Abstract
The nitrogen-metabolic phosphotransferase system, PTSNtr, consists of the enzymes INtr, NPr and IIANtr that are encoded by ptsP, ptsO, and ptsN, respectively. Due to the proximity of ptsO and ptsN to rpoN, the PTSNtr system has been postulated to be closely related with nitrogen metabolism. To define the correlation between PTSNtr and nitrogen metabolism, we performed ligand fishing with EIIANtr as a bait and revealed that D-glucosamine-6-phosphate synthase (GlmS) directly interacted with EIIANtr. GlmS, which converts D-fructose-6-phosphate (Fru6P) into D-glucosamine-6-phosphate (GlcN6P), is a key enzyme producing amino sugars through glutamine hydrolysis. Amino sugar is an essential structural building block for bacterial peptidoglycan and LPS. We further verified that EIIANtr inhibited GlmS activity by direct interaction in a phosphorylation-state-dependent manner. EIIANtr was dephosphorylated in response to excessive nitrogen sources and was rapidly degraded by Lon protease upon amino sugar depletion. The regulation of GlmS activity by EIIANtr and the modulation of glmS translation by RapZ suggest that the genes comprising the rpoN operon play a key role in maintaining amino sugar homeostasis in response to nitrogen availability and the amino sugar concentration in the bacterial cytoplasm.
Similar content being viewed by others
Introduction
The phosphotransferase system (PTS) is a systematic bacterial device composed of a cascade of enzymes that transfer phosphate moieties derived from phosphoenolpyruvate (PEP) in sequential order. The canonical PTS that occurs in a wide range of bacteria is a sugar PTS called the phosphoenolpyruvate:carbohydrate PTS, which is comprised of enzyme I (EI), histidine phosphocarrier protein (HPr), and enzyme II complexes (EIIA, EIIB, EIIC, and sometimes EIID). The EIIA, EIIB, and EIIC proteins usually form substrate-specific transfer cascades with one membrane-spanning protein, EIIC in general, in contact with their extracellular substrates1. In contrast, EI and HPr are universal cytoplasmic proteins, which take-up diverse carbohydrates, such as sugars and their derivatives1. In addition to the uptake and concomitant phosphorylation of many carbohydrates, the PTS also conducts diverse regulatory functions, sensing available carbon sources2,3,4,5. The PTS accomplishes regulatory roles either by phosphorylating target proteins or by directly interacting with their target proteins. Proteins containing a specific PTS-recognizable phosphorylation domain are phosphorylated by the components of the PTS phosphorylation cascade and their activities are modulated. The phosphorylatable target proteins include a variety of non-PTS transporters and transcription regulators5. In the latter case, phosphorylated or unphosphorylated forms of the PTS proteins directly interact with target proteins, leading to activation or repression of their functions mainly involved in transport and transcription regulation6,7. EIIAGlc-mediated catabolite repression is a well-established regulatory mechanism by the PTS8,9. Unphosphorylated EIIAGlc under glucose abundant conditions interacts with proteins necessary for the transport and metabolism of non-PTS carbohydrates, such as lactose, maltose, and glycerol, and inhibits their activities for PTS-catalyzed uptake of glucose as the preferred sugar. Interestingly, numerous Proteobacteria possess incomplete PTS cascades devoid of any known EIIB or EIIC proteins5. Thus, an incomplete PTS lacking substrate-specific EIIB and EIIC proteins is supposed to conduct regulatory functions by interacting with non-PTS substrates, instead of taking up sugar.
Extensive genome analysis revealed a paralog of the sugar PTS, called nitrogen PTS (PTSNtr) in many Proteobacteria10. However, the repertoire is incomplete. In parallel with the sugar PTS, the PTSNtr possesses EINtr (an EI paralog encoded by ptsP) and NPr (an HPr paralog encoded by ptsO), which catalyze phosphorylation of EIIANtr (an EIIAMtl paralog encoded by ptsN) with a PEP-derived phosphoryl group but the PTSNtr is devoid of any known counterparts of the membrane-bound EIIB and EIIC enzymes, which transport extracellular substrates10,11,12. Accordingly, the PTSNtr has been speculated to conduct regulatory functions exclusively using EIIANtr as the output regulator protein rather than a component of transport machinery13. An increasing number of data show that PTSNtr is associated with a plethora of cellular processes, including virulence7,14, nitrogen metabolism10,15, carbon metabolism16, K+ homeostasis6,17,18, and (p)ppGpp synthesis/hydrolysis19,20.
The ptsN gene encoding EIIANtr is located downstream of rpoN in many Proteobacteria21. An alternative σ54 sigma factor encoded by rpoN participates in the expression of diverse genes and operons exclusively associated with nitrogen utilization and metabolism22. A genetic approach was employed on the rpoN operon containing ptsN to unravel the primary role of PTSNtr conserved in many bacteria during evolution and revealed that the ancestral rpoN operon composed of at least 11 genes has evolved to retain rpoN, yhbH, ptsN, rapZ, and ptsO in many Gammaproteobacteria13,23, suggesting a conserved function for the remaining genes. Localization of the ptsN and ptsO PTSNtr genes in the rpoN operon suggests a role for PTSNtr associated with nitrogen metabolism. Due to the absence of a component responsible for uptake and concomitant phosphorylation of a specific substrate, it has been unclear what stimuli determine the phosphorylation status of the PTSNtr components. However, it was recently revealed that EINtr senses nitrogen availability through the glutamine (Gln) and α-ketoglutarate (α-KG) ratio and accordingly modulates the phosphorylation status of the EIIANtr output regulator in E. coli24. Nitrogen is prerequisite for producing proteins, nucleic acids, and cell wall constituents. Therefore, it is critical for bacteria to sense availability of cellular nitrogen sources and allot them depending on the circumstances. The functional relevance of the genes comprising the rpoN operon to nitrogen utilization for cell wall construction has been frequently demonstrated. NPr encoded by ptsO decreases biosynthesis of lipid A in the lipopolysaccharides (LPS) layer by inhibiting LpxD activity and blocking the inflow of UDP-GlcNAc into cell wall constituents25. RapZ encoded by rapZ negatively controls synthesis of the D-glucosamine-6-phosphate synthase (GlmS) by triggering decay of GlmZ, a small RNA facilitating glmS translation26. GlmS is a key enzyme in the LPS and peptidoglycan biosynthetic pathways. The association between σ54, a transcription factor encoded by rpoN, and bacterial exterior constitution has been observed in many bacteria27,28. Co-clustering of PTSNtr genes with rpoN and convergent negative roles of the rpoN operon genes during synthesis of cell envelope constituents have led to exploring a concordant role for EIIANtr in cell envelope formation in response to nitrogen abundance. In this study, we discovered direct protein-protein interaction between EIIANtr and GlmS and suggest a novel role for ptsN regulating amino sugar biosynthesis, which is contextually associated with adjacent genes.
Results
Specific interaction between Enzyme IIANtr and glucosamine-6-phosphate synthase (GlmS) in Salmonella Typhimurium
Although the general sugar PTS components exert regulatory functions by either phosphorylating or interacting with target proteins, PTSNtr EIIANtr seems to have a bias toward interaction-mediated regulation. Many studies have demonstrated direct interactions between unphosphorylated EIIANtr and diverse proteins involved in K+ transport6,17, virulence regulation7, and the ppGpp-mediated stringent response19,20. A common theme of these EIIANtr interactions is that unphosphorylated EIIANtr prevails during the protein-protein interaction. To search for target proteins interacting with EIIANtr, EIIANtr tagged with His6 at its C-terminus (EIIANtr-His6) was isolated in its unphosphorylated form and used as bait in a ligand-fishing strategy with Escherichia coli as a representative Gammaproteobacteria6,21. Here, to archive more diversity in binding partners, Salmonella enterica serovar Typhimurium, a member of Enterobacteriaceae with high genetic similarity to E. coli, was chosen. EIIANtr-His6 was incubated with a crude protein extract of Salmonella Typhimurium SL1344, and the proteins bound to EIIANtr-His6 were pulled down with a metal-affinity resin. The sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) analysis of isolated protein complexes from ligand-fishing experiments revealed a protein band of approximately 70 kDa that specifically co-precipitated with EIIANtr-His6 (Fig. 1a). Liquid chromatography-tandem mass spectrometry (LC-MS/MS) peptide mapping identified this protein as GlmS. Other proteins predicted together with GlmS in the LC-MS/MS analysis are listed in Supplementary Table S2. The specific interaction between EIIANtr and GlmS was further validated in vivo. His6-GlmS and EIIANtr were produced simultaneously under the lac promoter of the pETDuet plasmid in E. coli, and the crude protein extract was passed through a Ni-NTA resin column. EIIANtr was pulled down together with His6-GlmS (Fig. 1b), indicating direct interaction between EIIANtr and GlmS in vitro and in vivo.
EIIANtr specifically binds to glucosamine-6-phosphate synthase (GlmS).
(a) Ligand-fishing using EIIANtr-His6 as a bait. Total cell extract from S. Typhimurium SL1344 culture was incubated with EIIANtr-His6 or not. The proteins eluted from Ni-NTA resin (lane 1, whole cell extract only; lane 2, whole cell extract mixed with EIIANtr-His6) and purified EIIANtr-His6 (lane 3) were analyzed by SDS-PAGE. The protein band indicated by the arrow in lane 2 was identified as GlmS by LC-MS/MS analysis. Size markers (M) in kDa are aligned at left. (b) Co-purification of EIIANtr and GlmS in vivo. E. coli BL21 (DE3) harboring pETDuet-glmS·ptsN was geneticallymanipulated to produce EIIANtr and His6-tagged GlmS simultaneously by isopropyl β-D-1-thiogalactoside (IPTG) addition. EIIANtr bound to His6-GlmS was co-purified using Ni-NTA metal-affinity resin. Proteins analyzed in SDS-PAGE are aliquots from total cell extracts (T), supernatant after centrifuging the cell extracts (S), column flow-through (F), wash (W), and the five elution fractions (E1–E5). Molecular masses of standards are presented in kDa on the left.
Phosphorylation status of EIIANtr influences its binding affinity with GlmS
An intimate association between PTSNtr and nitrogen metabolism has been proposed undoubtedly through genetic localization of ptsN and ptsO in the rpoN operon and multiple experimental results. The intracellular Gln: α-KG ratio indicates the balance between nitrogen and carbon sources and controls the regulatory circuit assigned to assimilate extracellular ammonia into Gln29 and the phosphorylation status of PTSNtr. The EINtr GAF domain senses the abundance of nitrogen through a high Gln/α-KG ratio, dampens autophosphorylation of EINtr, and dephosphorylates EIIANtr in turn24. Other ptsO mutants unable to phosphorylate EIIANtr reduce the expression of genes required for nitrogen assimilation and glutamine synthetase, suggesting a negative role for unphosphorylated EIIANtr during nitrogen incorporation into Gln30,31. It is plausible that EIIANtr responding to nitrogen availability orchestrates the rate of nitrogen acquisition and its utilization in other cellular components. However, it remains unresolved which molecule interacts with phosphorylated or unphosphorylated EIIANtr in the nitrogen metabolic and utilization pathways. The finding of an interaction between EIIANtr and GlmS may provide a clue to define the role of PTSNtr in nitrogen metabolism, as GlmS (D-glucosamine-6-phosphate synthase, EC 2.6.1.16) exploits Gln to produce equimolar D-glucosamine-6-phosphate (GlcN6P), lowering intracellular nitrogen resources.
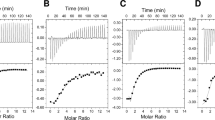
As demonstrated in other EIIANtr binding partners, we assumed that the unphosphorylated status of EIIANtr was preferred during complex formation with GlmS. Unphosphorylated EIIANtr under high Gln concentrations was likely to bind GlmS and stimulate its GlcN6P production with Gln consumption. However, to our surprise, the binding affinity of EIIANtr to GlmS was enhanced by phosphorylation. For this observation, Salmonella EIIANtr-His6 was modified to EIIANtr-His6 (K75D) (Fig. 2a) to exhibit a phosphorylation-dependent mobility shift (PDMS) on SDS-PAGE without changing its functional properties (Supplementary Fig. S1), as established in E. coli24. Salmonella EIIANtr-His6 (K75D) showed an upshift in mobility after PEP-derived phosphorylation (Fig. 2b) and was downshifted by adding Gln, indicating its unphosphorylated state under excessive nitrogen concentrations (Fig. 2c), as expected. When phosphorylated or unphosphorylated EIIANtr-His6 (K75D) was incubated with GlmS, phosphorylated EIIANtr recruited more GlmS than did the unphosphorylated form in a concentration-dependent manner but EIIAGlc used as a control did not interact (Fig. 3a and Supplementary Fig. S2A). The possibility of non-specific interaction of GlmS with anti-His6 antibody or Ni-NTA resin was ruled out as detailed in Supplementary Fig. S3A,B.
EIIANtr phosphorylation status is influenced by nitrogen abundance.
(a) EIIANtr modification strategy. The number of SDS molecules bound to a protein is affected by the charge of the amino acids around the phosphorylation site, which can change the mobility of the protein on SDS-PAGE24,54. A histidine (73) amino acid of EIIANtr was changed to alanine (73) to construct the unphosphorylated form of EIIANtr (H73A), and a lysine (75) was substituted for aspartic acid (75) to provide negative charge effects on EIIANtr. (b) Phosphorylation-dependent upshift of EIIANtr (K75D). The intact form of EIIANtr and its EIIANtr (K75D) mutant derivative were incubated with or without 1 mM PEP under phosphorylating conditions and then analyzed by SDS-PAGE. EIIANtr (K75D) showed excellent phosphorylation-dependent mobility shift (PDMS), whereas the intact form of EIIANtr showed comparable mobility independent of its phosphorylation. (c) Differential phosphorylation status of EIIANtr between metabolites. The phosphorylation-dependent mobility shift of EIIANtr-His6 (K75D) was assessed using different metabolites. EIIANtr-His6 (K75D) was incubated with PEP, His6-EINtr, and His6-NPr in the presence of glutamine (Gln) or α-ketoglutarate (α-KG). The phosphorylation levels of EIIANtr-His6 (K75D) were compared by SDS-PAGE.
Phosphorylation status of EIIANtr influences the binding affinity between EIIANtr and GlmS.
(a) Increased binding affinities of EIIANtr-His6 (K75D) to GlmS after phosphorylation. Phosphorylated and unphosphorylated forms of EIIANtr-His6 (K75D) were incubated with equivalent amounts of GlmS, and the levels of GlmS bound to phosphorylated (P) or unphosphorylated (U) EIIANtr-His6 (K75D) were compared (top). PEP-dependent phosphorylation of EIIANtr-His6 (K75D) was verified in parallel (bottom inlet). Line (N) does not contain either the EIIANtr-His6 (K75D) or the GlmS protein as a control. (b) Differential interaction between EIIANtr and GlmS depending on EIIANtr phosphorylation status in vivo. Protein-protein interactions between EIIANtr and GlmS were verified using a bacterial two-hybrid system. Plasmid pKT25 containing ptsN or ptsN (H73A) and plasmid pUT18 harboring glmS were introduced into a reporter strain respectively or in combination. The reporter strains were cultivated in LB broth supplemented with IPTG, and β-galactosidase activity was determined to examine the strength of the protein-protein interactions. This experiment was performed in triplicate.
The differential binding affinity of EIIANtr to GlmS due to its phosphorylation status was further verified in vivo using a bacterial two-hybrid system32. EIIANtr and GlmS fused to two different sub-domains of Bordetella pertussis adenylate cyclase bound each other and elevated cAMP levels. In accordance with the in vitro observations, EIIANtr (H73A), a derivative with a mutation at the phosphorylation site33, caused lower cAMP levels than that of intact EIIANtr (Figs 2a and 3b). This result was also confirmed by the different color intensities between bacterial cultures producing phosphorylatable or unphosphorylatable EIIANtr in the medium containing 5-bromo-4-chloro-3-indolyl-β-D-galactoside (X-gal) (Supplementary Fig. S2B).
EIIANtr inhibits GlmS activity
EIIANtr, unlike other interactions with TrkA6, KdpD17, E1 of pyruvate dehydrogenase33, and SsrB7, prefers the phosphorylation state to interact with GlmS. To decipher the link between PTSNtr and nitrogen or amino acid metabolism, the influence of EIIANtr on GlmS activity was studied. EIIANtr is known to exert both positive and negative effects on the roles of partner proteins17,33. GlmS converts D-fructose-6-phosphate (Fru6P) into GlcN6P by hydrolyzing Gln to glutamate (Glu). An enzymatic assay for GlmS was set up using Gln, Fru6P, and GlmS, and GlcN6P production was measured by high performance liquid chromatography (HPLC) (Supplementary Fig. S4A,B). Then, GlmS was incubated with the phospho- or unphospho-forms of EIIANtr-His6 (K75D) and GlcN6P production was compared (Fig. 4a). Adding EIIANtr-His6 (K75D) decreased GlcN6P production, and its phosphorylation caused a more drastic reduction of GlcN6P production, indicating a negative role of EIIANtr in GlmS activity.
EIIANtr inhibits GlmS-mediated GlcN6P production.
(a) Negative effects of EIIANtr on GlcN6P production. GlmS was incubated with different amounts of phosphorylated or unphosphorylated EIIANtr-His6 (K75D) (0–16 pmol) and GlcN6P production was measured by HPLC. The results from triplicates are plotted. (b) No effect of EIIANtr on the expression of genes involved in amino sugar metabolism. Wild-type and ΔptsN mutant strains were transformed with pWJ04 containing ptsN and its presumable promoter, and qRT-PCR was conducted to compare mRNA levels of glmS, glmU, glmM, and ptsN. All mRNA levels were normalized using gyrB, and the relative expression ratios were averaged from three independent measurements.
GlmS exerts negative feedback regulation in response to GlcN6P. Excessive GlcN6P production stimulates degradation of glmS mRNA26,34. To define the negative effect of EIIANtr on amino sugar production in detail, the levels of mRNAs relevant to amino sugar metabolism were evaluated in the presence or absence of EIIANtr. Quantitative reveres transcription-polymerase chain reaction (qRT-PCR) revealed that the mRNA levels of glmS, glmM, and glmU required to convert Fru6P to uridinediphospho-N-acetylglucosamine (UDP-GlcNAc), a main amino sugar substrate for cell wall structure, were not affected by EIIANtr (Fig. 4b), indicating that EIIANtr modulates GlmS activity by protein-protein interaction and not by transcriptional regulation.
EIIANtr affects the GlmS-mediated bacterial growth rate
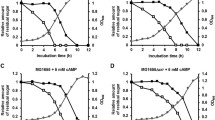
Inhibiting GlmS activity through the interaction with EIIANtr was further confirmed in vivo. GlmS is a key enzyme in the production of amino sugars, which are essential precursors for bacterial cell wall peptidoglycans and LPS in the outer membrane of Gram-negative bacteria. Therefore, bacteria deprived of functional GlmS suffer from attenuated growth due to a lack of building blocks for constructing the cell wall when exogenous amino sugars are depleted35,36. To examine whether growth of Salmonella is attenuated by the interaction between EIIANtr and GlmS, a balanced-lethal system was established in which the absence of chromosomal glmS was partially complemented by controlled expression of glmS on the pWJ10 plasmid containing the lac promoter (Supplementary Fig. S5). A ΔglmS mutant strain with a severe growth defect due to the lack of cell envelope components37 was supplemented with pWJ10 in the presence of 10 μM IPTG, showing approximately 50% growth of wild-type Salmonella (Fig. 5a). The additional deletion of the ptsN gene in the ΔglmS mutant containing pWJ10 restored growth to a level comparable to that of wild-type cells, suggesting that GlmS was relieved of the inhibition by EIIANtr. Accordingly, introducing plasmids producing intact EIIANtr or its derivative, EIIANtr (H73A), re-repressed the growth of the ΔglmS ΔptsN strain containing pWJ10. As expected, EIIANtr (H73A), which is incapable of phosphorylation in response to stimuli15, showed a less inhibitory effect than the intact form probably due to diminished binding affinity to GlmS (Fig. 5b). Examination of viable cell number under the same condition showed the consistent result with that of bacterial growth rate analysis described above (Supplementary Fig. S6).
Salmonella growth is influenced by the interaction between EIIANtr and GlmS.
(a) The absence of glmS is lethal to bacteria. Effects of GlmS and EIIANtr on bacterial growth were assessed individually or in combination. Introduction of pWJ10 (a pUHE21-2lacIq::glmS) providing GlmS in trans partially restored the growth defect in ΔglmS mutant. GlmS production level was titrated using 10 μM IPTG (Supplementary Fig. S5). (b) Modulation of growth rate by the interaction between EIIANtr and GlmS. Using the ΔglmS mutant complemented with pWJ10 as a parent strain, the effect of the EIIANtr interaction with GlmS on bacterial growth was evaluated by deleting the chromosomal ptsN gene and introducing pWJ04 and pWJ05, which provided EIIANtr and EIIANtr (H73A), respectively. All growth measurements in (a,b) were performed in triplicate, and the average optical densities at 600 nm are plotted.
These results verify that EIIANtr-mediated GlmS inhibition might lead to a growth defect attributable to the impaired integrity of the cell wall and further imply a role for EIIANtr controlling the production rate of amino sugars depending on nitrogen accessibility.
EIIANtr and GlmS interact to form a heterotrimeric complex
A cascade of enzymes, including GlmS, GlmM, and GlmU are required for bona fide amino sugar synthesis. GlmS, which is responsible for the first and rate-limiting step in the hexosamine pathway, converts Fru6P into GlcN6P. GlcN6P is subsequently modified to glucosamine-1-phosphate (GlcN1P) by GlmM and further converted to UDP-GlcNAc by GlmU38,39,40. However, in the presence of abundant exogenous N-acetylglucosamine (GlcNAc) transported via NagE and ManXYZ, the GlmS-catalyzed reaction is bypassed, whereas GlmM and GlmU are consistently required for its conversion into UDP-GlcNAc41. Hence, the activity of GlmS is selectively shut down in response to amino sugar availability. As glmS and glmU constitute the glmUS operon and their transcripts are strictly coupled42, there should be a regulatory mechanism differentiating synthesis and activity of GlmS from those of GlmU after transcription. For example, translation of glmS mRNA is separately controlled by the GlmY/GlmZ/RapZ system, which responds to the cellular concentration of GlcN6P26, as described in detail below. In addition, GlmS changes its structure from an active dimer to an inactive hexamer as its cellular concentration increases and GlcN6P is accumulated as the product43. To refine the interaction between EIIANtr and GlmS in greater detail, we determined the stoichiometric mass-action model in the EIIANtr and GlmS complex. We first assumed the architectural contribution of EIIANtr toward converting the GlmS structure into the inactive hexameric form: EIIANtr binding likely helps GlmS molecules conglomerate in an inactive form without repulsion or stabilizes the inactive polymeric structure to prevent its return to the active dimer form. To examine this possibility, EIIANtr and GlmS in solution were subjected to size exclusion chromatography with multi-angle light scattering (SEC-MALS) in combination or individually (Fig. 6a). GlmS peaked at about 128 kDa, which is equivalent to the mass of the dimeric form of GlmS, and EIIANtr was maintained as a monomer, showing a peak of around 20 kDa in solution. During SEC, the MALS detected a large molecule of approximately 148 kDa in the solution containing GlmS and EIIANtr, suggesting two molecules of GlmS obstructed by a molecule of EIIANtr. The peaks were further dissected by SDS-PAGE to define the protein composition (Fig. 6b). These results suggest that EIIANtr inhibits GlmS by binding to its active dimeric form but is less likely to contribute to conversion of GlmS into the inactive hexameric form.
Trimeric complex is formed between EIIANtr and GlmS.
(a) Stoichiometric analysis of the EIIANtr and GlmS complex. A solution of the EIIANtr and GlmS complex was analyzed by SEC-MALS. The molecular mass of GlmS alone in solution was 128 kDa, indicating the active dimer, and that of EIIANtr was 20 kDa, indicating the monomeric form. A macromolecule with a peak of 148 kDa appeared in the solution composed of EIIANtr and GlmS, and the size was accordant with a EIIANtr-GlmS2 heterotrimer composed of two GlmS molecules (128 kDa) and one EIIANtr (20 kDa). (b) Identifying the eluted fractions by SDS-PAGE. The SEC-MALS eluted fractions from the 148, 128, and 20 kDa proteins, which were presumably the EIIANtr-GlmS2 complex, GlmS dimer, and the EIIANtr monomer, respectively, were analyzed by SDS-PAGE and Coomassie Blue staining. Green, red, and blue lines indicate the 148, 128, and 20 kDa fractions, respectively.
EIIANtr stability is modulated by Lon in response to amino sugar availability
Maintaining GlcN6P homeostasis has been ascribed to the GlmY/GlmZ/RapZ system24. RapZ inhibits GlmS expression by accelerating the decay of the small RNA GlmZ, which facilitates glmS translation. However, bacteria sensing an amino sugar deficiency exploit GlmY to antagonize GlmZ degradation by RapZ. Co-localization of ptsN and rapZ in the rpoN operon reinforces the concordant role of these two genes in GlmS-mediated regulation of cell envelope integrity. We suspected that amino sugar was the signal triggering coordinated negative regulation of EIIANtr and RapZ on GlmS. However, ptsN and rapZ mRNA levels were not affected by amino sugar availability, whereas glmS transcription was negatively feedback regulated by ample amino sugars, as reported previously26,44 (Supplementary Figs S7A,B and S8A,B). Instead, intriguingly, the level of the EIIANtr protein decreased when GlcN6P production stopped due to the lack of GlmS and when no exogenous GlcNAc, a metabolite substitute for GlcN6P26,44,45 was supplied (Fig. 7a). In contrast, adding GlcNAc increased the levels of EIIANtr regardless of the presence of GlmS (Fig. 7a). These results indicate that intracellular amino sugar availability controls EIIANtr at the protein level.
EIIANtr degradation is controlled by Lon protease in response to amino sugar availability.
(a) Comparison of EIIANtr-FLAG protein levels between wild-type and ΔglmS mutant strains. Protein samples were isolated from wild-type and ΔglmS mutant strains in the presence or absence of GlcNAc 3 h post inoculation as described in Supplementary Figs S8A and S10, and subjected to Western blot analysis to compare the EIIANtr-FLAG levels between conditions. (b) Effect of amino sugar availability on stability of EIIANtr-FLAG. Chloramphenicol was added to the cultures of the wild-type and ΔglmS mutant strains at 3 h as described above, and total proteins harvested every 30 min were used to assess the EIIANtr and DnaK decay rates. (c) Comparison of the stability of EIIANtr-FLAG in the absence of Lon or ClpXP protease in the ΔglmS mutant strain. The ΔglmS, ΔglmS Δlon, and ΔglmS ΔclpXP Salmonella strains were cultured under conditions identical to those used above, and the level of EIIANtr-FLAG and DnaK was assessed in each strain using Western blotting every 30 min after adding chloramphenicol. In all experiments, equivalent amounts of total protein were loaded in each lane for SDS-PAGE and DnaK levels were measured in parallel to verify equivalent total protein amounts between lanes.
To further explore modulation of intracellular EIIANtr concentration depending on amino sugar availability, the rate of EIIANtr proteolysis was compared between amino sugar-abundant and -depleted conditions. To prevent de novo protein synthesis, chloramphenicol was added to the culture 3 h post-inoculation when amino sugar availability influenced bacterial growth rate (Supplementary Fig. S9)26,46. The ΔglmS mutant strain depleted of amino sugars rapidly degraded EIIANtr-FLAG but retained EIIANtr-FLAG at high levels for at least 2 h when provided with excessive GlcNAc, indicating regulation of the EIIANtr proteolysis rate by amino sugar availability (Fig. 7b). Lon and Clp proteases in E. coli and other bacteria are responsible for 70–80% of energy-dependent protein degradation47,48. We observed that deleting lon quenched the programmed proteolysis of EIIANtr-FLAG upon amino sugar depletion in the ΔglmS mutant strain (Fig. 7c). In this experiment, DnaK levels were measured in parallel to verify equivalent total protein amounts between lanes, and a similar test using RpoB, which was used as an additional control instead of DnaK, was also conducted (Supplementary Fig. S10). These results indicate that Lon accelerates EIIANtr degradation when Salmonella lacks amino sugar metabolites.
Discussion
Nitrogen is essential to every living organism, including bacteria; it is assimilated into amino acids and further into proteins, which constitute cellular components that conduct diverse biological activities. Nitrogen is also used in amino sugar compounds, the structural residues of bacterial cell walls. The nitrogen moiety of Gln is integrated into Fru6P by GlmS, producing GlcN6P. GlcN6P is further converted to UDP-GlcNAc by a cascade of cytoplasmic enzymes, such as GlmM and GlmU. UDP-GlcNAc is an essential structural building block for peptidoglycans and LPS in bacterial cell walls. Therefore, the balance between synthesis and decomposition of amino sugars is directly influenced by the availability of carbon and nitrogen.
Cells under excessive nitrogen convert α-KG to Glu and further to Gln49. Gln is utilized as a nitrogen source under nitrogen-limiting conditions50. Hence, the Gln: α-KG ratio is generally used to predict the cellular balance between nitrogen and carbon abundance: high Gln/α-KG values under nitrogen-rich conditions vs. low Gln/α-KG values under nitrogen-depleted conditions. Bacteria must promptly recognize detrimental changes, such as nutrient deprivation, and adjust metabolic processes for their adaptation. For example, it has been recently revealed that Caulobacter crescentus triggers the phosphorylation of EINtr, the first enzyme of PTSNtr in response to low glutamine concentrations and subsequently phosphorylates EIIANtr, which in turn inhibits the hydrolase activity of SpoT by directly binding to SpoT. Modulation of SpoT activity by PTSNtr influences the cellular accumulation of (p)ppGpp, an alarmone controlling bacterial cell cycle progression and growth20. A similar role of PTSNtr sensing the nitrogen availability and modulating cellular metabolism was disclosed in this study. GlmS is the rate-limiting enzyme that consumes a molecule of Gln to synthesize an equivalent amount of the amino sugar GlcN6P. Thus, GlmS activity is tightly controlled in response to the availability of cellular nitrogen and amino sugars. Based on our results, we propose a regulatory circuit wherein EIIANtr fine-tunes GlmS activity by assessing the abundance of nitrogen and amino sugars (Fig. 8). In the presence of sufficient nitrogen, dephosphorylated EIIANtr does not significantly compromise GlmS activity, which, in turn, provides abundant amino sugars for constructing cell walls. When cells suffer from depleted nitrogen, phosphorylated EIIANtr tightly binds to GlmS and inhibits the enzyme from consuming Gln to produce GlcN6P, which slows down LPS and peptidoglycan synthesis. Not only nitrogen availability but also amino sugar abundance modulates the influence of EIIANtr on GlmS activity. Bacteria supplemented with abundant amino sugars, such as GlcNAc, delay the decay of EIIANtr and subsequently decelerate bona fide amino sugar synthesis via GlmS. However, when the supply of exogenous amino sugars is suspended and the GlcN6P remnants are exhausted, Lon accelerates EIIANtr proteolysis and relieves GlmS to supplement the lack of cellular amino sugars.
Proposed regulatory model for the control of GlmS activity by EIIANtr in response to nitrogen and amino sugar availability.
At high cellular glutamine (Gln) concentrations, EIIANtr tends to be unphosphorylated and liberates the active form of GlmS that supplies GlcN6P and accelerates synthesis of the bacterial cell envelope (①). When GlcN6P is present in excess (②) or when Gln availability is restricted (③), phosphorylated EIIANtr binds GlmS with increased affinity and inhibits its activity. The depletion of pre-existing GlcN6P leads to EIIANtr degradation by Lon protease, freeing the active form of GlmS to supplement the lack of GlcN6P and maintain amino sugar homeostasis (④).
GlmS is a pivotal enzyme for maintaining cell wall integrity and employs multiple latch devices for regulation. GlmS maintains its own structural equilibrium between the active dimeric conformation and the inactive hexameric state by shifting toward the inactive hexameric form when its concentration increases and the GlcN6P product accumulates43. In addition to the conformational dynamics of GlmS, its translation is also controlled by the coordinated action between two regulatory GlmY and GlmZ small RNAs and the RapZ RNase adaptor protein. Following dissociation of the glmS monocistronic transcript from the glmUS cotranscript by RNaseE-mediated processing, GlmZ base-pairs with glmS mRNA and the aid of Hfq activates translation of glmS mRNA51. GlmZ turnover is determined by binding with RapZ, which recruits RNase E to the complex to facilitate GlmZ degradation. However, under low cellular GlcN6P concentrations, GlmY, with a secondary structure similar to GlmZ, increases and sequesters RapZ from GlmZ as a decoy, liberating GlmZ to activate glmS translation and subsequent GlcN6P production26. The results in our study established that GlmS is not only controlled at the post-transcriptional level by GlmY/GlmZ/RapZ but is also coordinated at the post-translational level by EIIANtr and Lon. PTSNtr senses accessibility of nitrogen and controls the phosphorylation status of EIIANtr. Phosphorylated or unphosphorylated EIIANtr hampers catalytic GlmS activity by binding to the active duplex form with different affinities. However, GlmS is released from the inhibition by Lon-mediated degradation of EIIANtr upon depletion of cellular amino sugars.
Because of the incomplete architecture of PTSNtr lacking the phosphate recipient coupled with EIIANtr, EIIANtr has been speculated to serve as a substantive regulator, and its pleiotropic regulation has been observed in diverse cellular processes, including nitrogen metabolism, K+ homeostasis, and interactions with host cells7,17,33. However, the pleiotropic regulatory effects of PTSNtr may not be completely attributable to the direct interactions between PTSNtr components and cellular targets; rather, many of the regulatory outputs might be secondarily caused by a few unmediated processes operated by EIIANtr. K+ homeostasis controlled by direct binding of EIIANtr to TrkA and KdpD would explain such a mechanistic link6,17. The cellular K+ concentration managed by EIIANtr could be a manifold signal leading to activation or inhibition of enzymes, transcriptional regulation, and pH homeostasis13.
Similarly, additional direct targets of EIIANtr may yet to be identified, and GlmS is one of them. Direct regulation of GlmS by EIIANtr discloses a sophisticated regulatory circuit balancing metabolite fluxes among carbon, nitrogen, and amino sugars. The possibility of a metabolic link between carbon and nitrogen has been undoubtedly raised. The sugar PTS and nitrogen PTS cross-talk by phosphorylating counterpart components10,52. α-KG has been identified as a key molecule balancing carbon and nitrogen assimilation and controlling EIIANtr regulatory activity. A challenge for bacteria living in fluctuating nutritional conditions is to notice the accessibility of essential nutrients, such as carbon and nitrogen, and determine the flux rates from nutrients to energy production and building biomass. Doucette et al. observed that a sudden increase in nitrogen availability led to an immediate glucose uptake and discovered that α-KG, which accumulates under nitrogen-depleted conditions, inhibits the autophosphorylation of enzyme I (EI) of sugar PTS and blocks the entry of glucose, synchronizing carbon uptake with nitrogen availability49. α-KG which is accumulated in nitrogen-limiting conditions, in turn, binds to the GAF domain of NifA, a transcriptional activator for nitrogen fixation (nif) genes, and triggers nitrogen uptake into the cell53. Cells fortified with nitrogen convert α-KG to Glu and further to Gln and subsequently resume glucose uptake by releasing the sugar PTS EI from α-KG inhibition. Intriguingly, the GAF domain responding to α-KG is also possessed by EINtr of PTSNtr and a nitrogen deficiency is directly delivered to EIIANtr 54, which dampens GlmS activity, thereby shutting down inflow of nitrogen into amino sugars. Thus, one could speculate that α-KG serves as an interface signal integrating cellular physiological status and coordinating carbon and nitrogen assimilation; physiological status is further transformed to EIIANtr across the PTSNtr phosphorelay system, which accommodates amino sugar biosynthesis on demand.
It was observed that the unphosphorylated form of EIIANtr is exclusively dominant during rapid growth of Pseudomonas putida, a member of Gammaproteobacteria, and suggested that EIIANtr phosphorylation status might be influenced by cellular physiological status representing the availability of carbon and nitrogen55. However, it is unknown how phosphorylation of EIIANtr alters bacterial fitness during environmental adaptation. Our results suggest that EIIANtr, which is phosphorylated under nitrogen-limiting conditions, compromises bona fide amino sugar biosynthesis by inhibiting GlmS and decelerates production of peptidoglycans and LPS. Gammaproteobacteria with the rpoN operon comprising the PTSNtr genes and rapZ have presumably evolved the rpoN operon to control amino sugar homeostasis in accordance with nitrogen availability.
Methods
Bacterial strains, plasmids, and culture conditions
Salmonella was genetically manipulated using the phage λ Red recombination system56 and phage P22-mediated transduction57 with Salmonella enterica serovar Typhimurium SL1344 as the parent strain. All bacterial strains and plasmids used in this study are listed in Supplementary Table S1. Bacteria were grown aerobically at 37 °C in LB or M9 minimal medium supplemented with nutrients as described. Antibiotics were used at the following concentrations: ampicillin, 50 μg/ml; chloramphenicol, 25 μg/ml; and kanamycin, 50 μg/ml. A detailed description strain and plasmid construction is provided in the Supplementary information.
Ligand-fishing to search for EIIANtr-His6 protein targets
Salmonella Typhimurium SL1344 cells were grown overnight in 300 ml LB broth at 37 °C with shaking and were harvested and resuspended in 10 ml lysis buffer [20 mM Tris-HCl (pH 8.0) and 300 mM NaCl]. The cells were disrupted by sonication on ice and then pelleted by centrifugation at 15,000 g for 1 h at 4 °C. The supernatant was mixed with 1 mg EIIANtr-His6 or not and was further incubated with 300 μl Ni-NTA resin for 1 h at 4 °C. The incubated mixture was loaded onto a Poly-Prep chromatography column (8 × 40 mm) (Bio-Rad, Hercules, CA, USA) and washed with 5 ml washing buffer [20 mM Tris-HCl (pH 8.0), 300 mM NaCl, and 5 mM imidazole], and the proteins bound to the resin were eluted with elution buffer [20 mM Tris-HCl (pH 8.0), 300 mM NaCl, and 250 mM imidazole]. Aliquots of the eluted protein samples were analyzed by SDS-PAGE and stained with Coomassie Brilliant Blue G. The protein that was specifically bound to EIIANtr-His6 was excised from the gel and subjected to in-gel digestion with trypsin and LC-MS/MS analysis as described previously58.
Immunoprecipitation of EIIANtr using His6-Glms in vivo
E. coli BL21 (DE3) cells, which produce GlmS tagged with His6 at its N-terminus (His6-GlmS) and EIIANtr on pETDuet-glmS·ptsN, were grown in 200 ml LB, and protein expression was induced by adding 1 mM IPTG at OD600 of 2.0 for 6 h. The cell suspension was disrupted by sonication and centrifuged, and the supernatant was mixed with 1 ml Ni-NTA metal-affinity resin. After the mixture was loaded onto a Poly-Prep chromatography column, the column was washed four times with 2 ml washing buffer [20 mM Tris-HCl (pH 8.0), 300 mM NaCl, and 20 mM imidazole]. The proteins bound to the column were eluted with 500 μl of elution buffer [20 mM Tris-HCl (pH 8.0), 300 mM NaCl, and 50–200 mM imidazole]. Aliquots from the total cell extract (T), the supernatant after the cell extract was centrifuged (S), column flow-through (F), wash from washing (W), and the eluted fractions (E) were separated on 12% SDS-PAGE, and the gels were analyzed after staining with Coomassie Brilliant Blue G.
Bacterial two-hybrid system for studying protein-protein interactions
The ptsN gene and its derivative and glmS gene were fused in-frame to the 3′-end of the cyaA gene fragments in pKT25 and pUT18C32,59, respectively, as described in the supplemental materials and methods. The E. coli reporter strain BTH101 was transformed with the bait and prey plasmids and cultured in LB broth containing IPTG (0.5 mM). To determine the interactions between proteins in the β-galactosidase assay, cell lysates in working buffer [70 mM Na2HPO4-H2O, 30 mM NaHPO4-H2O, 1 mM MgSO4, 0.2 mM MnSO4, and 100 mM β-mercaptoethanol] were incubated with o-nitrophenol-β-galactoside (4 mg/ml) as a substrate, and absorbance values at 420 nm and 550 nm were transformed into Miller units60.
In vitro phosphorylation assay
To measure the Phosphorylation-Dependent Mobility Shift (PDMS) of EIIANtr, EIIANtr-His6 (K75D) (2 μg) was incubated with His6-EINtr (1 μg) and His6-NPr (1 μg) in the presence or absence of PEP (1 mM) in a total volume of 20 μl containing 0.1 M Tris-HCl (pH 7.5), 2 mM MgCl2, 1 mM EDTA, 10 mM KCl, and 0.5 mM dithiothreitol22. After a 20 min incubation at 37 °C, the reaction was stopped by adding 5 μl 4× SDS-Sample Buffer (L1100-001; GeneDEPOT, Barker, TX, USA) [250 mM Tris-HCl (pH 6.8), 40% glycerol, 8% SDS, and 8% β-mercaptoethanol], and the aliquots were analyzed by 15% SDS-PAGE. The proteins were stained with Coomassie Brilliant Blue G. To examine the effect of various metabolites on PTSNtr phosphotransferase activity, EIIANtr-His6 (K75D) (3 μg) was incubated with PEP (1 mM), His6-EINtr (0.3 μg), and His6-NPr (0.3 μg) in the presence of Gln, α-KG, or GlcN6P at 5 mM each. After a 5 min incubation at 37 °C, the reactions were stopped and processed as described above.
In vitro protein-protein interactions between phosphorylated/unphosphorylated EIIANtr and GlmS
To compare the binding affinities of EIIANtr to GlmS depending on phosphorylation status, EIIANtr-His6 (K75D) (100 μg) was incubated with His6-EINtr (10 μg) and His6-NPr (10 μg) in the presence or absence of PEP (2 mM) in a total volume of 100 μl. PEP-dependent phosphorylation of EIIANtr-His6 (K75D) was verified by 15% SDS-PAGE analysis as described above, and the remainder (95 μl of each) that was not used in EIIANtr-His6 (K75D) PDMS was mixed with Ni-NTA resin (50 μl) in 1 ml binding buffer [20 mM Tris-HCl (pH 8.0), and 300 mM NaCl] and incubated at 4 °C for 30 min. Purified GlmS (100 μg) was cleaved by thrombin to detach the His6-tag from His6-GlmS and was added to each reaction and incubate for 30 min at 4 °C under the same conditions. To dissociate the bound proteins, elution buffer [20 mM Tris-HCl (pH 8.0), 300 mM NaCl, and 250 mM imidazole] was passed through the Ni-NTA resin several times. The eluent was subsequently analyzed by SDS-PAGE, followed by Coomassie Brilliant Blue staining.
Size exclusion chromatography with multi-angle light scattering
SEC-MALS experiments were performed using a fast protein liquid chromatography system (GE Healthcare) connected to a Wyatt MiniDAWN TREOS MALS instrument and a Wyatt Optilab rEX differential refractometer. A Superdex 200 10/300 GL (GE Healthcare) gel-filtration column pre-equilibrated with equilibrium buffer [20 mM Tris-HCl (pH 8.0) and 300 mM NaCl] was normalized using ovalbumin protein. Proteins (2 mg) were injected at a flow rate of 0.5 ml/min. Data were analyzed using the Zimm model for static light-scattering data fitting and graphed using EASI graph with a UV peak in ASTRA V software (Wyatt Technology Corp., Goleta, CA, USA).
Glucosamine-6-phosphate synthase activity assay
The following stock solutions were prepared before the GlmS activity assay: 0.5 M Tris-HCl (pH 7.5), 0.5 M KCl, 10 mM EDTA (pH 7.5), 60 mM Fru6P, and 50 mM L-Gln. A pre-warmed solution containing 6 mM Fru6P, 10 mM L-Gln in 50 mM Tris-HCl (pH 7.5), 50 mM KCl, and 1 mM EDTA was mixed with GlmS (0–16 pmol) and incubated at 37 °C for 30 min. GlmS activity was inactivated by heating at 80 °C for 20 min, and the GlcN6P product was measured by HPLC. To determine the effects of EIIANtr on GlmS activity, different amounts of EIIANtr-His6 (K75D) (0–16 pmol) were added to the reaction, and the levels of GlcN6P produced were compared. Reaction mixtures without GlmS or Fru6P were used as negative controls. A detailed description of the procedure can be found in ref. 61.
RNA isolation and quantitative real-time RT-PCR
Salmonella strains were grown in LB or W-salts medium24. W-salts medium was supplemented with 20 mM alanine and 0.26 mM histidine. Different carbon sources were added separately when required: 0.2% GlcNAc, 0.2% glucose or 0.2% glycerol. Total RNAs were isolated at the mid-log phase using the RNeasy mini kit (Qiagen, Valencia, CA, USA). After DNase treatment of the isolated total RNAs, cDNA was synthesized with the RNA to cDNA EcoDryTM Premix and random hexamers (Clontech, Palo Alto, CA, USA). The synthesized cDNA was mixed with 2 × iQ SYBR Green Supermix (Bio-Rad), and real-time PCR was performed using the CFX 3.1 (Bio-Rad). mRNA expression levels of target genes were normalized relative to that of the gyrB (DNA gyrase subunit B). All qRT-PCR primer sets used in this study are listed in the Supplementary Table S4.
SDS-PAGE and Western blotting analysis
Protein samples (purified or interaction complexes) or whole-cell fractions (cell extract after sonication or cell pellets) were dissolved in Laemmli sample buffer and boiled for 5 min. The protein samples loaded onto 12% or 15% SDS-PAGE gels were separated based on molecular weights, and transferred to a polyvinylidene difluoride membrane. The membrane was blocked with 5% nonfat dry milk in 1× Tris-buffered saline-Tween 20 (TBS-T) buffer and probed with anti-FLAG antibody (3:2,000 dilution, F1804; Sigma, St. Louis, MO, USA), anti-His6 antibody (3:2,000 dilution, sc-8036; Santa Cruz Biotechnology, Santa Cruz, CA, USA), anti-DnaK antibody (1:10,000 dilution, ADI-SPA-880; Enzo Life Science, Farmingdale, NY, USA) or anti-RpoB antibody (1:10,000 dilution, sc-56766; Santa Cruz Biotechnology, Santa Cruz, CA, USA) as primary antibodies. Anti-mouse IgG conjugated with peroxidase (3:5,000 dilution, sc-2005; Santa Cruz Biotechnology) was used as the secondary antibody in all Western blots. The chemiluminescent signals were developed with a West-Zol plus Western blot detection system (Intron Biotechnology, Seoul, South Korea).
Additional Information
How to cite this article: Yoo, W. et al. Fine-tuning of amino sugar homeostasis by EIIANtr in Salmonella Typhimurium. Sci. Rep. 6, 33055; doi: 10.1038/srep33055 (2016).
References
Postma, P. W., Lengeler, J. W. & Jacobson, G. R. Phosphoenolpyruvate:carbohydrate phosphotransferase systems of bacteria. Microbiol. Rev. 57, 543–594 (1993).
Görke, B. & Stülke, J. Carbon catabolite repression in bacteria: many ways to make the most out of nutrients. Nat. Rev. Microbiol. 6, 613–624 (2008).
Lengeler, J. W. & Jahreis, K. Bacterial PEP-dependent carbohydrate: phosphotransferase systems couple sensing and global control mechanisms. Contrib. Microbiol. 16, 65–87 (2009).
Escalante, A., Cervantes, A. S., Gosset, G. & Bolivar, F. Current knowledge of the Escherichia coli phosphoenolpyruvate-carbohydrate phosphotransferase system: peculiarities of regulation and impact on growth and product formation. Appl. Microbiol. Biotechnol. 94, 1483–1494 (2012).
Deutscher, J. et al. The bacterial phosphoenolpyruvate:carbohydrate phosphotransferase system: regulation by protein phosphorylation and phosphorylation-dependent protein-protein interactions. Microbiol. Mol. Biol. Rev. 78, 231–256 (2014).
Lee, C. R., Cho, S. H., Yoon, M. J., Peterkofsky, A. & Seok, Y. J. Escherichia coli enzyme IIANtr regulates the K+ transporter TrkA. Proc. Natl. Acad. Sci. USA 104, 4124–4129 (2007).
Choi, J. et al. Salmonella pathogenicity island 2 expression negatively controlled by EIIANtr-SsrB interaction is required for Salmonella virulence. Proc. Natl. Acad. Sci. USA 107, 20506–20511 (2010).
Harwood, J. P., Gazdar, C., Prasad, C. & Peterkofsky, A. Involvement of the glucose enzymes II of the sugar phosphotransferase system in the regulation of adenylate cyclase by glucose in Escherichia coli. J. Biol. Chem. 251, 2462–2468 (1976).
Feucht, B. U. & Saier, M. H. Jr. Fine control of adenylate cyclase by the phosphoenolpyruvate:sugar phosphotransferase systems in Escherichia coli and Salmonella typhimurium. J. Bacteriol. 141, 603–610 (1980).
Powell, B. S. et al. Novel proteins of the phosphotransferase system encoded within the rpoN operon of Escherichia coli. J. Biol. Chem. 270, 4822–4839 (1995).
Reizer, J. et al. Novel phosphotransferase-encoding genes revealed by analysis of the Escherichia coli genome: a chimeric gene encoding an Enzyme I homologue that possesses a putative sensory transduction domain. Gene 181, 103–108 (1996).
Peterkofsky, A., Wang, G. & Seok, Y. J. Parallel PTS systems. Arch. Biochem. Biophys. 453, 101–107 (2006).
Pflüger-Grau, K. & Görke, B. Regulatory roles of the bacterial nitrogen-related phosphotransferase system. Trends. Microbiol. 18, 205–214 (2010).
Higa, F. & Edelstein, P. H. Potential virulence role of the Legionella pneumophila ptsP ortholog. Infect. Immun. 69, 4782–4789 (2001).
Lee, C. R. et al. Requirement of the dephospho-form of enzyme IIANtr for derepression of Escherichia coli K-12 ilvBN expression. Mol. Microbiol. 58, 334–344 (2005).
Jahn, S., Haverkorn van Rijsewijk, B. R., Sauer, U. & Bettenbrock, K. A role for EIIANtr in controlling fluxes in the central metabolism of E. coli K12. BioChim. Biophys. Acta. 1833, 2879–2889 (2013).
Lüttmann, D. et al. Stimulation of the potassium sensor KdpD kinase activity by interaction with the phosphotransferase protein IIANtr in Escherichia coli. Mol. Microbiol. 72, 978–994 (2009).
Reaves, M. L. & Rabinowitz, J. D. Characteristic phenotype associated with ptsN-null mutants in Escherichia coli K-12 are absent in strains with functional ilvG. J. Bacteriol. 193, 4576–4581 (2011).
Karstens, K., Zschiedrich, C. P., Bowien, O., Stülke, J. & Görke, B. Phosphotransferase protein EIIANtr interacts with SpoT, a key enzyme of the stringent response in Ralstonia eutropha H16. Microbiol. 160, 711–722 (2014).
Ronneau, S., Petit, K., Bolle, X. D. & Hallez, R. Phosphotransferase-dependent accumulation of (p)ppGpp in response to glutamine deprivation in Caulobacter crescentus. Nat. Commun. 7, 11423 (2016).
Boёl, G. et al. Transcription regulators protentially controlled by HPr kinase-phosphorylase in gram-negative bacteria. J. Mol. Microbiol. Biotechnol. 5, 206–215 (2003).
Reitzer, L. & Schneider, B. L. Metabolic context and possible physiological themes of σ54-dependent genes in Escherichia coli. Microbiol. Mol. Biol. Rev. 65, 422–444 (2001).
Comas, I., González-Candelas, F. & Zúñiga, M. Unraveling the evolutionary history of the phosphoryl-transfer chain of the phosphoenolpyruvate:phosphotransferase system through phylogenetic analyses and genome context. BMC. Evol. Biol. 8, 147 (2008).
Lee, C. R. et al. Reciprocal regulation of the autophosphorylation of enzyme INtr by glutamine and alpha-ketoglutarate in Escherichia coli. Mol. Microbiol. 88, 473–485 (2013).
Kim, H. J., Lee, C. R., Kim, M., Peterkofsky, A. & Seok, Y. J. Dephosphorylated NPr of the nitrogen PTS regulates lipid A biosynthesis by direct interaction with LpxD. Biochem. Biophys. Res. Commun. 409, 556–561 (2011).
Göpel, Y., Papenfort, K., Reichenbach, B., Vogel, J. & Görke, B. Targeted decay of a regulatory small RNA by an adaptor protein for RNase E and counteraction by an anti-adaptor RNA. Genes. Dev. 27, 552–564 (2013).
Fisher, M. A. et al. Borrelia burgdorferi σ54 is required for mammalian infection and vector transmission but not for tick colonization. Proc. Natl. Acad. Sci. USA 102, 5162–5167 (2005).
Francke, C. et al. Comparative analyses imply that the enigmatic sigma factor 54 is a central controller of the bacterial exterior. BMC. Genomics. 12, 385 (2011).
Ninfa, A. J. & Atkinson, M. R. PII signal transduction proteins. Trends. Microbiol. 8, 172–179 (2000).
Merrick, M. J., Taylor, M., Saier, M. H. Jr. & Reizer, J. The role of genes downstream of the σN structural gene rpoN in Klebsiella pneumoniae. Nitrogen Fixation: Fundamentals and Applications 27, 189–194 (1995).
Merrick, M. J. & Coppard, J. R. Mutations in genes downstream of the rpoN gene (encoding σ54) of Klebsiella pneumoniae affect expression from σ54-dependent promoters. Mol. Microbiol. 3, 1765–1775 (1989).
Karimova, G., Pidoux, J., Ullmann, A. & Ladant, D. A bacterial two-hybrid system based on a reconstituted signal transduction pathway. Proc. Natl. Acad. Sci. USA 95, 5752–5756 (1998).
Pflüger-Grau, K., Chavarría, M. & Lorenzo, V. The interplay of the EIIANtr component of the nitrogen-related phosphotransferase system (PTSNtr) of Pseudomonas putida with pyruvate dehydrogenase. BioChim. Biophys. Acta. 1810, 995–1005 (2011).
Collins, J. A., Irnov, I., Baker, S. & Winkler, W. C. Mechanism of mRNA destabilization by the glmS ribozyme. Genes. Dev. 21, 3356–3368 (2007).
Sarvas, M. Mutant of Escherichia coli K-12 defective in D-glucosamine biosynthesis. J. Bacteriol. 105, 467–471 (1971).
Plumbridge, J. A. et al. Coordinated regulation of amino sugar-synthesizing and -degrading enzymes in Escherichia coli K-12. J. Bacteriol. 175, 4951–4956 (1993).
Kim, K. et al. A novel balanced-lethal host-vector system based on glmS. PLoS One 8, e60511 (2013).
Mengin-Lecreulx, D. & Van, H. J. Identification of the glmU gene encoding N-acetylglucosamine-1-phosphate uridyltransferase in Escherichia coli. J. Bacteriol. 175, 6150–6157 (1993).
Mengin-Lecreulx, D. & Van, H. J. Copurification of glucosamine-1-phosphate acetyltransferase and N-acetylglucosamine-1-phosphate uridyltransferase activities of Escherichia coli: Characterization of the glmU gene product as a bifunctional enzyme catalyzing two subsequent steps in the pathway for UDP-N-acetylglucosamine synthesis. J. Bacteriol. 176, 5788–5795 (1994).
Mengin-Lecreulx, D. & Van, H. J. Characterization of the essential gene glmM encoding phosphoglucosamine mutase in Escherichia coli. J. Biol. Chem. 271, 32–39 (1996).
Durand, P., Golinelli-Pimpaneau, B., Mouilleron, S., Badet, B. & Badet-Denisot, M. A. Highlights of glucosamine-6P synthase catalysis. Arch. Biochem. Biophys. 474, 302–317 (2008).
Plumbridge, J. Co-ordinated regulation of amino sugar biosynthesis and degradation: the NagC repressor acts as both an activator and a repressor for the transcription of the glmUS operon and requires two separated NagC binding sites. EMBO J. 14, 3958–3965 (1995).
Mouilleron, S. et al. Structural basis for morpheein-type allosteric regulation of Escherichia coli glucosamine-6-phosphate synthase: equilibrium between inactive hexamer and active dimer. J. Biol. Chem. 287, 34533–34546 (2012).
Kalamorz, F., Reichenbach, B., März, W., Rak, B. & Görke, B. Feedback control of glucosamine-6-phosphate synthase GlmS expression depends on the small RNA GlmZ and involves the novel protein YhbJ in Escherichia coli. Mol. Microbiol. 65, 1518–1533 (2007).
Kawada-Matsuo, M. et al. GlmS and NagB regulate amino sugar metabolism in opposing directions and affect Streptococcus mutans virulence. PLoS One 7, e33382 (2012).
Weisberger, A. S. Inhibition of protein synthesis by chloramphenicol. Annu. Rev. Med. 18, 483–494 (1967).
Maurizi, M. R. Proteases and protein degradation in Escherichia coli. Experimentia 48, 178–201 (1992).
Biran, D., Gur, E., Gollan, L. & Ron, E. Z. Control of methionine biosynthesis in Escherichia coli by proteolysis. Mol. Microbiol. 37, 1436–1443 (2000).
Doucette, C. D., Schwab, D. J., Wingreen, N. S. & Rabinowitz, J. D. alpha-ketoglutarate coordinates carbon and nitrogen utilization via enzyme I inhibition. Nat. Chem. Biol. 7, 894–901 (2011).
Magasanik, B. The regulation of nitrogen utilization in enteric bacteria. J. Cell. Biochem. 51, 34–40 (1993).
Görke, B. & Vogel, J. Noncoding RNA control of the making and breaking of sugars. Genes. Dev. 22, 2914–2925 (2008).
Rabus, R., Reizer, J., Paulsen, I. & Saier, M. H. Jr. Enzyme INtr from Escherichia coli. A novel enzyme of the phosphoenolpyruvate-dependent phosphotransferase system exhibiting strict specificity for its phosphoryl acceptor, NPr. J. Biol. Chem. 274, 26185–26191 (1999).
Little, R. & Dixon, R. The amino-terminal GAF domain of Azotobacter vinelandii NifA binds 2-oxoglutarate to resist inhibition by NifL under nitrogen-limiting conditions. J. Biol. Chem. 278, 28711–28718 (2003).
Lee, C. R., Park, Y. H., Kim, Y. R., Peterkofsky, A. & Seok, Y. J. Phosphorylation-dependent mobility shift of proteins on SDS-PAGE is due to decreased binding of SDS. Bull. Korean. Chem. Soc. 34, 2063–2066 (2013).
Pflüger-Grau, K. & Lorenzo, V. Growth-dependent phosphorylation of the PtsN (EIINtr) protein of Pseudomonas putida. J. Biol. Chem. 282, 18206–18211 (2007).
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl. Acad. Sci. USA 97, 6640–6645 (2000).
Chan, R. K., Botstein, D., Watanabe, T. & Ogata, Y. Specialized transduction of tetracycline resistance by phage P22 in Salmonella typhimurium. Virol. 50, 883–898 (1972).
Jeong, J. Y. et al. Expression of ptsG encoding the major glucose transporter is regulated by ArcA in Escherichia coli. J. Biol. Chem. 279, 38513–38518 (2004).
Karimova, G., Ullmann, A. & Ladant, D. A bacterial two-hybrid system that exploits a cAMP signaling cascade in Escherichia coli. Methods. Enzymol. 328, 59–73 (2000).
Miller, J. H. Experiments in Molecular Genetics (Cold Spring Harbor Laboratory Press, Plainview, NY) (1972).
Li, Y. et al. An enzyme-coupled assay for amidotransferase activity of glucosamine-6-phosphate synthase. Anal. Biochem. 370, 142–146 (2007).
Acknowledgements
This study was supported by a grant (14162MFDS972) from the Ministry of Food and Drug Safety, Korea in 2016.
Author information
Authors and Affiliations
Contributions
W.Y., H.Y. and S.R. designed experiments, prepared the Figures and wrote the manuscript. W.Y. performed experiments. W.Y., H.Y., Y.-J.S., C.-R.L., H.H.L. and S.R. analyzed the data and reviewed the manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Yoo, W., Yoon, H., Seok, YJ. et al. Fine-tuning of amino sugar homeostasis by EIIANtr in Salmonella Typhimurium. Sci Rep 6, 33055 (2016). https://doi.org/10.1038/srep33055
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep33055
- Springer Nature Limited
This article is cited by
-
Retrospective application of transposon-directed insertion-site sequencing to investigate niche-specific virulence of Salmonella Typhimurium in cattle
BMC Genomics (2019)
-
Determination of protein phosphorylation by polyacrylamide gel electrophoresis
Journal of Microbiology (2019)
-
Enzyme IIANtr Regulates Salmonella Invasion Via 1,2-Propanediol And Propionate Catabolism
Scientific Reports (2017)