Abstract
Establishment of implantation in pig is accompanied by a coordinated interaction between the maternal uterine endometrium and conceptus development. We investigated the expression profiles of endometrial tissue on Days 9, 12 and 15 of pregnancy and on Day 12 of non-pregnancy in Yorkshire and performed a comprehensive analysis of long non-coding RNAs (lncRNAs) in endometrial tissue samples by using RNA sequencing. As a result, 2805 novel lncRNAs, 2,376 (301 lncRNA and 2075 mRNA) differentially expressed genes (DEGs) and 2149 novel transcripts were obtained by pairwise comparison. In agreement with previous reports, lncRNAs shared similar characteristics, such as shorter in length, lower in exon number, lower at expression level and less conserved than protein coding transcripts. Bioinformatics analysis showed that DEGs were involved in protein binding, cellular process, immune system process and enriched in focal adhesion, Jak-STAT, FoxO and MAPK signaling pathway. We also found that lncRNAs TCONS_01729386 and TCONS_01325501 may play a vital role in embryo pre-implantation. Furthermore, the expression of FGF7, NMB, COL5A3, S100A8 and PPP1R3D genes were significantly up-regulated at the time of maternal recognition of pregnancy (Day 12 of pregnancy). Our results first identified the characterization and expression profile of lncRNAs in pig endometrium during pre-implantation phases.
Similar content being viewed by others
Introduction
Successful implantation of pregnancy is dependent on a complex molecular cross-talk between the conceptus development and the maternal uterus1,2. The embryo-maternal communication occurs approximately on Day 12 of pregnancy in pigs3 and the establishment of implantation in pig is accompanied by a dynamic production of estrogens, progesterone, prostaglandins, adhesion molecules and immunological factors. Porcine conceptuses secrete a large amount of estrogens, primarily 17β-estradiol (E2) on Days 11 and 12 and between Days 14 and 18 of pregnancy4,5,6. Estrogens enhance endometrial prostaglandin E2 (PGE2) production around Day 12 of pregnancy, resulting in sequestering of prostaglandin (PGF2α) within the uterine lumen and increasing the PGE2: PGF2α, which could promote the maintenance of corpus luteum (CL) and progesterone (P4) production7,8. P4 could stimulate the secretion of endometrial production required for conceptus development and implantation. Higher PGE2 secretion in uterine lumen coincides with the elevated expression of HOXA10 transcription factor which is critical for implantation. A stable adhesion between conceptus and endometrium requires reduction in mucin-1 on the apical surface of epithelium Furthermore, growth factors, cytokines and its receptors are all involved in embryo-maternal interactions9,10. In response to these factors, the uterine endometrial undergoes dramatic morphological and functional changes to be more easily receptive to the conceptus development11. Thus, to increase the rate of successful implantation, it is essential to understand the mechanism of the genetic regulation of endometrial changes.
Some of the identified genes are revealed to have expression changes of different levels in the uterine endometrial during early pregnancy at the time of estrogen release. For example, fibroblast growth factor 7 (FGF7)12,13 and secreted phosphoprotein 1 (SPP1)14,15 are abundantly expressed in uterine endometrial epithelial cells during early pregnancy. Interleukin-1 (IL-1) is identified as a key regulator of inflammatory response that modulates the communication between the maternal endometrium and embryo16,17. Legumain (LGMN) and its inhibitor, namely cystatins 6 (CST6) at the maternal-fetal interface, may play an important role in the establishment and maintenance of pregnancy in pigs18. In addition to the investigation of single-candidate genes and pathways, many studies have revealed a number of many different patterns of gene expression in uterine endometrial during the process of maternal recognition of implantation and pregnancy by using various technologies. Take for instance, microarrays have been used to describe endometrial gene expression on Days 1211,19,20, 1421, 1511,22 and 1619 of the estrous cycle and pregnancy and on Days 13, 18 and 24 of the attachment and non-attachment sites23. Similarly, digital gene expression profiling has been used to described differences in gene expression of endometrial samples collected from Erhualian sows and Landrace × Large White pigs on Day 12 of pregnancy24. In addition, systematic studies have been performed by RNA sequencing (RNA-seq) to describe the transcriptome changes in prepuberal gilts (crossbreed of German Landrace and Pitetrain) endometrium on Days 12 and 14 of the estrous cycle and pregnancy25,26. A further advantage of this whole transcriptome sequencing technology is able to describe unannotated transcriptional activity by identifying numerous noncoding transcripts27. Long non-coding RNAs (lncRNAs) have received much attention in the past several years and are found to play important functional roles in epigenetic regulation, chromatin modification, genomic imprinting, transcriptional control as well as pre- and post- translational mRNA processing28. RNA-seq technology has largely leveled up the discovery and analysis of non-coding RNA and differential methods have been developed to identify novel lncRNAs using RNA-seq data29,30. Despite the fact that many studies highlight the important roles of lncRNAs in different tissues30,31,32, little is known about the biological function and significance of lncRNAs in uterine endometrial during embryo implantation in pigs.
In this study, we investigated the expression profiles of endometrial tissue on Days 9 (YK9), 12 (YK12) and 15 (YK15) of pregnancy and on Days 12 (YK12K) of non-pregnancy in Yorkshire (YK) and conducted a comprehensive analysis of lncRNAs of endometrial tissue samples by using RNA-seq. The goals of the study were to determine how many and which transcripts (mRNA and lncRNA) are differentially expressed, the relationship between the selected lncRNA with its neighbor mRNA and to determine which biological processes and pathways are significantly changed by comparing tanscriptomic profiles of endometrial in different pregnant periods. To its credit transcripts information from pig endometrial transcriptomes can be used for further gene expression studies in endometrial tissue, which may help a better understanding of molecular and cellular events that occur in the endometrial during implantation period.
Results
RNA Sequencing and identification of mRNA and lncRNAs in Porcine Endometrial
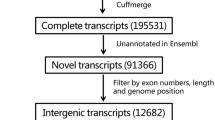
We analyzed the RNA-Seq data from 12 porcine endometrial samples in which 85 to 105 million raw reads and 82 to 99 million clear reads per sample were obtained. 4953 novel lncRNAs were assembled by Cufflinks33 and Scripture27 (Supplementary Fig. S1). As the identification of transcripts involved immature mRNA fragments, we used four tools, namely Coding Potential Calculator (CPC), Pfam-scan (PFAM), phylogenetic codon substitution frequency (phyloCSF), Coding-Non-Coding-Index (CNCI) to remove potential coding transcripts. Finally, 2805 putative non-coding transcripts were retained (Supplementary Fig. S2). Given a false discovery rate (FDR) of 4% and q-value (P-adjusted) <0.05, 2,376 (301 lncRNA and 2075 mRNA) differentially expressed genes (DGEs) were obtained from pairwise comparison of samples collected from Yorkshire pigs on Days 9, 12 and 15 of the pregnancy (i.e., YK9 vs YK12; YK12 vs YK15; YK9 vs YK15) and pairwise Day 12 of pregnancy with Day 12 of non-pregnancy (YK12 vs YK12K) (Table 1 and Supplementary Table S1). As shown in Fig. 1A, 16 DEGs were common among three comparisons (1 lncRNA and 15 mRNA). All of the obtained lncRNAs were not previously identified. In addition, a set of 2149 mRNA transcripts were found not to be annotated, over 39% of novel transcripts could be mapped with human NCBI Refseq database, nearly 14% of them mapped in mouse genome and 29% of them were classified as unknown transcripts (Supplementary Table S2). To validate the RNA-seq results, ten genes (TCONS_01729386, TCONS_01325501, FGF7, NMB, FGF9, VEGFC, VEGFA, Muc1, ESR1 and RBP4) were chosen for quantitative PCR (qPCR) (Supplementary Table S3). The selected lncRNAs were significantly different expressed at least in one comparison group and its predicted target genes have been reported involved in the implantation process, the selected mRNAs were involved in processes important during early embryo implantation. The results showed that expression patterns of these genes were in excellent agreement with the RNA-seq findings (Table 2).
Gene expression profiling and number of differentially expressed genes for endometrium during pre-implantation phase.
(A) Venn diagram of common differential expression genes in three comparison groups (YK12 vs YK9, YK15 vs YK12 and YK15 vs YK9). (B) A hierarchical heat map showing the transformaed expression values for transcript (mRNA and lncRNA). Yellow shows higher expression and blue shows lower espression.
Genomic features of lncRNAs
Many studies have shown that lncRNAs were shorter in length, less conserved than protein coding transcripts27. In agreement with previous studies, our results indicated that the predicated lncRNA are shorter in length than protein coding transcripts (Fig. 2A) and their genes tend to contain fewer exon (Fig. 2B). We found that lncRNAs in endometrium were longer than in skeletal muscle (1043bp on average), the number of exon was similar34. Interestingly, lncRNAs in pig endometrium are shorter in length than lncRNAs in human (1 kb on average), but longer than those in mouse (550nt on average) and zebrafish (1113nt on average) and contain fewer exon number than human (2.9 exon on average), mouse (3.7 exon on average) and zebrafish (2.8 exon on average)35,36. We also found that our predicted novel lncRNAs were less conserved than protein coding transcripts by using phastCon (Fig. 3), which was similar to that of the previous reports37. Furthermore, the identified lncRNAs in our dataset tend to be shorter in Orf length than protein coding genes (Fig. 2C). We annotated the traits of each putative novel lncRNA, such as chromatin state, proximity to coding genes and the relationship of location and its target genes (Supplementary Table S4).
Genomic features of predicted lncRNAs.
(A) Length distribution of 38222 coding transcripts (pink) and 2805 new predicted lncRNAs (blue). (B) Exon number distribution of coding transcripts and lncRNAs. (C) Orf length distribution of coding transcripts and lncRNAs. Dotted line represents the average value.
Differences in gene expression patterns between lncRNAs and protein coding transcripts
Recent studies suggested that lncRNAs may act in cis and affect the gene expression of their chromosomal neighborhood in 100k of upstream and downstream38. To investigate the relationship between lncRNAs and their neighboring coding genes, we analyzed gene pairs formed by lncRNAs and their neighboring genes and identified 11270 coding gene: coding gene pairs (1824 in divergent) and 1607 lncRNA: coding gene pairs (335 in divergent) (Fig. 4). Each category of gene pairs was mainly uni-direction pairs. For these coding genes near lncRNAs, pathway of “Regulation of actin cytoskeleton”, “Jak-STAT signling pathway”, “FoxO signaling pathway” and “MAPK signaling pathway” were enriched (Fig. 5). We observed a more correlated expression pattern of lncRNAs with their neighboring gene pairs (mean correlation: 0.297) than random coding gene pairs (mean correlation: 0.019) (P-value < 2.2e-16, Kolomogorv-Smirnov Test) and it exhibit a relatively higher correlated than coding gene pairs (mean correlation: 0.195) (P-value = 4.441e-16) (Fig. 6A). Notably, there was a significantly higher correlation between divergent lncRNAs: coding gene pairs (mean correlation: 0.294) than divergent (bidirectional) coding gene pairs (mean correlation: 0.247) (P-value = 0.0109) as well as random coding gene pairs (mean correlation: 0.019) (P-value < 2.2e-16) (Fig. 6B). This analysis suggested that the correlation between lncRNAs and their neighboring coding genes were higher than random gene pairs and coding gene pairs.
Correlation of expression patterns between pairs of neighboring genes.
(A) Shown are distributions of Pearson correlation coefficients in expression levels between either 11270 pairs of coding gene neighbors (red), 1607 pairs of long non-coding RNAs (lncRNAs) and their neighboring coding genes (green), or random pairs of genes (blue). (B) Shown are distribution of Pearson correlation coefficients calculated as in A, but only for 335 pairs of divergently transcribed pairs of lncRNA ans protein-coding genes (green) or 1824 pairs of divergently transcribed protein-coding genes (red). (C) Distribution of distance from one TSS (transcription start sites) to another, in all directions of lncRNA: coding gene pairs (green) and coding gene pairs (purple), divergent direction of lncRNA: coding gene pairs (red) and coding gene pairs (blue). (D) Box plots (showing the 15th, 25th, 50th, 75th and 95th percentiles) showing the expression feature of lncRNA and mRNA in each samples.
Further analysis illustrated that many lncRNAs were located with a 4-kb region surrounding the transcription start sites (TSSs) of coding genes and the majority of these lncRNAs originate from divergent transcription of lncRNAs: coding gene pairs (Fig. 6C). This conclusion was consistent with the previous results39,40. Our results indicated that lncRNAs tend to be expressed at lower levels than mRNA (Fig. 6D). We next examined the expression of selected lncRNAs in ten tissues of pregnancy and non-pregnancy (Fig. 7). Notably, lncRNAs tended to have high expression levels in endometrium than in other tissues.
Differential expression cluster analysis and Functional Prediction of LncRNAs in Endometrial Tissue Samples
To gain insight into the similarities of endometrium from Yorkshire of four ages at the transcriptome scale, data from all the differentially expressed genes in endometrium were used in a systematic cluster analysis. The heat map clearly suggested that YK12K and YK9 were initially clustered together because their expression profiles were similar, while YK12 and YK15 were clustered in another class (Fig. 1B). To further predict the function of lncRNAs in endometrial of pigs, we performed a Gene Ontology (GO) analysis with the selected mRNAs which neighbor lncRNAs or have high correlations with the expression of lncRNAs in 4 comparison groups. GO terms with the highest number of DGEs were related to remodeling of the endometrium, such as “binding”, “cellular process”, “immune system process” and “multicellular organismal process” (Supplementary Fig. S3). Furthermore, significantly enrichment GO terms (corrected p-Value < 0.05) were mainly including “multicellular organismal process” and “MHC class II protein complex” (Supplementary Table S5). The KEGG analysis revealed that the significantly enriched pathways during early pregnancy were “cytokine-cytokine receptor interaction”, “ribosome” and “neuroactive ligand-receptor interaction”, respectively. Interestingly, “estrogen signaling pathway” was the specific enrichment pathway in YK12 vs YK9 comparison group and “PI3K-Akt” and “Jak-STAT” were the common pathway in the four comparison groups (Supplementary Fig. S4). To find the specific enrichment functional terms for up-regulated and down-regulated lncRNAs, separated GO and KEGG analysis were performed. All up-regulated lncRNAs in four comparisons were related to steroidogenesis, immune function, cellular component biogenesis and tissue remodeling (Supplementary Table S6). As the number of down-regulated lncRNAs was small, there were no significant GO terms enriched in the DEGs in the four comparisons and the four most enriched KEGG pathways were “ribosome”, “allograft rejection”, “leukocyte transendothelial migration” and “glycerophospholipid metabolism” (Supplementary Table S6). In addition, the category related to “allograft rejection” term was found to be peculiar for down-regulated genes enrichment in YK15 vs YK9 group.
Functional analysis of mRNA in endometrial tissue samples
DEGs were enrichment in biological functions of immune response and extracellular matrix in uterine endometrium for YK15 vs YK9 and YK15 vs YK12 comparisons groups (Supplementary Table S7), but there were no GO terms enrichment for other two groups. To identify the signal transduction pathways in these four comparisons, we performed analysis on the basis of the KEGG pathway database. It was observed that the common signaling pathways was “PI3K-Akt signaling pathway” in YK12 vs YK9, YK15 vs YK12 and YK15 vs YK9 comparisons groups (Supplementary Fig. S5, S6).
Discussion
The majority of embryonic loss has occurred primarily around Day 11–12 and the maternal recognition of pregnancy signal around Day 12 of gestation41,42. At mRNA expression levels, a large number of high throughput platforms have been performed to analyze the endometrial mechanisms and pathways around Day 12 of gestation20,21. The analysis of gene expression merely on mRNA levels has become feasible. With the development of high-throughput technologies for large-scale expression feasible, RNA-seq has accelerated the discovery and characterization of lncRNAs, a new class of biologically-significant RNA transcripts30,35. However, to our knowledge, lncRNAs in endometrium tissue is little known. As for the first study of lncRNAs in endometrium of pigs, we identified approximately 2806 putative noncoding transcripts and 2,376 (301 lncRNA and 2075 mRNA) DEGs. 36 up-regulated and 38 down-regulated DGEs were obtained from YK12 compared with YK12K. However, previous study have found that 1335 with higher and 1258 with lower expression levels of DEGs from pregnant gilts compared to the non-pregnant controls. This may caused by difference between species.
The primary goal of this study was to identify non-coding RNA, mainly lncRNA. Our newly identified lncRNAs in pig endometrium shared many characteristics with those in other mammalian species. They are shorter, lower in exon number, lower in expression level and less conserved than protein coding transcripts. Furthermore, the conservation of lncRNAs in pig was modestly lower than in human and mouse. In particular, recent studies demonstrated that some lncRNAs can regulate gene expression of their chromosomal neighborhood in cis43,44. Non-coding RNA transcripts can also be derived from divergent transcripts in mammals45,46 and the RNAs produced by divergent transcription may serve a regulatory function for expression of the downstream genes47. According to the analysis of the gene pairs formed by lncRNAs and their neighboring genes, we found lncRNAs transcripted coordinately with neighboring genes and at a higher level than the neighboring coding gene pairs. The majority of lncRNAs are divergent transcription of active protein coding genes. As a contrast of this, some other researchers suggested that pairs of coding gene neighboring are slightly more correlated to each other than neighboring lincRNA: protein-coding gene pairs35.
In this study, lncRNAs mainly included long intergenic noncoding RNA (lincRNA), inronic lncRNA and anti-sense lncRNA. Previous studied have noted that lincRNAs can be classified as enhancer-associated (elncRNA) or promoter-associated (plncRNA)48,49. A notable feature of the lncRNAs is remarkable tissue-specific as compared with protein-coding genes50,51, both elncRNA and plncRNAs exhibit tissue specificity32. So we verified selected lncRNAs used for validating the RNA-seq results in difference tissues and got the similar conclusion, RNA expression profiling across pig tissues revealed that the transcript TCONS_01325501 was highly expressed in endometrium.
Most evidence suggests that the expression of lncRNAs can regulate and have high correlations with expression of neighboring mRNAs35,52. Based on this, we searched coding genes 10k/100k upstream and downstream of lncRNA as the cis target genes and predicted the function of lncRNA. Consequently, we found that many lncRNAs may exert their function through predicted mRNA which can play pivotal roles in pig endometrium during pre-implantation phase. For example, the sequence of lncRNA TCONS_01325501 matched with FGF7, FGF7 has been reported to stimulate cell proliferation, differentiation, migration and vascular angiogenesis12,53. This finding suggests that TCONS_01325501 may affect the expression of FGF7 and therefore be involved in the interaction between the uterus and conceptus. In this study, lncRNAs of TCONS_01729386 was the only identified different transcripts in YK12 vs YK9, YK15 vs YK12 and YK15 vs YK9 comparisons groups, its predicted target mRNA were FGF9 and IL-1. The role of IL-1 and FGF9 during pre-implantation phases is important for established of pregnancy. FGF9 was significantly higher expressed in pregnant animals, in line with the fact that FGF9 has previously been identified as a growth factor in pig endometrium54. IL-1 was identified as one such paracrine factor that modulates the communication between the maternal endometrium and embryo16,55. Therefore, we speculated that TCONS_01729386 may regulate the embryo implantation. However, these predicted functions of lncRNAs require experimental verification.
The most critical period during implantation in pigs is considered to be Day 12 of pregnancy, the time when maternal begins to recognize of pregnancy7. One of the most important finding of the study was that in the four comparison groups we identified 5 genes, namely LOC100153672, NMB, LOC100622067, S100A8 and JPH1, which were specific up-regulated expression genes in YK12 sample compared to YK9, YK15 and YK12K samples (see Supplementary Table S5, Supplementary Table S6). GO functional annotation analysis showed that they were all involved in the “protein binding” and “extracellular matrix” terms and were related to “PI3K-Akt signaling pathway”, “MAPK signaling pathway”, “Protein digestion and absorption” and “Insulin signaling pathway”. Other eight genes, namely FGF7, C7, COL5A3, ZFYVE28, USH1C, PPP1R3D and EPS8L3 were up-regulated in YK12 samples as compared to YK9 and YK15 samples, which enriched of GO terms related to protein binding and transcription factor activity. S100A8 is low molecular weight calcium binding protein and is found at high level in the extracellular milieu during inflammatory conditions56, while another study found that up-regulation of S100A8 is a key component of the early endometrial response to uterine infection57. PPP1R3D is clustered into the pathway of insulin signaling pathway, which is a possible influential factor of blood vessel development. Apposition and attachment of pig conceptuses need a maternal vascular support. Estrogens are pivotal for maternal recognition of pregnancy in pigs. FOXO1 is known to be regulated by steroid hormones, including estrogen and progesterone, which is up-regulated in the uterine endometrium on Day 12 compared with Day 9 of pregnancy58. Other important finding was “Estrogen signaling pathway” which as the specific pathway in YK12 vs YK9 group, this may due to the highest level on Day 12 of pregnancy than other periods. PIK3R3 and ESR1 were the DEGs in this pathway, also we found that PIK3R3 was in the other two pathways, “Jak-STAT” and “PI3-Akt”. IL6R, IL5 and IL10RB were enriched in “Jak-STAT” pathway, EGF and IL6R were enriched in “PI3-Akt” pathway.
All up-regulated lncRNAs in four comparisons were related with steroidogenesis in this study. Early pregnancy is accompanied by many immune reactions simultaneously occurring in the uterus16. GO annotation analysis indicated that the activation of immune system was the strongest at Day 15 of pregnancy. Previous study have shown that sows with lower immune system activation are prone to implantation59. Mitosis-related genes, such as FGF7, FGF9, IGFBP2, MET and MKI67, are up-regulated in endometrium on Day 12 of pregnancy. Previous study suggested that MUC4 was significantly up-regulated in the pregnant endometrium, which indicated that MUC4 played a vital role in protecting the porcine surface epithelium against invasion of the conceptuses. In this study, the expression of MUC4 was significantly higher on Day 12 than on Day 9 of pregnancy.
In conclusion, we first generated the expression profile of lncRNA in pig endometrium based on a transcriptome RNA-seq approach. We have identified lncRNA and mRNA expression profile for pig endometrium on Days of 9, 12, 15 of pregnancy and Day 12 of non-pregnancy in Yorkshire pigs. Importantly, we analyzed the genomic feature and expression profiles of all identified lncRNAs. Bioinformatic analysis suggests that some lncRNAs are involved in important biological processes associated with embryo implantation such as binding, cell adhesion and growth factor activity and may play an important role in regulating the gene expression of pre-implantation. As the role of lncRNAs in pigs have not yet been fully identified and understood, this study should provide valuable resource for further studies. This study also provided a resource for lncRNA studies in other noninvasive animal implantation.
Materials and Methods
Ethics Statement
All studies involving animals were conducted according to the regulation (No. 5 proclaim of the Standing Committee of Hubei People’s Congress) approved by the Standing Committee of Hubei People’s Congress, P. R. China. Sample collection was approved by the ethics committee of Huazhong Agricultural University. Animals were humanely sacrificed as necessary to ameliorate suffering.
Sample collection
Twenty Yorkshire gilts with similar age and genetic background from one commercial herd were selected. Animals were randomly assigned to cyclic (n = 3) and pregnant (n = 9) group. Gilts of pregnant group were artificially inseminated twice after estrus. Uteri were obtained from animals slaughtered on Days 12 (n = 3) of the estrous cycle or Days 9 (n = 3), 12 (n = 3) and 15 (n = 3) of pregnancy, each uterine horn was flushed with PBS (pH 7.4) and subsequently opened longitudinally at the inner side. Samples from the endometrium of the pregnant and non-pregnant sows were taken from three locations of each uterine horn: proximal, medial and distal. Tissue samples were frozen in liquid nitrogen and stored at −80 °C until RNA was isolated.
Total RNA isolation
Total RNA was isolated from each individual sample using TRIzol reagent (Invitrogen, USA). Purity and quantity of total RNA were measured by using Nanodrop equipment. Integrity of RNA was assessed using the RNA Nano6000 Assay Kit of the Bionalyzer 2100 system (Agilent Technologies, CA, USA)
Library preparation for lncRNA sequencing
A total amount of 3 μg RNA per sample was used as input material for the RNA sample preparations. Firstly, ribosomal RNA was removed by Epicentre Ribo-zero™ rRNA Removal Kit (Epicentre, USA) and rRNA free residue was cleaned up by ethanol precipitation. Subsequently, sequencing libraries were generated using the rRNA-depleted RNA by NEBNext® Ultra™ Directional RNA Library Prep Kit for Illumina® (NEB, USA) following manufacturer’s recommendations. First strand cDNA was synthesized using random hexamer primer and M-MuLV Reverse Transcriptase. Second strand cDNA synthesis was subsequently performed using DNA Polymerase I and RNase H. In the reaction buffer, dNTPs with dTTP were replaced by dUTP. After adenylation of 3′ ends of DNA fragments, NEBNext Adaptor with hairpin loop structure were ligated to prepare for hybridization. In order to select cDNA fragments of the preferred 150~200 bp in length, the library fragments were purified with AMPure XP system (Beckman Coulter, Beverly, USA). At last, products were purified (AMPure XP system) and library quality was assessed on the Agilent Bioanalyzer 2100 system. The libraries were sequenced at the Novogene Bioinformatics Institute (Beijing, China) on an Illumina Hiseq 2500 platform and 100 bp paired-end reads were generated.
Data analysis
Raw reads of fastq format were firstly processed through in-house perl scripts. In this step, clean data (clean reads) were obtained by removing reads that contain adapter or ploy-N and low quality reads from raw data. At the same time, Q20, Q30 and GC content of the clean data were calculated. All the downstream analyses were based on the clean data with high quality. Reads were mapped with Tophat (v2.0.9) to the porcine genome sequence assembly (Sscrofa 10.2). The mapped reads of each sample were assembled by both Scripture27 and Cufflinks33.
Coding potential analysis and Target gene prediction
We used CNCI, CPC, PFAM and phyloCSF to distinguish mRNA from lncRNA. CNCI profiles can effectively distinguish protein-coding and non-coding sequences independent of known annotations by adjoining nucleotide triplets60. CPC searches the sequences with known protein sequence database to clarify the coding and non-coding transcripts mainly through assessing the extent and quality of the ORF in a transcript61. Each transcript can be translated in all three possible frames and Pfam Scan (v1.3) used to identify occurrence of any of the known protein family domains documented in the Pfam database62. PhyloCSF examines evolutionary signatures characteristic in alignments with conserved coding regions, such as the high frequencies of synonymous codon substitutions and conservative amino acid substitutions and the low frequencies of other mis-sense and non-sense substitutions to distinguish protein-coding and non-coding transcripts63.
Transcripts without coding potential were our candidate set of lncRNAs. Then, we searched coding genes 10k/100k upstream and downstream of lncRNA as the cis target gene. Trans role of lncRNA is to identify each other by the expression level.
Conservative analysis
The phast software (v1.3) generally used for phylogenetic analysis and thus phastCons expression. PhastCons is a conservation scoring and identifying program of conserving elements. We used phyloFit to compute phylogenetic models for conserved and non-conserved regions among species and then set the model and HMM transition parameters for phastCons to compute the conservation scores of lncRNA and coding genes64.
GO and KEGG enrichment analysis
The quantification of both lncRNAs and coding genes in each sample was calculated by Cuffdiff (v2.1.1)27 and transcripts with a P-adjust < 0.05 were assigned as being differentially expressed. KEGG is a database resource for understanding high-level functions and utilities of the biological system, so we used KOBAS software to test the statistical enrichment of differential expression genes or lncRNA target genes in KEGG pathways.
Real-time RT-PCR
RNA samples from the 12 animals used for the RNA-seq experiment were analyzed by qPCR. Total cDNA was synthesized using reverse transcriptase Kit (TaKaRa, Dalian). QPCR were performed using LightCycler 480II Real-Time PCR System and SYBR® Green PCR Master Mix (TaKaRa, Dalian). Each real-time RT-PCR reaction (in 25 μL) involved 12.5 μL 2 × SYBR Green Realtime PCR Master Mix (TaKaRa, Dalian), 1 μL of each primer, 2 μL cDNA and 8.5 μL H2O. The cycling conditions included an initial single cycle (95 °C for 3 min) and followed by 40 cycles (95 °C for 15 s; 57 °C for 15 s; 72 °C for 20 s). All amplifications were followed by dissociation curve analysis of the amplified products. Specific primers were designed using the NCBI, specificities were confirmed with BLAST and gene expression levels were normalized with RPS20 to attain the relative expression by using 2 (−ΔΔCt) value methods (Table 1). The statistical difference in gene expression was analyzed by SAS during different endometrium development stages of pregnancy. The correlation between the results of RNA-seq and qPCR was calculated using correlation test.
Additional Information
How to cite this article: Wang, Y. et al. Analyses of Long Non-Coding RNA and mRNA profiling using RNA sequencing during the pre-implantation phases in pig endometrium. Sci. Rep. 6, 20238; doi: 10.1038/srep20238 (2016).
References
Geisert, R. D. & Yelich, J. V. Regulation of conceptus development and attachment in pigs. J Reprod Fertil Suppl 52, 133–149 (1997).
Vigano, P., Mangioni, S., Pompei, F. & Chiodo, I. Maternal-conceptus cross talk–a review. Placenta 24 Suppl B, S56–61 (2003).
Geisert, R. D., Zavy, M. T., Moffatt, R. J., Blair, R. M. & Yellin, T. Embryonic steroids and the establishment of pregnancy in pigs. J Reprod Fertil Suppl 40, 293–305 (1990).
Geisert, R. D., Renegar, R. H., Thatcher, W. W., Roberts, R. M. & Bazer, F. W. Establishment of pregnancy in the pig: I. Interrelationships between preimplantation development of the pig blastocyst and uterine endometrial secretions. Biol Reprod 27, 925–939 (1982).
Geisert, R. D., Brookbank, J. W., Roberts, R. M. & Bazer, F. W. Establishment of pregnancy in the pig: II. Cellular remodeling of the porcine blastocyst during elongation on day 12 of pregnancy. Biol Reprod 27, 941–955 (1982).
Geisert, R. D., Thatcher, W. W., Roberts, R. M. & Bazer, F. W. Establishment of pregnancy in the pig: III. Endometrial secretory response to estradiol valerate administered on day 11 of the estrous cycle. Biol Reprod 27, 957–965 (1982).
Bazer, F. W. & Thatcher, W. W. Theory of maternal recognition of pregnancy in swine based on estrogen controlled endocrine versus exocrine secretion of prostaglandin F2alpha by the uterine endometrium. Prostaglandins 14, 397–400 (1977).
Christenson, L. K., Farley, D. B., Anderson, L. H. & Ford, S. P. Luteal maintenance during early pregnancy in the pig: role for prostaglandin E2. Prostaglandins 47, 61–75 (1994).
Waclawik, A. Novel insights into the mechanisms of pregnancy establishment: regulation of prostaglandin synthesis and signaling in the pig. Reproduction 142, 389–399 (2011).
Ziecik, A. J. et al. Mechanisms for the establishment of pregnancy in the pig. Reprod Domest Anim 46 Suppl 3, 31–41 (2011).
Kim, M. et al. Microarray Analysis of Gene Expression in the Uterine Endometrium during the Implantation Period in Pigs. Asian-Australas J Anim Sci 25, 1102–1116, (2012).
Ka, H., Jaeger, L. A., Johnson, G. A., Spencer, T. E. & Bazer, F. W. Keratinocyte growth factor is up-regulated by estrogen in the porcine uterine endometrium and functions in trophectoderm cell proliferation and differentiation. Endocrinology 142, 2303–2310 (2001).
Ka, H. et al. Regulation of expression of fibroblast growth factor 7 in the pig uterus by progesterone and estradiol. Biol Reprod 77, 172–180 (2007).
Garlow, J. E. et al. Analysis of osteopontin at the maternal-placental interface in pigs. Biol Reprod 66, 718–725 (2002).
White, F. J. et al. Steroid regulation of cell specific secreted phosphoprotein 1 (osteopontin) expression in the pregnant porcine uterus. Biol Reprod 73, 1294–1301 (2005).
Simon, C., Mercader, A., Gimeno, M. J. & Pellicer, A. The interleukin-1 system and human implantation. Am J Reprod Immunol 37, 64–72 (1997).
Bankers-Fulbright, J. L., Kalli, K. R. & McKean, D. J. Interleukin-1 signal transduction. Life Sci 59, 61–83 (1996).
Shim, J. et al. Analysis of legumain and cystatin 6 expression at the maternal-fetal interface in pigs. Mol Reprod Dev 80, 570–580 (2013).
Kiewisz, J. et al. Global gene expression profiling of porcine endometria on Days 12 and 16 of the estrous cycle and pregnancy. Theriogenology 82, 897–909 (2014).
Gu, T. et al. Endometrial gene expression profiling in pregnant Meishan and Yorkshire pigs on day 12 of gestation. BMC genomics 15, 156 (2014).
Ostrup, E., Bauersachs, S., Blum, H., Wolf, E. & Hyttel, P. Differential endometrial gene expression in pregnant and nonpregnant sows. Biol Reprod 83, 277–285 (2010).
Franczak, A., Wojciechowicz, B. & Kotwica, G. Transcriptomic analysis of the porcine endometrium during early pregnancy and the estrous cycle. Biol Reprod 13, 229–237 (2013).
Chen, X. et al. Differential gene expression in uterine endometrium during implantation in pigs. Biol Reprod 92, 52 (2015).
Zhang, H. et al. Differential gene expression in the endometrium on gestation day 12 provides insight into sow prolificacy. BMC genomics 14, 45 (2013).
Samborski, A., Graf, A., Krebs, S., Kessler, B. & Bauersachs, S. Deep sequencing of the porcine endometrial transcriptome on day 14 of pregnancy. Biol Reprod 88, 84 (2013).
Samborski, A. et al. Transcriptome changes in the porcine endometrium during the preattachment phase. Biol Reprod 89, 134 (2013).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat Biotechnol 28, 511–515 (2010).
Kretz, M. et al. Suppression of progenitor differentiation requires the long noncoding RNA ANCR. Genes Dev 26, 338–343 (2012).
Prensner, J. R. et al. Transcriptome sequencing across a prostate cancer cohort identifies PCAT-1, an unannotated lincRNA implicated in disease progression. Nat Biotechnol 29, 742–749 (2011).
Weikard, R., Hadlich, F. & Kuehn, C. Identification of novel transcripts and noncoding RNAs in bovine skin by deep next generation sequencing. BMC genomics 14, 789 (2013).
Kretz, M. et al. Control of somatic tissue differentiation by the long non-coding RNA TINCR. Nature 493, 231–235 (2013).
Tsoi, L. C. et al. Analysis of long non-coding RNAs highlights tissue-specific expression patterns and epigenetic profiles in normal and psoriatic skin. Genome Biol 16, 24 (2015).
Guttman, M. et al. Ab initio reconstruction of cell type-specific transcriptomes in mouse reveals the conserved multi-exonic structure of lincRNAs. Nat Biotechnol 28, 503–510 (2010).
Zhao, W. et al. Systematic identification and characterization of long intergenic non-coding RNAs in fetal porcine skeletal muscle development. Sci Rep 5, 8957 (2015).
Cabili, M. N. et al. Integrative annotation of human large intergenic noncoding RNAs reveals global properties and specific subclasses. Genes Dev 25, 1915–1927 (2011).
Pauli, A. et al. Systematic identification of long noncoding RNAs expressed during zebrafish embryogenesis. Genome Res 22, 577–591 (2012).
Qu, Z. & Adelson, D. L. Bovine ncRNAs are abundant, primarily intergenic, conserved and associated with regulatory genes. PloS one 7, e42638 (2012).
Orom, U. A. et al. Long noncoding RNAs with enhancer-like function in human cells. Cell 143 (2010).
Sigova, A. A. et al. Divergent transcription of long noncoding RNA/mRNA gene pairs in embryonic stem cells. Proc Natl Acad Sci USA 110, 2876–2881 (2013).
Seila, A. C. et al. Divergent transcription from active promoters. Science 322, 1849–1851 (2008).
Yelich, J. V., Pomp, D. & Geisert, R. D. Detection of transcripts for retinoic acid receptors, retinol-binding protein and transforming growth factors during rapid trophoblastic elongation in the porcine conceptus. Biol Reprod 57, 286–294 (1997).
Geisert, R. D. et al. Maternal recognition of pregnancy signal or endocrine disruptor: the two faces of oestrogen during establishment of pregnancy in the pig. Soc Reprod Fertil Suppl 62, 131–145 (2006).
Ponjavic, J., Oliver, P. L., Lunter, G. & Ponting, C. P. Genomic and transcriptional co-localization of protein-coding and long non-coding RNA pairs in the developing brain. PLoS Genet 5, e1000617 (2009).
Pennisi, E. Long noncoding RNAs may alter chromosome’s 3D structure. Science 340, 910 (2013).
Core, L. J., Waterfall, J. J. & Lis, J. T. Nascent RNA sequencing reveals widespread pausing and divergent initiation at human promoters. Science 322, 1845–1848 (2008).
Preker, P. et al. RNA exosome depletion reveals transcription upstream of active human promoters. Science 322, 1851–1854 (2008).
Seila, A. C., Core, L. J., Lis, J. T. & Sharp, P. A. Divergent transcription: a new feature of active promoters. Cell cycle 8, 2557–2564 (2009).
Wang, B. et al. Reprogramming efficiency and quality of induced Pluripotent Stem Cells (iPSCs) generated from muscle-derived fibroblasts of mdx mice at different ages. PLoS Curr 3, RRN1274 (2011).
Marques, A. C. et al. Chromatin signatures at transcriptional start sites separate two equally populated yet distinct classes of intergenic long noncoding RNAs. Genome Biol 14, R131 (2013).
Mercer, T. R., Dinger, M. E. & Mattick, J. S. Long non-coding RNAs: insights into functions. Nat Rev Genet 10, 155–159 (2009).
Ponting, C. P., Oliver, P. L. & Reik, W. Evolution and functions of long noncoding RNAs. Cell 136, 629–641 (2009).
Guttman, M. et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 458, 223–227 (2009).
Ka, H., Spencer, T. E., Johnson, G. A. & Bazer, F. W. Keratinocyte growth factor: expression by endometrial epithelia of the porcine uterus. Biol Reprod 62, 1772–1778 (2000).
Tsai, S. J., Wu, M. H., Chen, H. M., Chuang, P. C. & Wing, L. Y. Fibroblast growth factor-9 is an endometrial stromal growth factor. Endocrinology 143, 2715–2721 (2002).
Simon, C. et al. Interleukin-1 system in the materno-trophoblast unit in human implantation: immunohistochemical evidence for autocrine/paracrine function. J Clin Endocrinol Metab 78, 847–854 (1994).
Nemeth, J. et al. S100A8 and S100A9 are novel nuclear factor kappa B target genes during malignant progression of murine and human liver carcinogenesis. Hepatology 50, 1251–1262 (2009).
Swangchan-Uthai, T. et al. Influence of energy balance on the antimicrobial peptides S100A8 and S100A9 in the endometrium of the post-partum dairy cow. Reproduction 145, 527–539 (2013).
Kyo, S. et al. Forkhead transcription factor FOXO1 is a direct target of progestin to inhibit endometrial epithelial cell growth. Clin Cancer Res 17, 525–537 (2011).
Wettemann, R. P., Bazer, F. W., Thatcher, W. W., Caton, D. & Roberts, R. M. Conceptus development, uterine response, blood gases and endocrine function of gilts exposed to increased ambient temperature during early pregnancy. Theriogenology 30, 57–74 (1988).
Sun, L. et al. Utilizing sequence intrinsic composition to classify protein-coding and long non-coding transcripts. Nucleic Acids Res 41, e166 (2013).
Kong, L. et al. CPC: assess the protein-coding potential of transcripts using sequence features and support vector machine. Nucleic Acids Res 35, W345–349 (2007).
Punta, M. et al. The Pfam protein families database. Nucleic Acids Res 40, D290–301 (2012).
Lin, M. F., Jungreis, I. & Kellis, M. PhyloCSF: a comparative genomics method to distinguish protein coding and non-coding regions. Bioinformatics 27, i275–282 (2011).
Siepel, A. et al. Evolutionarily conserved elements in vertebrate, insect, worm and yeast genomes. Genome Res 15, 1034–1050 (2005).
Acknowledgements
This work was supported by the National Basic Research Program of China (2012CB138504) and the National Porcine Industry Technology System (CAR-36).
Author information
Authors and Affiliations
Contributions
M.L. conceived and designed the experiments; Y.W. analyzed the data, prepared figure and contributed writing the manuscript; W.W., S.X., H.L., T.H. and J.Z. collected the samples; Y.W. and S.X. performed the experiments; L.L., X.L. and X.Q. reviewing the manuscript. All authors contributed to the manuscript at various stages.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Wang, Y., Xue, S., Liu, X. et al. Analyses of Long Non-Coding RNA and mRNA profiling using RNA sequencing during the pre-implantation phases in pig endometrium. Sci Rep 6, 20238 (2016). https://doi.org/10.1038/srep20238
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep20238
- Springer Nature Limited
This article is cited by
-
Physiological and Biochemical Mechanisms of Prunus sachalinensis Root for Coping with Short-term Waterlogging and Subsequent Recovery
Journal of Soil Science and Plant Nutrition (2024)
-
Proteomic profiles and the function of RBP4 in endometrium during embryo implantation phases in pigs
BMC Genomics (2023)
-
Identification and characterization of circRNAs in peri-implantation endometrium between Yorkshire and Erhualian pigs
BMC Genomics (2023)
-
Comprehensive analysis of lncRNAs involved in skeletal muscle development in ZBED6-knockout Bama pigs
BMC Genomics (2021)
-
Integrated analysis of lncRNA, miRNA and mRNA reveals novel insights into the fertility regulation of large white sows
BMC Genomics (2020)