3 is ADP-ribosylated by Adprt1a in vitro, when this site is lost E19 is instead modified.
E18 and E19 of H2Bv3 are ADP-ribosylated in response to DNA DSBs in vivo
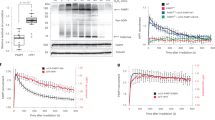
We next assessed whether the ability of Adprt1a to modify H2Bv3 at E18 and E19 in vitro is reflected in these amino acids being targets for DSB-induced ADP-ribosylation in vivo. In contrast to vertebrates, Dictyostelium contain single copies of all histone genes, with the exception of H4, of which there are two copies53,55. Therefore, we exploited gene replacement technology to manipulate the endogenous h2bv3 locus at the ADP-ribosylation sites identified in vitro (Fig. 4A and Appendix Figure S2) and assessed the impact of these mutations on DSB-induced H2Bv3 ADP-ribosylation. Accordingly, a gene replacement vector was designed containing the h2b gene that whilst under the control of its own promoter, contains E18A and E19A mutations (h2bv3E18AE19A). Given this strategy results in strains containing some residual vector sequence, as a control we generated an equivalent strain, but with a wild-type version of the h2Bv3 gene replacing the endogenous locus (h2bv3wt).
(A) Gene replacement strategy to replace the endogenous h2b locus with either wild-type h2b (h2bwt), or h2b containing E18AE19A mutations (h2bE18AE19A). (B and C) Two independent h2bwt and h2bE18AE19A strains were left untreated or exposed to 300 μg/ml phleomycin for 60 minutes as indicated. Histone-enriched acid extracts were prepared and western blotting performed using the indicated antibodies. Arrow heads indicate the position of ADP-ribosylated H2B protein in h2bwt cells. (D) Wild-type H2Bv3-Flag or H2Bv3-Flag with the indicated mutations were expressed in Ax2 cells. Whole cell or chromatin extracts were prepared and western blotting performed with the indicated antibodies. (E) Ax2 cells expressing wild-type H2Bv3-Flag or the indicated H2Bv3-Flag mutants were left untreated or exposed to 300 μg/ml of phleomycin for 1 hour as indicated. Histone-enriched acid extracts were prepared and western blotting performed with the indicated antibodies. ADP-ribosylated H2Bv3-Flag is indicated with an arrow.
The resulting h2bv3wt and h2bv3E18AE19A strains were exposed to phleomycin and ADP-ribosylation assessed using a reagent that detects both MARylation and PARylation. Using high resolution SDS-PAGE, three protein species of between 15–25 kDa are robustly ADP-ribosylated in response to DSBs in two independent h2bv3wt strains (Fig. 4B and C). The intensity of the middle of these three bands, which migrates with a similar molecular weight to H2Bv3, is dramatically reduced in two independent h2bv3E18AE19A strains, indicating loss of H2Bv3 ADP-ribosylation. Whilst these data clearly reveal that H2Bv3 is ADP-ribosylated at E18/E19 in vivo, they are unable to distinguish whether there is a preference for either E18 or E19 to be ADP-ribosylated in response to DSBs. To address this question we generated Ax2 strains that express H2Bv3-Flag containing E18A, E19A or E18AE19A mutations and assessed the ability of these proteins to be ADP-ribosylated in response to DSBs in vivo. All H2Bv3-Flag variants are expressed at similar levels and effectively incorporated into chromatin (Fig. 4D). Consistent with previous observations (Fig. 2), we observe robust DNA damage-induced ADP-ribosylation of a band in H2Bv3-Flag expressing cells that is absent from cells that do not express the protein, confirming the identity of this band as H2Bv3-Flag (Fig. 4E). Whilst the E19A mutation has little impact on ADP-ribosylation status of this protein, the E18A mutation slightly reduces its modification. In contrast, mutation of both E18 and E19 dramatically reduces ADP-ribosylation of H2Bv3-Flag (Fig. 4E). Taken together, these data indicate that similar to our observations in vitro, E18 is the preferred site on H2Bv3 ADP-ribosylated in response to DNA DSBs in vivo, although in its absence E19 is also able to be modified. Moreover, the almost complete loss of H2Bv3 ADP-ribosylation following mutation of E18 and E19 indicate no other sites on the protein are robustly ADP-ribosylated following DSBs.
Discussion
Dictyostelium Adprt1a mediates DSB-induced ADP-ribosylation to promote NHEJ43. Similarly, human PARP3 has also been implicated in regulating NHEJ by facilitating accumulation of APLF and Ku at damage sites24,26. However, the substrates modified by Adprt1a or PARP3 in vivo are unknown. The substrates targeted by other ARTs are similarly unclear. For example, recent advances in mass spectrometry have identified a number of proteins, including histones that are ADP-ribosylated in response to genotoxic agents31,32. Although modification of these substrates is disrupted by PARP inhibitors, these compounds target multiple ARTs68, making it difficult to attribute these events to a specific ART(s). Our data indicate histones are likely targets for ADP-ribosylation in response to DNA DSBs and we demonstrate that H2B is modified at E18 and E19 in response to this variety of DNA damage. We observe co-fractionation of several ADP-ribosylated proteins with histones during an optimised acid extraction procedure, and through manipulation of the endogenous h2b locus, identify that one of these modifications represents ADP-ribosylation of H2B at E18/E19. Given ADP-ribosylation of acid extracted histones is dependent on Adprt1a and Adprt2; we concluded that modification of H2B at E18/E19 in response to DSBs is dependent on these ARTs.
Histones are known targets for ADP-ribosylation. All core histones, in addition to H1, are modified in response to DNA alkylating agents39,40,69. However, whether similar patterns of histone ADP-ribosylation are observed in response to other DNA damage types and the sites modified is unclear. We only observe robust ADP-ribosylation of H2B at E18/E19 following exposure of cells to DNA DSBs, suggesting little modification of this site in the absence of genotoxic stress. Isolation of H2B from rat nuclei following incubation with radio-labelled NAD + identified E2 of H2B as an ADP-ribosylation site, although these experiments were performed in the absence of DNA damaging agents37. It is tempting to speculate that whilst E18 and E19 of Dictyostelium H2B are not absolutely conserved in vertebrates (Fig. 3E), E2 of human H2B might be analogous to these sites. Given the susceptibility of inducing DNA damage and ADP-ribosylation during cell extraction procedures31, it is possible the ADP-ribosylation of rat H2B observed previously is a reflection of DNA damage induced during sample preparation, or low levels of endogenous DNA damage present in cells in the absence of genotoxins. In further support of E2 of human H2B being equivalent to the ADP-ribosylation sites identified here, PARP3 specifically modifies H2B at this site in reconstituted nucleosomes in vitro62. Together, these observation may indicate that ADP-ribosylation sites do not have to be absolutely conserved to be functionally important, but rather reside in equivalent regions of the protein. Interestingly, we observe that disruption of E19 increases ADP-ribosylation of H2B by Adprt1a in vitro, presumably at E18 given the E18AE19A mutant is not significantly modified in these assays (Fig. 3F). Whilst it is interesting to speculate that E19 may quench ADP-ribosylation of H2B at E18, this observation is not apparent when assessing the ADP-ribosylation status of the equivalent mutants in vivo (Fig. 4E). Nevertheless, both in vitro and in vivo assays reveal that whilst mutation of E18 reduces ADP-ribosylation of H2B by Adprt1a, a significant reduction in its modification status is only observed when E18 and E19 are mutated in combination (Figs 3F and 4E). These observations underscore redundancy and further plasticity in ADP-ribosylation events that has implications regarding the design of protein mutations to study these modifications.
In further support of Adprt1a directly modifying H2B at E18/E19, we provide evidence that H2B is MARylated at E18/E19 in response to DNA DSBs. It is interesting to note, however, that a number of polypeptides in histones preparations are ADP-ribosylated following DNA DSBs in the h2bE18AE19A strains, suggesting that similar to other organisms several Dictyostelium histones other than H2B are ADP-ribosylated in response to DNA damage. ADP-ribosylation promotes the enrichment of chromatin remodelling and repair factors at DNA lesions through ADP-ribose interaction domains70. Additionally, different ADP-ribose interaction domains recognise different modification types. For example, whilst the PBZ domain recognises PAR chains, the macro domain is able to recognise MARylated proteins70,71. To date, it is unknown whether DNA damage induces MARylation or PARylation of histones, or a combination of the two at different sites to influence differential recruitment of repair factors to DNA lesions. In this regard, although Adprt1a and the PBZ domain of Ku70 are required for accumulation of Ku at DSBs and efficient NHEJ43, we do not observe sensitivity of the h2bE18AE19A mutants to DSBs during spore germination (Appendix Figure S3), a stage of the Dictyostelium life cycle where cells become reliant on this repair pathway46. Therefore, it will be important to establish whether ADP-ribosylation of different histone sites, or with a specific type of ADP-ribosylation event, is a determining factor in influencing which factors are recruited to a particular type of DNA lesion. The availability of Dictyostelium as a model in which histone ADP-ribosylation sites can be manipulated will help resolve these questions.
Materials and Methods
Cell culture and strain generation
Dictyostelium cells were grown according to standard procedures, either axenically or on SM agar plates in association with Klebsiella aerogenes. The adprt1a−, adprt2− and adprt1a−adprt2− strains have been previously described43.
To construct the targeting vector for h2bv3 gene replacement, h2bv3 coding sequence, in addition to regions flanking the gene (Chromosome 4; position 453224 to 456713), was amplified by PCR from AX2 genomic DNA using the primers 5′-CTTATACGATCGACTGATGCTGTAACAATAG-3′ and 5′-CCATATCATGGTGGATATTACCATGGTC-3′. The amplified fragment was first subcloned into pJET 1.2 (Thermo Scientific). The selection marker (BSR cassette) was excised from pLPBLP72 using SmaI, and inserted to a BsrGI site in the 3′ non-coding region downstream of h2bv3 gene. To eliminate unnecessary restriction sites of pJET 1.2 vector, the whole h2bv3 gene and flanking regions, including the selection marker, was cut using BspHI, treated by Klenow fragment to fill in ends, and then cut with NotI. This fragment was then inserted into pBluescriptII SK- (Agilent Technologies) at HindIII (treated by Klenow fragment), and NotI sites.
To introduce E18 and E19 to A mutations into the targeting vector, a short fragment containing 5′ non-coding region, and 5′ sequences of h2bv3 coding region was amplified by PCR using 5′-CAATAAAsATTACACTCCTAA-3′ and 5′-CAGAAGCTTCGGTGGCGATTCTGT-3′ primers, and then subcloned into pJET 1.2 vector. E18 and E19 to A mutation was introduced by QuickChange method (Agilent Technologies) using a 5′-CAAAGGTTCAACTCAATCCGGAGCAGCGAAAACCGCTTCAACCAC-3′ primer, and corresponding anti-sense primer. The mutagenesis primer also carried a BspEI restriction enzyme site introduced into the h2bv3 gene without changing its amino acid sequence to facilitate screening of mutants. The short AfeI and HindIII fragment carrying the mutation was used to replace the corresponding sequence in the targeting vector.
Both targeting vectors carrying wildtype and mutated h2bv3, were then transfected to AX2 cells by electroporation and selected under 10 μg/ml blasticidin for two weeks. Blasticidin resistant clones were isolated on a bacterial lawn. Gene replacement by targeting vectors was verified by PCR. Introduction of mutations was also verified by cutting PCR fragment with BspEI. The genomic region surrounding the h2bv3 locus was sequenced to confirm mutations.
To express H2Bv3-Flag in Dictyostelium the vector pDBSrH2B-3XFlag, which drives C-terminally tagged H2Bv3 from the constitutive actin 15 promoter on an extrachromosomal vector, was transfected into Ax2 cells and pools of blasticidin resistant colonies analysed. Mutation of E18A, E19A or both were introduced by replacing the coding sequence of the N-terminal region of H2Bv3 following PCR amplification of the sequences including the relevant mutations generated in pQE30 (see below).
Subcellular fractionation and immunoprecipitation
Chromatin enriched fractions and whole cell extracts were prepared as previously described43. Histone enriched acid extracts were prepared as described previously55. Briefly, untreated or Phleomycin-treated cells were washed in ice-cold KK2 buffer, and then resuspended in nuclear extraction buffer (50 mM Tris pH8.0, 10 mM NaCl, 3 mM MgCl2, 3 mM CaCl2, 0.5 M sorbitol, 0.6% Triton X 100, Complete protease inhibitor cocktail (Roche), phosphatase inhibitor cocktail 2 and 3 (Sigma), 10 μM Benzamide, and 200 μM DEA) at a cell density of 1 × 108/ml, incubated with rotation at 4 °C for 15 minutes, and the nuclei pelleted by centrifugation at 2300 xg for 5 minutes. Extracted nuclei were resuspended in nuclear extraction buffer containing 4 M urea, and 2% β-mercaptoethanol, agitated at 4 °C for 15 minutes, and pelleted again by centrifuging at 2300 × g for 5 minutes. Isolated nuclei were then resuspended in 0.4 N HCl at a density of 5 × 108/ml, and mixed overnight by rotation at 4 °C. Acid extracted histones were harvested by centrifugation at 16000 xg, and the supernatant precipitated by addition of x6.5 volume of acetone, incubating at −20 °C for 2 hours, and centrifuging at 16000xg for 15 minutes at 4 °C. The pellet was washed twice with ice-cold acetone, dried, and resuspended in 8 M Urea supplemented with 5% beta-mercaptoethanol for further analysis.
For immunoprecipitation, chromatin extracts from cells expressing Flag-tagged H2Bv3 were boiled in 2xSDS sample buffer (100 mM Tris-HCl pH6.8, 4% SDS) to disrupt protein-protein and protein-DNA interactions. Samples were diluted 10 times with IP buffer (50 mM Tris pH8, 200 mM NaCl, 1 mM EDTA, 1 mM DTT, 10% glycerol, 1% triton X-100, phosphatase inhibitor cocktail 2,3 [Roche], 200 μM DEA, 500 μM benzamide, protease inhibitor cocktail tablet [Roche]). Anti-Flag M2 beads (Sigma) were added and the mixture incubated on a rotating wheel for 1 h at 4 °C. Beads were collected by centrifugation band washed 3 times in IP buffer followed by Western blot analysis.
Sensitivity Assays
Sensitivity assays using germinated spores were performed as described previously46. In brief, fruiting bodies were suspended in KK2 containing 0.1% NP-40, and then passed through 19.5-gauge needle to liberate spores. Spores were washed twice by KK2 + 0.1% NP-40, and resuspended in KK2 at the density of 2 × 107 cells/ml. Germination of spores was induced by heat shock at 45 °C for 30 min. Hatched spores were then diluted to 106 cells/ml in a 1:5 ratio of HL5/KK2, and incubated in shaking suspension at 100 rpm for 18 hrs at 22 °C with the concentrations of phleomycin as indicated. 250 cells were plated onto three 140mm Petri dishes with K. aerogenes in association with SM agar, and incubated at 22 °C. Colonies on Petri dishes were counted after 3 to 5 days, and cell survival calculated by normalizing with the number of colonies without phleomycin treatment.
Antibodies
Full length His-tagged H2Bv3 was purified from bacteria under denaturing conditions of 8 M Urea using QIAexpressionist protein purification kit (QIAGEN). Purified protein used to inoculate sheep (Diagnostics Scotland, Edinburgh, Scotland) to produce anti-Dictyostelium H2Bv3 anti-serum.
Other antibodies employed in this study are PAR polyclonal antibody (Trevigen; 4336-BPC-100), poly-ADP-ribose binding reagent (Millipore; MABE1031), pan-ADP-ribose binding reagent (Millipore; MABE1016), histone H3 (Abcam; ab1791), γH2AX (Abcam; ab11174), Myc 9E10 (Santa Cruz; sc-40), actin C-11 (Santa Cruz; sc-1615), FLAG M2 (Sigma; F1804), penta-His (Qiagen; 34660), DdKu8043.
Protein expression and purification
H2Bv3 was cloned into pQE30 expression vector to generate N-terminally tagged His6-H2Bv3 and mutation of E18, E19 or both to alanine introduced by site-directed mutagenesis (QuikChange, Agilent Technologies). The primers used to introduce the mutations were ACTCAATCTGGTGCAGAGAAAACCGCTTCA (E18A), ACTCAATCTGGTGAAGCGAAAACCG (E19A) and ACTCAATCTGGTGCAGCGAAAACCG (E18AE19A). Truncation of the first 25 amino acids of H2Bv3 was performed by PCR amplification of the coding sequence without the first 25 amino acids using primer CCAGGATCCATGACCCCAAAAGTAACCAAAACC and reinserting, in frame, into pQE30. All constructs were verified by DNA sequencing.
Expression constructs were introduced into E. coli M15 and expression induced with 0.1 mM IPTG for 4 h at 22 °C. Cells were then harvested and resuspended in ice-cold column buffer (50 mM Tris-HCl pH8, 150 mM NaCl, 10% glycerol, 30 mM Imidazole, protease inhibitor cocktail tablet [Roche] and 1 mM PMSF [Sigma]). Cell lysis was performed by sonication. Soluble proteins were then collected following centrifugation of lysates at 15,000 rpm for 30 min and loaded onto a column containing prewashed Ni-NTA beads (Qiagen). After washing with 30 mM, 50 mM and 100 mM Imidazole, H2Bv3 was eluted using column buffer with 200 mM Imidazole, then dialyzed against storage buffer (50 mM Tris-HCl pH8, 10% glycerol), snap frozen in liquid nitrogen and stored at −80 °C.
To generate the catalytic-dead version of Adprt1a (Adprt1acd), H789 and Y823A in the conserved NAD + binding pocket of the catalytic domain were mutated to alanine by the QuickChange method (Agilent Technologies). His-tagged Adprt1a and Adprt1acd were purified in a similar way to other proteins (see above), although cell lysis was performed using Bugbuster (Merck) to avoid genomic DNA contamination. IMAC buffer (50 mM HEPES pH8, 500 mM NaCl, 10% glycerol, 1 mM DTT, 1 mM Imidazole) was used instead of the column buffer.
ADP-ribosylation assays
Recombinant His-tagged Adprt1a (100 ng) was mixed with 1.5 mM NAD+, 250 μM Biotinylated NAD+ (Trevigen) in 50 mM Tris-HCl pH8 and 2 mM MgCl2 either with or without 100 μg/ml of sheared salmon sperm DNA. Recombinant His-tagged Dictyostelium histone (100 ng) was added where required and the reaction performed at room temperature for 30 mins before termination by addition of 2xSDS loading buffer and boiling for 5 mins. Samples were resolved by SDS-PAGE and transferred to PVDF membrane and the ADP-ribosylation signal detected using HRP-conjugated Streptavidin, followed by ECL. For assays that employed radioactive NAD + where indicated, 100 nM 32P-labeled NAD+ was used instead of Biotinylated NAD+. The ADP-ribosylation of proteins was assessed by separating samples on SDS-PAGE gel, vacuum-drying and exposing to X-ray film.
For de-ADP-ribosylation assays, ADP-ribosylation reactions using Biotinylated NAD+ were terminated by adding 10 mM 3-aminobenzamide. Then, 3 μl of 2.4 μM MacroD1 protein or 9 nM PARG enzyme (Trevigen) were added to the reaction either undiluted or diluted 1:5 and 1:10. Mixtures were incubated at 37 °C for 30 mins and the reaction stopped by adding 2xSDS loading buffer and boiling for 5 min. The ADP-ribosylation status was then assessed by Western blot.
Additional Information
How to cite this article: Rakhimova, A. et al. Site-specific ADP-ribosylation of histone H2B in response to DNA double strand breaks. Sci. Rep. 7, 43750; doi: 10.1038/srep43750 (2017).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Hottiger, M. O., Hassa, P. O., Luscher, B., Schuler, H. & Koch-Nolte, F. Toward a unified nomenclature for mammalian ADP-ribosyltransferases. Trends Biochem Sci 35, 208–219, doi: 10.1016/j.tibs.2009.12.003 (2010).
Vyas, S. et al. Family-wide analysis of poly(ADP-ribose) polymerase activity. Nat Commun 5, 4426, doi: 10.1038/ncomms5426 (2014).
Gibson, B. A. & Kraus, W. L. New insights into the molecular and cellular functions of poly(ADP-ribose) and PARPs. Nat Rev Mol Cell Biol 13, 411–424, doi: 10.1038/nrm3376 (2012).
Caldecott, K. W. Single-strand break repair and genetic disease. Nat Rev Genet 9, 619–631, doi: 10.1038/nrg2380 (2008).
Krishnakumar, R. & Kraus, W. L. The PARP side of the nucleus: molecular actions, physiological outcomes, and clinical targets. Mol Cell 39, 8–24, doi: 10.1016/j.molcel.2010.06.017 (2010).
El-Khamisy, S. F., Masutani, M., Suzuki, H. & Caldecott, K. W. A requirement for PARP-1 for the assembly or stability of XRCC1 nuclear foci at sites of oxidative DNA damage. Nucleic Acids Res 31, 5526–5533 (2003).
Masson, M. et al. XRCC1 is specifically associated with poly(ADP-ribose) polymerase and negatively regulates its activity following DNA damage. Mol. Cell. Biol. 18, 3563–3571 (1998).
Okano, S., Lan, L., Caldecott, K. W., Mori, T. & Yasui, A. Spatial and temporal cellular responses to single-strand breaks in human cells. Mol Cell Biol 23, 3974–3981 (2003).
Bekker-Jensen, S. et al. Human Xip1 (C2orf13) is a novel regulator of cellular responses to DNA strand breaks. J Biol Chem 282, 19638–19643, doi: 10.1074/jbc.C700060200 (2007).
Kanno, S. et al. A novel human AP endonuclease with conserved zinc-finger-like motifs involved in DNA strand break responses. Embo J 26, 2094–2103, doi: 10.1038/sj.emboj.7601663 (2007).
Rulten, S. L., Cortes-Ledesma, F., Guo, L., Iles, N. J. & Caldecott, K. W. APLF (C2orf13) is a novel component of poly(ADP-ribose) signaling in mammalian cells. Mol Cell Biol 28, 4620–4628, doi: 10.1128/MCB.02243-07 (2008).
Iles, N., Rulten, S., El-Khamisy, S. F. & Caldecott, K. W. APLF (C2orf13) is a novel human protein involved in the cellular response to chromosomal DNA strand breaks. Mol Cell Biol 27, 3793–3803, doi: 10.1128/MCB.02269-06 (2007).
Bryant, H. E. et al. PARP is activated at stalled forks to mediate Mre11-dependent replication restart and recombination. Embo J 28, 2601–2615, doi: 10.1038/emboj.2009.206 (2009).
Sugimura, K., Takebayashi, S., Taguchi, H., Takeda, S. & Okumura, K. PARP-1 ensures regulation of replication fork progression by homologous recombination on damaged DNA. J Cell Biol 183, 1203–1212, doi: 10.1083/jcb.200806068 (2008).
Yang, Y. G., Cortes, U., Patnaik, S., Jasin, M. & Wang, Z. Q. Ablation of PARP-1 does not interfere with the repair of DNA double-strand breaks, but compromises the reactivation of stalled replication forks. Oncogene 23, 3872–3882, doi: 10.1038/sj.onc.1207491 (2004).
Audebert, M., Salles, B. & Calsou, P. Involvement of poly(ADP-ribose) polymerase-1 and XRCC1/DNA ligase III in an alternative route for DNA double-strand breaks rejoining. J Biol Chem 279, 55117–55126, doi: 10.1074/jbc.M404524200 (2004).
Robert, I., Dantzer, F. & Reina-San-Martin, B. Parp1 facilitates alternative NHEJ, whereas Parp2 suppresses IgH/c-myc translocations during immunoglobulin class switch recombination. J Exp Med 206, 1047–1056, doi: 10.1084/jem.20082468 (2009).
Wang, M. et al. PARP-1 and Ku compete for repair of DNA double strand breaks by distinct NHEJ pathways. Nucleic Acids Res 34, 6170–6182, doi: 10.1093/nar/gkl840 (2006).
Brown, M. L., Franco, D., Burkle, A. & Chang, Y. Role of poly(ADP-ribosyl)ation in DNA-PKcs- independent V(D)J recombination. Proc Natl Acad Sci USA 99, 4532–4537, doi: 10.1073/pnas.072495299 (2002).
Luijsterburg, M. S. et al. PARP1 Links CHD2-Mediated Chromatin Expansion and H3.3 Deposition to DNA Repair by Non-homologous End-Joining. Mol Cell 61, 547–562, doi: 10.1016/j.molcel.2016.01.019 (2016).
Pines, A., Mullenders, L. H., van Attikum, H. & Luijsterburg, M. S. Touching base with PARPs: moonlighting in the repair of UV lesions and double-strand breaks. Trends Biochem Sci 38, 321–330, doi: 10.1016/j.tibs.2013.03.002 (2013).
Boehler, C. et al. Poly(ADP-ribose) polymerase 3 (PARP3), a newcomer in cellular response to DNA damage and mitotic progression. Proc Natl Acad Sci USA 108, 2783–2788, doi: 10.1073/pnas.1016574108 (2011).
Loseva, O. et al. PARP-3 is a mono-ADP-ribosylase that activates PARP-1 in the absence of DNA. J Biol Chem 285, 8054–8060, doi: 10.1074/jbc.M109.077834 (2010).
Rulten, S. L. et al. PARP-3 and APLF function together to accelerate nonhomologous end-joining. Mol Cell 41, 33–45, doi: 10.1016/j.molcel.2010.12.006 (2011).
Fenton, A. L., Shirodkar, P., Macrae, C. J., Meng, L. & Koch, C. A. The PARP3- and ATM-dependent phosphorylation of APLF facilitates DNA double-strand break repair. Nucleic Acids Res 41, 4080–4092, doi: 10.1093/nar/gkt134 (2013).
Beck, C. et al. PARP3 affects the relative contribution of homologous recombination and nonhomologous end-joining pathways. Nucleic Acids Res 42, 5616–5632, doi: 10.1093/nar/gku174 (2014).
Daniels, C. M., Ong, S. E. & Leung, A. K. Phosphoproteomic approach to characterize protein mono- and poly(ADP-ribosyl)ation sites from cells. J Proteome Res 13, 3510–3522, doi: 10.1021/pr401032q (2014).
Daniels, C. M., Ong, S. E. & Leung, A. K. The Promise of Proteomics for the Study of ADP-Ribosylation. Mol Cell 58, 911–924, doi: 10.1016/j.molcel.2015.06.012 (2015).
Gagne, J. P. et al. Proteome-wide identification of poly(ADP-ribose) binding proteins and poly(ADP-ribose)-associated protein complexes. Nucleic Acids Res 36, 6959–6976, doi: 10.1093/nar/gkn771 (2008).
Isabelle, M., Gagne, J. P., Gallouzi, I. E. & Poirier, G. G. Quantitative proteomics and dynamic imaging reveal that G3BP-mediated stress granule assembly is poly(ADP-ribose)-dependent following exposure to MNNG-induced DNA alkylation. J Cell Sci 125, 4555–4566, doi: 10.1242/jcs.106963 (2012).
Jungmichel, S. et al. Proteome-wide identification of poly(ADP-Ribosyl)ation targets in different genotoxic stress responses. Mol Cell 52, 272–285, doi: 10.1016/j.molcel.2013.08.026 (2013).
Zhang, Y., Wang, J., Ding, M. & Yu, Y. Site-specific characterization of the Asp- and Glu-ADP-ribosylated proteome. Nat Methods 10, 981–984, doi: 10.1038/nmeth.2603 (2013).
Messner, S. & Hottiger, M. O. Histone ADP-ribosylation in DNA repair, replication and transcription. Trends Cell Biol 21, 534–542, doi: 10.1016/j.tcb.2011.06.001 (2011).
Burzio, L. O., Riquelme, P. T. & Koide, S. S. ADP ribosylation of rat liver nucleosomal core histones. J Biol Chem 254, 3029–3037 (1979).
Jump, D. B., Butt, T. R. & Smulson, M. Nuclear protein modification and chromatin substructure. 3. Relationship between poly(adenosine diphosphate) ribosylation and different functional forms of chromatin. Biochemistry 18, 983–990 (1979).
Riquelme, P. T., Burzio, L. O. & Koide, S. S. ADP ribosylation of rat liver lysine-rich histone in vitro . J Biol Chem 254, 3018–3028 (1979).
Ogata, N., Ueda, K. & Hayaishi, O. ADP-ribosylation of histone H2B. Identification of glutamic acid residue 2 as the modification site. J Biol Chem 255, 7610–7615 (1980).
Ogata, N., Ueda, K., Kagamiyama, H. & Hayaishi, O. ADP-ribosylation of histone H1. Identification of glutamic acid residues 2, 14, and the COOH-terminal lysine residue as modification sites. J Biol Chem 255, 7616–7620 (1980).
Kreimeyer, A., Wielckens, K., Adamietz, P. & Hilz, H. DNA repair-associated ADP-ribosylation in vivo. Modification of histone H1 differs from that of the principal acceptor proteins. J Biol Chem 259, 890–896 (1984).
Boulikas, T. At least 60 ADP-ribosylated variant histones are present in nuclei from dimethylsulfate-treated and untreated cells. Embo J 7, 57–67 (1988).
Messner, S. et al. PARP1 ADP-ribosylates lysine residues of the core histone tails. Nucleic Acids Res 38, 6350–6362, doi: 10.1093/nar/gkq463 (2010).
Eichinger, L. et al. The genome of the social amoeba Dictyostelium discoideum. Nature 435, 43–57 (2005).
Couto, C. A. et al. PARP regulates nonhomologous end joining through retention of Ku at double-strand breaks. J Cell Biol 194, 367–375, doi: 10.1083/jcb.201012132 (2011).
Hsu, D. W., Gaudet, P., Hudson, J. J., Pears, C. J. & Lakin, N. D. DNA damage signaling and repair in Dictyostelium discoideum. Cell Cycle 5, 702–708 (2006).
Hsu, D. W. et al. DNA double-strand break repair pathway choice in Dictyostelium. J Cell Sci 124, 1655–1663, doi: 10.1242/jcs.081471 (2011).
Hudson, J. J. et al. DNA-PKcs-dependent signaling of DNA damage in Dictyostelium discoideum. Curr Biol 15, 1880–1885, doi: 10.1016/j.cub.2005.09.039 (2005).
Pears, C. J. et al. The role of ADP-ribosylation in regulating DNA double-strand break repair. Cell Cycle 11, 48–56, doi: 10.4161/cc.11.1.18793 (2012).
Pears, C. J. & Lakin, N. D. Emerging models for DNA repair: Dictyostelium discoideum as a model for nonhomologous end-joining. DNA Repair (Amst) 17, 121–131, doi: 10.1016/j.dnarep.2014.01.008 (2014).
Zhang, X. Y. et al. Xpf and not the Fanconi anaemia proteins or Rev3 accounts for the extreme resistance to cisplatin in Dictyostelium discoideum. PLoS Genet 5, e1000645, doi: 10.1371/journal.pgen.1000645 (2009).
Rajawat, J., Vohra, I., Mir, H. A., Gohel, D. & Begum, R. Effect of oxidative stress and involvement of poly(ADP-ribose) polymerase (PARP) in Dictyostelium discoideum development. FEBS J 274, 5611–5618, doi: 10.1111/j.1742-4658.2007.06083.x (2007).
Mir, H., Alex, T., Rajawat, J., Kadam, A. & Begum, R. Response of Dictyostelium discoideum to UV-C and involvement of poly (ADP-ribose) polymerase. Cell Prolif 48, 363–374, doi: 10.1111/cpr.12182 (2015).
Couto, C. A. et al. Nonhomologous end-joining promotes resistance to DNA damage in the absence of an ADP-ribosyltransferase that signals DNA single strand breaks. J Cell Sci 126, 3452–3461, doi: 10.1242/jcs.128769 (2013).
Stevense, M., Chubb, J. R. & Muramoto, T. Nuclear organization and transcriptional dynamics in Dictyostelium. Dev Growth Differ 53, 576–586, doi: 10.1111/j.1440-169X.2011.01271.x (2011).
Dubin, M., Fuchs, J., Graf, R., Schubert, I. & Nellen, W. Dynamics of a novel centromeric histone variant CenH3 reveals the evolutionary ancestral timing of centromere biogenesis. Nucleic Acids Res 38, 7526–7537, doi: 10.1093/nar/gkq664 (2010).
Hsu, D. W., Chubb, J. R., Muramoto, T., Pears, C. J. & Mahadevan, L. C. Dynamic acetylation of lysine-4-trimethylated histone H3 and H3 variant biology in a simple multicellular eukaryote. Nucleic Acids Res 40, 7247–7256, doi: 10.1093/nar/gks367 (2012).
Chubb, J. R. et al. Developmental timing in Dictyostelium is regulated by the Set1 histone methyltransferase. Dev Biol 292, 519–532, doi: 10.1016/j.ydbio.2005.12.054 (2006).
Muller-Taubenberger, A., Bonisch, C., Furbringer, M., Wittek, F. & Hake, S. B. The histone methyltransferase Dot1 is required for DNA damage repair and proper development in Dictyostelium. Biochem Biophys Res Commun 404, 1016–1022, doi: 10.1016/j.bbrc.2010.12.101 (2011).
Sawarkar, R., Visweswariah, S. S., Nellen, W. & Nanjundiah, V. Histone deacetylases regulate multicellular development in the social amoeba Dictyostelium discoideum. J Mol Biol 391, 833–848, doi: 10.1016/j.jmb.2009.06.067 (2009).
Grundy, G. J. et al. PARP3 is a sensor of nicked nucleosomes and monoribosylates histone H2B(Glu2). Nat Commun 7, 12404, doi: 10.1038/ncomms12404 (2016).
D’Amours, D., Desnoyers, S., D’Silva, I. & Poirier, G. G. Poly(ADP-ribosyl)ation reactions in the regulation of nuclear functions. Biochemical Journal 342, 249–268 (1999).
Langelier, M. F., Riccio, A. A. & Pascal, J. M. PARP-2 and PARP-3 are selectively activated by 5′ phosphorylated DNA breaks through an allosteric regulatory mechanism shared with PARP-1. Nucleic Acids Res 42, 7762–7775, doi: 10.1093/nar/gku474 (2014).
Grundy, G. J. et al. The Ku-binding motif is a conserved module for recruitment and stimulation of non-homologous end-joining proteins. Nat Commun 7, 11242, doi: 10.1038/ncomms11242 (2016).
Slade, D. et al. The structure and catalytic mechanism of a poly(ADP-ribose) glycohydrolase. Nature 477, 616–620, doi: 10.1038/nature10404 (2011).
Barkauskaite, E. et al. Visualization of poly(ADP-ribose) bound to PARG reveals inherent balance between exo- and endo-glycohydrolase activities. Nat Commun 4, 2164, doi: 10.1038/ncomms3164 (2013).
Sharifi, R. et al. Deficiency of terminal ADP-ribose protein glycohydrolase TARG1/C6orf130 in neurodegenerative disease. Embo J, doi: 10.1038/emboj.2013.51 (2013).
Rosenthal, F. et al. Macrodomain-containing proteins are new mono-ADP-ribosylhydrolases. Nat Struct Mol Biol 20, 502–507, doi: 10.1038/nsmb.2521 (2013).
Jankevicius, G. et al. A family of macrodomain proteins reverses cellular mono-ADP-ribosylation. Nat Struct Mol Biol 20, 508–514, doi: 10.1038/nsmb.2523 (2013).
Wahlberg, E. et al. Family-wide chemical profiling and structural analysis of PARP and tankyrase inhibitors. Nat Biotechnol 30, 283–288, doi: 10.1038/nbt.2121 (2012).
Adamietz, P. & Rudolph, A. ADP-ribosylation of nuclear proteins in vivo. Identification of histone H2B as a major acceptor for mono- and poly(ADP-ribose) in dimethyl sulfate-treated hepatoma AH 7974 cells. J Biol Chem 259, 6841–6846 (1984).
Kleine, H. & Luscher, B. Learning how to read ADP-ribosylation. Cell 139, 17–19, doi: 10.1016/j.cell.2009.09.018 (2009).
Rack, J. G., Perina, D. & Ahel, I. Macrodomains: Structure, Function, Evolution, and Catalytic Activities. Annu Rev Biochem, doi: 10.1146/annurev-biochem-060815-014935 (2016).
Faix, J., Kreppel, L., Shaulsky, G., Schleicher, M. & Kimmel, A. R. A rapid and efficient method to generate multiple gene disruptions in Dictyostelium discoideum using a single selectable marker and the Cre-loxP system. Nucleic Acids Res 32, e143, doi: 10.1093/nar/gnh136 (2004).
Acknowledgements
MacroD1 protein was provided by Keith Caldecott (University of Sussex). NL’s laboratory is supported by Cancer Research UK (www.cancerresearch.org.uk; grant C1521/A12353), Medical Research Council (www.mrc.ac.uk; MR/L000164/1) and NC3Rs (www.nc3rs.org.uk; NC/K00137X/1). AR was supported by a Clarendon Award (University of Oxford). CP’s laboratory is supported by NC3Rs (www.nc3rs.org.uk; NC/M000834/1).
Author information
Authors and Affiliations
Contributions
ADP-ribosylation of chromatin associated proteins, S.U. and H.Y.W.; H2Bv3-FLAG experiments, A.R.; Adprt1a biochemical analysis, D.W.H. and A.R.; Generation of h2bv3wt and h2bv3E18AE19A strains and analysis, S.U. C.P. and N.L. directed the work. N.L. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Rakhimova, A., Ura, S., Hsu, DW. et al. Site-specific ADP-ribosylation of histone H2B in response to DNA double strand breaks. Sci Rep 7, 43750 (2017). https://doi.org/10.1038/srep43750
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep43750
- Springer Nature Limited
This article is cited by
-
Linking DNA repair and cell cycle progression through serine ADP-ribosylation of histones
Nature Communications (2022)
-
Role of epigenetics in unicellular to multicellular transition in Dictyostelium
Genome Biology (2021)
-
The regulatory landscape of the human HPF1- and ARH3-dependent ADP-ribosylome
Nature Communications (2021)
-
Knock-in and precise nucleotide substitution using near-PAMless engineered Cas9 variants in Dictyostelium discoideum
Scientific Reports (2021)