Abstract
Previous studies have investigated the association between osteopontin (OPN) gene polymorphisms, rs17524488 (−156 GG/G), rs11730582 (−443 T/C) and rs9138 (C/A) and cancer risk in the Chinese population. However, the results are controversial and indefinite. We therefore carried out a meta-analysis to derive a more precise estimation of these associations. The PubMed database was systematically searched to identify potentially eligible reports. Crude odds ratios (OR) and 95% confidence intervals (CI) were used to assess the strength of associations between 3 OPN gene polymorphisms and cancer risk in a Chinese population. A total of 10 articles involving 2,391 cases and 3,007 controls were evaluated. The pooled OR indicated that OPN rs17524488 (−156 GG/G) polymorphism was significantly associated with cancer risk in Chinese population. In a stratified analysis by source of control, significant associations were also observed among rs17524488 (−156 GG/G) and rs11730582 (−443 T/C) polymorphisms and cancer. In addition, a stronger association was observed between rs9138 (C/A) polymorphism and cancer risk. In conclusion, this meta-analysis suggests that OPN rs17524488 (−156 GG/G), rs11730582 (−443 T/C) and rs9138 (C/A) polymorphisms may be associated with cancer susceptibility in the Chinese population. Nevertheless, further investigation on a larger population covering different ethnicities are warranted.
Similar content being viewed by others
Introduction
Cancer is a serious disease threatening public health worldwide. The estimates of newly diagnosed cancer cases/cancer-related deaths in worldwide and China were approximately 14.1/8.2 million in 2012 and 3.37/2.11 million in 2011, respectively1,2. The crude incidence was 235.23/105 (268.65/105 in males, 200.21/105 in females)3 in China. Predisposition to cancer may be attributed to certain genetic polymorphisms that arise from single nucleotide polymorphisms (SNPs). In fact, numerous genome-wide studies of common cancers suggest a number of loci within the genome that, in spite of low-penetrance, may increase an individual’s susceptibility to cancer4,5,6.
Osteopontin (OPN) is a phosphorylated acidic glycoprotein with several functions in different physiological and pathological processes, including bone turnover, wound healing, ischemia, inflammation, autoimmune responses and tumorigenesis, mediated by stimulation of certain signaling pathways via binding to cellular receptors, integrins and CD44 variants7,8,9.
Overexpression of OPN has been described in several conditions in which basic inflammatory processes are activated, such as arthritis10, myocardial remodeling after infarction11, kidney interstitial fibrosis after obstructive uropathy and other renal insults12, wound healing13 and several types of cancer. This is because OPN is a metastasis-related gene that contributes to the progression of over 30 types of cancer14,15.
The gene encoding OPN, also known as secreted phosphoprotein 1 (SPP1), is mapped on human chromosome 4q21-q25, together with other members of the SIBLING family of proteins, bone sialoprotein and dentin matrix protein-1, which share some structural characteristics16.
More than 10 SNPs have been identified in the OPN promoter. These polymorphisms may affect the transcriptional activity of OPN and some of them are thought to be genetic risk factors for disease susceptibility17,18,19. Several polymorphisms in the human gene encoding OPN have been identified in the Chinese population, of which the rs17524488 (−156 GG/G), rs11730582 (−443 T/C) and rs9138 (C/A) polymorphisms are most frequently studied.
Considering the impact of the cancer risk potentially resulting from OPN gene, a number of studies have explored the association between these polymorphisms and cancer. However, individual studies have yielded inconsistent or conflicting findings, possibly caused by limitations associated with an individual study. To shed light on these contradictory results and to more precisely evaluate the relationship among OPN gene polymorphisms and cancer risk, we performed a meta-analysis of 10 published studies20,21,22,23,24,25,26,27,28,29, the original regions of which were all from China, no other ethnicities or regions existed.
Methods
Search strategy and inclusion criteria
We did our best to include all case–control studies published until date, regarding the association between OPN rs17524488 (−156 GG/G), rs11730582 (−443 T/C) and rs9138 (C/A) polymorphisms and cancer risk. Eligible studies were found by searching the PubMed database for relevant reports published between 2010 and 2014. The search terms were “Osteopontin”, “polymorphism” and “cancer”. In addition, the references of all retrieved articles were also manually searched for additionally relevant publications. The inclusion criteria were as follows: (1) evaluating the association between OPN rs17524488 (−156 GG/G), rs11730582 (−443 T/C) and rs9138 (C/A) polymorphisms and cancer risk in a Chinese population; (2) case–control study; (3) and sufficient information (GG/GG, GG, GGG for rs17524488; TT, CC, TC for rs11730582; and CC, AA, CA for rs9138) for calculating the pooled odds ratios (OR) with 95% confidence intervals (CI).
Data extraction
Data included the following: first author, publication year, country, cancer type, source of control, each genotype frequency of the case and control groups, genotype methods and the Hardy–Weinberg equilibrium (HWE) value in the control group.
Statistical analysis
Odds ratios (OR) with 95% confidence intervals (CI) were assessed for determining the relationship between OPN rs17524488 (−156 GG/G), rs11730582 (−443 T/C) and rs9138 (C/A) polymorphisms and cancer. The pooled OR was estimated for rs17524488 (−156 GG/G) by homozygous (GG vs. GG/GG) and recessive models [GG vs. (GGG + GG/GG)] as well as the allele model (G vs. GG); rs11730582 (−443 T/C) by homozygous (CC vs. TT) and recessive models [CC vs. (CT + TT)] as well as the allele model (C vs. T); and rs9138 (C/A) by homozygous (AA vs. CC) and recessive models [AA vs. (AC + CC)] as well as the allele model (A vs. C).
Heterogeneity was evaluated using a chi-square-based Q-test30 and the summary OR was determined with the Z-test. If P > 0.10 for the Q-test, a lack of heterogeneity among studies was found, meanwhile the fixed effects model should be used, otherwise, the random effects model should be used31,32. The HWE was assessed by a chi-square test in controls; P < 0.05 was considered significant. Sensitivity analysis was performed on excluded individual studies to assess the stability of the results. Publication bias was assessed by both Egger’s test and Begg’s test33. All statistical tests were used by Stata software (version 11.0; StataCorp LP, College Station, TX).
Results
Characteristics of Studies
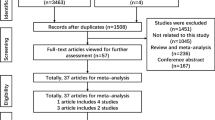
Figure 1 and Tables 1, 2 show the study selection process and main characteristics of included studies, respectively. A total of 24 articles were retrieved based on the search criteria. Among them, 10 articles were excluded because they did not provide information about OPN gene polymorphism. An additional 4 articles, without control group data, were excluded. Thus, a total of 10 articles with 2,391 cases and 3,007 controls were included in the meta-analysis20,21,22,23,24,25,26,27,28,29. For the rs17524488 (−156 GG/G) polymorphism, 7 studies were available, including a total of 2,130 cases and 2,644 controls. For the rs11730582 (−443 T/C) polymorphism, 8 studies involved a total of 2,241 cases and 2,857 controls. For the rs9138 (C/A) polymorphism, 2 studies involved a total of 418 cases and 424 controls. Among these, 2 studies focused on gastric cancer. The distribution of genotypes among controls was consistent with HWE in all but 2 studies20,21.
Quantitative data synthesis
Results of OPN rs17524488 (−156 GG/G), rs11730582 (−443 T/C) and rs9138 (C/A) polymorphisms and cancer risk are presented in Table 3 and Figs 2, 3, 4, 5. For rs17524488 (−156 GG/G) polymorphism, significant association was observed in all cancer-type combined studies (GG vs. GGG + GG/GG: OR = 0.81, 95% CI: 0.66–0.99, P = 0.028 for heterogeneity, P = 0.043). Subgroup analysis by source of control showed that statistically significant associations were present in PB (GG vs. GGG + GG/GG: OR = 0.81, 95% CI: 0.72–0.91, P = 0.558 for heterogeneity, P = 0.000; GG vs. GG/GG: OR = 0.84, 95% CI: 0.77–0.91, P = 0.103 for heterogeneity, P = 0.000; G vs. GG: OR = 0.75, 95% CI: 0.67−0.85, P = 0.232 for heterogeneity, P = 0.000). Two studies20,21 were not satisfied with the HWE, to make our analysis more powerful, we excluded these two studies and re-analysis. To our regret, no association was found, which indicated that the heterogeneity may exist in this SNP.
Forest plot of cancer risk associated with the OPN rs17524488 (−156 GG/G) polymorphism (GG vs. GGG+GG/GG) in the whole.
The squares and horizontal lines correspond to the study-specific OR and 95% CI. The area of the squares reflects the weight (inverse of the variance). The diamond represents the summary OR and 95% CI.
Forest plot of cancer risk associated with the OPN rs17524488 (−156 GG/G) polymorphism (G vs. GG) in the PB subgroup.
The squares and horizontal lines correspond to the study-specific OR and 95% CI. The area of the squares reflects the weight (inverse of the variance). The diamond represents the summary OR and 95% CI.
Forest plot of cancer risk associated with the OPN rs11730582 (−443 T/C) polymorphism (CC vs. TT) in the PB subgroup.
The squares and horizontal linescorrespond to the study-specific OR and 95% CI. The area of the squares reflects the weight (inverse of the variance). The diamond represents the summary OR and 95% CI.
Forest plot of cancer risk associated with the OPN rs9138 (C/A) polymorphism (AA vs. CC) in the whole.
The squares and horizontal lines correspond to the study-specific OR and 95% CI. The area of the squares reflects the weight (inverse of the variance). The diamond represents the summary OR and 95% CI.
For the rs11730582 (−443 T/C) polymorphism, significant association with cancer risk was observed in PB subgroup (CC vs. CT + TT: OR = 0.63, 95% CI: 0.49–0.82, P = 0.903 for heterogeneity, P = 0.000; CC vs. TT: OR = 0.46, 95% CI: 0.35–0.62, P = 0.200 for heterogeneity, P = 0.000). For the rs9138 (C/A) polymorphism, significant relationship was detected overall (AA vs. AC + CC: OR = 1.62, 95% CI: 1.02–2.57, P = 0.883 for heterogeneity, P = 0.041; AA vs. CC: OR = 2.16, 95% CI: 1.28–3.63, P = 0.565 for heterogeneity, P = 0.004). To our regret, no association was found between gastric cancer and OPN rs17524488 (−156 GG/G) or rs11730582 (−443 T/C) polymorphism.
Sensitivity analysis and publication bias. Sensitivity analysis was used to determine whether modification of the inclusion criteria affected the final results. The sensitivity analysis did not influence the results excessively by omitting any single study for rs17524488 (−156 GG/G) (Fig. 6). However, for rs11730582 (−443 T/C), a single study named Mu et al.23 may influence the whole results (Fig. 7). Because only two studied of rs9138 (C/A), the sensitivity analysis was not examed. Egger’s and Begg’s tests were performed to assess publication bias and the funnel plot symmetry was examined. Finally, no proof of publication bias was obtained (Table 4, Figs 8, 9).
Discussion
The overall goal of a meta-analysis is to combine the results of previous studies to arrive at a summary conclusion about a body of research. It is most useful in summarizing prior research when individual studies are too small to yield a valid conclusion. In this study, we analyzed the associations between OPN rs17524488 (−156 GG/G), rs11730582 (−443 T/C) and rs9138 (C/A) polymorphisms and cancer risk using a meta-analysis to obtain a powerful conclusion. To the best of our knowledge, this is the first meta-analysis providing comprehensive insights into the effects of the OPN rs17524488 (−156 GG/G), rs11730582 (−443 T/C) and rs9138 (C/A) polymorphisms and risk associated with all types of cancer in a Chinese population. Our meta-analysis included 2,391 cases and 3,007 controls20,21,22,23,24,25,26,27,28,29.
For the rs17524488 (−156 GG/G) polymorphism in the OPN promoter region, the overall results suggested that the subjects with G allele showed decreased susceptibility to cancer in a Chinese population. Moreover, individuals carrying either −156 GG or −443 CC genotype may have lower cancer susceptibility. However, people with AA genotype may have decreased cancer risk than CC-, CT, or CC + CT carriers. Considering that the previous single-institution study for cancer had a small sample size and may not justify the significance of current work, further studies are needed to clarify the effect of the 3 polymorphisms on the risk of cancer. A possible explanation for this phenomenon is that different polymorphisms may exert different effects on gene function, subsequently resulting in varying cancer susceptibility. Furthermore, a single gene or a single environmental factor is not likely to have a large effect on cancer susceptibility. Complex interactions between several genetic and environmental factors may be involved in cancer development.
Meta-analysis has been recognized as an effective method to answer a wide variety of clinical questions by summarizing and reviewing previously published, quantitative research. However, some limitations in our meta-analysis should be mentioned. First, our results were based on unadjusted estimates; more accurate outcomes would result from adjustments for other confounders such as gender, age, body mass index, lifestyle and so on. Second, the studies included in this analysis were insufficient, especially in terms of a subgroup analysis. Thus, potential publication bias is very likely to exist, in spite of no evidence obtained from our statistical tests. Third, language of studies was limited to English, which may result in potential language bias. Fourth, a comparison of mRNA expression levels of the OPN gene between cancer and normal tissue should have been reported and included, which would better explain genetic function. Next, inter-gene and gene-environment interactions had not been evaluated owing to the absence of original data. Fifth, our study focused on Chinese people, other ethnicities should be reported and included. Sixth, a single study23 may influence the whole results in the sensitivity analysis for the rs11730582 (−443 T/C) polymorphism, which suggested our study may be poorly powerful and stable. Finally, we also could not integrate different studies to look at the association between these three OPN SNPs and one specific cancer type, owning to insufficient publications. If there have a number of studies related different cancers in the future, this work may be carried out. In contrast, some advantages should also be highlighted. Our analysis comprehensively and systematically sheds light on the relationship between OPN rs17524488 (−156 GG/G), rs11730582 (−443 T/C) and rs9138 (C/A) polymorphisms and the susceptibility to cancer in the Chinese population. Additionally, due to the larger sample size, our meta-analysis increases the power and plausibility of our conclusion when compared with previous, individual studies. Finally, the studies included in this analysis were published between 2010 and 2014; thus, these studies are quite recent.
In summary, our meta-analysis suggests that OPN rs17524488 (−156 GG/G), rs11730582 (−443 T/C) and rs9138 (C/A) polymorphisms are associated with cancer risk in the Chinese population. Larger sample sizes of different ethnic populations are required to confirm our findings.
Additional Information
How to cite this article: Mi, Y. et al. OPN gene polymorphisms, rs17524488 GG/G, rs11730582 T/C and rs9138 C/A and cancer risk in a Chinese population. Sci. Rep. 5, 14164; doi: 10.1038/srep14164 (2015).
References
Torre, L. A. et al. Global cancer statistics, 2012. CA Cancer J Clin 65, 87–108 (2015).
The National Cancer Center, the Ministry of Health Bureau of disease control and Prevention. China cancer registry annual report 2015. http://www.moh.gov.cn/jkj/. Accessed April 13, 2015.
Chen, W. Q. et al. Report of Cancer Incidence and Mortality in China, 2010. Chin Cancer 23, 1–10 (2014).
Dong, L. M. et al. Genetic susceptibility to cancer: the role of polymorphisms in candidate genes. JAMA 299, 2423–2436 (2008).
Wokolorczyk, D. et al. A range of cancers is associated with the rs6983267 marker on chromosome 8. Cancer Res 68, 9982–9986 (2008).
Harismendy, O. & Frazer, K. A. Elucidating the role of 8q24 in colorectal cancer. Nat Genet 41, 868–869 (2009).
Denhardt, D. T. et al. Osteopontin-induced modifications of cellular functions. Ann N Y Acad Sci 760, 127–142 (1995).
Weber, G. F., Ashkar, S. & Cantor, H. Interaction between CD44 and osteopontin as a potential basis for metastasis formation. Proc Assoc Am Physicians 109, 1–9 (1997).
Wang, K. X. & Denhardt, D. T. Osteopontin: role in immune regulation and stress responses. Cytokine Growth Factor Rev 19, 333–345 (2008).
Yamamoto, N. et al. Essential role of the cryptic epitope SLAYGLR within osteopontin in a murine model of rheumatoid arthritis. J Clin Invest 112, 181–188 (2003).
Tamura, A. Osteopontin is released from the heart into the coronary circulation in patients with a previous anterior wall myocardial infarction. Circ J 67, 742–744 (2003).
Xie, Y. et al. Expression, roles, receptors and regulation of osteopontin in the kidney. Kidney Int 60, 1645–1657 (2001).
Rittling, S. R. & Denhardt, D. T. Osteopontin function in pathology: lessons from osteopontin-deficient mice. Exp Nephrol 7, 103–113 (1999).
Weber, G. F., Lett, G. S. & Haubein, N. C. Osteopontin is a marker for cancer aggressiveness and patient survival. Br J Cancer 103, 861–869 (2010).
Weber, G. F. The cancer biomarker osteopontin: combination with other markers. Cancer Genomics Proteomics 8, 263–288 (2011).
Fisher, L. W. & Fedarko, N. S. Six genes expressed in bones and teeth encode the current members of the SIBLING family of proteins. Connect Tissue Res 44, 33–40 (2003).
Giacopelli, F. Polymorphisms in the osteopontin promoter affect its transcriptional activity. Physiol Genomics 20, 87–96 (2004).
Liu, C. C. et al. The impact of osteopontin promoter polymorphisms on the risk of calcium urolithiasis. Clin Chim Acta 411, 739–7343 (2010).
Hendig, D. et al. SPP1 promoter polymorphisms: identification of the first modifier gene for pseudoxanthoma elasticum. Clin Chem 53, 829–836 (2007).
Zhao, F. et al. Genetic polymorphisms in the osteopontin promoter increases the risk of distance metastasis and death in Chinese patients with gastric cancer. BMC Cancer 12, 477 (2012).
Chen, Y. et al. Osteopontin genetic variants are associated with overall survival in advanced non-small-cell lung cancer patients and bone metastasis. J Exp Clin Cancer Res 32, 45 (2013).
Fan, Y. et al. The polymorphisms of osteopontin gene and plasma osteopontin protein levels with susceptibility to colorectal carcinoma. DNA Cell Biol 32, 594–600 (2013).
Mu, G., Wang, H., Cai, Z. & Ji, H. OPN −443C>T genetic polymorphism and tumor OPN expression are associated with the risk and clinical features of papillary thyroid cancer in a Chinese cohort. Cell Physiol Biochem 32, 171–179 (2013).
Wang, J. L. et al. Correlation between OPN gene polymorphisms and the risk of nasopharyngeal carcinoma. Med Oncol 31, 20 (2014).
Xu, Q. et al. OPN gene polymorphisms are associated with susceptibility and clinicopatholigical characteristics of cervical cancer in a Chinese cohort. Cancer Biomark 10, 233–239 (2011).
Lee, T. Y. et al. Osteopontin promoter polymorphisms are associated with susceptibility to gastric cancer. J Clin Gastroenterol 47, e55–59 (2013).
Wang, J. et al. Association of osteopontin polymorphisms with nasopharyngeal carcinoma risk. Hum Immunol 75, 76–80 (2014).
Chiu, Y. W. et al. The implication of osteopontin (OPN) expression and genetic polymorphisms of OPN promoter in oral carcinogenesis. Oral Oncol 46, 302–306 (2010).
Chen, J. et al. SPP1 promoter polymorphisms and glioma risk in a Chinese Han population. J Hum Genet 55, 456–461 (2010).
Li, J. et al. Identification of high-quality cancer prognostic markers and metastasis network modules. Nat Commun 13, 1–34 (2010).
Mantel, N. & Haenszel, W. Statistical aspects of the analysis of data from retrospective studies of disease. J Natl Cancer Inst 22, 719–748 (1959).
DerSimonian, R. & Laird, N. Meta-analysis in clinical trials. Control Clin Trials 7, 177–188 (1986).
Hayashino, Y., Noguchi, Y. & Fukui, T. Systematic evaluation and comparison of statistical tests for publication bias. J Epidemiol 15, 235–243 (2005).
Acknowledgements
This work was supported by the Joint research project of Hospital management center in Wuxi (No. YGZX1203, 1318).
Author information
Authors and Affiliations
Contributions
Y.M. and F.D. devised the concept, designed the experiment and wrote the manuscript. F.D. and K.R. performed the experiments and analyzed the data. L.Z. and N.F. supervised the project. All authors discussed the results and contributed in this manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Mi, Y., Ren, K., Dai, F. et al. OPN gene polymorphisms, rs17524488 GG/G, rs11730582 T/C and rs9138 C/A and cancer risk in a Chinese population. Sci Rep 5, 14164 (2015). https://doi.org/10.1038/srep14164
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep14164
- Springer Nature Limited