Abstract
In flowering plants, three genomes (nuclear, mitochondrial and plastid) coexist and intracellular horizontal transfer of DNA is prevalent, especially from the plastid to the mitochondrion genome. However, the plastid genomes are generally conserved in evolution and have long been considered immune to foreign DNA. Recently, the opposite direction of DNA transfer from the mitochondrial to the plastid genome has been reported in two eudicot lineages. Here we sequenced 6 plastid genomes of bamboos, three of which are neotropical woody species and three are herbaceous ones. Several unusual features were found, including the duplication of trnT-GGU and loss of one copy of rps19 due to contraction of inverted repeats (IRs). The most intriguing was the ~2.7 kb insertion in the plastid IR regions in the three herbaceous bamboos. Furthermore, the insertion was documented to be horizontally transferred from the mitochondrial to the plastid genome. Our study provided evidence of the mitochondrial-to-plastid DNA transfer in the monocots, demonstrating again that this rare event does occur in other angiosperm lineages. However, the mechanism underlying the transfer remains obscure and more studies in other plants may elucidate it in the future.
Similar content being viewed by others
Introduction
Horizontal DNA transfer, the transfer of genetic material between donor and recipient organisms by asexual means, has played a major role in the evolution of bacteria1,2. Recently, the prevalence and importance of this process in the evolution of eukaryotes is being increasingly recognized, especially in plants3,4,5. In addition to the plant-to-plant horizontal DNA transfer, a large amount of these cases reported involved the transfer of DNA within a cell in the same plant species6,7,8.
In flowering plants, three distinct genomes of nuclear, mitochondrial and plastid coexist within each individual cell, facilitating the intracellular DNA exchange. However, the direction of DNA transfer between the three genomes is highly uneven9. For example, the nuclear genome usually harbors lots of nucleotide sequences derived from the mitochondrial and plastid DNA7,8,10. Similarity, the mitochondrial genome can also accumulate large amounts of nuclear DNA and particularly plastid DNA11,12,13, although the size of mitochondrial genome is generally much smaller than the nuclear genome in angiosperms. By contrast, the plastid genome is nearly immune from the promiscuous DNA and the transfer of DNA from the nuclear and mitochondrial to the plastid genome is very scarce7,9. There are only two examples of dicots reported in the angiosperms: one from the carrot (Daucus carota)14,15 and the other from the common milkweed (Asclepias syriaca)16. Both of them are the transfer of DNA from mitochondrion to plastid and the transferred segment is ~1.5 kb and ~2.4–4.7 kb in length in the plastid genomes of carrots and milkweeds14,15,16, respectively. Nevertheless, little is known about the exact mechanisms underlying the transfer9, although the non-LTR retrotransposon in the carrots15 or homologous recombination through double-strand break repair in the milkweeds16 was proposed as the possible mechanism in explaining the transfer.
The plastid genomes of angiosperms are highly conserved in evolution17,18,19. The plastid genomes usually have a circular genome organization, with a pair of inverted repeats (IRs) dividing the whole genome into two single-copy regions: large-single copy (LSC) and small-single copy (SSC) regions. The angiosperms plastid genomes range generally from 120 to 160 kb in size, encoding approximately 80 unique protein-coding genes that are mainly responsible for photosynthesis. And the gene order in the plastid genomes is generally conserved across angiosperm species. Furthermore, the rates of DNA sequence substitution are slow in the plastid genomes and about threefold lower than the nuclear DNA17,20.
The approximately 1430 species of bamboos around the world comprise of the Bambusoideae, a subfamily in the grass family (Poaceae), which can be divided into three main lineages: temperate woody bamboos (tribe Arundinarieae), tropical woody bamboos (tribe Bambuseae) and herbaceous bamboos (tribe Olyreae)21,22. The tropical woody bamboos can be further divided into neotropical and paleotropical woody bamboos21,22. Three subtribes (Buergersiochloinae, Parianinae and Olyrinae) are recognised within Olyreae, consisting of 21 genera and 122 species21. There were 30 complete plastid genomes of bamboos sequenced when the manuscript was submitted and 29 of them were from the woody bamboos23,24,25. All the sequenced bamboos have the same plastid genome organization and are very similar to each other in nearly every aspect of genome features, only with limited DNA sequence divergence23,24,25.
Results
Sequencing of six bamboo plastid genomes
We reported here the complete plastid genome sequences of six bamboos through sequencing of total genomic DNA or PCR-amplified plastid DNA using Illumina technology. Among these plastid genomes, three were from the herbaceous bamboo genus Pariana and the other three were the neotropical woody bamboos: Chusquea sp., C. circinata and Otatea glauca21 (Table 1). The C. circinata and O. glauca were sequenced on PCR-amplified plastid DNA with 2.45 and 1.96 million quality filtered paired-end reads of 90 bp obtained and an estimated sequencing depth of 4638.7× and 3057.7×, respectively. The remaining four bamboos were sequenced on total DNA and 1.99, 2.12, 3.07 and 3.12 million quality filtered paired-end reads of 90 bp with an estimated sequencing depth of 41.0×, 23.7×, 14.4× and 30.1× were obtained for Chusquea sp., Pariana sp., P. radiciflora and P. campestris, respectively. The four junctions between IRs and LSC/SSC in the sequenced plastid genomes were all validated by Sanger resequencing24.
The six newly sequenced plastid genomes of bamboos are very similar to those of previously published bamboos23,24,25 (Table 1). The presence or absence of genes in the genome is the same in these bamboos, except that Chusquea sp. has two copies of trnT-GGU and all three members of Pariana encode only one copy of the rps19 gene. The 72-bp trnT-GGU is duplicated in Chusquea sp. associated with a 94-bp tandem repeat, but not in other bamboos. The rps19 gene near the junctions between IRs and LSC regions is always confined to the IR regions in the sequenced bamboo plastid genomes23,24,25, while in Pariana it lies entirely within the LSC region due to shift of the IRs/LSC junctions. In terms of genome size, the lineage of neotropical woody bamboos shows a tendency of reduction in the Bambusoideae and the reduction is especially evident in the IRs size of O. glauca (~20.3 kb versus an average ~21.8 kb in other bamboos) (Table 1). By contrast, the IRs size undergoes expansion (~22.5 kb versus an average ~21.8 kb in other bamboos) in Pariana with an overall genome size larger than 140 kb. Although the evolution of the IR regions is very conserved17, these newly sequenced plastid genomes were variable in the IRs size. Further investigation revealed that the variation could be attributed to the variability in the length of the intergenic spacer between trnI-CAU and trnL-CAA.
Insertion of mitochondrion-derived DNA in the trnI-CAU-trnL-CAA intergenic spacer
The trnI-CAU-trnL-CAA intergenic spacer located in the IR regions is very similar in size among the 30 sequenced plastid genomes of bamboos, ranging narrowly from 2471 to 2525 bp. The remnants of the pseudogenized ycf2 gene that is encoded in most other angiosperms could be identified in this intergenic spacer in grasses26. However, sequencing of the six bamboo plastid genomes here revealed extensive variation of this intergenic spacer in length: Pariana had the longest (3811 bp) and O. glauca the shortest (950 bp) (Fig. 1). By contrast, the size of other intergenic spacers of these bamboos in the IR regions remained fairly constant as in other bamboos. In O. glauca, the trnI-CAU-trnL-CAA intergenic spacer was much reduced to 950 bp because of a large deletion of 1542 bp between nucleotide positions 234 to 1775 when using C. circinata as a reference. This spacer also experienced sequence deletion at nearly the same position as in O. glauca (positions 421 to 1799 in C. circinata) in all three species of Pariana (Fig. 1) and the size of the deletion was estimated to be 1379 bp. However, the 1379-bp deletion in Pariana was accompanied by a 2706-bp insertion with the total size of the trnI-CAU-trnL-CAA intergenic spacer expanding to 3811 bp (Fig. 1). In addition, the sequences of insertion differed by only one nucleotide site (A to C substitution) among the three sequenced species of Pariana. Although the similarity of sequences surrounding the insertion in the trnI-CAU-trnL-CAA intergenic spacer was very high (>98%) between Pariana and the other sequenced bamboos, the sequence of insertion did not show similarity to any other sequences in the bamboo plastid genomes, suggesting that it perhaps originated from foreign DNA.
Variability in the trnI-CAU-trnL-CAA intergenic spacer in the plastid genomes of Bambusoideae with the size indicated in parentheses.
The insertion without plastid DNA homology in the plastid genomes of Pariana is shown in blue and the other plastid DNA sequences are shown in green. Sequence deletions larger than 25 bp relative to other species are indicated by black lines.
To identify the origin of the insertion, the inserted sequence in the plastid genome of P. campestris was used as query in a Blastn search against all the sequences deposited in the NCBI nucleotide database. A total of 16 significant hits with >70% similarity to the query over 100 bp were identified and 12 of these with ≥85% similarity were listed in Table 2. These hits were all derived from the mitochondrial DNA sequences of angiosperms, except that a 201-bp segment of the insertion had matching sequences from both mitochondrial and nuclear genomes of Triticum aestivum (Table 2). This result strongly suggested that the inserted sequence in plastid genome of Pariana originated from the mitochondrial DNA. Furthermore, the hit receiving the highest Blastn score was from the mitochondrial genome of the bamboo Ferrocalamus rimosivaginus (Arundinarieae)27, covering ~97% sequence of the insertion. The second one was also from another bamboo Bambusa oldhamii (Bambuseae), covering ~29% sequence of the insertion. It should here be noted that the F. rimosivaginus and B. oldhamii are the only two bamboos with mitochondrial genome sequences available in the NCBI nucleotide database to date. The remaining hits all covered less than 20% sequence of the insertion (Table 2). Notably, the first 75 bp sequence of the insertion lacked detectable similarity with any of the plastid, mitochondrial, or nuclear sequences available in the NCBI nucleotide database.
Validation of the mitochondrion-to-plastid DNA transfer
Given that the plastid genomes are highly conserved in bamboos23,24,25, the large insertion being of mitochondrial DNA origin is unexpected. It is needed to confirm that the insertion was not a product of misassembly. As we used total genomic DNA of Pariana for Illumina sequencing, the reads were a mixture of DNA sequences from plastid, mitochondrial and nuclear genomes. However, the plastid genome is usually the smallest one among these three genomes in angiosperms and the assembled plastid DNA sequences can be easily distinguished from mitochondrial and nuclear DNA sequences by sequencing depth. As shown in Fig. 2, the inserted region in P. campestris had an average sequencing depth of 56.9×, which was very similar to the surrounding regions with an average sequencing depth of 61.8× and 58.8× before and after the insertion, respectively. Although different sequencing depths of 29.8× and 28.7× were revealed for the inserted regions in Pariana sp. and P. radiciflora, respectively, the constant sequencing depth in the surrounding regions was also observed in these two bamboos (Supplementary Fig. 1). On the other hand, we mapped the Illumina reads of P. campestris to three randomly selected intronless genes from the F. rimosivaginus mitochondrial genome27, ccmFN, matR and rrn26 and obtained much lower sequencing depth of 5.6×, 7.1× and 11.9×, respectively. The sequencing depth for the three genes was also low in the other two Pariana species, ranging from 3.5× to 6.8×. These results indicated that the inserted regions had similar sequencing depth to the surrounding plastid regions, rather than the mitochondrial genome, even in considering that reads originating from the mitochondrial genome may have added a similar sequencing depth as inferred from the three mitochondrial genes to the inserted regions due to sequence similarity. The nuclear genome is much larger than mitochondrial genome and the sequencing depth would be even lower. In all, the analysis of sequencing depth demonstrated that the insertion in the plastid genomes of Pariana was unlikely due to misassembly.
We further used PCR amplification and Sanger sequencing to confirm the insertion. PCR amplification using two pairs of primers (Supplemental Table 1) derived from the regions flanking the endpoints of the insertion yielded the predicted junction fragments in all three Pariana species but not in the other representative bamboos in Fig. 1. We also performed PCR amplification that was only expected for the bamboos without the insertion. There were no specific amplicons obtained in the three species of Pariana, just as predicted, whereas amplifications were successful when using the representative bamboos (Fig. 1) without the insertion as templates. Furthermore, we designed additional 4 pairs of primers to sequence the whole insertion region (Supplemental Table 1). Sanger sequencing of the PCR products from these 6 pairs of primers yielded a total of 3327 bp sequences for each of the three Pariana species, which was separately identical to the assembled plastid genomes. Finally, we used PCR to screen P. parvispica and Eremitis parviflora from the sister genus within the same subtribe Parianinae21,28 with available DNA materials for the presence of the insertion. However, PCR products were only obtained with 4 pairs of primers (approximately from positions 84,798 to 87,222 in P. campestris in Fig. 2 spanning one junction of the insertion) and 2367 and 2430 bp of sequences were generated for P. parvispica and E. parviflora, respectively. These sequences shared 99% similarity with their counterparts in P. campestris (Supplemental Fig. 2).
Transfer of mitochondrial DNA to the plastid genome in bamboos
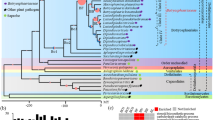
To place the mitochondrion-to-plastid DNA transfer in an evolutionary context, we built a phylogenetic tree for the representative species from the three bamboo tribes. Maximum likelihood analyses of the complete plastid genome sequences as a whole and partitioned by the LSC, SSC and IR regions yielded the same topology with nearly identical bootstrap values (Fig. 3 and Supplemental Fig. 3). Each of the three tribes were monophyletic and Olyreae was sister to Bambuseae, supported by 100% bootstrap value (Fig. 3), as in previous studies22,23. This intracellular DNA transfer was restricted to Pariana and its sister genus Eremitis, indicating that it likely occurred in their common ancestor.
The mVISTA29 alignment of the mitochondrial DNA sequence of F. rimosivaginus and the inserted sequences of 5 Parianinae species revealed a high global level of conservation with sequence identities only falling below 90% between positions approximately 600 and 750 (Fig. 4). Taken P. campestris as example, the inserted segment and the F. rimosivaginus mitochondrial genome differed by only 25 substitutions (transitions : transversions = 13 : 12) and 9 indels with a total length of 55 bp in an alignment of 2666 bp. Nevertheless, the sequences within 5000 bp upstream and downstream of the matching sequence in the F. rimosivaginus mitochondrial genome did not show any similarity to the plastid DNA sequences. The matching sequence is located in the intergenic region between the genes rps7 and atp6 in the mitochondrial genome of F. rimosivaginus27.
The mVISTA similarity plot of the insertion sequences in the plastid genomes of Pariana and Eremitis compared with the mitochondrial DNA sequence of F. rimosivaginus.
The blank areas downstream from ~2.1 kb in E. parviflora and P. parvispica indicate undetermined sequence of insertion in these bamboos.
To investigate the possible underlying mechanism of integration of mitochondrial DNA into the plastid genome, we performed sequence analysis of the insertion. Similar to the IR regions, the insertion had a GC content that was higher than the LSC/SSC regions of the plastid genome (44.7% compared to 36.2% in Pariana). Using ORF (open reading frame) finder in the NCBI (http://www.ncbi.nlm.nih.gov/gorf/gorf.html), we identified a total of 4 ORFs larger than 200 bp with the largest one encoding 116 amino acids. However, these ORFs had no significant similarity to any genes or proteins in the NCBI database. Finally, the insertion was found to be totally devoid of repeated sequences, as well as the 200-bp surrounding sequences in the plastid genome.
Discussion
The evolution of the reported plastid genomes is fairly conserved among the bamboos23,24,25. In our study, sequencing of six plastid genomes from neotropical woody bamboos and herbaceous bamboo subtribe Parianinae revealed identical genome organization and conserved evolution in bamboos again, but also limited variations. Firstly, the number of encoding genes is different in Chusquea sp. and Pariana. In Chusquea sp., the plastid genome contains two copies of trnT-GGU due to a 94-bp tandem repeat. This phenomenon has been repeatedly observed in plant plastid genomes30,31,32, suggesting that repeated sequences may play an important role in generating new genetic material in their evolution. In Pariana, the rps19 gene that is originally located in the IR regions in other bamboos was found to be relocated in the LSC region and thus lose one copy. The junctions between the IR and SSC regions in the plastid genomes are highly variable in grasses with the termini of two genes ndhF and ndhH repeatedly migrating into and out of the adjacent IR regions33. Here we provide evidence that the junctions between the IR and LSC regions can also shift greatly to a degree with the whole rps19 gene migrating into the LSC region. Secondly and more unexpectedly, there is a large deletion in the IR regions accompanied by insertion derived from the mitochondrial DNA sequence in Pariana. And more importantly the insertion was confirmed by analysis of sequencing depth as well as PCR amplification and Sanger sequencing. It is noteworthy that another independent study of plastid genomes of 17 bamboo species including P. radiciflora and Eremitis sp. also revealed this insertion of horizontally transferred mitochondrial DNA34.
In the GenBank, there are presently 446 complete plastid genomes sequenced from angiosperms (last accessed November 11th, 2014). However, only plastid genomes from two eudicot lineages have been demonstrated to harbor the horizontal transferred DNA from the mitochondrion14,15,16, even though other published genomes may also contain undiscovered mitochondrion derived DNA sequences9. The study of Wysocki et al.34 and ours focusing on bamboos represent the first example of mitochondrion-to-plastid DNA transfer in monocots (the third in angiosperms), expanding the phylogenetic representation of angiosperms for this phenomenon and confirming again that this direction of intracellular DNA transfer, though very rarely, does occur.
The insertion of mitochondrial DNA origin is located in the rps2-rpoC2 intergenic spacer in the plastid LSC region in the milkweeds16 while in the rps12-trnV-GAC intergenic spacer in the plastid IR regions in the carrots14,15. In the bamboos, it is also located in the intergenic spacer of trnI-CAU-trnL-CAA in the plastid IR regions. The integration of horizontal transferred DNA into the intergenic regions rather than the coding regions is expected, as the insertion would disrupt gene functions. The insertion occurred in the IR regions in the carrots and bamboos while in the LSC regions in the milkweeds. The high GC content of the transferred DNA (44.7% in Pariana and 44.0% in carrots versus 40.3% in milkweeds), which is more similar to the IR regions than to the single-copy regions in the plastid genome19, may give an explanation. In addition, the insertion is likely to be accompanied by deletion of plastid sequence at the same region as observed in the carrots (339 bp)14,15 and here (1379 bp). However, we could not exclude the possibility that the deletion and insertion took place independently in Pariana as in the plastid genome of O. glauca a deletion of 1542 bp also occurred at almost the same region while without the insertion (Fig. 1). The size of the insertion is ~1.5 kb in the carrots14,15 and varies between ~2.4–4.7 kb in the milkweeds16. The insertion has an identical size of 2.7 kb in the three sequenced bamboos of Pariana but we also revealed potential variations in another Pariana species and species from the sister genus Eremitis as in Wysocki et al.34. Given that the insertion is likely to be originated from a single event in the common ancestor14,15,16, these results suggest that the transferred sequence may be evolving rapidly. The insertions have been speculated to be transferred as a contiguous sequence from the mitochondria to the plastid genome in the two previously reported examples14,15,16, although they are fragmented into two or three pieces in the corresponding mitochondrial genomes. Our example in which the insertion derived from an intact segment in the mitochondrial genome corroborates this speculation.
While as organelle genomes coexisting in the cell, the ability to incorporating foreign DNA is definitely different between the plastid and mitochondrial genomes in angiosperms7,8,9. Presently, the underlying mechanism is unclear about the integration of foreign DNA into the plastid genome9. In the case of Pariana, a distinguished genus within Olyreae characterized by showy spike-like inflorescences implying possible insect pollination21,35, little is known about its genetics and genomics. Although the retrotransposition or homologous recombination has been proposed in previous studies15,16, we found no evidence in supporting either of these hypotheses in the bamboos. The plastid-to-mitochondrion transferred DNA, such as the multiple tRNA genes of plastid origin36, can play an important role in the mitochondrial genome. The role, if any, of the mitochondrion-to-plastid transferred DNA in the evolution of plastid genome otherwise needs more studies to demonstrate in the future.
Methods section
Sequencing and assembly of the bamboo plastid genomes
Fresh leaf material was collected from the six bamboos sampled in this study. Among them, P. campestris and P. radiciflora were collected in the field in French Guiana, Pariana sp. was cultivated and kindly provided by Prof. Nian-He Xia (South China Botanical Garden) and the other three bamboos were all cultivated at the Kunming Botanical Garden of the Kunming Institute of Botany. The voucher numbers for the sampled bamboos were DZL14021 for P. campestris, DZL14016 for P. radiciflora, ZMY026 for Pariana sp., ZXZ12008 for Chusquea sp., KMBG1217 for C. circinata and KMBG1223 for O. glauca.
Total genomic DNA was extracted using the CTAB method37 from 0.2–0.5 g fresh material. Total genomic DNA of four bamboos (three Pariana species and Chusquea sp.) was used for Illumina sequencing. For C. circinata and O. glauca, the plastid genome was amplified in overlapping fragments using long-range PCR primers from Yang et al.38 and PCR fragments were pooled together in roughly equal mass as previously described for subsequent sequencing25. The DNA samples were sheared to approximately 500 bp at BGI-Shenzhen, followed by library preparation. Paired-end sequencing of 90 bp was conducted on Illumina HiSeq 2000 at BGI-Shenzhen following the manufacturer’s protocol.
Illumina raw reads were first quality trimmed at BGI-Shenzhen. In which reads with adaptor contamination and more than 50% base pairs with a Phred quality score lower than or equal to 5 and having more than 1% ambiguous Ns were all discarded. Subsequently, a combination of de novo assembly using SOAPdenovo v.1.0539 and reference-assisted mapping was used to assemble the plastid genomes. The plastid genome of the closely related species B. emeiensis24 and Olyra latifolia23 was used as reference for assembly of neotropical woody bamboos and herbaceous bamboos, respectively. SOAPdenovo assemblies were generated with k-mer values of 31, 47, or 81 depended on different data sets. We discarded scaffolds and contigs <1000 bp and the rest were mapped to the reference genome using Blastn search in NCBI in the default settings. These mapped scaffolds and contigs that had >90% sequences similarity to the reference genome were collected for subsequent genome finishing. The small gaps and/or ambiguous nucleotides in the assembly were resolved by read mapping (see next section). However, the regions representing ~66–83 kb and ~22–33 kb in the reference genome of B. emeiensis24 could not be assembled for C. circinata and O. glauca, respectively, due to failure of long-range PCR amplification. Alternatively, we sequenced these regions using PCR primers in previous studies24,25 or designing new ones if necessary (Supplemental Table S2) by Sanger sequencing. The four junctions between the single-copy and IR regions of the plastid genome were ascertained by Sanger sequencing as previously described24. The gene content of the assembled plastid genomes was identified by DOGMA40 and manual adjustment was applied if necessary. The annotated plastid genome sequences were deposited in GenBank under accession numbers KP319241-KP319246.
Analysis of sequencing depth
We mapped the filtered Illumina reads using BWA v.0.7.741 in the default settings to the assembled plastid genomes with one copy of the IRs removed. We only considered these mapped reads that are in properly paired with SAMtools v.0.1.742 and used them to calculate the sequencing depth. For the insertion and surrounding regions, we determined the sequencing depth by averaging the depth at each base extracted using SAMtools. To determine the sequencing depth for the potential assembled mitochondrial DNA sequences, we also mapped the reads to the three mitochondrial genes (ccmFN, matR and rrn26) of F. rimosivaginus27 and calculated the depth in the same way.
Validation of the insertion by PCR amplification
The presence or absence of the insertion in the plastid genome was surveyed by PCR in the three Pariana species and in representatives of the major bamboo lineages21,22 (Fig. 1). Another Pariana species (P. parvispica) and E. parviflora that is sister to Pariana21,28 were also screened. We designed two primer pairs in sequences which are flanking the insertion endpoints and thus diagnostic for the presence of the insertion and also performed PCR amplification with two primer pairs specific to the plastid DNA sequence without the insertion25 (Supplemental Table S1). We confirmed both the endpoints of the insertion in the plastid genomes of Pariana sp., P. campestris and P. radiciflora. In addition, we also designed 4 PCR primer pairs (Supplemental Table S1) based on the assembled inserted sequence to obtain the whole region.
Sequence analysis
To determine the origin of the insertion, we conducted Blastn search against the NCBI nucleotide database online. Multiple sequences of the trnI-CAU-trnL-CAA intergenic spacer were aligned in MEGA543 and manual adjustments were made when necessary. The repetitive sequences were identified by the use of Blastn searches with a minimum identity of 90%.
Phylogenetic analyses
The complete plastid genome sequences from two species of subfamily Pooideae as outgroups and 16 bamboos (Fig. 3) were aligned in MAFFT v.7.21544 with default parameters. A few poorly aligned regions were manually adjusted in MEGA543. We performed unpartitioned and partitioned maximum likelihood analyses of the matrix using RAxML v.8.0.2045. In partitioned analysis, the dataset was divided into three partitions corresponding to the LSC, SSC and IR regions of the plastid genome. The maximum likelihood tree was constructed with the combined rapid bootstrap (500 replicates) and search for the best tree in a single run. The GTR + G model was used as suggested in the RAxML manual.
Additional Information
How to cite this article: Ma, P.-F. et al. Evidence for horizontal transfer of mitochondrial DNA to the plastid genome in a bamboo genus. Sci. Rep. 5, 11608; doi: 10.1038/srep11608 (2015).
References
Treangen, T. J. & Rocha, E. P. Horizontal transfer, not duplication, drives the expansion of protein families in prokaryotes. PLoS Genet. 7, e1001284 (2007).
Oren, Y. et al. Transfer of noncoding DNA drives regulatory rewiring in bacteria. Proc. Natl. Acad. Sci. USA 111, 16112–16117 (2014).
Richardson, A. O. & Palmer, J. D. Horizontal gene transfer in plants. J. Exp. Bot. 58, 1–9 (2007).
Bock, R. The give-and-take of DNA: horizontal gene transfer in plants. Trends Plant Sci. 15, 11–22 (2010).
Li, F. W. et al. Horizontal transfer of an adaptive chimeric photoreceptor from bryophytes to ferns. Proc. Natl. Acad. Sci. USA 111, 6672–6677 (2014).
Kleine, T., Maier, U. G. & Leister, D. DNA transfer from organelles to the nucleus: the idiosyncratic genetics of endosymbiosis. Annu. Rev. Plant Biol. 60, 115–138 (2009).
Smith, D. R. Extending the limited transfer window hypothesis to inter-organelle DNA migration. Genome Biol. Evol. 3, 743–748 (2011).
Smith, D. R., Crosby, K. & Lee, R. W. Correlation between nuclear plastid DNA abundance plastid number supports the limited transfer window hypothesis. Genome Biol. Evol. 3, 365–371 (2011).
Smith, D. R. Mitochondrion-to-plastid D. N. A. transfer: it happens. New Phytol. 202, 736–738 (2014).
Yoshida, T., Furihata, H. Y. & Kawabe, A. Patterns of genomic integration of nuclear chloroplast DNA fragments in plant species. DNA Res. 21, 127–140 (2014).
Alverson, A. J. et al. Insights into the evolution of mitochondrial genome size from complete sequences of Citrullus lanatus and Cucurbita pepo (Cucurbitaceae). Mol. Biol. Evol. 27, 1436–1448 (2009).
Goremykin, V. V., Salamini, F., Velasco, R. & Viola, R. Mitochondrial DNA of Vitis vinifera and the issue of rampant horizontal gene transfer. Mol. Biol. Evol. 26, 99–110 (2009).
Sloan, D. B. & Wu, Z. History of plastid DNA insertions reveals weak deletion and AT mutation biases in angiosperm mitochondrial genomes. Genome Biol. Evol. 6, 3210–3221 (2014).
Iorizzo, M. et al. De novo assembly of the carrot mitochondrial genome using next generation sequencing of whole genomic DNA provides first evidence of DNA transfer into an angiosperm plastid genome. BMC Plant Biol. 12, 61 (2012).
Iorizzo, M. et al. Against the traffic: the first evidence for mitochondrial DNA transfer into the plastid genome. Mob. Genet. Elements 2, 261–266 (2012).
Straub, S. C. K., Cronn, R. C., Edwards, C., Fishbein, M. & Liston, A. Horizontal transfer of DNA from the mitochondrial to the plastid genome and its subsequent evolution in milkweeds (Apocynaceae). Genome Biol. Evol. 5, 1872–1885 (2013).
Palmer, J. D. Comparative organization of chloroplast genomes. Annu. Rev. Genet. 19, 325–354 (1985).
Jansen, R. K. et al. Methods for obtaining and analyzing whole chloroplast genome sequences. Methods Enzymol. 395, 348–384 (2005).
Wicke, S., Schneeweiss, G. M., dePamphilis, C. W., Müller, K. F. & Quandt, D. The evolution of the plastid chromosome in land plants: gene content, gene order, gene function. Plant Mol. Biol. 76, 273–297 (2011).
Wolfe, K. H., Li, W. H. & Sharp, P. M. Rates of nucleotide substitution vary greatly among plant mitochondrial, chloroplast and nuclear DNAs. Proc. Natl. Acad. Sci. USA 84, 9054–9058 (1987).
Bamboo Phylogeny Group, An updated tribal and subtribal classification for the Bambusoideae (Poaceae). Proceedings of the 9th World Bamboo Congress [ Gielis, J. & Potters, G. (eds)] [3–27] (Antwerp, Belgium, 2012).
Kelchner, S. A. & Bamboo Phylogeny Group, Higher level phylogenetic relationships within the bamboos (Poaceae: Bambusoideae) based on five plastid markers. Mol. Phylogenet. Evol. 67, 404–413 (2013).
Burke, S. V., Clark, L. G., Triplett, J. K., Grennan, C. P. & Duvall, M. R. Biogeography and phylogenomics of new world Bambusoideae (Poaceae), revisited. Am. J. Bot. 101, 886–891 (2014).
Zhang, Y. J., Ma, P. F. & Li, D. Z. High-throughput sequencing of six bamboo chloroplast genomes: phylogenetic implications for temperate woody bamboos (Poaceae: Bambusoideae). PLoS ONE 6, e20596 (2011).
Ma, P. F., Zhang, Y. X., Zeng, C. X., Guo, Z. H. & Li, D. Z. Chloroplast phylogenomic analyses resolve deep-level relationships of an intractable bamboo tribe Arundinarieae (Poaceae). Syst. Biol. 63, 933–950 (2014).
Diekmann. K. et al. Complete chloroplast genome sequence of a major allogamous forage species, perennial ryegrass (Lolium perenne L.). DNA Res. 16, 165–176 (2009).
Ma, P. F., Guo, Z. H. & Li, D. Z. Rapid sequencing of the bamboo mitochondrial genome using Illumina technology and parallel episodic evolution of organelle genomes in grasses. PLoS ONE 7, e30297 (2012).
Oliveira, R. P. et al. A molecular phylogeny of Raddia and its allies within the tribe Olyreae (Poaceae, Bambusoideae) based on noncoding plastid and nuclear spacers. Mol. Phylogenet. Evol. 78, 105–117 (2014).
Mayor, C. et al. VISTA: visualizing global DNA sequence alignments of arbitrary length. Bioinformatics 16, 1046–1047 (2000).
Lee, H. L., Jansen, R. K., Chumley, T. W. & Kim, K. J. Gene relocations within chloroplast genomes of Jasminum and Menodora (Oleaceae) are due to multiple, overlapping inversions. Mol. Biol. Evol. 24, 1161–1180 (2007).
Lin, C. P., Wu, C. S., Huang, Y. Y. & Chaw, S. M. The complete chloroplast genome of Ginkgo biloba reveals the mechanism of inverted repeat contraction. Genome Biol. Evol. 4, 374–381 (2012).
Do, H. D. K., Kim, J. S. & Kim, J. H. A. trnI_CAU triplication event in the complete chloroplast genome of Paris verticillata M.Bieb. (Melanthiaceae, Liliales). Genome Biol. Evol. 6, 1699–1706 (2014).
Davis, J. I. & Soreng, R. J. Migration of endpoints of two genes relative to boundaries between regions of the plastid genome in the grass family (Poaceae). Am. J. Bot. 97, 874–892 (2010).
Wysocki, W. P., Clark, L. G., Attigala, L., Ruiz-Sanchez, E. & Duvall, M. R. Evolution of the bamboos (Bambusoideae; Poaceae): a full plastome phylogenomic analysis. BMC Evol. Biol. 15, 50 (2015).
Soderstrom, T. R. & Calderón, C. E. Insect pollination in tropical rain forest grasses. Biotropica 3, 1–16 (1971).
Richardson, A. O., Rice, D. W., Young, G. J., Alverson, A. J. & Palmer, J. D. The “fossilized” mitochondrial genome of Liriodendron tulipifera: ancestral gene content and order, ancestral editing site and extraordinarily low mutation rate. BMC Biol. 11, 29 (2013).
Doyle, J. J. & Doyle, J. L. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem. Bull. 19, 11–15 (1987).
Yang, J. B., Li, D. Z. & Li, H. T. Highly effective sequencing whole chloroplast genomes of angiosperms by nine novel universal primer pairs. Mol. Ecol. Resour. 14, 1024–1031 (2014).
Li, R. et al. De novo assembly of human genomes with massively parallel short read sequencing. Genome Res. 20, 265–272 (2010).
Wyman, S. K., Jansen, R. K. & Boore, J. L. Automatic annotation of organellar genomes with DOGMA. Bioinformatics 20, 3252–3255 (2004).
Li, H. & Durbin, R. Fast and accurate long-read alignment with Burrows-Wheeler transform. Bioinformatics 26, 589–595 (2010).
Li, H. et al. The sequence alignment/map (SAM) format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Tamura, K. et al. MEGA5: molecular evolutionary analysis using maximum likelihood, evolutionary distance and maximum parsimony methods. Mol. Biol. Evol. 28, 2731–2739 (2011).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Acknowledgements
This study was supported by the National Natural Science Foundation of China (grants U1136603, 31170204 and 31300184). We thank Meng-Yuan Zhou and Xian-Zhi Zhang for laboratory support and Lu Lu for help with obtaining bamboo samples.
Author information
Authors and Affiliations
Contributions
P.F.M. and D.Z.L. conceived and designed the research. Y.X.Z., Z.H.G. and D.Z.L. collected the plant material. P.F.M. performed the experiments and analyzed the data. All the authors discussed the results. P.F.M. and D.Z.L. wrote the paper and all the authors read and approved the final manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Ma, PF., Zhang, YX., Guo, ZH. et al. Evidence for horizontal transfer of mitochondrial DNA to the plastid genome in a bamboo genus. Sci Rep 5, 11608 (2015). https://doi.org/10.1038/srep11608
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep11608
- Springer Nature Limited
This article is cited by
-
The mitochondrial genome sequence of Abies alba Mill. reveals a high structural and combinatorial variation
BMC Genomics (2022)
-
Comparative plastomes and phylogenetic analysis of seven Korean endemic Saussurea (Asteraceae)
BMC Plant Biology (2022)
-
Efficiency verification of CRISPR–Cas9-mediated mutagenesis of target gene sgRNA using soybean protoplasts
Plant Biotechnology Reports (2022)
-
Comparative chloroplast genomes: insights into the evolution of the chloroplast genome of Camellia sinensis and the phylogeny of Camellia
BMC Genomics (2021)
-
Comparative analysis of nuclear, chloroplast, and mitochondrial genomes of watermelon and melon provides evidence of gene transfer
Scientific Reports (2021)