Abstract
Alzheimer’s disease (AD) is characterized by excessive production and deposition of amyloid-beta (Aβ) proteins as well as synapse dysfunction and loss. While soluble Aβ oligomers (AβOs) have deleterious effects on synapse function and reduce synapse number, the underlying molecular mechanisms are not well understood. Here we screened synaptic organizer proteins for cell-surface interaction with AβOs and identified a novel interaction between neurexins (NRXs) and AβOs. AβOs bind to NRXs via the N-terminal histidine-rich domain (HRD) of β-NRX1/2/3 and alternatively-spliced inserts at splicing site 4 of NRX1/2. In artificial synapse-formation assays, AβOs diminish excitatory presynaptic differentiation induced by NRX-interacting proteins including neuroligin1/2 (NLG1/2) and the leucine-rich repeat transmembrane protein LRRTM2. Although AβOs do not interfere with the binding of NRX1β to NLG1 or LRRTM2, time-lapse imaging revealed that AβO treatment reduces surface expression of NRX1β on axons and that this reduction depends on the NRX1β HRD. In transgenic mice expressing mutated human amyloid precursor protein, synaptic expression of β-NRXs, but not α-NRXs, decreases. Thus our data indicate that AβOs interact with NRXs and that this interaction inhibits NRX-mediated presynaptic differentiation by reducing surface expression of axonal β-NRXs, providing molecular and mechanistic insights into how AβOs lead to synaptic pathology in AD.
Similar content being viewed by others
Introduction
Alzheimer’s disease (AD) is characterized by the accumulation of toxic amyloid-β (Aβ)-peptides, the principal constituent of plaques in the brains of AD patients1,2. In addition, loss of synapses in the brain is an early pathological feature of AD and is the best correlate of cognitive impairment3,4. Experimentally, Aβ oligomers (AβOs) cause synapse loss and synapse dysfunction both in vitro and in mouse models of AD4,5,6. The in vitro experiments show that the application of soluble AβOs to hippocampal slices or cultured neurons decreases immunoreactivity for pre- and post-synaptic proteins7,8,9,10,11,12 and the density of dendritic spines9,13,14,15,16. Further, Aβ treatment of hippocampal slices distorts synaptic plasticity4,5,6,17. Specifically, Aβ treatment blocks long-term potentiation (LTP) and enhances long-term depression (LTD). Thus, synapses exhibit vulnerability to Aβ. However, little is known about the molecular mechanisms underlying this vulnerability.
Synaptic organizing complexes are trans-synaptic adhesion molecules with an ability to promote pre- and/or post-synaptic organization (hereinafter synaptogenic activity) and are thought to function as essential molecular signals for synapse formation, maturation, maintenance, and plasticity18,19,20,21,22. The neuroligin (NLG)-neurexin (NRX) complex is one of the most well-studied synaptic organizing complexes, and mutations in this complex are genetic determinants predisposing to cognitive disorders such as autism and schizophrenia18,19,21. Recent studies have identified several other synaptic organizing complexes including TrkC-PTPσ23, Slitrk-PTPδ24,25, the leucine-rich repeat transmembrane protein (LRRTM) 1/2/3-NRX26,27,28,29,30, calsyntenin3-α-NRX31, GluRδ-cerebellin-NRX32,33,34, NGL3-LAR35,36, and IL1RAPL1/IL1RacP-PTPδ37,38,39. Thus there are many synaptic organizing proteins, any of which could be targets for Aβ in synapse disruption.
Indeed, there is recent evidence that Aβ pathogenic processes may affect synaptic organizers. Several organizers including NRXs40,41, leukocyte antigen-related tyrosine phosphatase (LAR)42,43 and NLG144 are cleaved by proteinases involved in the generation of Aβ, indicating that their function may be altered coordinately with Aβ production. Also, AβOs bind to soluble NLG1 ectodomain45. These results emphasize the importance of understanding whether and how AβOs affect the physiological roles of synaptic organizing complexes. But, there has been no published study that has systematically tested for physical and functional interactions of synaptic organizing complex proteins with AβOs.
In this study, we performed cell surface binding assays using soluble AβOs to screen for their interaction with synaptic organizers expressed at the cell surface. We found that AβOs interact with NRX family members and defined their AβO-binding domains. AβOs diminish NRX-mediated presynaptic organization by decreasing β-NRX expression on the axonal surface, although this does not affect NRX-NLG1 interaction or NRX-LRRTM2 interaction. In a transgenic mouse line with increased production of Aβ species, synaptic expression of β-NRXs is decreased. Together, our results indicate that AβOs interact with NRXs and that this interaction disrupts NRX-based synapse organization by destabilizing surface β-NRX on axons.
Results
A candidate screen isolates β-neurexins as Aβ oligomer-interacting proteins
To test whether there are any synaptic organizers that interact with Aβ oligomers, we performed cell surface binding assays in which soluble oligomers of amyloid-β (1–42) peptide conjugated with biotin (biotin–Aβ42) were added onto COS-7 cells expressing each synaptic organizer. We first confirmed that the biotin–Aβ42 peptides were properly oligomerized into low and high molecular weight oligomers by western blot analysis (Fig. 1a) and also that the biotin–Aβ42 oligomers bound COS-7 cells expressing the known Aβ42 oligomer receptors, the paired immunoglobulin-like receptor B (PirB)46 and the cellular prion protein (PrP c)47 (Fig. 1b). We screened a total of 22 synaptic organizers. We found no significant binding of biotin–Aβ42 oligomers on COS-7 cells expressing NLG1 or NLG2 (Fig. 1c) although a previous study reported an interaction between AβOs and NLG145. Instead, we detected significant binding of biotin–Aβ42 oligomers on COS-7 cells expressing (NRX1βS4(−)) (Fig. 1c). Interestingly, we did not detect any binding signals on COS-7 cells expressing any of the other synaptic organizers that we tested including type IIa receptor-type protein tyrosine phosphatases (RPTPs: PTPσ, PTPδ, and LAR), LRRTM2, TrkC, and Slitrk family members (Fig. 1c).
(a) Immunoblotting with an anti-β-Amyloid 1–16 antibody (6E10) confirms the formation of soluble oligomers of untagged amyloid-β (1–42) peptide (Aβ) and of biotin-tagged Aβ42 peptides (biotin-Aβ). The preparations include both low and high molecular weight (HMW) oligomers. The preparation without an oligomerization incubation step (Fresh) does not include HMW oligomers. Full gel blots for the cropped blots (a) are shown in the Supplementary Fig. 4. (b) The biotin-Aβ42 oligomers bind to COS-7 cells expressing the N-terminal extracellular HA-tagged known Aβ42 oligomer receptors, paired immunoglobulin-like receptor B (HA-PirB) and prion protein (HA-PrPc), but not those expressing HA-CD4 as a negative control. Surface HA was immunostained to verify expression of these constructs on the COS-7 cell surface. (c) Representative images showing cell surface binding assays testing for interaction between biotin-Aβ42 oligomers (250 nM, monomer equivalent) and known synaptic organizers. Biotin-Aβ42 oligomers were added to COS-7 cells expressing the indicated construct. Note that biotin-Aβ42 oligomers bind to COS-7 cells expressing HA-neurexin (NRX)1βS4(−), but not to those expressing any of the other organizers including HA-neuroligin1 (HA-NLG1). For the N-terminal extracellular HA-tagged constructs, surface HA was immunostained to verify expression of the construct on the COS-7 cell surface. Scale bars represent 30 μm (b,c).
We next checked for a biochemical interaction between NRX1β and AβOs in a pull-down assay using untagged Aβ42 oligomers. High-molecular weight Aβ42 oligomers and 5-mer Aβ42 oligomers, but not Aβ42 monomers, were coprecipitated with purified NRX1βS4(−)-Fc proteins pre-immobilized on Protein G magnetic beads (Fig. 2a). In contrast, either untagged Aβ42 oligomers or monomers were not coprecipitated with purified Fc proteins pre-immobilized on Protein G magnetic beads. These results suggest that AβOs can interact directly with the extracellular domain of NRX1βS4(−). We next determined the binding affinity by saturation analysis in cell-surface binding assays (Fig. 2b,c). The binding of biotin–Aβ42 oligomers to NRX1βS4(−) increased and became saturated with increasing amounts of biotin–Aβ42 oligomers. A Scatchard plot of the binding data revealed that the apparent dissociation constant (Kd) value is 183.5 nM monomer equivalent. Thus the interaction between NRX1β and biotin–Aβ42 oligomers is within the typical nanomolar range for biologically significant ligand-receptor interactions. Together, these results indicate that Aβ oligomers bind directly with nanomolar affinity to NRX1β.
(a) Pull-down assay of untagged Aβ42 oligomers with purified NRX1βS4(−)-Fc proteins. Full gel blots for the cropped blots (a) are shown in the Supplementary Fig. 4. (b) Saturable binding of biotin-Aβ42 oligomers to COS-7 cells expressing HA-NRX1βS4(−). Data are presented as mean ± SEM. (c) Scatchard plot of binding data shown in (b). The Kd = 183.5 nM monomer equivalent. (n = 30 cells for each plot).
Neurexins interact with Aβ oligomers via the N-terminal histidine-rich domain of β-neurexin1/2/3 and an insert at alternative-splicing site 4 of α/β-neurexin1/2
Given that the NRX family is composed of many different isoforms such as α/β-isoforms and alternative splicing site 4 (S4)-positive or S4-negative isoforms18, which have differing binding affinity and selectivity for NRX-interacting proteins, we next tested which NRX isoforms interact with AβOs (Fig. 3). In the case of S4-negative isoforms, biotin–Aβ42 oligomers interacted with NRX1β, 2β and 3β at a similar level but not with NRX1α, 2α or 3α, indicating that Aβ42 oligomers interact with β-NRX-specific domains in the absence of an insert at S4. Given that the N-terminal histidine-rich domain (HRD; amino acids 50–83 in NRX1β) is unique to the β-isoforms48, we next tested the binding of biotin–Aβ42 oligomers to COS-7 cells expressing NRX1β lacking the HRD (HA-NRX1β∆HRD) and detected no binding (Fig. 3a,b). Further, COS-7 cells expressing HA-NRX2β∆HRD or HA-NRX3β∆HRD also displayed no binding signal (Fig. 3a,b). These results indicate that the HRD of β-NRX1/2/3 is one of the domains responsible for Aβ42 oligomer binding. Next, we investigated Aβ42 oligomer-binding to S4-positive NRX isoforms. Biotin–Aβ42 oligomers interacted with S4-positive NRX1α and 2α but not 3α (Fig. 3a,b), indicating that the inserts at the S4 site of NRX1 and NRX2 interact with Aβ42 oligomers. Indeed, S4-positive NRX1β and 2β displayed stronger binding of Aβ42 oligomers than S4-negative NRX1β and 2β, respectively (Fig. 3a,b). The enhancement of binding by the S4 insert is similar to the difference in the binding signals of S4-negative and S4-positive NRX1α and 2α (Fig. 3b), suggesting that Aβ42 oligomer-binding to the S4 insert additively increases the binding of Aβ42 to NRX1β and 2β. Together, these data indicate that the HRDs of NRX1β, 2β and 3β and the S4 inserts of NRX1α/β and 2α/β are responsible for Aβ42 oligomer interaction.
(a) Representative images showing the binding of biotin-Aβ42 oligomers (250 nM, monomer equivalent) to COS-7 cells expressing the indicated isoform of extracellularly HA-tagged NRX. S4(−) and S4(+) indicate without and with an insert at splicing site 4, respectively. HA fluorescent signals correspond to surface HA. Note that in assays with S4-negative isoforms, there is no bound biotin-Aβ42 oligomer signal on cells expressing α-NRX1, 2, or 3 whereas cells expressing β-NRX1, 2, and 3 all have significant bound biotin-Aβ42 oligomer signal. Cells expressing β-NRX1, 2, and 3 lacking the histidine-rich domain (∆HRD) have no biotin-Aβ42 oligomer binding signal. Cells expressing S4-positive isoforms of α-NRX1 or 2 or β-NRX1, 2, or 3 have significant bound biotin-Aβ42 oligomers. Scale bar represents 30 μm. (b) Quantification of bound biotin-Aβ42 oligomers for each NRX construct. n = 30 cells for each construct from three independent experiments, one-way ANOVA, P < 0001. *P < 0.01 and ***P < 0.0001 compared with HA-CD4 and §P < 0.0001 between S4(−) and S4(+) by Bonferroni multiple comparisons tests. Data are presented as mean ± SEM.
Aβ treatment diminishes neurexin-mediated presynaptic differentiation
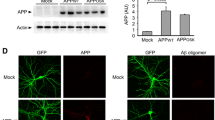
NRXs mediate the presynaptic induction activity of NLGs and LRRTM1/2/3 in hippocampal neurons18,19,20,21,22,27,28,29,30. We thus tested whether Aβ42 oligomers affect NRX-mediated presynaptic differentiation in coculture-based artificial synapse formation assays (Fig. 4). As reported previously, HEK293 (hereinafter HEK) cells expressing NLG1, NLG2, or LRRTM2 induced the accumulation of the excitatory presynaptic marker VGLUT1. NLG1- or NLG2-expressing HEK cells further induced the accumulation of the inhibitory presynaptic marker VGAT in cocultured hippocampal neurons. Treatment with Aβ42 oligomers significantly decreased VGLUT1 accumulation induced by NLG1, NLG2, or LRRTM2 (Fig. 4a,b). Interestingly, treatment with Aβ42 oligomers had no effect on VGAT accumulation induced by NLG1 or NLG2 (Fig. 4a,c). These data indicate that Aβ treatment diminishes excitatory, but not inhibitory, presynaptic differentiation induced by NRX-interacting synaptic organizers. Notably, treatment with Aβ42 oligomers did not affect TrkC-induced or Slitrk2-induced VGLUT1 accumulation (Fig. 4a,b), which is mediated by RPTPs in cocultured hippocampal neurons22,23,24,25, indicating that Aβ treatment diminishes NRX-mediated, but not RPTP-mediated, presynaptic differentiation. Aβ42 oligomers did not affect VGAT accumulation induced by HEK cells expressing Slitrk2 (Fig. 4a,c). The observed phenotypes induced by Aβ42 oligomers are not due to the reduction of surface expression levels of the tested organizers on HEK cells because Aβ42 oligomers did not alter their surface expression (Supplementary Fig. 1). Together these results indicate that Aβ42 oligomers selectively diminish NRX-mediated excitatory presynaptic differentiation and also that RPTP-mediated presynaptic differentiation is insensitive to Aβ42 oligomers.
(a) Representative images of triple immunolabeling for VGLUT1, VGAT and surface HA in HEK293 cells expressing the indicated extracellularly HA-tagged construct cocultured with cultured hippocampal neurons and treated with Aβ42 oligomers (AβOs, 500 nM, monomer equivalent) or vehicle. HA-CD4 is used as a negative control protein as it lacks synaptogenic activity. AβOs seem not to affect VGLUT1 accumulation induced by an HA-TrkC non-catalytic isoform (HA-TrkC) or VGLUT1 or VGAT accumulation induced by HA-Slitrk2. In contrast, AβOs seem to decrease VGLUT1 accumulation induced by HA-NLG1, HA-NLG2, or HA-LRRTM2. AβOs seem not to affect VGAT accumulation induced by HA-NLG1 or HA-NLG2. Scale bar represents 20 μm. (b,c) Quantification of the total intensity of VGLUT1 (b) and VGAT (c) puncta on HEK293 cells expressing the indicated HA-tagged proteins divided by HEK293 cell area. n = 30 cells for each construct from three independent experiments, one-way ANOVA, P < 0.0001. #P < 0.05 and *P < 0.01 for the indicated comparisons between vehicle control and AβO treatment by Bonferroni multiple comparisons tests. n.s., not significant. Data are presented as mean ± SEM.
Aβ treatment has no effect on neurexin interaction with neuroligin1 or LRRTM2
We next investigated cellular mechanisms by which Aβ42 oligomers diminish NRX-mediated presynaptic differentiation. One possibility could be that Aβ42 oligomers interfere with the interaction of NRX with NLG1, NLG2, and/or LRRTM2. To test this, we performed cell surface binding assays using recombinant Fc-tagged NLG1 or LRRTM2 ectodomain proteins (NLG1-Fc or LRRTM2-Fc, respectively). Consistent with previous studies, NLG1-Fc bound to COS-7 cells expressing NRX1βS4(−) or (NRX1βS4(+)) (Fig. 5a,b) but not to those expressing a form with a point mutation that completely abolishes NLG interaction, NRX1βS4(−)D137A49 (Fig. 5a,b). LRRTM2-Fc bound to COS-7 cells expressing NRX1βS4(−), but not NRX1βS4(+) or NRX1βS4(−)D137A, as reported previously29 (Fig. 5c,d). Interestingly, the application of biotin–Aβ42 oligomers had no significant effect on the binding of NLG1-Fc or LRRTM2-Fc to any of the NRX1β constructs (Fig. 5b,d). These data indicate that the diminishment of NRX-mediated presynaptic differentiation by Aβ42 oligomers is not due to Aβ interference in the interaction of NRXs with NLG1, NLG2, or LRRTM2.
(a,c) Representative images of triple labeling for bound Fc proteins, bound Aβ42 peptides and surface HA on COS-7 cells expressing the indicated extracellularly HA-tagged neurexin (NRX)1β construct. Recombinant neuroligin1-Fc (NLG1-Fc; 20 nM) binds to COS-7 cells expressing HA-NRX1βS4(+) or HA-NRX1βS4(−) but not to those expressing HA-NRX1βS4(−)D137A (a), and recombinant LRRTM2-Fc (20 nM) binds to COS-7 cells expressing HA-NRX1βS4(−) but not to those expressing HA-NRX1βS4(+) or HA-NRX1βS4(−)D137A (c). Treatment with Aβ42 oligomers (AβOs, 500 nM, monomer equivalent) does not seem to affect the binding in any of these cases. Scale bars represent 30 μm. (b,d) Quantification of recombinant NLG1-Fc (b) or LRRTM2-Fc (d) bound to COS-7 cells expressing the indicated HA-NRX1β constructs treated with vehicle control or 500 nM Aβ42 oligomers. n = 30 cells for each condition from three independent experiments, one-way ANOVA, P < 0.0001. n.s., not significant by Bonferroni multiple comparisons test. Data are presented as mean ± SEM.
Aβ treatment decreases surface expression of neurexin1β on axons
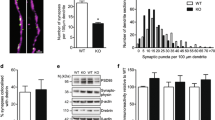
We next tested another possible mechanism: Aβ42 oligomers could decrease the surface expression of NRXs on axons and thereby diminish NRX-mediated presynaptic differentiation. We cotransfected hippocampal neurons with mCherry (for imaging neuronal morphology) and NRX1β extracellularly tagged with super-ecliptic pHluorin (SEP) (SEP-NRX1β), and then performed time-lapse imaging of SEP-NRX1β expressed on mCherry-positive axons (Fig. 6). SEP is a pH-sensitive GFP variant that yields fluorescence at neutral pH (e.g. on the cell surface) but is quenched at low pH (e.g. inside cytoplasmic vesicles), thus it allows for monitoring the surface expression level of tagged proteins50. In mCherry-expressing axons, cotransfected SEP-NRX1βS4(+), SEP-NRX1βS4(−), or SEP-NRX1βS4(−) lacking the HRD (SEP-NRX1βS4(−)∆HRD) had punctate distributions (Fig. 6a–c). Our immunocytochemistry further confirmed that these SEP-NRX1β puncta colocalized with VGLUT1 puncta (Supplementary Fig. 2), suggesting that the SEP-NRX1β proteins likely accumulated at presynaptic boutons. The application of Aβ42 oligomers into the extracellular solution decreased the SEP fluorescent signal of SEP-NRX1βS4(+) and SEP-NRX1βS4(−) with similar decay (Fig. 6d). The similarity of the decay suggests that the binding of Aβ42 oligomers to the S4 insert is not essential for AβO-induced reduction of NRX surface expression. On the other hand, the application of Aβ42 oligomers did not affect the SEP signal of SEP-NRX1βS4(−)∆HRD (Fig. 6d), which does not bind Aβ42 oligomers (Supplementary Fig. 3). Thus, the binding of Aβ42 oligomers to the HRD is essential for AβO-induced reduction of NRX surface expression. Together with the results of our coculture assays (Fig. 4), these data suggest that Aβ42 oligomers diminish NRX-mediated presynaptic differentiation by decreasing surface expression of β-NRXs on axons.
(a–c) Representative time-lapse images of axons expressing extracellularly super-ecliptic pHluorin (SEP)-tagged NRX1βS4(+) (a), SEP-NRX1βS4(−) (b) and SEP-NRX1βS4(−) lacking the histidine-rich domain (SEP-NRX1βS4(−)∆HRD) (c) treated with Aβ42 oligomers (500 nM, monomer equivalent) at t = 0 min. Scale bars represent 5 μm. (d) Quantification of SEP intensity signals at 5 min before and 10, 30 and 60 min after Aβ42 oligomer treatment. n = 29 for SEP-NRX1βS4(+), n = 26 for SEP-NRX1βS4(−), and n = 33 for SEP-NRX1βS4(−)ΔHRD puncta from 9 cells for each condition from three independent experiments, two-way repeated measures ANOVA, F(3, 255) = 42.94, P < 0.0001 for time and F(2, 85) = 18.95, P < 0.0001 for construct. **P < 0.001, and ***P < 0.0001 compared with SEP signal at 5 min before the treatment, and ***P < 0.0001 compared with SEP-NRX1βS4(−)ΔHRD by Bonferroni multiple comparisons tests. n.s., not significant. Data are presented as mean ± SEM.
Synaptic expression of β-neurexin is decreased in a mouse model of Alzheimer’s disease
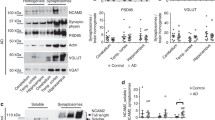
Finally, we tested whether AβOs affect synaptic expression of endogenous NRXs in vivo (Fig. 7). We prepared synaptosomal fractions from the hippocampus and the cortex of J20 APP mice, transgenic mice expressing a mutant form of human amyloid precursor protein (APP) that have progressively increasing AβO expression and amyloid deposition51. When compared to those of wild-type littermates, the hippocampal and cortical synaptosomes from J20 APP mice both had significantly reduced levels of β-NRX proteins, but not α-NRX proteins (Fig. 7a,b). These data indicate that there is selective reduction of endogenous β-NRXs in synapses in the J20 transgenic AD mouse model.
(a) Representative immunoblots of neurexins (NRXs) in synaptosomes from the hippocampus and from the cerebral cortex of J20 APP mice and wild-type (WT) littermates at 6 months of age. The labels 1, 2, and 3 indicate samples from different mice. Full gel blots for the cropped blots (a) are shown in the Supplementary Fig. 4. (b) Quantification of synaptic expression of β-NRXs (bands indicated by a lower right square bracket in (a) and α-NRXs (bands indicated by upper right square bracket in (a) normalized to β-actin protein expression in synaptosomes from the hippocampus and the cortex, expressed relative to WT. n = 5 samples per genotype for hippocampus, with each n representing pooled hippocampi from two mice. n = 6 samples per genotype for cortex, with each n representing a cortex from one mouse. Unpaired t tests, #P < 0.05 for β-NRXs and P = 0.15 for α-NRXs in hippocampus, and **P < 0.001 for β-NRXs and P = 0.31 for α-NRXs in cortex. n.s., not significant. Data are presented as mean ± SEM.
Discussion
In this study, we uncovered a direct interaction of Aβ42 oligomers with NRXs. We further determined that the HRDs of NRX1β, 2β and 3β and S4 inserts of NRX1 and NRX2 are responsible for the interaction between Aβ42 oligomers and NRXs. Aβ42 oligomers diminish NRX-mediated presynaptic organization by decreasing surface expression of β-NRXs on axons. Further, synaptic expression of endogenous β-NRXs is selectively decreased in a line of transgenic mice with increased production of Aβ peptides. Together, our findings demonstrate that NRX is an interactor of Aβ42 oligomers and plays a role in AβO-induced synapse pathology.
It has been well known that AβOs have postsynaptic adverse effects such as the inhibition of LTP, the enhancement of LTD and the loss of dendritic spines (see the review of refs 4 and 6 and references therein). These effects are mediated by Aβ-interacting postsynaptic membrane proteins such as prion47,52,53, PirB46 and EphB254, whose binding to Aβ alters the function of postsynaptic NMDA-type glutamate receptor and/or metabotropic glutamate receptor 5 and consequently affects synaptic plasticity and dendritic spine density. However, Aβ accumulates at presynaptic terminals as well as at postsynaptic sites, and Aβ can also distort presynaptic structure and function6,9,11. The mechanisms underlying the presynaptic actions of Aβ are not well understood. NRXs are presynaptic cell-adhesion molecules crucial for presynaptic organization and functions18,19. Thus the most significant finding of this study is the identification of NRXs as novel AβO-interacting presynaptic membrane proteins. Our data show that Aβ42 oligomers interact with NRXs and that this interaction leads to a decrease in NRX expression on the axon surface. NRXs regulate synapse organization through interacting with multiple postsynaptic adhesion molecules including NLG1-418,19, LRRTM1/2/326,27,28,29,30, and calsyntenin-331. Thus, the NRX family serves as a presynaptic molecular hub to integrate and/or coordinate multiple trans-synaptic organizing signals22. Our findings therefore suggest that Aβ42 oligomers may dampen the presynaptic hub function of NRXs and thereby disrupt the balance of the multiple synaptic organizing complexes. The dysregulation of NRXs by Aβ42 oligomers would therefore be an important mechanism underlying Aβ vulnerability of synapses.
Like members of the NRX family, RPTP family members including PTPσ, PTPδ, and LAR act as presynaptic hubs by mediating trans-synaptic interactions with multiple organizers such as TrkC, Slitrks, NGL-3, and IL1RAPL122. Our binding screen shows that RPTP family members do not interact with Aβ42 oligomers. Further, our coculture data suggest that Aβ42 oligomers diminish NRX-mediated, but not RPTP-mediated, presynaptic differentiation. Thus, NRX-based synaptic organizing complexes are sensitive to Aβ42 oligomers whereas RPTP-based complexes are not. Although synapses are vulnerable to Aβ, the majority of synapses remain after the treatment of cultured neurons with AβOs13,15 and even at the late stage of AD3,55. This suggests that synapses exhibit two conflicting properties: Aβ vulnerability and tolerance. Thus, Aβ vulnerability and tolerance of synapses may be partly determined by two different presynaptic hubs: an Aβ-sensitive hub (NRX) and an Aβ-insensitive hub (RPTP).
In our coculture assays, Aβ42 oligomers suppress excitatory presynaptic differentiation induced by HEK cells expressing multiple NRX-interacting synaptic organizer proteins: NLG1, NLG2, and LRRTM2. Our results demonstrating disruption of the hub protein NRX represent one mechanism underlying this effect. Direct interaction of AβOs with these NRX interactors could be another possible mechanism. A previous study reported a direct interaction between Aβ42 peptides and the NLG1 ectodomain45. However, our cell surface binding assay showed no binding of Aβ42 oligomers to NLG1-expressing fibroblasts. This discrepancy could result from a difference in the production of the target proteins: the previous study used soluble recombinant NLG1 ectodomain proteins whereas we used membrane-bound NLG1 expressed on the cell surface. The same previous study45 and our binding assay show that NLG2 does not interact with Aβ42 oligomers and we also show that they do not bind LRRTM2 in cell surface binding assays. These data support the idea that the molecular mechanism by which Aβ42 oligomers suppress the synaptogenic activity of NLG1 and other synaptogenic NRX interactors is by binding and decreasing NRXs on the axon surface, rather than by directly binding multiple synaptogenic NRX interactors.
The effects of AβOs may not be limited to synapse organization but may also impact on synaptic function as NRX proteins can regulate neurotransmitter release: α-NRX isoforms regulate Ca2+-triggered neurotransmitter release by functionally coupling calcium channels to presynaptic machinery56 whereas β-NRX isoforms regulate endocannabinoid-dependent glutamate release probability57. A previous study has shown that Aβ increases the presynaptic release probability of glutamate58. Thus, the interaction between NRXs and Aβ42 oligomers that we have uncovered here could mediate Aβ-dependent glutamate release by changing calcium channel activation and/or endocannabinoid signaling pathways. Future studies using α-NRX1/2 double knockout mice and/or β-NRX1/2/3 triple knockout mice could test this possibility.
Aβ seems to selectively affect glutamatergic (excitatory) presynaptic terminals. A previous study showed that Aβ has no effect on GABA release probability58 and our coculture data show that Aβ42 oligomers diminish excitatory, but not inhibitory, presynaptic differentiation induced by NLG1/2. Since our data has also shown that NRX is central to Aβ vulnerability, the mechanism underlying differential sensitivity of glutamatergic and GABAergic axons to Aβ likely involves NRX. Our binding assays show that the binding of Aβ42 oligomers to NRXs depends on the isoform type (α versus β) and S4 site insertion, and a recent study using single-cell mRNA profiling has shown that there is cell-type-specific expression of NRX isoforms59. Thus, comparison of the expression profiles of α- and β-isoforms and S4 splicing in glutamatergic and GABAergic neurons could yield insight into the differing Aβ sensitivity of their axons.
We have shown here that the molecular mechanism of the Aβ-NRX interaction involves two domains: the S4 inserts of NRX 1/2 and the HRDs of β-NRX forms. NRX S4 inserts are critical for determining the binding affinity of NLGs with NRXs and the binding selectivity of LRRTMs with NRXs18,19,29. Although Aβ42 oligomers bind to the S4 inserts of NRX1 and NRX2, Aβ42 oligomers have no effect on the binding of NLG1 or LRRTM2 to NRX1βS4(+) and also have similar effects on surface expression of SEP-NRX1βS4(+) and SEP-NRX1βS4(−). These findings suggest that the binding of Aβ42 oligomers to the S4 insert is not essential for AβO-induced diminishment of NRX-mediated presynaptic differentiation. Instead, our binding assays show that the HRDs of NRX1β, 2β, and 3β are necessary for the binding of Aβ42 oligomers to β-NRXs and our time-lapse imaging study demonstrates that the HRD of β-NRXs is crucial for AβO-induced reduction of NRX surface expression on axons. The importance of the HRD for axonal expression of NRX is additionally supported by our in vivo finding that β-NRXs (which possess an HRD), but not α-NRXs (which lack HRDs), are decreased significantly in synaptosomes of J20 APP mice. Therefore, the HRD likely contributes to the stabilization of surface β-NRXs on axons under normal physiological conditions.
Our binding assays demonstrate that β-NRX HRDs and NRX1/2 S4 inserts are the domains responsible for Aβ42 oligomer binding of NRXs. This is helpful to develop new therapeutic strategies aimed at preventing Aβ-induced synapse pathology, in particular presynaptic dysfunction since NRXs function presynaptically18,56,57. For example, neutralizing antibodies and/or small peptides that block NRX-AβO interactions could normalize presynaptic glutamate release distorted by AβOs. Thus, our findings provide new molecular insights into how Aβ-induced synapse pathology could be prevented and/or reduced.
Materials and Methods
Plasmids
To generate a series of extracellularly HA-tagged neurexin (HA-NRX) constructs, cDNA encoding the mature form of each NRX isoform was subcloned into spNRX1β-HA-C1, a vector containing a CMV promoter upstream of the N-terminal signal peptide sequence of NRX1β (spNRX1β) followed by HA and a multiple cloning site. The following NRX vectors were used as a PCR template for the subcloning: intracellular CFP-tagged mouse NRX1βS4(+), 1βS4(−), 1αS4(+), 1αS4(−), 2αS4(+), 2αS4(−), 3αS4(+), and 3αS4(−) (kindly provided by Dr. Ann Marie Craig (University of British Columbia)) and intracellular V5-tagged mouse NRX2βS4(+), 2βS4(−), 3βS4(+), and 3βS4(−) (kindly provided by Dr. Takeshi Uemura (Shinshu University)). For β-NRX constructs lacking their N-terminal histidine-rich domain (HRD), the coding sequence for the mature forms of NRX1β lacking HRD (aa 50–83), NRX2β lacking HRD (aa 54–87) and NRX3β lacking HRD (aa 48–81) were subcloned into spNRX1β-HA-C1 following the NRX1β signal sequence and HA. For extracellularly super-ecliptic pHluorin (SEP)-tagged neurexin1β (SEP-NRX1β) constructs, the coding sequence for the mature form of each NRX1β was subcloned into spNRX1β-SEP-C1, a vector containing a CMV promoter upstream of spNRX1β followed by the SEP coding region and a multiple cloning site. All constructs were verified by DNA sequencing. Further details are described in Supplementary Methods.
Animals
All animal experiments were carried out in accordance with the Canadian Council on Animal Care guidelines and approved by the IRCM Animal Care Committee and the McGill University Animal Care Committee. We used heterozygous transgenic adult C57BL/6 mice (6 months old, mixed sex) expressing the human amyloid precursor protein (hAPP) carrying the Swedish (K670N, M671L) and Indiana (V717F) familial AD mutations driven by the platelet-derived growth factor (PDGF) β-chain promoter (APP mice, J20 line)51 and age-matched wild-type (WT) littermates.
Preparation of Aβ42 oligomers
Aβ(1–42) (r-peptide, A-1002–2, 1 mg) and biotin-tagged Aβ(1–42) (Anaspec, AS-23523-05, 0.5 mg) were used to generate oligomeric forms essentially as described previously60. Full details of the Aβ42 oligomer preparation are provided in Supplementary Methods. The preparations were stored at −80 °C or used in experiments immediately. Individual Aβ oligomer stocks were never thawed and re-frozen. To confirm oligomer formation, the preparation was run on a 4–20% TGX precast gel (Biorad) and immunoblotted with anti-β-Amyloid 1–16 (1:5000; mouse IgG1; clone 6E10; Covance).
Neuron culture, coculture-based artificial synapse formation assay and immunocytochemistry
Cultures of rat hippocampal neurons, COS-7 cells, HEK293 cells, coculture-based artificial synapse formation assays, and immunocytochemistry were performed essentially as reported previously23,24. Transfections into COS-7 and HEK293 cells were performed using TransIT-LT1 (Mirus Bio. LLC). For transfections into hippocampal neurons, the ProFection Mammalian Transfection System (Promega) was used. For artificial synapse formation assays, transfected HEK293 cells were co-cultured with rat hippocampal neurons. Cultures were fixed with parafix solution (4% paraformaldehyde and 4% sucrose in PBS (pH 7.4)) for 12 minutes followed by permeabilization with PBST (PBS + 0.2% Triton X-100). They were incubated with blocking solution (PBS + 3% bovine serum albumin (BSA) and 5% normal goat serum) for 1 hour at room temperature, then with primary antibodies in blocking solution (overnight, 4 °C) and secondary antibodies (1 hour, room temperature). Images were acquired as 12-bit grayscale and prepared using Adobe Photoshop CS5. For quantification, sets of cells were stained simultaneously and imaged with identical settings. Further details are described in Supplementary Methods.
Cell surface binding assay
For testing for binding of biotin-Aβ42 oligomers, COS-7 cells on coverslips were transfected with the indicated expression vectors and maintained for 24 hours. The transfected cells were washed with extracellular solution (ECS) containing 168 mM NaCl, 2.4 mM KCl, 20 mM HEPES (pH 7.4), 10 mM D-glucose, 2 mM CaCl2, and 1.3 mM MgCl2 with 100 μg/ml BSA (ECS/BSA) and then incubated with ECS/BSA containing 250 nM biotin-Aβ42 oligomers (monomer equivalent) for 1 hour at 4 °C to prevent endocytosis. The cells were washed in ECS, fixed with parafix solution for 12 min at room temperature, incubated with blocking solution for 1 hour at room temperature, followed by the immunolabeling of surface HA as described above, and then incubated with Alexa594-conjugated streptavidin (1:4000; Jackson ImmunoResearch) and Alexa488-conjugated anti-rabbit IgG (H+L) (1:500; Invitrogen) for 1 hour at room temperature to label bound biotin-Aβ42 oligomers and surface HA, respectively. Further details are described in Supplementary Methods.
Pull-down assays
Purified soluble recombinant human NRX1βS4(−) ectodomain fused to human Fc (NRX1βS4(−)-Fc, 5268-NX-050, R&D systems) or human Fc (a negative control) generated from the pc4-sp-Fc vector23 were used for the pull-down assays. NRX1β-Fc or Fc proteins were pre-immobilized with Protein G magnetic beads (Dynabeads Protein G, Life Technology) in 20 mM sodium phosphate buffer (pH 7.0) for 2 hours at 4 °C. The pre-immobilized NRX1β-Fc or Fc proteins were then incubated with untagged Aβ oligomers in binding solution (20 mM HEPES (pH 7.4), 2 mM CaCl2, and 1.3 mM MgCl2) for 1 hour at 4 °C. Subsequently, the bead suspensions were washed five times with binding solution. Bound peptides and proteins were eluted with 100 mM glycine-HCl. Eluted samples were diluted in SDS sample buffer without boiling, separated on a 4–20% gradient SDS-PAGE gel and analyzed by western blotting with anti-β-Amyloid 1–16 (1:5000; mouse IgG1; clone 6E10; Covance) or horseradish peroxidase (HRP)-conjugated anti-human Fc (1:10,000; Jackson ImmunoResearch) antibodies.
Time-lapse imaging
For time-lapse imaging, hippocampal neurons cultured on 18-mm coverslips were cotransfected with a SEP-NRX construct and mCherry at 10 days in vitro (DIV) and used for imaging at 20–22 DIV. During imaging, the live transfected neurons were mounted in a Chamlide CMB magnetic chamber (Live Cell Instrument) and maintained in ECS at 37 °C controlled by a Tempcontrol 37–2 device (Pecon Germany) without perfusion. Aβ42 oligomers (500 nM, monomer equivalent) were manually added into ECS in the chamber 5 minutes after taking the first image. Fluorescent imaging was performed using a Leica DMIRE2 inverted microscope (Leica Germany) equipped with an Orca ER CCD camera (Hamamatsu Japan) and a 63 × 1.4 NA oil objective lens. All images were acquired by Volocity software (Perkin Elmer) at 1344 × 1024 resolution with 12 bits/pixel.
Fluorescence quantification
All imaging and image analysis were done while blind to the experimental condition. Analysis was performed by using Metamorph 7.8 software (Molecular Devices), Microsoft Excel, and GraphPad Prism 6. For binding of biotin-Aβ42 oligomers and Fc-fusion proteins, the average intensity of bound protein per COS-7 cell area minus off-cell background was normalized to the average intensity of the surface HA signal on COS-7 cells expressing the indicated HA-tagged proteins. For cocultures, fields for imaging were chosen using only the HA and phase contrast channels to locate HA-positive HEK293 cells in neurite-rich regions. The VGLUT1 or VGAT channel was thresholded and the total intensity of the puncta within HA-positive HEK293 cell regions was measured. For time-lapse imaging, the average background intensity of the image before Aβ treatment was measured, and this value was subtracted from the intensity of each frame of the time-lapse image sequences. The axons of transfected neurons were defined based on the morphology of mCherry-expressing neurons. In the image before Aβ treatment, the areas corresponding to puncta of SEP-NRX1β in mCherry-positive axons were manually traced as regions of interest (ROIs) using Metamorph 7.8. The average intensity of SEP and mCherry signals in these ROIs in each frame was measured. To quantify the effects of Aβ treatment on NRX surface expression, the SEP signal was normalized to the mCherry signal. Correction of the image shift in the x–y plane was done by comparing mCherry and SEP images. Pseudo-color images were created based on the fluorescence intensity range of the image prior to the Aβ treatment by Metamorph 7.8.
Synaptosome preparation and Immunoblotting
Preparation of synaptosome fractions from mice was performed essentially as described previously26. For all samples, protein concentrations were measured in DC Protein Assays (Biorad). After normalizing protein concentration, samples were run on 10% polyacrylamide gels. For immunoblotting NRXs, unboiled samples were used. Signals were developed using Immobilon Western Chemiluminescent HRP Substrate (Millipore) and captured by an ImageQuant LAS 4000 instrument (GE healthcare). Band signal intensity was measured using Metamorph 7.8 software and normalized to β-actin signal intensity for quantification. Further details are described in Supplementary Methods.
Statistical analysis
Statistical tests were performed using GraphPad Prism 6. Data distribution was assumed to be normal. Statistical comparisons were done by Student’s unpaired t test, one-way ANOVA and two-way repeated measures ANOVA with post hoc Bonferroni multiple comparisons tests, as indicated in the figure legends. All data are represented as the mean ± standard error of the mean (SEM) from three independent experiments and statistical significance was defined as P < 0.05.
Additional Information
How to cite this article: Naito, Y. et al. Amyloid-β Oligomers Interact with Neurexin and Diminish Neurexin-mediated Excitatory Presynaptic Organization. Sci. Rep. 7, 42548; doi: 10.1038/srep42548 (2017).
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
Hardy, J. & Selkoe, D. J. The amyloid hypothesis of Alzheimer’s disease: progress and problems on the road to therapeutics. Science 297, 353–356 (2002).
Holtzman, D. M., Morris, J. C. & Goate, A. M. Alzheimer’s disease: the challenge of the second century. Sci Transl Med 3, 77sr71 (2011).
Scheff, S. W. & Price, D. A. Synaptic pathology in Alzheimer’s disease: a review of ultrastructural studies. Neurobiol Aging 24, 1029–1046 (2003).
Sheng, M., Sabatini, B. L. & Sudhof, T. C. Synapses and Alzheimer’s disease. Cold Spring Harb Perspect Biol 4, a005777 (2012).
Selkoe, D. J. Alzheimer’s disease is a synaptic failure. Science 298, 789–791 (2002).
Mucke, L. & Selkoe, D. J. Neurotoxicity of amyloid β-protein: synaptic and network dysfunction. Cold Spring Harb Perspect Med 2, a006338 (2012).
Snyder, E. M. et al. Regulation of NMDA receptor trafficking by amyloid-β. Nat Neurosci 8, 1051–1058 (2005).
Roselli, F. et al. Soluble β-amyloid1–40 induces NMDA-dependent degradation of postsynaptic density-95 at glutamatergic synapses. J Neurosci 25, 11061–11070 (2005).
Calabrese, B. et al. Rapid, concurrent alterations in pre- and postsynaptic structure induced by naturally-secreted amyloid-β protein. Mol Cell Neurosci 35, 183–193 (2007).
Roselli, F., Hutzler, P., Wegerich, Y., Livrea, P. & Almeida, O. F. Disassembly of shank and homer synaptic clusters is driven by soluble β-amyloid1–40 through divergent NMDAR-dependent signalling pathways. PLoS One 4, e6011 (2009).
Russell, C. L. et al. Amyloid-β acts as a regulator of neurotransmitter release disrupting the interaction between synaptophysin and VAMP2. PLoS One 7, e43201 (2012).
Ripoli, C. et al. Effects of different amyloid β-protein analogues on synaptic function. Neurobiol Aging 34, 1032–1044 (2013).
Shrestha, B. R. et al. Amyloid β peptide adversely affects spine number and motility in hippocampal neurons. Mol Cell Neurosci 33, 274–282 (2006).
Wei, W. et al. Amyloid beta from axons and dendrites reduces local spine number and plasticity. Nat Neurosci 13, 190–196 (2010).
Shankar, G. M. et al. Natural oligomers of the Alzheimer amyloid-β protein induce reversible synapse loss by modulating an NMDA-type glutamate receptor-dependent signaling pathway. J Neurosci 27, 2866–2875 (2007).
Pozueta, J. et al. Caspase-2 is required for dendritic spine and behavioural alterations in J20 APP transgenic mice. Nat Commun 4, 1939 (2013).
Shankar, G. M. et al. Amyloid-β protein dimers isolated directly from Alzheimer’s brains impair synaptic plasticity and memory. Nat Med 14, 837–842 (2008).
Sudhof, T. C. Neuroligins and neurexins link synaptic function to cognitive disease. Nature 455, 903–911 (2008).
Craig, A. M. & Kang, Y. Neurexin-neuroligin signaling in synapse development. Curr Opin Neurobiol 17, 43–52 (2007).
Siddiqui, T. J. & Craig, A. M. Synaptic organizing complexes. Curr Opin Neurobiol 21, 132–143 (2011).
Bemben, M. A., Shipman, S. L., Nicoll, R. A. & Roche, K. W. The cellular and molecular landscape of neuroligins. Trends Neurosci 38, 496–505 (2015).
Takahashi, H. & Craig, A. M. Protein tyrosine phosphatases PTPδ, PTPσ, and LAR: presynaptic hubs for synapse organization. Trends Neurosci 36, 522–534 (2013).
Takahashi, H. et al. Postsynaptic TrkC and presynaptic PTPσ function as a bidirectional excitatory synaptic organizing complex. Neuron 69, 287–303 (2011).
Takahashi, H. et al. Selective control of inhibitory synapse development by Slitrk3-PTPδ trans-synaptic interaction. Nat Neurosci 15, 389–398 (2012).
Yim, Y. S. et al. Slitrks control excitatory and inhibitory synapse formation with LAR receptor protein tyrosine phosphatases. Proc Natl Acad Sci USA 110, 4057–4062 (2013).
Linhoff, M. W. et al. An unbiased expression screen for synaptogenic proteins identifies the LRRTM protein family as synaptic organizers. Neuron 61, 734–749 (2009).
de Wit, J. et al. LRRTM2 interacts with Neurexin1 and regulates excitatory synapse formation. Neuron 64, 799–806 (2009).
Ko, J., Fuccillo, M. V., Malenka, R. C. & Sudhof, T. C. LRRTM2 functions as a neurexin ligand in promoting excitatory synapse formation. Neuron 64, 791–798 (2009).
Siddiqui, T. J., Pancaroglu, R., Kang, Y., Rooyakkers, A. & Craig, A. M. LRRTMs and neuroligins bind neurexins with a differential code to cooperate in glutamate synapse development. J Neurosci 30, 7495–7506 (2010).
Um, J. W. et al. LRRTM3 Regulates Excitatory Synapse Development through Alternative Splicing and Neurexin Binding. Cell Rep 14, 808–822 (2016).
Pettem, K. L. et al. The Specific α-Neurexin Interactor Calsyntenin-3 Promotes Excitatory and Inhibitory Synapse Development. Neuron 80, 113–128 (2013).
Matsuda, K. et al. Cbln1 is a ligand for an orphan glutamate receptor δ2, a bidirectional synapse organizer. Science 328, 363–368 (2010).
Matsuda, K. & Yuzaki, M. Cbln family proteins promote synapse formation by regulating distinct neurexin signaling pathways in various brain regions. Eur J Neurosci 33, 1447–1461 (2011).
Uemura, T. et al. Trans-synaptic interaction of GluRδ2 and Neurexin through Cbln1 mediates synapse formation in the cerebellum. Cell 141, 1068–1079 (2010).
Woo, J. et al. Trans-synaptic adhesion between NGL-3 and LAR regulates the formation of excitatory synapses. Nat Neurosci 12, 428–437 (2009).
Kwon, S. K., Woo, J., Kim, S. Y., Kim, H. & Kim, E. Trans-synaptic adhesions between netrin-G ligand-3 (NGL-3) and receptor tyrosine phosphatases LAR, protein-tyrosine phosphatase δ (PTPδ), and PTPσ via specific domains regulate excitatory synapse formation. J Biol Chem 285, 13966–13978 (2010).
Valnegri, P. et al. The X-linked intellectual disability protein IL1RAPL1 regulates excitatory synapse formation by binding PTPδ and RhoGAP2. Hum Mol Genet 20, 4797–4809 (2011).
Yoshida, T. et al. IL-1 receptor accessory protein-like 1 associated with mental retardation and autism mediates synapse formation by trans-synaptic interaction with protein tyrosine phosphatase δ. J Neurosci 31, 13485–13499 (2011).
Yoshida, T. et al. Interleukin-1 receptor accessory protein organizes neuronal synaptogenesis as a cell adhesion molecule. J Neurosci 32, 2588–2600 (2012).
Saura, C. A., Servian-Morilla, E. & Scholl, F. G. Presenilin/γ-secretase regulates neurexin processing at synapses. PLoS One 6, e19430 (2011).
Bot, N., Schweizer, C., Ben Halima, S. & Fraering, P. C. Processing of the synaptic cell adhesion molecule neurexin-3β by Alzheimer disease α- and γ-secretases. J Biol Chem 286, 2762–2773 (2011).
Haapasalo, A. et al. Presenilin/γ-secretase-mediated cleavage regulates association of leukocyte-common antigen-related (LAR) receptor tyrosine phosphatase with β-catenin. J Biol Chem 282, 9063–9072 (2007).
Viswanathan, J. et al. Ubiquilin-1 modulates γ-secretase-mediated ε-site cleavage in neuronal cells. Biochemistry 52, 3899–3912 (2013).
Suzuki, K. et al. Activity-dependent proteolytic cleavage of neuroligin-1. Neuron 76, 410–422 (2012).
Dinamarca, M. C., Weinstein, D., Monasterio, O. & Inestrosa, N. C. The synaptic protein neuroligin-1 interacts with the amyloid β-peptide. Is there a role in Alzheimer’s disease? Biochemistry 50, 8127–8137 (2011).
Kim, T. et al. Human LilrB2 is a beta-amyloid receptor and its murine homolog PirB regulates synaptic plasticity in an Alzheimer’s model. Science 341, 1399–1404 (2013).
Lauren, J., Gimbel, D. A., Nygaard, H. B., Gilbert, J. W. & Strittmatter, S. M. Cellular prion protein mediates impairment of synaptic plasticity by amyloid-β oligomers. Nature 457, 1128–1132 (2009).
Reissner, C., Runkel, F. & Missler, M. Neurexins. Genome biology 14, 213 (2013).
Graf, E. R., Kang, Y., Hauner, A. M. & Craig, A. M. Structure function and splice site analysis of the synaptogenic activity of the neurexin-1β LNS domain. J Neurosci 26, 4256–4265 (2006).
Miesenbock, G., De Angelis, D. A. & Rothman, J. E. Visualizing secretion and synaptic transmission with pH-sensitive green fluorescent proteins. Nature 394, 192–195 (1998).
Mucke, L. et al. High-level neuronal expression of Aβ1–42 in wild-type human amyloid protein precursor transgenic mice: synaptotoxicity without plaque formation. J Neurosci 20, 4050–4058 (2000).
Um, J. W. et al. Metabotropic glutamate receptor 5 is a coreceptor for Alzheimer Aβ oligomer bound to cellular prion protein. Neuron 79, 887–902 (2013).
Hu, N. W. et al. mGlu5 receptors and cellular prion protein mediate amyloid-β-facilitated synaptic long-term depression in vivo . Nat Commun 5, 3374 (2014).
Cisse, M. et al. Reversing EphB2 depletion rescues cognitive functions in Alzheimer model. Nature 469, 47–52 (2011).
Scheff, S. W., DeKosky, S. T. & Price, D. A. Quantitative assessment of cortical synaptic density in Alzheimer’s disease. Neurobiol Aging 11, 29–37 (1990).
Missler, M. et al. α-neurexins couple Ca2+ channels to synaptic vesicle exocytosis. Nature 423, 939–948 (2003).
Anderson, G. R. et al. β-Neurexins Control Neural Circuits by Regulating Synaptic Endocannabinoid Signaling. Cell 162, 593–606 (2015).
Abramov, E. et al. Amyloid-β as a positive endogenous regulator of release probability at hippocampal synapses. Nat Neurosci 12, 1567–1576 (2009).
Fuccillo, M. V. et al. Single-Cell mRNA Profiling Reveals Cell-Type-Specific Expression of Neurexin Isoforms. Neuron 87, 326–340 (2015).
Caetano, F. A. et al. Amyloid-beta oligomers increase the localization of prion protein at the cell surface. J Neurochem 117, 538–553 (2011).
Acknowledgements
We thank Ayako Takahashi for excellent preparation of neuron cultures and Dr. Cristina Vasuta and Youssouf Soumounou for technical supports. We thank Dr. Lennart Mucke (Gladstone Institute of Neurological Disease and Department of Neurology, UCSF, CA) and the J. David Gladstone Institutes for the transgenic hAPPSwe,Ind (J20 line) mouse breeders. This work was supported by the Alzheimer Society Research Program Biomedical Young Investigator Grant, Canadian Institutes of Health Research (CIHR) grant MOP-133517, the Fonds de la recherche du Québec Research Scholars (Junior 2) and a Scottish Rite Charitable Foundation of Canada Research Grant to H.T., CIHR grant MOP-126001 to E.H., the Alzheimer Society Research Program Biomedical Doctoral Awards to Y.N. and the 2016-IRCM-Michel-Bélanger Scholarship to A.K.L.
Author information
Authors and Affiliations
Contributions
Y.N. performed a majority of the experiments including cell surface binding screen and assays, pull-down experiments, coculture experiments and time-lapse imaging. Y.T. prepared synaptosomal fractions of J20 APP mice and western blot analysis. A.K.L. performed binding assays. E.H. prepared J20 APP mice for synaptome experiments. H.T. supervised the project. Y.N. and H.T. conceived the project and prepared the manuscript with critical input from E.H. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Naito, Y., Tanabe, Y., Lee, A. et al. Amyloid-β Oligomers Interact with Neurexin and Diminish Neurexin-mediated Excitatory Presynaptic Organization. Sci Rep 7, 42548 (2017). https://doi.org/10.1038/srep42548
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep42548
- Springer Nature Limited
This article is cited by
-
A Peptide Motif Covering Splice Site B in Neuroligin-1 Binds to Aβ and Acts as a Neprilysin Inhibitor
Molecular Neurobiology (2024)
-
A novel rhein-huprine hybrid ameliorates disease-modifying properties in preclinical mice model of Alzheimer’s disease exacerbated with high fat diet
Cell & Bioscience (2023)
-
Finding memo: versatile interactions of the VPS10p-Domain receptors in Alzheimer’s disease
Molecular Neurodegeneration (2022)
-
1-(7-Chloroquinolin-4-yl)-N-(4-Methoxybenzyl)-5-Methyl-1H-1,2, 3-Triazole-4- carboxamide Reduces Aβ Formation and Tau Phosphorylation in Cellular Models of Alzheimer’s Disease
Neurochemical Research (2022)
-
Neurexin 3 transmembrane and soluble isoform expression and splicing haplotype are associated with neuron inflammasome and Alzheimer’s disease
Alzheimer's Research & Therapy (2019)