Abstract
Enterohemorrhagic E. coli (EHEC) causes diarrhea and hemorrhagic colitis with life-threatening complications, such as hemolytic uremic syndrome. Their major virulence factor is Shiga toxin (Stx), which is encoded by bacteriophages. Of the two types of Stx, the production of Stx2, particularly that of Stx2a (a subtype of Stx2), is a major risk factor for severe EHEC infections, but the Stx2 production level is highly variable between strains. Here, we define four major and two minor subtypes of Stx2a-encoding phages according to their replication proteins. The subtypes are correlated with Stx2a titers produced by the host O157 strains, suggesting a critical role of the phage subtype in determining the Stx2a production level. We further show that one of the two subclades in the clade 8, a proposed hyper-virulent lineage of O157, carries the Stx2 phage subtype that confers the highest Stx2 production to the host strain. The presence of this subclade may explain the proposed high virulence potential of clade 8. These results provide novel insights into the variation in virulence among O157 strains and highlight the role of phage variation in determining the production level of the virulence factors that phages encode.
Similar content being viewed by others
Introduction
The evolution of pathogenic bacteria is often promoted by the horizontal gene transfer (HGT) of virulence-associated genetic determinants1,2. Although most, if not all, HGT events are mediated by mobile genetic elements (MGEs), such as bacteriophages (simply called phages), plasmids and transposons, these MGEs are often regarded as mere vehicles carrying the virulence determinants. However, MGEs themselves evolve rapidly and display remarkable genetic variation, even between closely related elements. A particularly high degree of genetic variation is exhibited by phages, which have shorter generation times and can easily recombine with other phages, as evidenced by the widespread genetic mosaicism observed in phage genomes3,4,5. Such variations among phages may directly or indirectly affect the evolution of the host bacteria to which the phage transduces virulence determinant(s); however, the biological significance of genetic variation among phages has not yet been fully elucidated. Although infection and lysogenization by toxin-encoding phages converts otherwise non-pathogenic bacteria into highly virulent pathogens6,7,8,9,10, the production levels of such toxins often vary among bacterial strains11,12,13,14. Variations in toxin-encoding phages may be linked to such differences in toxin production levels, but this possibility has not yet been fully explored in any pathogenic bacteria.
Shiga toxin (Stx) is the key virulence factor of enterohemorrhagic Escherichia coli (EHEC), which causes diarrhea and hemorrhagic colitis with life-threatening complications, such as hemolytic uremic syndrome (HUS), although EHEC possesses a broad range of virulence factors, many of which are needed for the full virulent phenotype15. Stx, a member of the AB-type toxin family, inhibits protein synthesis in host cells by depurinating a specific adenine residue of the 28S rRNA15. Stxs are classified as Stx1 and Stx2, both of which contain several subtypes16. EHEC strains produce one or more Stx subtypes15. Strains of serotype O157 (EHEC O157), the most prevalent EHEC worldwide, produce Stx1, Stx2a and Stx2c, alone or in combination. The level of Stx2c production by EHEC O157 is generally thought to be very low17,18,19. Importantly, Stx2-producing strains cause more severe infections than Stx1-producing strains20. In fact, purified Stx2 is 1,000 times more toxic to human renal endothelial cells than Stx121.
The stx genes are encoded by lambdoid phages and are located downstream of the phage late gene promoter22. Although stx1 is under the control of the iron-regulated authentic promoter, the expression of stx2 depends strongly on the late promoter23,24,25,26,27,28. When Stx2-encoding phages (Stx2 phages hereafter) are integrated into the host chromosome (lysogenic state), stx2 is repressed by the phage CI repressor. However, once the SOS response is provoked in the host cells treated with DNA-damaging agents, such as mitomycin C (MMC), the activated RecA protein stimulates the autocleavage of the CI repressor, resulting in phage induction and replication. Upon induction, early genes are first expressed, after which the late promoter-dependent expression of late phage genes is induced by the action of the antiterminator protein Q. Protein Q is produced during early gene expression and subsequently acts on the transcription terminator, which is located immediately downstream of the late promoter. Therefore, Stx2 production is tightly coupled with phage induction and replication, which suggests that variations among Stx2 phages could affect the Stx2 production level of each EHEC. In fact, the level of Stx2 production varies remarkably even between EHEC strains of the same serotype (i.e., EHEC O157)11,12,17,18,19,29,30. Considerable genomic diversity has also been found among phages encoding the same Stx2 subtype (i.e., Stx2a phages)19,31,32.
Here, we present the results of a comprehensive analysis of 123 Stx2-positive O157 strains and their Stx2 phages. Our data indicate that Stx2c phages are highly homogeneous. However, most Stx2a phages are classified into four major subtypes according to their replication proteins and these subtypes are correlated with the Stx2a production levels by host strains. We further show that EHEC O157 strains belonging to clade 8, which has been proposed to be a highly pathogenic clade of O15719,33,34,35, can be separated into two subclades by a high-resolution phylogenetic analysis. One of these two subclades carries the subtype of the Stx2a phage that confers the highest Stx2a production to the host strain. The presence of this subclade may explain the proposed high virulence potential of clade 8.
Results
Phylogenetic clades, stx2 subtypes and Stx2 phage integration sites of 123 O157 strains
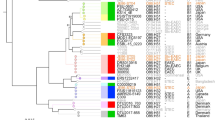
The 123 stx2-positive O157 strains analyzed in this study were all human strains isolated in various regions of Japan between the 1990s and 2006 (Supplementary Table S1), with the exception of EDL933, which was isolated in the United States in 198236. The strain set included 15 strains from asymptomatic carriers. A phylogenetic clade analysis was conducted via a rapid differentiation method based on the sequences of eight single nucleotide polymorphism (SNP) sites35,37. This analysis revealed that 4, 11, 20, 2, 6, 64 and 12 strains belonged to clades 1, 2, 3, 4/5, 6, 7 and 8, respectively and 4 strains were untypeable (Table 1). A restriction fragment-length polymorphism (RFLP) analysis of the stx2 genes38 showed that 49 and 59 of the strains contained either stx2a or stx2c alone, respectively, whereas the remaining 15 strains contained both stx2a and stx2c (Table 1).
The chromosomal integration sites of the Stx2 phages in each strain were determined using a long PCR-based method (see Methods). All the Stx2 phages were found to be integrated into one of three known integration sites: 39 Stx2a phages were integrated into the wrbA locus, 25 Stx2a phages were integrated into the argW locus and all 74 of the Stx2c phages were integrated into the sbcB locus (Table 1). Intriguingly, most of the strains carrying an Stx2a phage in wrbA belonged to clades 1, 2 and 3 (35/39) and were stx1-positive (36/39) (Table 1). In contrast, all 12 strains belonging to clade 8 carried an Stx2a phage in the argW locus, whereas four of them carried an Stx2c phage as well. None of the assessed clade 8 strains were stx1-positive. The majority of the strains in the other clades (clades 4 to 7) carried the Stx2c phage alone in the sbcB locus (59/72) or together with the Stx2a phage in argW (11/72). Thus, the distribution of the Stx2 phages appears to be clade dependent.
Stx2 production levels
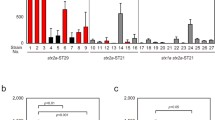
From the 123 strains, we selected 65 that represented various combinations of Stx2 subtypes, Stx2 phage integration sites and phylogenetic clades and we measured their Stx2 production levels in the presence and absence of MMC by means of a reversed passive latex agglutination (RPLA) assay. Even in the absence of MMC, Stx2 production was detected in all the strains. A remarkable strain-to-strain variation in toxin titer (which ranged from 2 to 1,600 in RPLA titer) was also observed (Fig. 1 and Supplementary Table S1). Consistent with previous reports17,18,19, MMC treatment did not strongly enhance Stx2 production in the strains carrying stx2c alone. In contrast, Stx2 production was remarkably enhanced by MMC in almost all the strains carrying stx2a, and the observed Stx2 production levels were highly variable among the strains. The levels of MMC-induced Stx2 production by these strains were significantly correlated with those in the absence of MMC (Supplementary Fig. S1), but there was no significant relation between Stx2 production and the clade of the strain or the integration site of the Stx2a phage. Recently, Neupane et al. showed that clade 8 strains overexpressed stx2 in both RNA and protein levels compared to clade 1–3 strains11. In this study, however, only four clade 8 strains were analyzed and one of them produced a very low level of Stx2. Thus, although the authors concluded that the increased stx2 expression is characteristics of clade8 strains, it is not exclusive to this clade and not all strains within clade 8 overexpress stx2. The Stx2 production patterns of stx2a/2c double-positive strains were similar to those of stx2a single-positive strains. This is consistent with a previous study by Kawano et al.18 and suggests that the Stx2 production levels of the stx2a/2c double-positive strains are largely representative of the levels of Stx2a production by each strain.
Highly variable Stx2 production levels among O157 strains.
The Stx2 production levels of the 65 O157 strains are shown. The supernatants of the polymixin B-treated cultures were prepared from the 65 O157 strains without or with the MMC treatment (panels (A,B), respectively). The Stx2 concentration in each supernatant was measured using the VTEC-Reversed Passive Latex Agglutination (VTEC-RPLA) assay kit and the resulting RPLA titers are shown. The Stx2 prophages of the strains indicated by the dots were sequenced in this study. The strains were categorized according to their stx2 subtypes and Stx2 phage integration sites.
Genomic diversity of Stx2 phages in O157
We selected 10 O157 strains that contained only one Stx2 phage (Table 2) and determined the genome sequences of their Stx2 prophages using a fosmid-based method. These strains produced different levels of Stx2 and their Stx phages carried either stx2a or stx2c and were integrated into wrbA, argW or sbcB (three strains had the Stx2a phage in wrbA, three had the Stx2a phage in argW and four had the Stx2c phage in sbcB). A dot-plot analysis of the genome sequences of these 10 phages and of Sp5 (the Stx2a phage of the O157 strain Sakai39, also known as VT2-Sakai) and 933W (the Stx2a phage of the O157 strain EDL93340) revealed that the Stx2a phages exhibited overall sequence similarities to each other, irrespective of their integration sites (wrbA or argW), whereas the Stx2c phages were highly divergent from the Stx2a phages (Supplementary Fig. S2). A more detailed comparison of the Stx2a phages revealed a notable degree of intra-group genomic diversity in their early regions (upstream of stx2) but only minor structural variations in their late regions (Fig. 2). As previously suggested33,41, the Stx2c phages of the O157 strains were highly homogeneous. The genomic structures of the four Stx2c phages sequenced in this study were well conserved, with only a few small replacements and strain-specific insertion sequence (IS) insertions being detected (Fig. 2).
Genome comparison of Stx2 phages.
The genome structures of the six Stx2a and four Stx2c phages sequenced in this study are drawn to scale. Two previously sequenced Stx2a phages of O157 strains Sakai and EDL933 (Sp5 and 933W, respectively) are also included. Homologous regions are indicated by shading and the sequence identities are indicated by different colors. The VT2-WGPS9 phage contains a large deletion encompassing the sequence from int to bet; this deletion was likely generated by an IS-mediated event. Integration sites of each phage and subtypes of Stx2a phages (α, β, γ and δ; see the main text and Fig. 3 for more details) are also indicated.
Subtyping of the Stx2a phages of O157 and comparison of the Stx2 production levels among the strains carrying each subtype of the Stx2a phage
Genomic variations in the early region of the Stx2a phages (Fig. 2) may be linked to the variation in Stx2 production levels among the stx2a-positive strains (Fig. 1). In this regard, the variation of replication proteins between the Stx2a phages is particularly interesting; we identified four types of replication proteins among the eight Stx2a phages examined (Fig. 2). We therefore grouped these phages into four subtypes, ϕStx2a_α (Sp5), ϕStx2a_β (VT2-F403 and VT2-F451), ϕStx2a_γ (933W, VT2-F422, VT2-F723 and VT-WGPS9) and ϕStx2a_δ (VT2-F765), according to the type of replication proteins. The replication genes completely differed in sequence between the four subtypes (Supplementary Fig. S3). We therefore developed a two-step PCR-based subtyping system for the Stx2a phages using an stx2a-specific primer (stx2-R) and primers specific to the replication genes of each subtype (Fig. 3A; primer sequences are available in Supplementary Table S2). Using this system, the Stx2a phages from 55 of the 64 stx2a-positive strains (including the stx2a/2c double-positive strains) were categorized into one of the four subtypes (15 ϕStx2a_α, 14 ϕStx2a_β, 16 ϕStx2a_γ or 10 ϕStx2a_δ); however, the 9 remaining stx2a-positive strains were untypeable (Supplementary Table S1). We therefore determined the sequences of early regions of all the untypeable Stx2a phages. From the comparison of the sequences, these untypeable phages were categorized into two groups (ϕStx2a_ε and ϕStx2a_ζ) (Supplementary Fig. S4 and Table S1). These two groups possessed similar replication genes. But they were very different from those of the four major subtypes and the 5′-half of the O gene homologs completely differed between the two subtypes (Supplementary Fig. S3). Therefore, two-step PCR systems to subtype these minor groups were also constructed using the unique sequences in their replication genes (Fig. 3A).
Stx2a phage subtypes and the comparison of Stx2 production levels among the strain groups carrying each subtype of the Stx2a phage.
(A) The six Stx2a phage subtypes that were identified and a two-step PCR-based subtyping system are shown here. Based on structural variations in the phage early region, the Stx2a phages were classified into six subtypes (α, β, γ, δ, ε and ζ). Representative genomic structures of each subtype are shown. The locations of the primers used in the first and second PCRs to subtype the Stx2a phages are also indicated. (B) The Stx2 production levels (RPLA titers) of the 54 stx2a-positive O157 strains with or without MMC treatment (shown in the lower and upper panels, respectively) were determined and compared between the six strain groups carrying each subtype of the Stx2a phage. The experiments were repeated at least three times per strain and representative data are shown. The statistical significance was calculated using an ANOVA followed by Tukey’s multiple-comparisons procedure (*P < 0.05). Note that, among the strains containing the ϕStx2a_γ phage, a strain indicated by open circle corresponds to strain WGPS9, the Stx2a phage of which could not be excised from the host chromosome due to the deletion of its int and xis genes.
We then compared the Stx2 production levels of the stx2a-positive strains and found significant differences in the Stx2 production levels among the four major groups of strains carrying each subtype of the Stx2a phage (Fig. 3B). In both the presence and absence of MMC, the ϕStx2a_γ group produced more Stx2 than the other groups (note that a ϕStx2a_γ subtype phage indicated by an open circle could not be excised from the chromosome due to the deletion of its int and xis genes). The ϕStx2a_α group also produced more Stx2 than the other two groups (ϕStx2a_β and ϕStx2a_δ) in the presence of MMC, although no significant differences were observed in the absence of MMC. This result suggests that strains carrying phage ϕStx2a_γ or ϕStx2a_α (particularly the former) can produce higher Stx2 levels than strains carrying other Stx2a phage subtypes. Among the strains carrying two minor subtypes of Stx2a phage, those carrying ϕStx2a_ζ may also produce a higher level of Stx2. However, more strains need to be examined to perform a reliable statistic analysis.
High-resolution phylogenetic analysis and Stx2a phage subtype distribution in O157 clade 8 strains
Clade 8 has been proposed to be a highly pathogenic clade of O15734,35. Although the mechanism(s) underlying the high pathogenicity of the strains in clade 8 has not yet been well elucidated, these strains may produce a higher level of Stx211. However, whereas 6 of the 12 clade 8 strains examined in this study were found to carry ϕStx2a_γ, the remaining six strains carried ϕStx2a_δ (Supplementary Table S1). To determine the phylogenetic relationship between these two types of clade 8 strains, we performed a whole genome sequence-based, high-resolution phylogenetic analysis of the clade 8 strains using high-quality draft genome sequences obtained from the Illumina MiSeq sequencer (Fig. 4 and Supplementary Table S3). We included three previously sequenced O157 strains in this analysis as a control: two clade 8 strains (TW14359 and EC4115, both isolated in the “2006 spinach outbreak” in the United States19,33) and one clade 3 strain (Sakai). Our results revealed that the clade 8 strains were divided into two distinct subclades (referred to as subclades 8a and 8b). Importantly, the subclade 8a strains, which included TW14359 and EC4115, carried ϕStx2a_γ exclusively and all the subclade 8b strains carried ϕStx2a_δ (Fig. 4). Furthermore, only subclade 8a strains (six of the eight strains, indicated by asterisks in Fig. 4) contained Stx2c phages in addition to Stx2a phages. It should also be noted that, in both subclades, the Stx2a phages were integrated into the same locus (argW); however, it is currently unknown whether the two subclades acquired their different Stx2a phage subtypes independently or whether the exchange of phage subtypes occurred in either or both subclade(s) after their separation. It is also unknown whether (or to what extent) the difference in genetic background between the two sublineages affects the Stx2a production level and potential virulence of the strains in each sublineage.
High-resolution phylogenetic analysis of the O157 clade 8 strains.
Based on 975 SNP sites on the chromosomal backbone, which were identified using a genome sequence comparison among the K-12 MG1655, O157 Sakai and 14 O157 clade 8 strains, a maximum likelihood phylogenetic tree was generated in RAxML ver. 7.2.8 using the GTR-GAMMA model. The reliabilities of the tree’s internal branches were assessed using bootstrapping with 1,000 pseudoreplicates. The scale bar represents the number of substitutions per site. Strain TW14359 (a completely sequenced clade 8 strain isolated from the spinach-associated outbreak in the USA in 2006) was used as a reference. Strain EC4115 (another completely sequenced clade 8 strain isolated from the spinach-associated outbreak) and strain Sakai (clade 3) were also included in this analysis. The other 12 strains were sequenced in this study. The subtypes of the Stx2a phage in each strain are indicated in parentheses. Asterisks indicate the strains carrying an Stx2c phage in addition to the Stx2a phage.
Discussion
As confirmed in this study (Fig. 1), the Stx2 production levels of the O157 strains were highly variable. However, the genetic determinants that specify these production levels have long remained unknown. Because Stx2 is a major risk factor for severe EHEC infections, including those caused by E. coli O15720,21, the factors responsible for high Stx2 production are of great interest. Through a comprehensive analysis of 123 stx2-positive O157 strains and their Stx2 phages, including the genome sequencing and comparison of 12 representative Stx2 prophages, we showed that the Stx2a phage subtype is a critical factor that determines the Stx2 production level in O157.
Based on the genome sequences of the six Stx2a phages determined in this study and two previously determined, we defined four major Stx2a subtypes, each having distinct replication proteins. The complete genome sequences of additional Stx2a phages, such as VT2phi_272 (HQ424692), Min27 (EU311208), Xuzhou21 (CP001925), SS17 (CP008805) and the Stx2a prophages from the “spinach outbreak” strains TW14359 and EC4115, are available in public databases19,33,42,43,44. However, these phages were either very similar or nearly identical to at least one of the six Stx2a phages analyzed in this study (Supplementary Fig. S5). In fact, most Stx2a phages of the strains tested could be categorized as one of these four major Stx2a phage subtypes using the PCR-based subtyping system developed in this study (Supplementary Table S1). Although sequence analysis of the nine untypeable phages further identified two minor Stx2a phage subtypes (Supplementary Fig. S4), it is likely that most of the major subtypes of Stx2a phages of the O157 strains have been identified. In addition, we showed that the Stx2c phages of O157, whose genome structures were distinct from those of the Stx2a phages, were highly homogeneous (Fig. 2).
Our data from the Stx2a phage subtyping and the determination of the Stx2 production levels of the O157 strains indicated a significant correlation between the Stx2a phage subtype and the Stx2 production level by O157. Other factors, such as genetic backgrounds associated with the lineages of host strains, could also affect the Stx2 production level by O157, but the Stx2a phage subtype may be one of the critical factors to determine it and the Stx2 production levels of each O157 strain appear to be predictable by subtyping their Stx2a phages. Of the four major subtypes, ϕStx2a_γ conferred the ability to produce the highest level of Stx2 in the host strains and strains carrying ϕStx2a_α produced more Stx2 than strains carrying the other two subtypes of Stx2 phages.
The efficiency of phage induction is thought to be critical in determining the level of stx2 expression, which depends on the phage late gene promoter27,28,45. However, variations of the cI repressor genes, which encode a master regulator of phage induction and the q genes, which encode an antiterminator that allows the stx2 gene expression from the phage late promoter, cannot account for these differences (Supplementary Fig. S6). The early regions of ϕStx2a_β and ϕStx2a_γ phages exhibited the greatest similarity among the four subtypes; these subtypes differed only in the types of replication genes, the presence of genes encoding Ren (a phage exclusion protein) and several hypothetical proteins (Figs 2 and 3A). However, strains carrying ϕStx2a_β and ϕStx2a_γ exhibited remarkable differences in Stx2 production (Fig. 3B). These divergent genes (or gene sets) are thus candidate factors that determine the efficiency of phage induction and therefore the level of Stx2 production.
Regarding the relationship between phage subtype and Stx2 production levels, the homogeneity of the Stx2c phages may also be important; this homogeneity could be linked to the low level of stx2c gene expression that is a general feature of this Stx2 subtype (Fig. 1). The relationship between Stx2 phage type and Stx2 production levels in non-O157 EHEC strains is also an issue of medical and scientific importance that requires additional research, particularly in the case of EHEC serotypes O26, O111, O103, O121, O145 and O45, all of which are considered to pose a particularly high risk46,47.
Another important finding of this study is the identification of a sublineage of clade 8 (named subclade 8a), which has acquired ϕStx2a_γ. It appears likely that the presence of this subclade could, at least partly, account for the proposed high virulence potential of clade 819,33,34. A large-scale statistical analysis of the relationship between the subclade (i.e., the Stx2a phage subtype) and the symptoms and disease severities of patients infected by clade 8 strains is required to verify this hypothesis. However, strains TW14359 and EC4115, which were isolated in the “2006 spinach outbreak” that resulted in remarkably high rates of hospitalization (51%) and HUS (16%)19,33,34, were both included in subclade 8a (Fig. 4). Our preliminary analysis of single nucleotide polymorphisms (SNPs) specific to each subclade indicated that subclade 8b corresponds to the previously described 87–14 lineage in Cluster 148,49 and the major branch of subclade 8a corresponds to the TW14359 lineage in the same cluster, whereas the minor branch comprised of strains 990057 and F416 exhibited a chimeric feature of the 87–14 and TW14359 lineages in terms of SNP composition.
Finally, it should also be mentioned that ϕStx2a_γ is also distributed in clade 3 and other clades (Supplementary Table S1). Further high-resolution phylogenetic analyses, combined with Stx2a phage subtyping and measurements of Stx2 production levels, may reveal additional highly pathogenic lineages of O157. In such lineages, the potential role of Stx1 and its bacteriophage in toxin expression or the attenuation of virulence may also be re-evaluated50,51.
In conclusion, we defined that the Stx2a phage subtypes of EHEC O157, which may be a critical factor to determine the Stx2 production level. Differences in phage induction efficiency among the subtypes, which appear to be specified by the types of replication proteins, may account for the differences in Stx2 production levels among the host strains. The proposed high virulence potential of clade 8 could be explained, at least partly, by the presence of subclade 8a, which has acquired ϕStx2a_γ, a phage that can confer the highest Stx2 production levels to the host strains. Thus, our data contribute to a better understanding of the virulence potential of O157 and provide novel insights into the medical importance of bacteriophage variation, which is closely linked to the production levels of the virulence factors encoded by these phages.
Materials and Methods
Bacterial strains, culture conditions and isolation of the genomic DNA
Of the 123 O157 strains examined (Supplementary Table S1), the Sakai and EDL933 strains were isolated from large outbreaks that occurred in 1996 and 1982 in Japan and the United States, respectively36,52. The remaining 121 strains were human isolates collected from various regions of Japan between 1990 and 2006. These strains included seven strains (WGPS3 to WGPS9) that we previously analyzed using the Whole Genome PCR Scanning method53. The strain set also contained 16 strains from asymptomatic carriers, which were isolated from food handlers and workers in day-care centers or some other occupations that are obligated to undergo periodic stool examinations. All the strains were routinely grown on Lysogeny Broth (LB) agar or in LB medium at 37 °C. The genomic DNA was purified from 2 ml of an overnight culture of each strain using the Genomic-tip 100/G and Genomic DNA buffer set (QIAGEN, Valencia, CA) according to the manufacturer’s instructions.
Detection and subtyping of stx and phage integration site determination
The detection of the stx1 and stx2 genes via PCR and the subtyping of the stx2 gene via a RFLP analysis have been previously described38,54. The integration sites of the Stx2 phages were determined using long PCR with the stx2-R primer and a combination of primers specific to each flanking region of the four known Stx2 phage integration sites (the wrbA, argW, sbcB and yehV loci). Long PCR was performed using the LA Taq PCR kit (Takara Shuzo, Kyoto, Japan), with 1 ng of genomic DNA used as the template for a two-step amplification program involving 30 cycles of 20 sec at 98 °C and 16 min at 69 °C (or at 66 °C if unsuccessful at 69 °C). All the primers used in this study are listed in Supplementary Table S2.
Determination of the Stx2 production level
The bacterial cells were inoculated into 2 ml of CAYE medium (Denka Seiken Co., Ltd., Tokyo, Japan) and were grown to the mid-log phase at 37 °C with shaking. MMC was added to the culture (to a final concentration of 500 ng/ml). After 3 h of incubation, polymixin B (Denka Seiken; final concentration 10,000 U/ml) was added and the incubation continued for 1 h at 37 °C with mixing every 10 min. The culture supernatants were then obtained by centrifugation (7,700 ×g for 10 min at 37 °C) and were used for the assay. The MMC-untreated samples were similarly prepared but without the addition of MMC. The supernatants were serially diluted in 96-well V-bottom microtiter plates using the dilution buffer contained in the VTEC-Reversed Passive Latex Agglutination (VTEC-RPLA) assay kit (Denka Seiken). The Stx2 titer of each sample was determined using the VTEC-RPLA assay kit according to the manufacturer’s instructions.
Sequencing of the Stx2 prophage genomes
Fosmid libraries were constructed using pCC1 (Epicentre, Madison, WI, USA) as previously described55. From each fosmid library, stx2-containing clones were screened using PCR with stx2A-specific primers. Of the stx2-positive clones, two clones that encompassed the first and second halves of the Stx2 prophage genome, respectively, were selected for each of the examined strains; these clones were sequenced using shotgun sequencing followed by primer walking. The sequences thus obtained were annotated using the Microbial Genome Annotation Pipeline (MiGAP)56 and were manually curated using IMC-GE software (In Silico Biology, Inc., Kanagawa, Japan). The genome sequences of the 10 Stx2 phages sequenced in this study have been deposited in the DDBJ/EMBL/GenBank databases (accession numbers; AP012529–AP012540).
Subtyping of the Stx2a phages
The Stx2a phages were subtyped using two-step PCR (Fig. 3A). The first PCR was performed using primers targeting the stx2A gene (stx2-R) and the replication proteins specific to each subtype. The same PCR condition as that used for long PCR to detect phage integration sits was employed. The second PCR was performed using the first PCR product (10,000 times diluted) as a template and KAPATaq polymerase (Kapa Biosystems, Woburn, MA, USA) with 30 cycles of 20 sec at 98 °C, 30 sec at 58 °C and 1 min at 72 °C; the primers for this step targeted the genes for the replication proteins specific to each subtype. Sequences of all primers used for Stx2a phage subtyping are shown in Supplementary Table S2.
High-resolution phylogenetic analysis of clade 8 strains
To perform genome sequencing on the Illumina MiSeq sequencing platform (Illumina, San Diego, CA, USA), genomic DNA libraries were prepared from each of the 12 clade 8 strains using the Nextera DNA Sample Prep kit (Illumina). The pooled libraries were subjected to multiplexed paired-end sequencing (2 rounds of 151 cycles). The sequence reads were assembled using the Velvet version 1.2.05 software57. Contig sequences from each clade 8 strain and the chromosome sequences of K-12 MG165558, O157 Sakai52 and O157 EC411533 were aligned with that of O157 TW1435919 using MUMmer version 3.2259 to identify the conserved backbone of these strains and its SNP sites. We found that a 2,976,129-bp sequence of the TW14359 chromosome (its total length) was conserved among all the strains examined, with >98% sequence identity and >1,000-bp alignment length. To exclude any SNPs in recombinogenic regions from the analysis, SNP clusters (>2 SNPs within 100 bp) were removed. Finally, RAxML ver. 7.2.860 was used to construct a maximum likelihood phylogenetic tree inferred from the concatenated alignment of 975 SNP sites located on the conserved backbone (the GTR-GAMMA model of nucleotide substitution, 10 replicate runs and 1,000 bootstraps).
Statistical analyses
Statistical significance was calculated using an analysis of variance (ANOVA) followed by Tukey’s multiple-comparisons procedure (Fig. 3B). Statistical correlations were analyzed using Pearson’s correlation coefficient (r) (Supplementary Fig. S1).
Additional Information
How to cite this article: Ogura, Y. et al. The Shiga toxin 2 production level in enterohemorrhagic Escherichia coli O157:H7 is correlated with the subtypes of toxin-encoding phage. Sci. Rep. 5, 16663; doi: 10.1038/srep16663 (2015).
References
Fortier, L. C. & Sekulovic, O. Importance of prophages to evolution and virulence of bacterial pathogens. Virulence 4, 354–365 (2013).
Pallen, M. J. & Wren, B. W. Bacterial pathogenomics. Nature 449, 835–842 (2007).
Hatfull, G. F. & Hendrix, R. W. Bacteriophages and their genomes. Current opinion in virology 1, 298–303 (2011).
Ohnishi, M., Kurokawa, K. & Hayashi, T. Diversification of Escherichia coli genomes: are bacteriophages the major contributors? Trends Microbiol 9, 481–485 (2001).
Brüssow, H., Canchaya, C. & Hardt, W. D. Phages and the evolution of bacterial pathogens: from genomic rearrangements to lysogenic conversion. Microbiology and molecular biology reviews: MMBR 68, 560–602 (2004).
Boyd, E. F. & Brüssow, H. Common themes among bacteriophage-encoded virulence factors and diversity among the bacteriophages involved. Trends Microbiol 10, 521–529 (2002).
Freeman, V. J. Studies on the virulence of bacteriophage-infected strains of Corynebacterium diphtheriae. J Bacteriol 61, 675–688 (1951).
Huang, A., Friesen, J. & Brunton, J. L. Characterization of a bacteriophage that carries the genes for production of Shiga-like toxin 1 in Escherichia coli. J Bacteriol 169, 4308–4312 (1987).
O’Brien, A. D. et al. Shiga-like toxin-converting phages from Escherichia coli strains that cause hemorrhagic colitis or infantile diarrhea. Science 226, 694–696 (1984).
Waldor, M. K. & Mekalanos, J. J. Lysogenic conversion by a filamentous phage encoding cholera toxin. Science 272, 1910–1914 (1996).
Neupane, M. et al. Shiga toxin 2 overexpression in Escherichia coli O157:H7 strains associated with severe human disease. Microb Pathog 51, 466–470 (2011).
Wagner, P. L., Acheson, D. W. & Waldor, M. K. Isogenic lysogens of diverse shiga toxin 2-encoding bacteriophages produce markedly different amounts of shiga toxin. Infect Immun 67, 6710–6714 (1999).
Son, M. S., Megli, C. J., Kovacikova, G., Qadri, F. & Taylor, R. K. Characterization of Vibrio cholerae O1 El Tor biotype variant clinical isolates from Bangladesh and Haiti, including a molecular genetic analysis of virulence genes. J Clin Microbiol 49, 3739–3749 (2011).
Siddique, A. K. et al. El Tor cholera with severe disease: a new threat to Asia and beyond. Epidemiol Infect 138, 347–352 (2010).
Croxen, M. A. et al. Recent advances in understanding enteric pathogenic Escherichia coli. Clin Microbiol Rev 26, 822–880 (2013).
Scheutz, F. et al. Multicenter evaluation of a sequence-based protocol for subtyping Shiga toxins and standardizing Stx nomenclature. J Clin Microbiol 50, 2951–2963 (2012).
de Sablet, T. et al. Differential expression of stx2 variants in Shiga toxin-producing Escherichia coli belonging to seropathotypes A and C. Microbiology 154, 176–186 (2008).
Kawano, K., Okada, M., Haga, T., Maeda, K. & Goto, Y. Relationship between pathogenicity for humans and stx genotype in Shiga toxin-producing Escherichia coli serotype O157. Eur J Clin Microbiol Infect Dis 27, 227–232 (2008).
Kulasekara, B. R. et al. Analysis of the genome of the Escherichia coli O157:H7 2006 spinach-associated outbreak isolate indicates candidate genes that may enhance virulence. Infect Immun 77, 3713–3721 (2009).
Boerlin, P. et al. Associations between virulence factors of Shiga toxin-producing Escherichia coli and disease in humans. J Clin Microbiol 37, 497–503 (1999).
Louise, C. B. & Obrig, T. G. Specific interaction of Escherichia coli O157:H7-derived Shiga-like toxin II with human renal endothelial cells. J Infect Dis 172, 1397–1401 (1995).
Waldor, M. K. & Friedman, D. I. Phage regulatory circuits and virulence gene expression. Curr Opin Microbiol 8, 459–465 (2005).
Calderwood, S. B. & Mekalanos, J. J. Iron regulation of Shiga-like toxin expression in Escherichia coli is mediated by the fur locus. J Bacteriol 169, 4759–4764 (1987).
Shimizu, T. et al. The serine 31 residue of the B subunit of Shiga toxin 2 is essential for secretion in enterohemorrhagic Escherichia coli. Infect Immun 75, 2189–2200 (2007).
Vareille, M., de Sablet, T., Hindre, T., Martin, C. & Gobert, A. P. Nitric oxide inhibits Shiga-toxin synthesis by enterohemorrhagic Escherichia coli. Proc Natl Acad Sci USA 104, 10199–10204 (2007).
Shimizu, T., Ohta, Y. & Noda, M. Shiga toxin 2 is specifically released from bacterial cells by two different mechanisms. Infect Immun 77, 2813–2823 (2009).
Tyler, J. S., Mills, M. J. & Friedman, D. I. The operator and early promoter region of the Shiga toxin type 2-encoding bacteriophage 933W and control of toxin expression. J Bacteriol 186, 7670–7679 (2004).
Shimizu, T., Ohta, Y., Tsutsuki, H. & Noda, M. Construction of a novel bioluminescent reporter system for investigating Shiga toxin expression of enterohemorrhagic Escherichia coli. Gene 478, 1–10 (2011).
Baker, D. R. et al. Differences in virulence among Escherichia coli O157:H7 strains isolated from humans during disease outbreaks and from healthy cattle. Appl Environ Microbiol 73, 7338–7346 (2007).
Zhang, Y. et al. Lineage and host source are both correlated with levels of Shiga toxin 2 production by Escherichia coli O157:H7 strains. Appl Environ Microbiol 76, 474–482 (2010).
Johansen, B. K., Wasteson, Y., Granum, P. E. & Brynestad, S. Mosaic structure of Shiga-toxin-2-encoding phages isolated from Escherichia coli O157:H7 indicates frequent gene exchange between lambdoid phage genomes. Microbiology 147, 1929–1936 (2001).
Ogura, Y. et al. Complexity of the genomic diversity in enterohemorrhagic Escherichia coli O157 revealed by the combinational use of the O157 Sakai OligoDNA microarray and the Whole Genome PCR scanning. DNA research 13, 3–14 (2006).
Eppinger, M., Mammel, M. K., Leclerc, J. E., Ravel, J. & Cebula, T. A. Genomic anatomy of Escherichia coli O157:H7 outbreaks. Proc Natl Acad Sci USA 108, 20142–20147 (2011).
Manning, S. D. et al. Variation in virulence among clades of Escherichia coli O157:H7 associated with disease outbreaks. Proc Natl Acad Sci USA 105, 4868–4873 (2008).
Iyoda, S. et al. Phylogenetic Clades 6 and 8 of Enterohemorrhagic Escherichia coli O157:H7 With Particular stx Subtypes are More Frequently Found in Isolates From Hemolytic Uremic Syndrome Patients Than From Asymptomatic Carriers. Open Forum Infect Dis 1, ofu061, 10.1093/ofid/ofu061 (2014).
Perna, N. T. et al. Genome sequence of enterohaemorrhagic Escherichia coli O157:H7. Nature 409, 529–533 (2001).
Riordan, J. T., Viswanath, S. B., Manning, S. D. & Whittam, T. S. Genetic differentiation of Escherichia coli O157:H7 clades associated with human disease by real-time PCR. J Clin Microbiol 46, 2070–2073 (2008).
Piérard, D., Muyldermans, G., Moriau, L., Stevens, D. & Lauwers, S. Identification of new verocytotoxin type 2 variant B-subunit genes in human and animal Escherichia coli isolates. J Clin Microbiol 36, 3317–3322 (1998).
Makino, K. et al. Complete nucleotide sequence of the prophage VT2-Sakai carrying the verotoxin 2 genes of the enterohemorrhagic Escherichia coli O157:H7 derived from the Sakai outbreak. Genes Genet Syst 74, 227–239 (1999).
Plunkett, G., 3rd, Rose, D. J., Durfee, T. J. & Blattner, F. R. Sequence of Shiga toxin 2 phage 933W from Escherichia coli O157:H7: Shiga toxin as a phage late-gene product. J Bacteriol 181, 1767–1778 (1999).
Strauch, E. et al. Bacteriophage 2851 is a prototype phage for dissemination of the Shiga toxin variant gene 2c in Escherichia coli O157:H7. Infect Immun 76, 5466–5477 (2008).
Su, L. K. et al. Lysogenic infection of a Shiga toxin 2-converting bacteriophage changes host gene expression, enhances host acid resistance and motility. Mol Biol 44, 60–73 (2010).
Xiong, Y. et al. A novel Escherichia coli O157:H7 clone causing a major hemolytic uremic syndrome outbreak in China. PLoS One 7, e36144, 10.1371/journal.pone.0036144 (2012).
Cote, R. et al. Comparative analysis of super-shedder strains of Escherichia coli O157:H7 reveals distinctive genomic features and a strongly aggregative adherent phenotype on bovine rectoanal junction squamous epithelial cells. PLoS One 10, e0116743, 10.1371/journal.pone.0116743 (2015).
Wagner, P. L. et al. Role for a phage promoter in Shiga toxin 2 expression from a pathogenic Escherichia coli strain. J Bacteriol 183, 2081–2085 (2001).
Brooks, J. T. et al. Non-O157 Shiga toxin-producing Escherichia coli infections in the United States, 1983-2002. J Infect Dis 192, 1422–1429 (2005).
Coombes, B. K. et al. Molecular analysis as an aid to assess the public health risk of non-O157 Shiga toxin-producing Escherichia coli strains. Appl Environ Microbiol 74, 2153–2160 (2008).
Leopold, S. R. et al. A precise reconstruction of the emergence and constrained radiations of Escherichia coli O157 portrayed by backbone concatenomic analysis. Proc Natl Acad Sci USA 106, 8713–8718, 10.1073/pnas.0812949106 (2009).
Turabelidze, G. et al. Precise dissection of an Escherichia coli O157:H7 outbreak by single nucleotide polymorphism analysis. J Clin Microbiol 51, 3950–3954, 10.1128/JCM.01930-13 (2013).
Donohue-Rolfe, A., Kondova, I., Oswald, S., Hutto, D. & Tzipori, S. Escherichia coli O157:H7 strains that express Shiga toxin (Stx) 2 alone are more neurotropic for gnotobiotic piglets than are isotypes producing only Stx1 or both Stx1 and Stx2. J Infect Dis 181, 1825–1829, 10.1086/315421 (2000).
Cornick, N. A., Jelacic, S., Ciol, M. A. & Tarr, P. I. Escherichia coli O157:H7 infections: discordance between filterable fecal shiga toxin and disease outcome. J Infect Dis 186, 57–63, 10.1086/341295 (2002).
Hayashi, T. et al. Complete genome sequence of enterohemorrhagic Escherichia coli O157:H7 and genomic comparison with a laboratory strain K-12. DNA research 8, 11–22 (2001).
Ohnishi, M. et al. Genomic diversity of enterohemorrhagic Escherichia coli O157 revealed by whole genome PCR scanning. Proc Natl Acad Sci USA 99, 17043–17048 (2002).
Pollard, D. R., Johnson, W. M., Lior, H., Tyler, S. D. & Rozee, K. R. Rapid and specific detection of verotoxin genes in Escherichia coli by the polymerase chain reaction. J Clin Microbiol 28, 540–545 (1990).
Ogura, Y. et al. Systematic identification and sequence analysis of the genomic islands of the enteropathogenic Escherichia coli strain B171-8 by the combined use of whole-genome PCR scanning and fosmid mapping. J Bacteriol 190, 6948–6960 (2008).
Sugawara, H., Ohyama, A., Mori, H. & Kurokawa, K. Microbial Genome Annotation Pipeline (MiGAP) for diverse users. The 20th International Conference on Genome Informatics (GIW2009) Poster and Software Demonstrations (Yokohama), S001-001-002. (2009).
Zerbino, D. R. & Birney, E. Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res 18, 821–829 (2008).
Blattner, F. R. et al. The complete genome sequence of Escherichia coli K-12. Science 277, 1453–1462 (1997).
Kurtz, S. et al. Versatile and open software for comparing large genomes. Genome Biol 5, R12 (2004).
Stamatakis, A. RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22, 2688–2690 (2006).
Acknowledgements
We thank A. Yoshida, Y. Inoue, M. Shimbara, N. Kanemaru, N. Kawano, N. Sakamoto and H. Iguchi for providing technical assistance. This work was supported by KAKENHI on the “Kiban-B: 20310116” (to T.H.), “Wakate-B: 23790478” (to Y.O.) and “Grant-in-Aid for Scientific Research on Innovative Areas: 25125718” (to Y.O.) from the Ministry of Education, Culture, Sports, Science and Technology of Japan.
Author information
Authors and Affiliations
Contributions
Y.O. and T.H. designed the research. Y.O., S.I.M., M.R.I., T.M., K.A., K.K., T.O., Y.G., K.M. and M.O. performed the experiments. Y.O. and T.H. wrote the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Ogura, Y., Mondal, S., Islam, M. et al. The Shiga toxin 2 production level in enterohemorrhagic Escherichia coli O157:H7 is correlated with the subtypes of toxin-encoding phage. Sci Rep 5, 16663 (2015). https://doi.org/10.1038/srep16663
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep16663
- Springer Nature Limited