Abstract
Pegaspargase combined with gemcitabine have greatly improved the outcomes of advanced extranodal NK/T cell lymphoma (ENKL). However, patients frequently undergo recurrent disease due to chemoresistance, and few predictive parameters are available. The present study explored potential biomarkers to predict the therapeutic response of advanced ENKL treated with pegaspargase/gemcitabine and evaluate the prognostic significance. Through serum proteomic analysis, we identified 61 upregulated and 22 downregulated proteins in nonresponders compared with responders. We further validated that patients with unfavourable treatment outcomes displayed higher levels of S100A9 and ORM1 via enzyme-linked immunosorbent assay (ELISA). Moreover, the sensitivity and specificity for detecting refractory patients were 81.5% and 71.4% for S100A9 > 62.0 ng/ml, 85.2% and 77.1% for ORM1 > 1436 ug/ml, 100% and 57.1% for S100A9 combined with ORM1. Furthermore, in multivariate analysis elevated levels of S100A9 were associated with poor 2-year OS (40.2% vs. 76.6%, RR = 2.92, p = 0.005) and 2-year PFS (33.1% vs. 61.1%, RR = 2.61 p = 0.011). High ORM1 also predicted inferior 2-year OS (38.7% vs.76.1, RR = 2.46, p = 0.023) and 2-year PFS (18.4% vs. 73.2%, RR = 2.86, p = 0.009). Our results indicated that S100A9 and ORM1 could serve as reliable predictors of therapeutic response and independent prognostic factors of survival in advanced ENKL patients treated with pegaspargase/gemcitabine.
Similar content being viewed by others
Introduction
Extranodal NK/T cell lymphoma (ENKL) represents a highly aggressive lymphoid malignancy characterized by dismal survival. As an Epstein-Barr Virus (EBV)-associated disease, populations from Asian countries and South America are more frequently affected1. Gene expression profiling (GEP) analysis suggested that activation of JAK-STAT, NF-κB and MAPK signaling pathways contributes to the pathogenesis of ENKL2. Due to the rarity of this disease, standard therapies for ENKL have not been established. Most localized patients could achieve remission after receiving radiotherapy and chemotherapy3. As to advanced ENKL patients, high-dose chemotherapy containing pegaspargase, L-asparaginase, gemcitabine are recommended. The SMILE (methotrexate, ifosfamide, L-asparaginase, dexamethasone and etoposide) regimen and DDGP (gemcitabine, pegaspargase, cisplatin, and dexamethasone) regimen have showed promising efficacy4,5. Unfortunately, some patients still experience refractory disease and undergo disease progression. Currently, few parameters except EBV-DNA copy numbers are reported to be correlated with therapeutic response in ENKL6,7. For those nonresponders and high risk patients, modified therapy strategy would prolong their survival. Investigation of reliable biomarkers for predicting treatment response are urgently needed.
Proteomic technology has emerged as a valuable tool to screen alterations in protein profilings of serum8. In cancer development, crucial proteins from cancer cells and tumor microenvironment are secreted into serum. Serum samples provide us a great much information. Biochemical analysis of serum may serve as a noninvasive method for cancer diagnosis and prognosis. Through quantitative proteomic analysis, specific protein signatures for meningiomas with different grades were identified9. We previously employed proteomic techniques and identified a protein that could distinguish healthy individuals and lymphoma patients10.
Here, we utilized proteomic approach to identify differentially expressed proteins between chemosensitive- and chemoresistant- advanced ENKL patients. Nonresponders presented higher levels of S100A9 and ORM1 than responders. Elevated levels of S100A9 and ORM1 were associated with shorter PFS and OS. Our results indicated that serum S100A9 and ORM1 could serve as powerful predictors of chemoresponsiveness and prognostic factors of survival in advanced ENKL treated with pegaspargase/gemcitabine.
Materials and Methods
Patients
For isobaric tags for relative and absolute quantitation (iTRAQ) and LC-MS/MS (liquid chromatography tandem mass spectrometry) analysis, pretreatment blood samples from 32 advanced ENKL patients were collected. For validation set, another cohorts of 62 advanced ENKL patients were enrolled. Blood samples were placed at room temperature for 1 hour to allow coagulation, and then centrifuged at 3500 rpm for 10 min at 4 °C. Serums were then aliquoted and stored at −80 °C until further experiment. All patients received DDGP (gemcitabine, pegaspargase, cisplatin, and dexamethasone) chemotherapy regimen. Assessment of treatment response was based on Revised Cheson’s standard response criteria. Complete remission (CR), partial response (PR), stable disease (SD), progressive disease (PD) were defined as previously described5. According to response and PFS, patients were divided into responders and nonresponders. Responders were those who achieved CR or PR and did not undergo disease progression within 6 months after therapy. Nonresponders were defined as those who achieved SD/PD after initial therapy or those who obtained temporary remission but relapsed within 6 months.
This study was approved by the ethics committee of the First Affiliated Hospital of Zhengzhou University. All experiments were performed in accordance with approved guidelines and regulations. All patients signed an informed consent when enrolled.
iTRAQ labeling and strong cation exchange (SCX) chromatography
Equal amounts of serum from each sample were acquired and pooled. Multiple Affinity Removal LC Column-Human 14 (Agilent) was employed to remove high abundance serum proteins. Bradford assay was performed to determine the concentrations of proteins. Then, proteins were digested into peptides with trypsin and desalted with C18-SD Extraction Disk Cartridge (66872-U sigma). Peptides were labeled with reagents following the iTRAQ kit (AB SCIEX) instructions and then mixed. For LC-MS/MS analysis, the complexity of peptides sample was firstly reduced utilizing strong cation exchange (SCX) chromatography. The labeled peptides were separated using a Polysulfoethyl column (4.6 × 100 mm, 5 um, 200 Å, PolyLCInc, Maryland, USA) on the AKTA Purifier 100 system (GE Healthcare). The peptide fractions were collected at an interval of one minute and grouped into 12 components according to SCX chromatograms. Components were lyophilized in a vacuum concentrator and desalted using C18 columns.
LC-MS/MS
Each component was further analyzed using a Q Exactive mass spectrometer (Thermo Finnigan) connected to the Easy nano Liquid Chromatography system. Briefly, dried fraction was resuspended in buffer A (0.1% formic acid) and packed with C18-reversed phase column (2 cm × 100 μm, 5 μm, Thermo Scientific). Peptides were eluted onto an analytical C18 column (75 um × 100 mm, 3 μm, Thermo Scientific) with a gradients of buffer B (84% acetonitrile and 0.1% Formic acid) at a flow rate of 300 nl/min over 60 min. The mass spectrometer was performed at a data-dependent mode. In a full MS scan range of m/z 300–1800, the ten most abundant precursor ions were choosed for high-energy collisional dissociation (HCD) fragmentation. Survey scans were acquired at a resolution of 70,000 at m/z 200 and resolution for HCD spectra was set to 17,500 at m/z 200.
Protein identification and bioinformatic analysis
For peptide and protein identification, tandem mass spectra were searched against a nonredundant International Protein Index arabidopsis sequence database v3.85 using Mascot2.2 (Matrix Science, London, UK; version 2.2) and proteome Discoverer1.4 (Thermo). The MASCOT search results for each SCX elution were further processed using the ProteomicsTools (version 3.05). All reported data were based on 99% confidence for protein identification as determined by false discovery rate (FDR) ≤1%. Quantification of proteins were performed according to intensity of report ions in each identified peptide. The differentially expressed proteins were searched against SwissProt database to find homologue sequences followed by gene ontology (GO) annotation and mapping. These proteins were also blasted against KEGG GENES and mapped to pathways in KEGG.
Enzyme-linked immunosorbent assay (ELISA)
Among the differentially expressed proteins between responders and nonresponders, S100A9 and ORM1 were selected as candidate biomarkers for further validation in another cohorts of 62 advanced ENKL patients. Serum levels of S100A9 and ORM1 were measured using commercially available ELISA Kit (Uscn Life Science, Wuhan, China) according to the manufacturer’s instructions.
Statistical analysis
The median levels of S100A9 and ORM1 from responders and nonresponders were compared by Mann Whitney tests. Receiver-operating-characteristics (ROC) analysis was employed to determine the cutoff concentration of S100A9 and ORM1 and evaluate the predictive accuracy. Overall survival (OS) was calculated from the date of the initial therapy until the date of death or the last follow-up. PFS was calculated from the date of the initial therapy until the date of disease progression or death from any cause. Survival analysis was performed using the Kaplan-Meier method and the log-rank test. Multivariate analysis using Cox regression model was conducted using SPSS 17.0.
Results
Identification of differentially expressed proteins in serums between responders and nonresponders
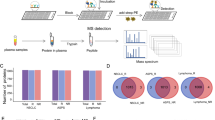
To explore potential biomarkers associated with treatment response in advanced ENKL patients, pretreatment serums from 16 responders and 16 nonresponders were collected. The clinicopathologic characteristics of the two groups were matched. The workflow of the discovery phase was shown in Fig. 1. After digested with trypsin and labeled with iTRAQ regents, the peptides were pooled. SCX chromatography was performed to reduce sample complexity. The fractions were then analyzed by a Q Exactive mass spectrometer connected to the Easy nano Liquid Chromatography. Comparative proteomic analysis identified 22 downregulated and 61 upregulated proteins with local false discovery rate (FDR) <5% in nonresponders (Supplementary 1). The upregulated proteins in nonresponders included S100A9, ORM1, alpha-1-antitrypsin, 14-3-3 protein zeta/delta. The downregulated proteins included platelet factor 4, vitamin D-binding protein, thrombospondin-1 and gelsolin. These differentially expressed proteins were mainly involved in the process of inflammatory/acute phase response, cell cycle and DNA damge (Supplementary 2). KEGG pathway analysis indicated that the dysregulated proteins were enriched in chemokine signaling pathway, hippo signaling pathway and PI3K-Akt signaling pathway.
Validation of the candidate biomarkers
From the above datas, we found that serum levels of S100A9 and ORM1 were significantly elevated in patients who obtained unfavourable outcomes. Since inflammation and EBV infection were deeply involved in the development of ENKL, we assumed that inflammatory factors such as S100A9 and ORM1 may be associated with treatment response and the prognosis of ENKL. ELISA assay was performed to validate S100A9 and ORM1 in another cohort of 62 patients. The baseline characteristics of patients were shown in Table 1. Serum levels of S100A9 and ORM1 were measured respectively. As shown in Fig. 2A, serum levels of S100A9 were significantly higher in nonresponders than that in responders. In Fig. 2B, nonresponders also presented higher levels of ORM1 than responders. The absolute quantification of S100A9 and ORM1 by ELISA allowed us to determine a threshold to differentiate low-S100A9 group and high-S100A9 group, low-ORM1 group and high-ORM1 group. ROC curve analysis indicated that serum levels of S100A9 at the concentration of 62.0 ng/ml possessed a sensitivity of 81.5% and a specificity of 71.4%. The cutoff point for ORM1 was 1436 ug/ml, with a sensitivity of 85.2% and a specificity of 77.1%. When S100A9 in combined with ORM1, the sensitivity increased to 100% (Table 2).
Correlation of S100A9 and ORM1 with overall survival and progression-free survival
To address the clinical relevance, serum levels of S100A9 and ORM1 were correlated with OS and PFS. The median follow-up is 17 months (range, 3–46 months). As shown in Fig. 3A, the 2-year OS for low-S100A9 group and high-S100A9 group were 76.6% and 40.2% (P = 0.0036), respectively. High-S100A9 group also predicted poor 2-year PFS in comparision with low-S100A9 group (33.1% vs 61.1%, p = 0.0019) (Fig. 3B). In terms of ORM1, high levels of ORM1 were associated with inferior 2-year OS (38.7% vs.76.1, p = 0.0012) and 2-year PFS (18.4% vs. 73.2%, p < 0.0001) (Fig. 4A,B). Furthermore, multivariate analysis using Cox regression model was conducted to explore potential prognostic factors for advanced ENKL patients. S100A9 > 62 ng/ml (RR = 2.92, 95% CI: 1.37–6.22, P = 0.005) and ORM1 > 1436 ug/ml (RR = 2.46, 95% CI: 1.14–5.32, P = 0.023) were independent predictors of overall survival. Elevated S100A9 (RR = 2.46, 95% CI: 1.14–5.32, P = 0.023) and ORM1 (RR = 2.86, 95% CI = 1.30–6.27, P = 0.009) were also adverse factors for PFS (Table 3). As shown in Fig. 5A, patients with high S100A9/high ORM1 had poor 2-year OS than those with high S100A9/low ORM, low S100A9/high ORM1, low S100A9/low ORM1 (19.05% vs. 66.67%, 34.28, 83.57%, p < 0.005). Moreover, the 2-year PFS for patients with high S100A9/high ORM1 and high S100A9/low ORM, low S100A9/high ORM1, low S100A9/low ORM1 were 14.29% and 65.63%, 26.67%, 77.49%, respectively (P < 0.005) (Fig. 5B). These results showed that high S100A9/high ORM1 predicted poor prognosis in advanced ENKL patients.
Discussion
Chemotherapy regimens containing pegaspargase, l-asparaginase, gemcitabine have shown remarkable efficacy in advanced ENKL. However, some patients still experience treatment failure and recurrent disease. Currently, few parameters are available to discriminate patients who would achieve unfavorable responses.
In the present study, we firstly conducted proteomic analysis to identify differentially expressed proteins in serum from responders and nonresponders. S100A9 and ORM1 were upregulated in nonresponders compared to responders. ELISA assay further confirmed that refractory patients displayed higher levels of S100A9 and ORM1. S100A9 in combined with ORM1 could effectively detect refractory patients with a sensitivity of 100%. Moreover, high-S100A9 and high-ORM1 were associated with inferior OS and PFS. Mutivariate analysis indicated that both S100A9 and ORM1 are independent prognostic factors of survival in advanced ENKL.
S100A9 is a calcium binding protein implicated in tumor growth. It interacts with toll-like receptor 4 (TLR4) or receptor for advanced glycation end products (RAGE) on the surface of cancer cells, which mediates activation of MAPK, NF-Kb, and AKT signalings11,12. Moreover, S100A9 can recruit myeloid derived suppressor cells (MDSC) in tumor sites and result in invasive phenotype of cancer13. Increased expression of S100A9 was observed in colitis-associated colon cancer, gastric cancer and lung cancer14. It binds to the RAGE or TLR on tumor cells, and thus activates oncogenic pathways and mediates inflammatory cascades15,16. In vitro studies indicated S100A9 at low concentration promoted the proliferation of hepatocellular carcinoma cells and metastasis of breast cancer17,18. Moreover, S100A9 can recuit MDSC in tumor sites, which suppresses T cell function and promote tumor metastasis. Growth of lymphoma tumor in S100A9-deficient mice was slower than that in wild-type mice, predominantly due to reduced recruitment of MDSCs19. Overexpression of S100A9 confers to chemoresistance. Advanced muscle invasive bladder cancer patients who displayed higher levels of S100A9 were insensitive to cisplatin-based chemotherapy and more frequently experienced disease progression20. For early-stage oral cancer, elevated S100A9 expression in tumor stroma increases macrophage recruitment and functions as an early recurrence marker21. In the current study, levels of S100A9 were elevated in chemoresistant ENKL patients.
ORM1, an acute phase protein, increases in serum as a response to inflammation. In addition, elevation of ORM1 were discovered in certain cancer patients22. Most cancer patients displayed increased levels of ORM1 in comparision with healthy controls. The study by Ganz PA and colleagues found that plasma levels of ORM1 were elevated in 89% of the solid tumor patients and 87% of the patients with hematologic neoplasms22. The prognostic values of ORM1 have also been demonstrated. In non-small cell lung cancer patients treated with docetaxel, the response rate for high baseline ORM1 group was 14%, while for low ORM1 group it was 44%. Patients with high ORM1 had a striking shorter survival than low-ORM1 patients23. Serum ORM1 also predicts the responses to neoadjuvant chemotherapy in advanced breast cancers24. Our results found that advanced ENKL patients who were sensitive to pegaspargase/gemcitabine treatment displayed lower ORM1 in serum. ENKL patients with low ORM1 had superior OS and PFS than high ORM1 patients.
Inflammation and EBV infection are deeply implicated in the pathogenesis of ENKL. However, the exact mechanisms underlying the role of EBV infection and inflammation in the development of ENKL remain largely unknown. The prognostic values of EBV-DNA and inflammatory factors in ENKL patients have been demonstrated previously. Ito Y and colleagues showed that pretreatment EBV-DNA copy number in whole blood was helpful to predict tumor response and survival7. Recent studies suggested that serum ferritin, interleukin-10 (IL-10) and c-reactive protein (CRP) could also serve as independent prognostic factors in ENKL25,26,27. Both S100A9 and ORM1 are inflammatory factors with multiple acyivities. Further studies would focus on how S100A9 and ORM1 affect the invasive type and drug sensitivity of ENKL cells.
In conclusion, we firstly identified S100A9 and ORM1 as novel serum biomarkers to predict the therapeutic response of advanced ENKL patients to pegaspargase/gemcitabine treatment. S100A9 and ORM1 could also serve as independent prognostic factors of survival in advanced ENKL.
Additional Information
How to cite this article: Zhou, Z. et al. S100A9 and ORM1 serve as predictors of therapeutic response and prognostic factors in advanced extranodal NK/T cell lymphoma patients treated with pegaspargase/gemcitabine. Sci. Rep. 6, 23695; doi: 10.1038/srep23695 (2016).
References
Tse, E. & Kwong, Y. L. Nasal NK/T-cell lymphoma: RT, CT, or both. Blood 126, 1400–1401, doi: 10.1182/blood-2015-07-655191 (2015).
Huang, Y. et al. Gene expression profiling identifies emerging oncogenic pathways operating in extranodal NK/T-cell lymphoma, nasal type. Blood 115, 1226–1237, doi: 10.1182/blood-2009-05-221275 (2010).
Tse, E. & Kwong, Y. L. How I treat NK/T-cell lymphomas. Blood 121, 4997–5005, doi: 10.1182/blood-2013-01-453233 (2013).
Yamaguchi, M. et al. Phase II Study of SMILE Chemotherapy for Newly Diagnosed Stage IV, Relapsed, or Refractory Extranodal Natural Killer (NK)/T-Cell Lymphoma, Nasal Type: The NK-Cell Tumor Study Group Study. J Clin Oncol 29, 4410–4416, doi: 10.1200/Jco.2011.35.6287 (2011).
Zhou, Z. et al. Effectiveness of gemcitabine, pegaspargase, cisplatin, and dexamethasone (DDGP) combination chemotherapy in the treatment of relapsed/refractory extranodal NK/T cell lymphoma: a retrospective study of 17 patients. Ann Hematol 93, 1889–1894, doi: 10.1007/s00277-014-2136-7 (2014).
Wang, L. et al. Post-treatment plasma EBV-DNA positivity predicts early relapse and poor prognosis for patients with extranodal NK/T cell lymphoma in the era of asparaginase. Oncotarget 6, 30317–30326, doi: 10.18632/oncotarget.4505 (2015).
Ito, Y. et al. Pretreatment EBV-DNA copy number is predictive of response and toxicities to SMILE chemotherapy for extranodal NK/T-cell lymphoma, nasal type. Clin Cancer Res 18, 4183–4190, doi: 10.1158/1078-0432.CCR-12-1064 (2012).
Ludvigsen, M., Hamilton-Dutoit, S. J., d’Amore, F. & Honore, B. Proteomic approaches to the study of malignant lymphoma: analyses on patient samples. Proteomics Clin Appl 9, 72–85, doi: 10.1002/prca.201400145 (2015).
Sharma, S., Ray, S., Moiyadi, A., Sridhar, E. & Srivastava, S. Quantitative Proteomic Analysis of Meningiomas for the Identification of Surrogate Protein Markers. Sci Rep-Uk 4, 7140, doi: 10.1038/Srep07140 (2014).
Zhang, M. Z. et al. Analysis of serum proteome profiles of non-Hodgkin lymphoma for biomarker identification. J Proteomics 72, 952–959, doi: 10.1016/j.jprot.2009.03.009 (2009).
Riva, M. et al. Induction of nuclear factor-kappa B responses by the S100A9 protein is Toll-like receptor-4-dependent. Immunology 137, 172–182, doi: 10.1111/j.1365-2567.2012.03619.x (2012).
Narumi, K. et al. Proinflammatory Proteins S100A8/S100A9 Activate NK Cells via Interaction with RAGE. J Immunol 194, 5539–5548, doi: 10.4049/jimmunol.1402301 (2015).
Wang, L. et al. Increased myeloid-derived suppressor cells in gastric cancer correlate with cancer stage and plasma S100A8/A9 proinflammatory proteins. J Immunol 190, 794–804, doi: 10.4049/jimmunol.1202088 (2013).
Markowitz, J. & Carson, W. E. Review of S100A9 biology and its role in cancer. Bba-Rev Cancer 1835, 100–109, doi: 10.1016/j.bbcan.2012.10.003 (2013).
Gebhardt, C. et al. RAGE signaling sustains inflammation and promotes tumor development. J Exp Med 205, 275–285, doi: 10.1084/jem.20070679 (2008).
Kallberg, E. et al. S100A9 interaction with TLR4 promotes tumor growth. PLoS One 7, e34207, doi: 10.1371/journal.pone.0034207 (2012).
Wu, R. et al. S100A9 promotes human hepatocellular carcinoma cell growth and invasion through RAGE-mediated ERK1/2 and p38 MAPK pathways. Exp Cell Res 334, 228–238, doi: 10.1016/j.yexcr.2015.04.008 (2015).
Gumireddy, K. et al. ID1 promotes breast cancer metastasis by S100A9 regulation. Mol Cancer Res 12, 1334–1343, doi: 10.1158/1541-7786.MCR-14-0049 (2014).
Leanderson, T. & Ivars, F. S100A9 and tumor growth. Oncoimmunology 1, 1404–1405, doi: 10.4161/onci.21027 (2012).
Kim, W. T. et al. S100A9 and EGFR gene signatures predict disease progression in muscle invasive bladder cancer patients after chemotherapy. Ann Oncol 25, 974–979, doi: 10.1093/annonc/mdu037 (2014).
Fang, W. et al. Elevated S100A9 expression in tumor stroma functions as an early recurrence marker for early-stage oral cancer patients through increased tumor cell invasion, angiogenesis, macrophage recruitment and interleukin-6 production. Oncotarget 6, 28401–28424, doi: 10.18632/oncotarget.4951 (2015).
Ganz, P. A., Shell, W. E. & Tokes, Z. A. Evaluation of a radioimmunoassay for alpha 1-acid glycoprotein to monitor therapy of cancer patients. J Natl Cancer Inst 71, 25–30 (1983).
Bruno, R. et al. Alpha-1-acid glycoprotein as an independent predictor for treatment effects and a prognostic factor of survival in patients with non-small cell lung cancer treated with docetaxel. Clin Cancer Res 9, 1077–1082 (2003).
Hyung, S. W. et al. A serum protein profile predictive of the resistance to neoadjuvant chemotherapy in advanced breast cancers. Mol Cell Proteomics 10, M111 011023, doi: 10.1074/mcp.M111.011023 (2011).
Li, Y. et al. Serum C-reactive protein (CRP) as a simple and independent prognostic factor in extranodal natural killer/T-cell lymphoma, nasal type. PLoS One 8, e64158, doi: 10.1371/journal.pone.0064158 (2013).
Wang, H. et al. Increased serum levels of interleukin-10 predict poor prognosis in extranodal natural killer/T-cell lymphoma patients receiving asparaginase-based chemotherapy. Onco Targets Ther 8, 2589–2599, doi: 10.2147/OTT.S91077 (2015).
Yamazaki, E. et al. Serum ferritin level is prognostic of patient outcome in extranodal NK/T cell lymphoma, nasal type. Med Oncol 31, 149, doi: 10.1007/s12032-014-0149-7 (2014).
Acknowledgements
This work was supported by the National Natural Science Foundation of China [grant number 81172118].
Author information
Authors and Affiliations
Contributions
Z.Z. designed and conducted the study; Z.L. and Z.S. collected and analyzed the datas; Z.Z. and X.Z. wrote the draft manuscript; Y.W. searched the related papers; L.L. prepared the Figures and tables and M.Z. designed this study and revised the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Zhou, Z., Li, Z., Sun, Z. et al. S100A9 and ORM1 serve as predictors of therapeutic response and prognostic factors in advanced extranodal NK/T cell lymphoma patients treated with pegaspargase/gemcitabine. Sci Rep 6, 23695 (2016). https://doi.org/10.1038/srep23695
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep23695
- Springer Nature Limited
This article is cited by
-
Orosomucoid 1 promotes colorectal cancer progression and liver metastasis by affecting PI3K/AKT pathway and inducing macrophage M2 polarization
Scientific Reports (2023)
-
Evaluation of IFIT3 and ORM1 as Biomarkers for Discriminating Active Tuberculosis from Latent Infection
Current Medical Science (2022)
-
Overexpression of S100A9 in tumor stroma contribute to immune evasion of NK/T cell lymphoma and predict poor response rate
Scientific Reports (2021)
-
miR-362-3p Targets Orosomucoid 1 to Promote Cell Proliferation, Restrain Cell Apoptosis and Thereby Mitigate Hypoxia/Reoxygenation-Induced Cardiomyocytes Injury
Cardiovascular Toxicology (2021)