Abstract
The target of rapamycin (TOR) positively controls cell growth in response to nutrients such as amino acids. However, research on the specific nutrients sensed by TOR is limited. Glutamine (Gln), a particularly important amino acid involved in metabolism in organisms, is synthesised and catalysed exclusively by glutamine synthetase (GS) and our previous studies have shown that Gln may regulate fecundity in vivo levels of the brown planthopper (BPH) Nilaparvata lugens. Until now, it has remained unclear whether Gln activates or inhibits the TOR signalling pathway. Here, we performed the combined analyses of iTRAQ (isobaric tags for relative and absolute quantification) and DGE (tag-based digital gene expression) data in N.lugens at the protein and transcript levels after GS RNAi and we found that 52 pathways overlap, including the TOR pathway. We further experimentally demonstrate that Gln activates the TOR pathway by promoting the serine/threonine protein kinase AKT and inhibiting the 5′AMP-activated protein kinase AMPK phosphorylation activity in the pest. Furthermore, TOR regulates the fecundity of N. lugens probably by mediating vitellogenin (Vg) expression. This work is the first report that Gln activates the TOR pathway in vivo.
Similar content being viewed by others
Introduction
In the early 1990s, the target of rapamycin (TOR) was initially discovered in Saccharomyces cerevisiae1. Since then, TOR has been identified in many other organisms ranging from yeast to mammals2,3. TOR is an atypical serine/threonine kinase of the phosphatidylinositol kinase-related kinase (PIKK) family and is a major regulator of growth in eukaryotes. TOR is found in two conserved complexes termed the TOR complex 1 (TORC1) and the TOR complex 2 (TORC2)4. The TOR pathway is activated by many extra- and intracellular signals such as nutrients (amino acids) and growth factors (insulin). Conversely, TOR is inhibited by numerous stress conditions such as cellular energy (ATP) depletion and hypoxia stress5,6,7.
TOR positively controls cell growth in response to nutrients. However, research on the specific nutrients sensed by TOR is limited8. Arginine (Arg) and Leucine (Leu) have an essential role in the phosphorylation of ribosomal S6 kinase (S6 K) and the eukaryotic initiation factor 4E binding protein 1 (4EBP1) mediated by TORC19,10. Glutamine (Gln) is the most abundant nonessential amino acid and its synthesis is catalysed exclusively by glutamine synthetase (GS). Gln regulates the proliferation of diverse cell types, which is a precursor for α-ketoglutarate of the tricarboxylic acid (TCA) cycle and provides nitrogen for protein and nucleotide synthesis11. However, the role of Gln in TOR activation remains poorly defined12. In S. cerevisiae, Gln controls TOR readouts (8). Similarly in mammalian cells, Gln activates TORC1 signalling13, Leu and Gln activate glutaminolysis and mTORC112 and L-glutamine flux regulates mTOR, translation and autophagy to coordinate cell growth and proliferation14. In contrast, the transcription factor FOXO3-induced GS activity inhibits mTOR activation by blocking the translocation of mTOR to the lysosomal surface15. Thus, it remains unclear whether Gln activates or inhibits the TOR signalling pathway.
Here, we demonstrate that Gln activates the TOR pathway in the brown planthopper (BPH) Nilaparvata lugens (Stål), a serious pest of rice crops with robust fecundity16. We also show that Gln promotes the serine/threonine protein kinase AKT and inhibits the 5′AMP-activated protein kinase AMPK phosphorylation activity. Furthermore, TOR regulates the fecundity of N. lugens probably by mediating vitellogenin (Vg) expression. This study is the first report to show that Gln activates the TOR pathway in vivo.
Results
Global changes at the protein and transcript levels after GS RNAi
To ensure that GS knockdown was effective in the 2 samples, the mRNA expression of the GS gene was detected by qRT-PCR (Supplementary Figure 1). Afterwardsthe iTRAQ Mass-Tagging mass spectrometric results from the two repetitions of the GS-RNAi and GFP-RNAi experiments were analysed using the ProteinPilotTM software 2.0.1 (Applied Biosystems, USA). N. lugens lacks a reference genome; therefore, the annotation of its proteins primarily depends on its EST databases and other insect databases17. This analysis produced a quantitative proteome of 958 proteins for the GS knockdown experiment (Table 1). The GS knockdown experiment resulted in 124 proteins that were statistically significant responders (p < 0.01), which is approximately 12.94% of the identified quantitative proteome. Among these proteins, 49 and 75 proteins were up- and down-regulated, respectively (Fig. 1A, Supplementary Table 1). Regarding the down-regulated proteins, we found the RNAi target protein GS (NLU012724.1) and fecundity-related protein vitellogenin (Vg) (NLU019204.1) (Fig. 1B), indicating that the RNAi was effective. Several TOR pathway proteins of interest, including two key upstream activators of Rheb (NLU021861.1) and 3′-phosphoinositide-dependent kinase (PDK1) (NLU017432.3), were also down-regulated.
Combined analyses of the iTRAQ and DGE data after GS RNAi.
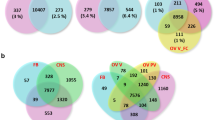
First day brachypterous female adults were injected with dsGS or dsGFP. Samples were used for DGE and iTRAQ 48 h and 72 h post-injection, respectively. (A) No. of differentially expressed proteins identified by iTRAQ, the conditions for protein spots were ≥1.2-fold or ≤0.8 (p < 0.05) for up-regulated or down-regulated proteins in dsGS compared to dsGFP. (B) The representative MS/MS spectra of down-regulated proteins, GS (NLU012724.1) and Vg (NLU019204.1). (C) No. of differentially expressed unigenes identified by DGE, the conditions for unigenes was FDR ≤0.001 and |log2Ratio| ≥ 1. (D) GO analysis of differentially expressed genes. (E) Venn diagrams of differentially expressed genes/proteins from iTRAQ and DGE analyses. (F) Cluster of pathways for iTRAQ and DGE. (G) Venn diagrams of regulated pathways from iTRAQ and DGE analyses.
The differentially expressed genes following RNAi treatment were analysed using the Digital Gene Expression (DGE) method. The results revealed 293 genes with significantly differential expression between the GS-RNAi and GFP-RNAi libraries, including 151 up- and 142 down-regulated genes (Fig. 1C). In the down-regulated genes, we also found GS and Vg in addition to several TOR pathway genes such as TOR (NLU007095.1), Rheb and S6 K (NLU007225.2). According to the GO classification, most of the differentially expressed genes correlated to biological processes, including the reproductive process and various developmental processes (Fig. 1D).
With the iTRAQ and DGE high-quality data sets, we performed an integrated analysis of the proteome and transcriptome. The overall analysis of the robustly regulated genes at the protein and mRNA levels found 5 genes that overlap in the up-regulated group and 7 genes in the down-regulated group (Fig. 1E, Supplementary Table 2). Pathway enrichment analysis for the proteins and transcripts was also performed using the KEGG database. In addition to the “unable to classify” group, the protein metabolism, carbohydrate metabolism and amino acid metabolism groups were ranked in the top 3 most enriched pathway ontology groups (Fig. 1F). Fifty-two overlapping pathways were found between iTRAQ and DGE (Fig. 1G), including the TOR signalling pathway.
Glutamine activates the TOR pathway in vivo
To validate the above result suggesting that GS knockdown influences the TOR signalling pathway, several experiments were conducted. After studying the feature of GS (Supplementary Figure 2), we depleted GS for 48 h and detected the transcript change of the TOR pathway-associated genes. The RNAi was effective because the GS mRNA and protein levels as well as enzymatic activity decreased (Fig. 2A,B). Three TOR pathway-associated genes, Rheb, TOR and S6 K, had significantly lower expression levels after GS RNAi compared to the control (dsGFP) (Fig. 2A), indicating that a positive relationship exists between GS and TOR.
Effect of GS knockdown on gene expression and enzyme activity.
First day brachypterous female adults were injected with 50 nL dsGS (5 ng/nL), dsGFP (5 ng/nL), ddH2O or MSX (10 mM), respectively. (A) The transcript levels of genes after injection by qRT-PCR. Data represent mean values±S.E.M (n = 3), *p < 0.05, **p < 0.01. (B) The relative activity of GS at 48 h post-injection. Data represent mean values±S.E.M (n = 3) and those in the columns followed by the different letters mean significant difference (p = 0.05, Duncan’s multiple range test). Inset shows Western blotting analysis of GS protein levels injected with either dsGFP or dsGS; the β-actin gene was used as an internal control. (C) The transcript levels of Vg at 48 h post-injection by qRT-PCR. Data represent mean values±S.E.M (n = 3) and those in the columns followed by the different letters mean significant difference (p = 0.05, Duncan’s multiple range test). Inset shows Western blotting analysis of Vg protein levels injected with either dsGFP or dsGS; the β-actin gene was used as an internal control.
Secondly, we detected the phosphorylation of S6 K, a molecular marker of TOR pathway activation18. After injecting 250 ng dsTOR or 10 mM rapamycin into the BPH adults, the phosphorylation of S6 K decreased (Supplementary Figure 3). Similar results were observed with the injection of dsGS or L-methionine sulfoximine (MSX), a GS specific inhibitor8 (Fig. 3A, lane 2 and 5). When Gln (the GS specific product) was added into dsGS or MSX, S6 K phosphorylation increased (Fig. 3A, lane 3 and 6). When feeding the BPH nymphs with the artificial diet (with Gln)19, S6 K phosphorylation increased compared to the diet without Gln (Fig. 3B, lane 2 and 4). When dsGS was added into the diet, S6 K phosphorylation also decreased (Fig. 3B, lane 1). These results show that Gln activates the TOR pathway via the phosphorylation of S6 K in vivo. In addition, the Vg mRNA and protein levels were down-regulated after the injection of dsGS and MSX (Fig. 2C), indicating that GS regulates BPH fecundity20.
Western blots showing S6 K phosphorylation levels and GS and Vg protein abundance under various nutritional conditions at 48 h post-injection.
(A) First day brachypterous female adults were injected with 50 nL dsGFP (5 ng/nL), dsGS, dsTOR, MSX (10 mM), Gln (20 mM) and ddH2O. Shown are representative blots (n = 3). (B) The 3rd to the 5th instar nymphs were fed an artificial diet (w/w Gln) together with dsGS or dsGFP (0.5 μg/μl). Shown are representative blots (n = 3).
TOR regulates fecundity in BPH
After the injection of dsTOR into the BPH adults, the TOR mRNA level significantly decreased by 59.66%, 68.66% and 66.68% at 24 h, 48 h and 72 h, respectively, compared to the animals injected with dsGFP (Fig. 4A). The expression level of the Vg gene was also significantly down-regulated by 47.66%, 52.74% and 59.68% after dsTOR injection. When the BPH adults were sampled 48 h after injectedion of rapamycin or dsTOR, significant reductions in the Vg protein level was observed (Fig. 4B, Supplementary Figure 3). As a result, emerged brachypterous BPH adults had fewer offspring (53.22 per pair, a 71.21% decrease). Most dsTOR-treated adults had less than 100 offspring; however, 200-300 nymphs were observed for the majority of the dsGFP-treated pairs (Fig. 4C). These results suggest that TOR regulates the fecundity of N. lugens probably by mediating Vg expression.
The effect of TOR on fecundity in N. lugens.
First day brachypterous female adults were injected with dsTOR or dsGFP. (A) The transcript levels of TOR after injection with dsRNA. (B) The transcript levels of Vg after RNAi. Data represent the mean values ± S.E.M of three replicates, ‘*’ indicates statistically significant difference (t-test, p < 0.05). The inset shows Western blotting analysis of Vg protein levels and the β-actin gene was used as an internal control. (C) The frequency distribution of N. lugens offspring.
Gln promotes AKT and inhibits AMPK
To further study the molecular mechanism of the activation of the TOR pathway via Gln, we chose two key upstream genes of the TOR pathway, AKT and AMPK. After silencing GS through RNAi, both the enzyme activity and phosphorylation of AKT decreased. When Gln was added, the enzyme activity and phosphorylation of AKT were up-regulated (Fig. 5A,B), suggesting that Gln promoted AKT. However, the opposite trend was observed when AMPK was measured (Fig. 5C,D), indicating that Gln inhibited AMPK.
Proposed mechanism of glutamine-mediated activation of the TOR signalling pathway
. (A) The relative activity of AKT at 48 h post-injection. Data represent the mean values±S.E.M (n = 3) and the values in the columns followed by different letters note a significant difference (p = 0.05, Duncan’s multiple range test). (B) Western blots showing S6 K phosphorylation levels and GS protein abundance. Shown are representative blots (n = 3). (C) The relative activity of AMPK at 48 h post-injection. Data represent the mean values±S.E.M (n = 3) and the values in the columns followed by the different letters note a significant difference (p = 0.05, Duncan’s multiple range test). (D) Western blots showing S6K phosphorylation levels and GS protein abundance. Shown are representative blots (n = 4). (E) The mRNA expression levels at 48 h after injection of dsAKT. Data represent the mean values±S.E.M (n = 3), *p < 0.05, **p < 0.01. (F) The relative activity of GS at 48 h after injection of dsAKT. Data represent the mean values±S.E.M (n = 3), *p < 0.05. (G) Proposed model of the activation of the TOR signalling pathway by Gln by promoting AKT and inhibiting AMPK in N. lugens.
When AKT was knocked down using RNAi, the TOR mRNA level significantly decreased (Fig. 5E). More importantly, GS activity decreased by 60.65% (Fig. 5F); however, its mRNA level did not significantly change (Fig. 5E), implying that AKT has a positive feedback mechanism that acts on GS.
In conclusion, GS regulates the content of Gln, which activates the TOR signalling pathway by promoting AKT and inhibiting AMPK. Furthermore, TOR regulates the fecundity of N. lugens probably by mediating Vg expression (Fig. 5G).
Discussion
Targeting a gene via RNAi is often used to study its function.However, the reports on the global changes of mRNA and protein profile that occur after RNAi targeting of a specific gene have been limited21,22. For example, after knock down of ISWI in Drosophila SL2 cells, Bonaldi et al. studied the global changes of transcriptome and proteome using microarray and SILAC-MS technologies and found that there is a very little correlation between mRNA and protein level23. In cultured male-like tissue cells of D. melanogaster, the expression level of many genes on the X chromosome decreased after RNAi targeting MSL complex; however, the gene expression on autosomes remained largely unchanged24. In our study, the combinative analyses of iTRAQ proteomic and DGE transcriptomic data after GS RNAi revealed 52 overlapping pathways, including the TOR pathway (Fig. 1). We identified 12 genes in which 5 were up-regulated and 7 were down-regulated at both mRNA and protein level (Supplementary Table 2). Similar to the previous report23, the fold of change in mRNA and protein levels was different.
Recently, accumulated evidences showed that the TOR pathway regulates protein synthesis and cell proliferation through integrating multiple upstream signals25. In N. lugens, the knockdown of TOR leads to decreased fecundity that is correlated with lower Vg expression (Fig. 4), implying that TOR regulates its fecundity probably by mediating Vg expression. However, TOR could also affect fecundity by other mechanisms. While in A. aegypti, TOR pathway is essential for the activation of Vg expression26,27, TOR-dependent activation of S6 kinase is a crucial step in the transduction of nutritional information during egg development of mosquitoes28. Moreover, in Drosophila, TOR regulates ovary size29 and is a determinant of follicle number by promoting cell survival30. In Apis mellifera, TOR governs diphenic development31. Both phenotypic and mechanistic studies are desired in order to explain roles of TOR in regulating fecundity of N. lugens.
TOR is a well known nutrient sensor in many organisms8,32. In multiple types of cancer cells, a positive correlation between Gln and TOR was observed12,13,14. Consistent with this, we found that Gln activates the TOR pathway via phosphorylating S6 K (Fig. 3). However, a different result was reported when Ba/F3 cell line was used15. Possible explanation is that the effect of Gln on TOR signalling may be cell type dependent15. Another possibility concerns a threshold effect of Gln concentration in cells. Since all experiments of this study were conducted in vivo, the findings may be more physiologically relevant.
Interestingly, we also found that Gln apparently promotes AKT activity, while inhibiting AMPK activity in N. lugens (Fig. 5G). This finding provides evidence that Gln regulates TOR not by a single pathway but by a network composed of growth (insulin) and energy pathways. However, it is still not clear how Gln stimulates AKT phosphorylation/activation. In mammalian systems, Gln stimulates synthesis and secretion of insulin-like growth factor 2 (IGF2) as well as IGF2-dependent AKT phosphorylation in beta cell33. In addition, Gln, Leu and Pro increase AKT phosphorylation in HepG2 liver cells34, presumably, through the inhibition of NF-κB. Also, how Gln inhibits AMPK is an open question. It is thus interesting and important to understand the mechanism underlying Gln-mediated AKT activation and AMPK inhibition.
Methods
Insect Rearing
A N. lugens laboratory strain that was originally obtained from Guangdong Academy of Agricultural Sciences (GDAAS; Guangdong, China) in September 2007 was used and this strain was reared in a continuous laboratory culture on BPH-susceptible rice plants (Huang Hua Zhan, bought from GDAAS). The insects were maintained in the laboratory at 26 ± 2 °C with 80 ± 10% humidity and a light-dark cycle of L16:D8 h20.
iTRAQ Analysis
To enrich the total proteins, two repetitions of the samples 72 h post-injection of dsGS were separately ground into powder in liquid nitrogen, homogenised in RIPA lysis buffer (Beyotime, China) and centrifuged at 12,000 rpm at 4 °C. The supernatant was precipitated in 10% trichloroacetic acid (TCA)/acetone, centrifuged at 12,000 rpm at 4 °C and redissolved in dissolution buffer (0.5 M triethylammonium bicarbonate, 1% sodium deoxycholate) with 5 min heating at 90 °C and probe sonication. The protein concentration was measured with a Bradford assay protein assay kit (Bio-Rad, USA). The 4-plex iTRAQ Labeling, Strong cation exchange (SCX) and RP HPLC-MS/MS were performed by Fitgene Biological Technology Co. Ltd (FITGENE, Guangzhou, China). The peptides were identified by the Peptide Prophet algorithm35.
DGE Analysis
The total RNA was extracted using the E.Z.N.A.® Total RNA Kit II (Omega) according to the manufacturer’s protocol. To obtain ideal gene expression information after RNAi, 48 hours post-injection of dsGS or dsGFP, the samples were used for transcriptome analysis. The genes differentially expressed between the two samples were identified using an algorithm as previously described36. The cDNA library was constructed and sequenced by the Beijing Genomics Institute (BGI, Shenzhen, China) on the Illumina sequencing platform (HiSeq™ 2000).
Western Blotting
Western blotting analyses were performed using standard techniques. The proteins were separated on a 12% SDS-PAGE gel and transferred to PVDF membranes (0.4 μm, Millipore) and the membranes were immunoblotted with AKT, phospho-S473 AKT, phospho-pT183+T172 AMPK (1:2000, Abcam, UK), phospho-T389 S6 K (1:1000, CST, USA), GS, Vg and S6 K (1:5000) serum (prepared by our lab). IgG goat anti-mouse and goat anti-rabbit antibodies conjugated with HRP were used for secondary antibodies (1:5000, Abcam, UK) and the membranes were visualised by ECL (enhanced chemiluminescence).
Quantitative Real-Time PCR Analysis
The primers used for real-time PCR are listed in Table 1. The synthesised first-strand cDNA was amplified by PCR in 10 μL reaction mixtures using a Light Cycler 480 system (Roche, USA) and the β-actin gene was used as an internal standard37. After the amplifications, a melting curve analysis was performed in triplicate and the results were averaged. The quantitative variation was calculated using three independent biological samples by a relative quantitative method (2−ΔΔCT)38.
RNA Interference and Sampling
The dsRNA of a target gene was produced using the T7 RiboMAX™ Express RNAi System (Promega, USA). After synthesis, the dsGS (439 bp), dsTOR (457 bp), dsAKT (447 bp) and dsGFP (688 bp) were quantified by ultramicro-spectrophotometers (NanoDrop 2000, Thermo) and were maintained at −80 °C until use. The sequence was verified by sequencing (Invitrogen Company, Shanghai, China). To deliver dsRNA into the body of BPHs, the dsRNA was added to the artificial diet39 or directly injected40. Before injection, the dsRNA and phenol red solution were mixed for observation. The purified dsRNA or ddH2O were slowly injected using 3.5 Drummond needles and the NARISHIGE IM-31 (Nikon, JAPAN).
AKT and AMPK Activity Quantitative Test
AKT and AMPK quantitative enzymatic activity test kits (Genmed Scientifics, Inc.) were used to conduct to detect the activity. According to the manufacturer’s protocol steps, the absorbance was read at 340 nm. Finally, the AKT and AMPK active quantitative indexes were calculated using separate formulas41.
Glutamine Synthetase Activity Assay
Glutamine synthetase (GS) was assayed using the method with a modification42. The assay medium included 230 μl of reaction mixture (100 mM Tris-HCl pH 7.4, 80 mM MgCl2, 20 mM cysteine, 20 mM sodium L-glutamate, 2 mM EGTA and 80 mM hydroxylamine hydrochloride). The reaction was started with the addition of 100 μl of enzyme extract and 100 μl 40 mM ATP, incubated for 30 min at 37 °C, stopped by the addition of 143 μl of a chromogenic agent (0.37 M FeCl3, 0.2 M TCA and 0.6 M HCl) and placed at room temperature after a moment. After centrifugation, the absorbance of the supernatant was read at 540 nm against a reagent blank.
Statistical analysis
The results are expressed as the means±S.E.M. SPSS 13.0 software was used to perform t-tests to identify significant differences at a 95% or 99% confidence level.
Additional Information
How to cite this article: Zhai, Y. et al. Activation of the TOR Signalling Pathway by Glutamine Regulates Insect Fecundity. Sci. Rep. 5, 10694; doi: 10.1038/srep10694 (2015).
References
Heitman, J., Movva, N. R. & Hall, M. N. Targets for cell cycle arrest by the immunesuppressant rapamycin in yeast. Science 253, 905–909 (1991).
Soulard, A., Cohen, A. & Hall, M. N. TOR signaling in invertebrates. Curr. Opin. Cell Biol. 21, 825–836 (2009).
Zoncu, R., Efeyan, A., & Sabatini, D. M. mTOR: from growth signal integration to cancer, diabetes and ageing. Nat. Rev. Mol. Cell Biol. 12, 21–35 (2011).
Wullschleger, S., Loewith, R. & Hall, M. N. TOR signaling in growth and metabolism. Cell 124, 471–484 (2006).
Hara, K. et al. Amino acid sufficiency and mTOR regulate p70 S6 kinase and eIF-4E BP1 through a common effector mechanism. J. Biol. Chem. 273, 14484–14494 (1998).
Dennis, P. B. et al. Mammalian TOR: a homeostatic ATP sensor. Science 294, 1102–1105 (2001).
Martin, D. E. & Hall, M. N. The expanding TOR signaling network. Curr. Opin. Cell Biol. 17, 158–166 (2005).
Crespo, J. L. et al. The TOR-controlled transcription activators GLN3, RTG1 and RTG3 are regulated in response to intracellular levels of glutamine. Proc. Natl. Acad. Sci. 99, 6784–6789 (2002).
Ban, H. et al. Arginine and Leucine regulate p70 S6 kinase and 4E-BP1 in intestinal epithelial cells. Int. J. Mol. Med. 13, 537–543 (2004).
Norton, L.E. et al. The leucine content of a complete meal directs peak activation but not duration of skeletal muscle protein synthesis and mammalian target of rapamycin signaling in rats. J. Nutr. 139, 1103–1109 (2009).
Reitzer, L. J., Wice, B. M. & Kennell, D. Evidence that glutamine, not sugar, is the major energy source for cultured HeLa cells. J. Biol. Chem. 254, 2669–2676 (1979).
Durán, R. V. et al. Glutaminolysis activates Rag-mTORC1 signaling. Mol. Cell 47, 349–358 (2012).
Wise, D. R. & Thompson, C. B. Glutamine addiction: a new therapeutic target in cancer. Trends Biochem. Sci. 35, 427–433 (2010).
Nicklin, P. et al. Bidirectional transport of amino acids regulates mTOR and autophagy. Cell 136, 521–534 (2009).
van der Vos, K. E. et al. Modulation of glutamine metabolism by the PI(3)K-PKB-FOXO network regulates autophagy. Nat. Cell Biol. 14, 829–837 (2012).
Heinrichs, E. A. Impact of Insecticides on the Resistance and Resurgence of Rice Planthoppers. In Planthoppers: Their Ecology and Management; Denno, R. F., Perfect, T. J., Eds.; Chapman and Hall Press: New York, pp 571–614 (1994).
Noda, H. et al. Annotated ESTs from various tissues of the brown planthopper Nilaparvata lugens: a genomic resource for studying agricultural pests. BMC Genomics 9, 117 (2008).
Kimura, N. et al. A possible linkage between AMP-activated protein kinase (AMPK) and mammalian target of rapamycin (mTOR) signalling pathway. Genes Cells 8, 65–79 (2003).
Fu, Q. et al. A chemically defined diet enables continuous rearing of the brown planthopper, Nilaparvata lugens (Stål) (Homoptera: Delphacidae). Appl. Entomol. Zool. 36, 111–116 (2001).
Zhai, Y. F. et al. Proteomic and transcriptomic analyses of fecundity in the brown planthopper Nilaparvata lugens (Stål). J. Proteome Res. 12, 5199–5212 (2013).
Wang, S. W. et al. Global role for polyadenylation-assisted nuclear RNA degradation in posttranscriptional gene silencing. Mol. Cell Biol. 28, 656–665 (2008).
Keller, C. et al. Proteomic and functional analysis of the noncanonical poly(A) polymerase Cid14. RNA 16, 1124–1129 (2010).
Bonaldi, T. et al. Combined use of RNAi and quantitative proteomics to study gene function in Drosophila. Mol. Cell 31, 762–772 (2008).
Hamada, F. N. et al. Global regulation of X chromosomal genes by the MSL complex in Drosophila melanogaster. Genes Dev. 19, 2289–2294 (2005).
Yang, Q. & Guan, K. L. Expanding mTOR signaling. Cell Res. 17, 666–681 (2007).
Hansen, I. A. et al. Target of rapamycin-mediated amino acid signaling in mosquito anautogeny. Proc. Natl. Acad. Sci.USA. 101, 10626–10631(2004).
Attardo, G. M. et al. Nutritional regulation of vitellogenesis in mosquitoes: implications for anautogeny. Insect Biochem Mol Biol. 35, 661–675 (2005).
Hansen, I. A. et al. Target of rapamycin-dependent activation of S6 kinase is a central step in the transduction of nutritional signals during egg development in a mosquito. J. Biol. Chem. 280, 20565–20572 (2005).
Zhang, Y. et al. Drosophila target of rapamycin kinase functions as a multimer. Genetics 172, 355–362 (2006).
LaFever,L. et al. Specific roles of Target of rapamycin in the control of stem cells and their progeny in the Drosophila ovary. Development 137, 2117–2126 (2010).
Patel, A. et al. The Making of a Queen: TOR Pathway Is a Key Player in Diphenic Caste Development. PLoS One 2, e509 (2007).
Wang, X. & Proud, C. G. Nutrient control of TORC1, a cell-cycle regulator. Trends Cell Biol. 19, 260–267 (2009).
Modi, H. et al. Glutamine Stimulates Biosynthesis and Secretion of Insulin-like Growth Factor 2 (IGF2), an Autocrine Regulator of Beta Cell Mass and Function. J. Biol. Chem. 289, 31972–31982 (2014).
van Meijl, L. E. et al. Amino acids stimulate Akt phosphorylation and reduce IL-8 production and NF-κB activity in HepG2 liver cells. Mol Nutr Food Res. 54, 1568–1573 (2010).
Keller, A. et al. Empirical statistical model to estimate the accuracy of peptide identifications made by MS/MS and database search. Anal. Chem. 74, 5383–5392 (2002).
Audic, S. & Claverie, J. M. The significance of digital gene expression profiles. Genome Res. 7, 986–995 (1997).
Chen, J. et al. Feeding-based RNA interference of a trehalose phosphate synthase gene in the brown planthopper, Nilaparvata lugens. Insect Mol. Biol. 19, 777–786 (2010).
Livak, K.J. & Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods 25, 402–408 (2001).
Dong, X. L. et al. Fork head transcription factor is required for ovarian mature in the brown planthopper, Nilaparvata lugens (Stål). BMC Mol. Biol. 12, 53 (2011).
Liu, S. H. et al. Gene knockdown by intro-thoracic injection of double-stranded RNA in the brown planthopper, Nilaparvata lugens. Insect Biochem. Mol. Biol. 40, 666−671 (2010).
Zhen, Z. et al. Anti-diabetic effects of a Coptis chinensis containing new traditional Chinese medicine formula in type 2 diabetic rats. Am. J. Chin. Med. 39, 53−63 (2011).
Meister, A. Glutamine synthetase from mammalian tissues. Methods Enzymol. 113, 185–199 (1985).
Acknowledgements
We are grateful to Fitgene Biological Technology Co. Ltd (FITGENE, Guangzhou, China) for iTRAQ related experiments. This work was supported by the National Natural Science Foundation of China (30930061) and the National Basic Research Program of China (973 Program) (No. 2010CB126200). We declare that we have no financial and personal relationships with other people or organizations that can inappropriately influence our work.
Author information
Authors and Affiliations
Contributions
W.Q.Z. designed the conception and reviewed the manuscript. Y.F.Z. performed the research and wrote the manuscript. Z.X.S. and J.Q.Z. performed development of methodology. K.K. acquired data (provided animals, provided facilities, etc.). J.C. performed analysis and interpretation of data (e.g., statistical analysis, biostatistics, computational analysis).
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0
About this article
Cite this article
Zhai, Y., Sun, Z., Zhang, J. et al. Activation of the TOR Signalling Pathway by Glutamine Regulates Insect Fecundity. Sci Rep 5, 10694 (2015). https://doi.org/10.1038/srep10694
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep10694
- Springer Nature Limited
This article is cited by
-
Capsaicin decreases fecundity in the Asian malaria vector Anopheles stephensi by inhibiting the target of rapamycin signaling pathway
Parasites & Vectors (2022)
-
Ammonia promotes the proliferation of bone marrow-derived mesenchymal stem cells by regulating the Akt/mTOR/S6k pathway
Bone Research (2022)
-
WNT/β-catenin Pathway: a Possible Link Between Hypertension and Alzheimer’s Disease
Current Hypertension Reports (2022)
-
Target of rapamycin (TOR) gene is vital for whitefly survival and reproduction
Journal of Biosciences (2021)
-
Development of fly tolerance to consuming a high-protein diet requires physiological, metabolic and transcriptional changes
Biogerontology (2020)