Abstract
Acinetobacter baumannii is an important human pathogen due to its multi-drug resistance. In this study, the genome of an ST10 outbreak A. baumannii isolate LAC-4 was completely sequenced to better understand its epidemiology, antibiotic resistance genetic determinants and potential virulence factors. Compared with 20 other complete genomes of A. baumannii, LAC-4 genome harbors at least 12 copies of five distinct insertion sequences. It contains 12 and 14 copies of two novel IS elements, ISAba25 and ISAba26, respectively. Additionally, three novel composite transposons were identified: Tn6250, Tn6251 and Tn6252, two of which contain resistance genes. The antibiotic resistance genetic determinants on the LAC-4 genome correlate well with observed antimicrobial susceptibility patterns. Moreover, twelve genomic islands (GI) were identified in LAC-4 genome. Among them, the 33.4-kb GI12 contains a large number of genes which constitute the K (capsule) locus. LAC-4 harbors several unique putative virulence factor loci. Furthermore, LAC-4 and all 19 other outbreak isolates were found to harbor a heme oxygenase gene (hemO)-containing gene cluster. The sequencing of the first complete genome of an ST10 A. baumannii clinical strain should accelerate our understanding of the epidemiology, mechanisms of resistance and virulence of A. baumannii.
Similar content being viewed by others
Introduction
Recognized as one of the most problematic bacterial pathogens due to emergence of multidrug-resistant (MDR) strains1, Acinetobacter baumannii has been responsible for a significant proportion of world-wide healthcare-acquired infections, including ventilator-associated pneumonia, surgical site and urinary tract infections and septicemia2,3,4,5. Additionally, MDR clinical isolates of this species have been found to infect military personnel wounded in combat zones6,7.
Hospital outbreaks of A. baumannii infection are often associated with multidrug resistance in the causative strains8,9. Besides their intrinsic resistance to certain antibiotics due to the presence of native β-lactamase genes, poor permeability and efflux systems, A. baumannii strains have acquired a large array of antibiotic resistance mutations and genes, located either on the chromosome or plasmids10,11. Furthermore, clinical strains of A. baumannii resistant to carbapenems, the last resort antibiotics to treat infections caused by A. baumannii, have been found to possess acquired bla genes encoding several groups of carbapenem-hydrolyzing β-lactamases, such as blaOXA-2312,13, blaOXA-24/33/4014,15 and blaOXA-5816. These bla genes appear to have been transferred via mobile genetic elements such as insertion sequences, transposons or plasmids12,13,15,16.
Despite the clinical significance of A. baumannii infections, the molecular basis on the virulence and acquisition of multidrug resistance by A. baumannii remains largely unknown. To better understand the genome plasticity, natural history, epidemiology and acquisition of resistance and pathogenicity islands/genes of A. baumannii, the complete genomes of 21 A. baumannii strains, including LAC-4, became available as of December 31, 201417,18,19,20,21,22,23,24,25,26,27,28,29,30. In addition, the genomes of hundreds of A. baumannii strains have been sequenced to scaffold or contig levels (http://www.ncbi.nlm.nih.gov/genome/genomes/403). Most of these sequenced A. baumannii genomes are divided into 31 groups on basis of their sequence similarity. These efforts and other incomplete Whole Genome Shotgun (WGS) projects involving A. baumannii strains and strains of other Acinetobacter species31,32 will likely offer additional insights on epidemiology, phylogenetics, evolution of pathogenic strains and gene flows among Acinetobacter species including A. baumannii.
Several years ago, we described antibiotic resistance patterns and clonal relationships of 20 MDR clinical isolates of A. baumannii obtained from nosocomial outbreaks in Los Angeles County hospitals (LAC-1 to LAC-20)5. Our pulsed-field gel electrophoresis (PFGE) fingerprinting analysis indicated that these isolates appeared to have originated from eight epidemiologically distinct lineages5. More significantly, we identified the LAC-4 strain as hypervirulent in a mouse model of intranasal infection33 in comparison to other clinical isolates and laboratory strains of A. baumannii, including the eight representative LAC isolates and the widely studied clinical strain AYE22,34. The LAC-4 strain reliably reproduces the most relevant features of human pulmonary A. baumannii infection, including significant extrapulmonary dissemination and bacteremia33. Subsequent studies showed that LAC-4 exhibits high serum resistance, expresses a highly efficient heme utilization system35 and contains some unique structure and composition in its surface polysaccharide36, which may contribute to its hypervirulence. However, the precise mechanism of the hypervirulence of LAC-4 remains to be determined. Here we describe the complete genome sequence of the LAC-4 strain with special emphasis on the comparative genomics analyses to identify genomic regions that may contribute to the acquisition of antibiotic resistance and establishment of superior colonization and invasion by this hypervirulent strain.
Results and discussion
Phylogenetic lineages based on trilocus multiplex PCR and MLST
To understand the epidemiology and phylogenetics of 20 clinical isolates of A. baumannii (including LAC-4) obtained from four apparent nosocomial outbreaks, we first attempted to determine the clonal relationships among these isolates of A. baumannii by Trilocus multiplex PCR (TLM-PCR) analyses. Our results indicate that we can only type four isolates (LAC-11, LAC-12, LAC-13 and LAC-14) belonging to Global Clone (GC) II (Table 1). Since the TLM-PCR method failed to resolve phylogenetic relationships of most of these A. baumannii isolates, multilocus sequence typing (MLST) scheme based on Pasteur Institute approach was subsequently employed. Our results showed that previously non-typable isolates belong to unusual ST types (such as ST10, ST241 and ST417) (Table 1). Previously, PFGE profiling divided these 20 outbreak isolates into eight distinct clonal groups: LAC-1 to LAC-3; LAC-4; LAC-5 and LAC-8; LAC-6; LAC-7, LAC-9 and LAC-10; LAC-11 to LAC-14; LAC-15; and LAC-16 to LAC-205. In accordance with the PFGE grouping5, MLST typed LAC-5 and LAC-8 to a rare ST241 (Table 1). Since these two strains were isolated during two separate outbreaks in a single hospital, it appears that the ST241 lineage persisted for at least four years (1997–2001) in the same facility. More interestingly, we found that LAC-4, which was much more virulent than other LAC isolates in mice33, belongs to ST10 (Table 1). Most importantly, two series of outbreaks were caused by ST10 strains in two separate hospitals (LAC-1 to LAC-4 in Hospital A; LAC-16 to LAC-20 in Hospital C) in Los Angeles County, California, during the late 1990s (Table 1), suggesting that ST10 strains were quite dominant in causing nosocomial outbreaks in Los Angeles County at the time, with LAC-4 being their representative. While LAC-1 to LAC-4 were all typed to ST10, the PFGE profile of LAC-4 diverged from those of LAC-1 to LAC-3 sufficiently to be grouped as a separate clone5, indicative of possible divergent evolution of LAC-4 from its original clone.
There have been few reports describing clinical isolates of A. baumannii belonging to ST10. Among 1237 A. baumannii strains with assigned STs in Pasteur Institute's MLST database as of Oct 23 2014 (the most recent update), only three A. baumannii ST10 strains were listed. Recently, an ST10 strain of A. baumannii was isolated from a wound of a Canadian patient who had been previously hospitalized in India37. This strain and a strain of Klebsiella pneumoniae were transmitted to five other patients in an Edmonton, Alberta hospital, resulting in the colonization of four patients and the death of one patient due to septic shock caused by the OXA-23-producing A. baumannii strain37. Furthermore, four of five MDR strains of A. baumannii (LAC-6, LAC-7, LAC-9 and LAC-10) isolated during another outbreak in Hospital A in 2001 were typed to an uncommon ST417, indicating ST417 became dominant in this hospital in 2001 (Table 1)5. These results show that there was a clonal succession of outbreak strains in that hospital during a five-year span (1996–2001), transitioning from ST10 dominance in 1996–1999 to ST417 dominance in 2001. Previously reported clinical outbreaks were frequently caused by strains belonging to ST1 (Global Clone I)38,39,40, ST2 (Global Clone II)40,41 or ST1540. To our knowledge, in this report we describe the first nosocomial outbreaks caused by unusual ST10 and ST417 A. baumannii strains (Table 1).
LAC-4 genome sequences and general features
Our MLST analysis showed that LAC-4 and eight other outbreak LAC strains belong to the unusual ST10 (Table 1). Previous studies have shown that LAC-4 exhibits several distinctive attributes (iron utilization and unusual repeating unit composition of surface polysaccharides) that may contribute to its hypervirulence in mice33,36. Since none of the A. baumannii strains whose genomes were completely sequenced belongs to ST10, we decided to completely sequence the genome of LAC-4 using a combination of Roche 454 FLX+, Illumina Hiseq2000 and Illumina Miseq platforms, with gaps/uncertainties filled or clarified by genomic PCR and sequencing of PCR products. Our analysis showed that the LAC-4 genome consists of a circular chromosome of 3,954,354 base pairs and two circular plasmids, one with 8,006 base pairs while the other with 6,076 base pairs. The general features of the LAC-4 genome are summarized in Table 2.
The LAC-4 genome contains multiple copies of five types of IS elements: ISAba1, ISAba13, ISAba125 and novel ISAba25 and ISAba26 whose names were recently assigned (see Table 2 and Supplementary Table S1 online). There are 19 copies of ISAba1 in the LAC-4 genome. Three pairs of ISAba1 constitute three novel transposons (Tn6250, Tn6251 and Tn6252) because identical target site duplication (TSD) sequences are found next to the external (outward facing) inverted repeats (see Fig. 1A; Supplementary Table S1 online), while one ISAba1 element is found to be linked to an ampC gene (Fig. 1B). In addition, we identified in the LAC-4 genome 12 copies of a novel IS element (ISAba25) whose closest hit from the IS Finder database is ISAba16 (see Supplementary Table S1 online). This IS element has 2,491 base pairs in length with three ORFs, while ISAba16 has 2,552 bp. Importantly, the largest ORF of ISAba25 has a 63-bp deletion compared to ISAba16, resulting in a predicted polypeptide of 527 amino acids, 21 amino acids shorter than its counterpart in ISAba16 (which has 548 amino acids). Two copies of ISAba25 are associated with genes of a RND type of efflux pump system (AdeIJK) (Fig. 1B), probably contributing to LAC-4's resistance to a number of antimicrobial agents. Furthermore, 14 copies of another novel IS element (ISAba26) are located on the LAC-4 chromosome (see Supplementary Table S1 online). This IS is 1,318 bp in length and contains one gene encoding a transposase with 402 amino acids. While transposase protein sequence BLAST analysis indicates ISAba26 belongs to the mutator family of transposases, IS Finder database BLASTp search identifies ISEc39 as its closest homologue, sharing 72% transposase amino acid identities. ISAba26 produces TSD of 8 bases in length (see Supplementary Table S1 online) and its inverted repeats (IRL and IRR) are 26 bp in lengths. Interestingly, two copies of ISAba26 flank a large number of continuous loci (15 genes) which are predicted to be involved in copper resistance (Fig. 1B). Moreover, 22 copies of ISAba13 are scattered around the chromosome of LAC-4 (see Supplementary Table S1 online). Among these elements, 14 copies were found to create TSD sequences of 9 bases in length as expected, while one element has an 8-base TSD (see Supplementary Table S1 online). Finally, we found 14 ISAba125 elements in LAC-4 genome, each with a TSD of 3 bases (see Supplementary Table S1 online).
Select examples of insertion sequences and transposons in LAC-4 genome.
Direction of transcription of genes is shown by the direction of the arrows. Dark arrows denote ORFs for transposase genes. Gray shaded arrows mark genes potentially conferring resistance to antimicrobial agents or copper. Genes involved in conjugation (tra genes) are marked by striped arrows, while genes coding for proteins of unknown or other functions are shown in open arrows. Due to space limitation, certain loci are labeled with abbreviated locus-tags without the full name (e.g., ABLAC_01150 is abbreviated as 01150). Sizes of genes and intergenic distances are not drawn to scale. (A) Composite transposons (Tn6250, Tn6251 and Tn6252) formed by ISAba1 elements. Target site duplication (TSD) of 9 bases in length was noted only next to the external inverted repeats (facing outwards). An asterisk (*) next to ISVsa3 indicates the partial nature of the element (likely not functional). (B) Gene or gene clusters flanked by IS elements. A delta symbol (Δ) next to ISAba1 denotes an apparently nonfunctional copy of the element.
The LAC-4 genome also includes two plasmids (Table 2). Plasmid pABLAC1 contains nine predicted ORFs (ABLAC_p100010–ABLAC_p100090). Among these loci, ABLAC_p100020 (rep) encodes a replicase (Aci3) belonging to group 3 of replicases defined42. Fifty-six base pairs upstream of the rep gene, four copies of iterons were found with a sequence of 5′- TAAAACGAGGTTTACCTTGCAT-3′, which is identical to those observed in replicons Ab203-Aci3 and Ab736-Aci742. Iterons have been shown to be involved in controlling plasmid replication via interacting with replicase proteins43. Additionally, an AT-rich sequence (5′-AAAAATAT-3′) identical to that found in the pRAY plasmid44 was located 37 bp downstream of the fourth iteron and 11 bp upstream of rep start codon. The AT-rich element and the iterons presumably serve as the oriV site of the plasmid. Furthermore, ABLAC-p100050 and ABLAC_p100060 were found to encode homologues of RelE toxin of toxin/antitoxin system and Cro/C1 transcriptional regulator, respectively. However, no homologue of antitoxin partner gene was found on the plasmid. Many of predicted proteins of plasmid pABLAC1 loci (ABLAC_p100020 through ABLAC_p10002060) share identical amino acid sequences with those encoded by genes harbored by the plasmid (pD1279779) in A. baumannii strain D127977923. On the other hand, plasmid pABLAC2 shares nearly 100% with a series of plasmids related to pRAY44,45, in particular pRAY*. Of particular note, two pABLAC2 loci (ABLAC_p200050 and ABLAC_p200060) encode homologues of MobA and MobC, respectively, which are known to be involved in plasmid mobilization; it is tempting to speculate that these mobilization genes may contribute to the transfer of the aminoglycoside resistance gene (ABLAC_p200010, ant(2″)-Ia or aadB) carried on this plasmid. Moreover, two copies of AT-rich sequence 5′-AAAAATAT-3′ were found within the coding region of a predicted ORF (ABLAC_p200080). However, no potential replication gene (rep) or iterons were found on pABLAC2. Similar to pRAY series of plasmids44,45, no plasmid partitioning or restriction/modification systems were identified in both plasmids of LAC-4.
Comparative genomics of LAC-4 with 20 other completely sequenced A. baumannii genomes
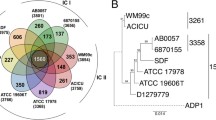
All protein-coding genes (CDSs) of LAC-4 were analyzed by mGenomeSubtractor-based in silico subtractive hybridization for presence of homologues against 20 other completely sequenced A. baumannii genomes. Our results indicate that the numbers of homologous CDSs (H-value > 0.42) among these genomes are 2,276 (Fig. 2A), representing 60% of the total CDSs in LAC-4. The LAC-4 chromosome is also presented in a continuous linear format showing the locations of all of its CDSs each with a degree of “blackness” reflecting relative conservation among the other 20 A. baumannii genomes (Fig. 2B). In particular, the LAC-4 genome exhibits the highest level of sequence identities to the BJAB0715 genome with 3,451 conserved genes (H-value > 0.42) (see Fig. 2A, 2B; Supplementary Fig. S1 online). When Acinetobacter baumannii genome groups are considered, LAC-4 was found to belong to Genome Group 3, represented by the genome of BJAB7015, as verified by a phylogenetic tree composed of 45 completely or partially sequenced A. baumannii genomes (see Supplementary Fig. S1 online).
The mGenomeSubtractor-based in silico subtractive hybridization of the LAC-4 genome against genomes of twenty other completely sequenced A. baumannii isolates.
The twenty subject A. baumannii chromosomes include: ATCC 17978 (NCBI accession no. NC_009085), SDF (NC_010400), AYE (NC_010410), ACICU (NC_010611), AB0057 (NC_011586), AB307-0294 (NC_011595), 1656-2 (NC_017162), MDR-ZJ06 (NC_017171), TCDC-AB0715 (NC_017387), MDR-TJ (NC_017847), TYTH-1 (NC_018706), D1279779 (NC_020547), BJAB07104 (NC_021726), BJAB0868 (NC_021729), BJAB0715 (NC_021733) and ZW85-1 (NC_023028) available at NCBI RefSeq project; and AB031 (GenBank accession no. CP009256), AC29 (CP007535), AB030 (CP009257) and AbH12O-A2 (CP009534) available at GenBank. (A) Histogram of BLASTn-based H-values for all 3,788 annotated protein-coding genes in the LAC-4 chromosome against all 20 subject chromosome sequences (color-coded). The H-value reflects the degree of similarity in terms of the length of match and the degree of identity at a nucleotide level between the matching gene in the subject genome and the query gene examined. The conserved genes were identified based on each of the obtained H-values great than 0.42. The genome of LAC-4 shows the most significant sequence identity to the BJAB0715 genome with 3,451 conserved genes (H-value > 0.42) among all the other 20 completely sequenced genomes; whereas, the lowest sequence identity to the SDF genome with 2,511 conserved genes (H-value > 0.42). (B) Chromosome map of LAC-4 with gene black/white-shade-coded based on the number of comparator A. baumannii genomes identified as harboring a nucleotide sequence-conserved homologue. Genes shown in absolute black (‘20’) are conserved across all 20 A. baumannii comparator genomes, with genes shown in decreasing shades of black being conserved in lower numbers of A. baumannii comparator genomes, while at the other extreme those shown in white (‘0’) are unique to LAC-4. Non-coding regions are shown as gaps. The genomic island-like hyper-variable regions (also see Table 3) are marked by red rectangles.
A total of 12 genomic islands were identified (Fig. 2B and Table 3). The GI1 (Table 3) consists of a novel transposon (Tn6250) composed of two ISAba1 elements at the ends and an IS1006 internally next to a partial ISVsa3 element (Fig. 1A). Additionally, this genomic island harbors resistance genes for streptomycin (strA and strB) and for sulphonamides (sul2), thus this GI is a resistance island (Table 3). The 34 kb GI2 contains two novel IS elements (ISAba26) near the ends (Table 3 and Fig. 1B). Notably, these two IS elements sandwich a long 15-gene cluster (ABLAC_05330 to ABLAC_05470) coding for various proteins involved in copper resistance (Fig. 1B), representing the second resistance island found in LAC-4. Most of the genes of this gene cluster were also present in the genome of A. baumannii ATCC 17978 and to a lesser extent, in those of A. baumannii strains AB0057 and AYE (Fig. 3A). Outside one of the ISAba26 elements (ABLAC_05500), another novel IS element (ISAba25) and a phage integrase gene (ABLAC_05570) are found (Table 3 and Fig. 3A), implicating the roles of mobile elements in the transfer of this GI. Furthermore, GI3 is bounded by two copies of a novel IS element (ISAba25; transposase ORFs: ABLAC_07480-07500 and ABLAC_0758-07600) and it harbors a partial ISAba1 element and genes for the AdeIJK, which are components of an RND-type efflux pump system involved in resistance phenotypes of multiple classes of antibiotics (Fig. 1B).
Alignment maps of the two representative A. baumannii LAC-4 genomic islands against related gene clusters among completely sequenced A. baumannii genomes.
The BLASTp searches + hit collocation approach was used to generate the alignment, with matching genes shown as color-matched. (A) The 34-kb GI2 carries the copper resistance gene cluster (highlighted by a red line). (B) GI11 harbors genes predicted to encode enzymes for detoxification of xenobiotic compounds such as formaldehyde.
Several GIs (GI5, GI6, GI7, GI8, GI9 and GI10) contain many genes encoding phage-derived proteins of unknown functions; the significance of these GIs remains to be elucidated (Table 3). Among these GIs, nearly all of the phage-derived genes of the GI8 have no homologues in any of the 20 other completely sequenced A. baumannii genomes (except several transposase genes and Zn-dependent protease gene) (data not shown). Moreover, GI11 contains a number of genes encoding putative proteins/enzymes with functions of detoxification of a variety of xenobiotic compounds (Fig. 3B); for example, glutathione S-transferase (ABLAC_32340, gstB), S-formylglutathione hydrolase (ABLAC_32430, frmB) and S-(hydroxymethyl)glutathione dehydrogenase (ABLAC_32440, frmA) are involved in resistance to and detoxification of formaldehyde and chlorine46. Interestingly, other than the frmA gene, most of the GI11 genes are unique to LAC-4 among the 21 completely sequenced A. baumannii genomes (data not shown). Finally, the GI12 contains the K (capsule) locus which encodes enzymes responsible for biosynthesis of the surface polysaccharide of LAC-4 (Fig. 4). Gene product homology analyses indicate that the K locus gene cluster harbors genes encoding two series of enzymes (FnlA, FnlB, FnlC and Leg1–6) which apparently catalyze biosynthesis of α-L-frucosamine and α-8-epi-legionaminic acid, respectively. These two sugars are the two precursor sugars (other than α-D-glucosamine, which is supplied by cellular metabolism) for the biosynthesis of three-sugar repeating unit of LAC-4 surface polysaccharide recently determined to consist of α-L-frucosamine, α-D-glucosamine and α-8-epi-legionaminic acid36. Interestingly, six ORFs (orfK1-6, ABLAC_36970–ABLAC_37020) in the middle of the gene cluster encode protein products (two of which are oxidoreductases) sharing no homology with any proteins in the 20 completely sequenced A. baumannii genomes (Fig. 4). Among the 20 A. baumannii strains whose genomes were completely sequenced, several have acquired resistance islands of various sizes18,25,34,47. For example, strain AYE harbors an 86-kb resistance island in which 45 resistance genes are located34. Similarly, MDR-TJ contains a 42-kb resistance island with four drug resistance genes47, while MDR-ZJ06 has a 38-kb resistance island (AbaR22) containing 40 genes, five of which are involved in antimicrobial resistance25.
LAC-4 K locus gene organization and functional assignment.
GI12 harbors genes of K locus in LAC-4. Color coding scheme follows that of Hu et al63. Two of the three-sugar repeating unit recently determined for LAC-4 surface polysaccharide36 can be produced via reactions catalyzed by gene products of the genes in the cluster (bottom). The product of wafP gene probably catalyzes the formation of the 1 → 3 glycosyl bond between α-L-FucNAC and α-D-GlcNAc. The wbpV gene (marked by *) is a pseudogene (with two separate ORFs) as a result of a point mutation.
Antibiotic resistance of LAC-4 and resistance genetic determinants
The recent emergence and rapid dissemination of multidrug and pandrug resistant A. baumannii has caused significant burdens in clinical management of infections world wide. While we previously described antimicrobial susceptibility profile of LAC-4 towards 17 antibiotics, minimal inhibitory concentration (MIC) values were not presented5. Our MIC results of 23 antimicrobial drugs indicate that this strain is resistant to 13 of the 23 antibiotics (six of eight classes) tested, including ampicillin, carbenicillin, piperacillin, most of the cephalosporin analogs, three of four aminoglycosides, nalidixic acid and ciprofloxacin, trimethoprim and chloramphenicol (see Supplementary Table S2 online). Compared to the most highly resistant strains (AYE, ACICU, MDR-TJ and MDR-ZJ06) among the 20 A. baumannii strains whose genomes were completely sequenced, the MDR LAC-4's level of resistance is moderate (e.g., susceptible to imipenem and meropenem, intermediate to amikacin), which reflects the relatively early isolation of this strain (in 1997) from a hospital in Los Angeles. Nevertheless, the availability of complete genome sequences of A. baumannii strains with varied degrees of antimicrobial resistance (including the moderately resistant LAC-4) is highly useful for the scientific research community, especially from the standpoints of emergence and dissemination of antimicrobial resistance genes.
To better understand the repertoire of MDR genetic determinants and organization, in-depth analysis of the genome sequences of LAC-4 was performed. Consistent with the fact that LAC-4 was not among the most resistant strains analyzed in our 2008 report5, the LAC-4 genome harbors a moderate number of genetic determinants, some of which are linked to mobile genetic elements (IS or Tn), with a potential to encode resistance functions observed in this bacterial strain (Table 4). For example, genes potentially encoding for all four classes of β-lactamases (Classes A, B, C and D) have been found (Table 4). Two of such loci (ampC and blaOXA-236) are closely associated with ISAba1 elements (Fig. 1B and 1A, respectively) which may provide exogenous promoter functions, consistent with LAC-4 being resistant to piperacillin, older versions of the penicillins and several cephalosporin antibiotics (see Supplementary Table S2 online). ISAba1 has also been found in the genomes of other A. baumannii strains associated with antibiotic resistance genes (including ampC), most likely driving robust expression of the resistance phenotypes17,47,48. While a blaOXA-51-like gene (ABLAC_23600) is found in the LAC-4 genome (Table 4) as expected, the absence of blaOXA-23-like, blaOXA-40-like and blaOXA-58-like genes (data not shown) in the genome explains its susceptible phenotypes to imipenem and meropenem (see Supplementary Table S2 online). In contrast, in MDR strains AB0057, MDR-TJ, MDR-ZJ06, BJAB07104, BJAB0868 and BJAB0715, the presence of blaOXA-23 could account for their carbapenem resistance17,18,25,47. In addition, presence of aminoglycoside 6-phosphotransferase [aph(6)-Id, i.e., strB] and/or aminoglycoside 3″-phosphotransferase [aph(3″)-Ib, i.e., strA] were known to contribute resistance to streptomycin49. The existence of several aminoglycoside modification enzyme genes [ABLAC_01120 (strA), ABLAC_01130 (strB) and ABLAC_p200010 (ant(2″)-Ia)] in the LAC-4 genome (Table 4) and an association of an ISAba1 with two of these loci (strA and strB) in the context of a novel transposon Tn6250 (Fig. 1A) correlate well with the strain's resistance to streptomycin, gentamicin, kanamycin and tobramycin (see Supplementary Table S2 onlinie). Finally, genes encoding a series of proteins for major facilitator superfamily (MFS), resistance-nodulation-division (RND) family and multidrug and toxic compound extrusion (MATE) family of efflux pumps were also located in the LAC-4 genome (Table 4). Notably, there are at least five complete sets of RND type of efflux pump systems in LAC-4 (Table 4). Besides the AdeIJK genes linked with several IS elements described above in GI3 (Table 3 and Fig. 1B), four other sets of RND type efflux pump systems include AdeABC (encoded by ABLAC_26430–26450), AdeFGH (encoded by ABLAC_11920–11940), CzcABC (encoded by ABLAC_02860–02880) and homologues of CzcABC (Table 4). The last two sets have been shown to participate in transporting toxic metals such as cobalt, zinc and cadmium50,51. These efflux pump systems and other membrane-associated transporters (Table 4) confer resistance to diverse classes of antibiotics and other toxic compounds/metals in bacteria.
Many A. baumannii isolates have been reported to possess a series of efflux pump gene clusters in their genomes, which encode several types of chemical extrusion apparatuses rendering cells resistant to a large variety of antibiotics and toxic compounds52,53. For instance, there are three sets of RND efflux pump systems in A. baumannii strains (AdeABC, AdeIJK and AdeFGH)54,55,56. While LAC-4 genome harbors three gene clusters each of which separately encodes one of the above RND type efflux pumps, no apparent two-component system gene homologues adeRS was found upstream of adeABC gene. Such two-component systems (AdeRS) were found to be critical in regulating expression of AdeABC efflux pumps in several strains of A. baumannii57,58. Interestingly, Sun and colleagues observed that the insertion of an ISaba1 into the adeS gene resulted in over-expression of adeABC operon hence the non-susceptibible phenotypes of clinical strains to tigecycline59. Additionally, AdeIJK appears to be regulated by AdeN, a TetR type of regulator in some strains60. On the contrary, the LAC-4's adeIJK genes exist in an unusual genomic context. These genes are not linked with any recognizable regulatory gene(s). Upstream of adeIJK genes, there exist a locus of unknown function (ABLAC_07540) and an apparent nonfunctional copy of ISAba1 element (Fig. 1B). All these genes are flanked by two copies of novel IS element ISAba25 (Fig. 1B). To our knowledge, this is the first report of a gene cluster encoding an RND efflux pump system being flanked by two ISAba25 elements, which may modify the expression of the efflux pump genes, thus conferring resistance to a number of antimicrobial agents.
Potential virulence factors
Because our recent studies have shown that the LAC-4 strain is much more virulent than other A. baumannii strains in the mouse model of intranasal infection33, the LAC-4 genome was searched for genes coding for potential virulence factors against entries from VFDB. A total of 615 gene hits were generated which could facilitate the identification, evaluation and validation of virulence factors in this strain and the species (see Supplementary Table S3 online). Potential virulence factor loci that are unique for LAC-4 include ABLAC_03200 (which may encode a fimbrial protein, PilE), ABLAC_37000 and ABLAC_37010 [two genes of K locus gene cluster (Fig. 4)] (see Supplementary Table S3 online). In addition, several other potential virulence factor loci are found in only 1 to 4 genomes of the 20 completely sequenced A. baumannii genomes: ABLAC_05370 (encoding a transcriptional activator protein CusR/CopR), ABLAC_05450 (encoding an Fe2+ transport system protein FeoB) and K locus gene cluster genes ABLAC_36940–39960 [encoding UDP-N-acetylglucosamine 2-epimerase (FnlC), reductase (FnlB) and 4,6-dehydratase,3-and 5-epimerizase (FnlA)](see Supplementary Table S3 online and Fig. 4). Additional genes of the K locus gene cluster (ABLAC_36850 and ABLAC_36870–ABLAC_36910) were also identified as potential virulence factor genes (see Supplementary Table S3 online). Recently, Russo and colleagues identified, within the K locus gene cluster of A. baumannii strain AB307-0294, two loci ptk and epsA (encode a putative protein tyrosine kinase and a putative polysaccharide export outer membrane protein, respectively) that are required for capsule-positive phenotype, for survival in human serum and survival in a rat soft tissue infection model61; gene knock-out mutants of either gene were completely cleared from animals in the in vivo experiments. The encoded proteins of these two genes shared 90% and 95% identity over the entire protein lengths with the products of loci ABLAC_37120 and ABLAC_37100, respectively, suggesting importance of K locus gene cluster genes to the virulence of A. baumannii. More interestingly, as described above (Fig. 4), LAC-4's K locus gene cluster also contains a series of genes (leg1-leg6, ABLAC_37030–37080) apparently encoding enzymes necessary for biosynthesis of legionaminic acid, the precursor for an uncommon sugar (α-8-epi-legionaminic acid) found in the repeating unit of the surface polysaccharide of this strain36 and others62,63. It has been proposed that since legionaminic acid structurally resembles sialic acid on mammalian host cell's surface glycoproteins, bacterial pathogens may utilize this unusual sugar to mimic host cell surface, thus escaping from host immune surveillance to facilitate their colonization and invasion64,65,66. Since the legionaminic acid biosynthesis genes are not common among the 20 completely sequenced A. baumannii genomes (well conserved in only three of the 20 genomes), it is tempting to hypothesize that these genes may contribute to LAC-4's hypervirulence in mice. Furthermore, there are six ORFs (orfK1-6, ABLAC_36970–ABLAC_37020) in LAC-4's K locus that are absent in the other 20 completely sequenced genomes. It would be exciting to determine if any of these ORFs is involved in pathogenesis of this strain.
The ability of bacterial pathogens to obtain scarce iron from their mammalian hosts is critical for survival and infectivity67,68. Consequently, proteins that participate in uptake and utilization of iron or heme have been recognized as crucial virulence factors69,70,71,72. Not surprisingly, additional loci that were identified as putative virulence factor genes include genes from heme utilization cluster 1 (ABLAC_24390 and ABLAC_24450) and the hemO cluster (ABLAC_16780, ABLAC_16790 and ABLAC_16800) (see Supplementary Table S3 online). Also identified as potential virulence factor genes are many linked genes (ABLAC_10700, ABLAC_10710, ABLAC_10730, ABLAC_10750–ABLAC_10810, ABLAC_10840–ABLAC_10880) which are major constituents of a large 20-gene cluster (ABLAC_10690–ABLAC_10880) for the biosynthesis and transport of the acinetobactin siderophore in LAC-4 strain (see Supplementary Table S3 online); this gene cluster shares identical gene organization as described for the same gene clusters in several other A. baumannii genomes73. Similarly, a second siderophore biosynthesis and transport gene cluster (ABLAC_24780–ABLAC_24890) was identified from LAC-4 genome based on the presence of six of the loci within the gene cluster (ABLAC_24780–ABLAC_24820, ABLAC_24850 and ABLAC_24890) being on the list of potential virulence genes in LAC-4 (see Supplementary Table S3 online). The proteins encoded by the genes of this cluster and gene organization are identical to the cluster 5 described previously73 for a putative hydroxamate siderophore.
Our previous results35 and those of others73 suggested that a heme utilization gene cluster including a gene encoding for heme oxygenase (thus called hemO cluster) was present only in some A. baumannii strains and may account for their enhanced virulence. Bioinformatics analysis confirmed the presence of a hemO gene cluster in the LAC-4 genome (ABLAC_16780 to ABLAC_16850). Furthermore, comparative genomics analyses using BLASTp search + hit colocation analysis showed that 11 of the other 20 complete genomes of A. baumannii strains also contain the hemO cluster (see Supplementary Fig. S2 online). Another heme utilization gene cluster (without hemO gene; to be called heme utilization cluster 1) was reported to be present in all strains of A. baumannii tested73. As predicted, LAC-4 also harbors heme utilization cluster 1 (ABLAC_24350–ABLAC_24460), which is present in all the other 20 A. baumannii genomes (see Supplementary Fig. S3 online). The presence of hemO cluster in LAC-4 and 19 other outbreak isolates5 was further investigated by PCR. All 20 outbreak isolates (including LAC-4) were found to contain the hemO cluster (Table 1; see Supplementary Fig. S4 online), suggesting the relatively common presence of this gene cluster in this collection of clinical isolates. In this regard, our previous studies have shown that LAC-5, LAC-7, LAC-11 and LAC-16 are no more virulent than ATCC 17978, ATCC 17961 or clinical isolate AYE in the mouse model of intranasal infection33. Taken together, our data suggest that the presence of hemO cluster per se may not entirely account for the hypervirulence of LA-4. Indeed, the LAC-4 genome analysis identified four gene clusters for iron/heme utilization (the hemO cluster, the heme utilization cluster 1 and two gene clusters for siderophore biosynthesis and transport). It is highly likely that these gene clusters provide redundant function to ensure efficient acquisition of iron for cellular use. Further genome-wide molecular genetics studies (such as transposon mutagenesis) will be needed to decipher the relative contribution of genes or gene clusters of the iron/heme utilization and other processes in the virulence of this strain.
In summary, here we reported the complete genome sequence of a hypervirulent, multidrug resistant clinical outbreak isolate (LAC-4) of A. baumannii with an extremely rare MLST sequence type ST10 (MLST Pasteur Scheme). Among the 20 strains of A. baumannii whose complete genomes were available before this report, there are three strains of ST1, nine strains of ST2, two of ST79, one each for ST17, ST23, ST267, ST437, ST638 and ST639 (see Supplementary Table S4 online). Thus, the LAC-4 genomics study reported here represents the first complete genome sequence of an ST10 A. baumannii strain, making an important addition to the growing list of complete A. baumannii genomes for the scientific research community world-wide. Additionally, molecular tests and comparative genomics analyses offer insight in the mechanisms of resistance and virulence in this important bacterium.
Methods
Bacterial strains
The 20 outbreak strains (LAC-1 to LAC-20) were collected from four apparent clinical nosocomial outbreaks from 3 hospitals in Los Angeles County between 1996 and 2004 and obtained from Los Angeles County Public Health Laboratory5. The antimicrobial susceptibility and genetic profiles of these non-duplicate isolates were described in details previously5. Specifically, strains LAC-1 through LAC-5 were isolated from an outbreak in Hospital A that lasted for several years (1996–1999), while LAC-6 to LAC-10 were from the 2001 outbreak in the same hospital5. Separately, strains LAC-11 to LAC-15 were obtained from an outbreak in Hospital B during 2003–2004. Furthermore, there was an outbreak in Hospital C spanning 1997 and 1998 where LAC-16 to LAC-20 were archived5.
Clonal relationships and sequence typing
Tri-locus multiplex PCR as described by Turton and colleagues74 was used to determine clonal relationships of these A. baumannii clinical isolates [Global Clones (GC) I, II or III, also known as International Clones I, II or III]. Multi-locus sequence typing (MLST) was performed based on methods of Diancourt et al75. PCR amplification was performed in separate reactions of a final volume of 50 μl. After amplification, aliquots of the PCR reactions were subject to agarose gel electrophoresis analysis. If successful, PCR products were purified using a QIAquick PCR purification kit (Qiagen, Valencia, CA) and then sequenced on an ABI 3730 automated fluorescent sequencer. Determination of the sequence type was carried out using details on the MLST scheme at www.pasteur.fr/mlst. To obtain MSLT ST assignments for A. baumannii strains whose genomes are complete, genome FASTA sequences were retrieved from NCBI and uploaded in the “batch sequence query” mode of MLST (Pasteur) database on the Acinetobacter baumannii MLST Databases website (http://pubmlst.org/abaumannii/), which will generate allelic profiles and STs.
Genomic DNA sequencing, assembly and gap closing
The LAC-4 strain was grown overnight at 37°C to stationary phase in LB medium and total DNA was isolated from harvested cells. The genome sequence of LAC-4 was first determined using Roche 454 FLX+ and assembled with the GS De Novo Assembler, resulting in sequences with 43.5-fold coverage. Then 105 large contigs (>500 kb) were obtained by a combination of re-sequencing by Illumina Hiseq2000 (paired-end sequencing for 400-bp library, with 255.6-fold coverage) and Illumina Miseq (mate-pair sequencing for 3000-bp library, with 408-fold coverage). Finally, gaps between these contigs were closed by genomic PCR and sequencing of PCR products using conventional Sanger method (Applied Biosystems 3730 Genetic Analyzer). An integrity check that performs a BLASTp analysis on neighboring pairs of proteins identified 37 pairs of proteins, which may represent single genes that either have gained mutations or have split into two open reading frames (ORFs) due to sequencing errors. Additional PCR and DNA sequencing analysis have either corrected sequencing mistakes (likely as a result of 454 sequencing errors around homopolymer nucleotides) or combined gene pairs into single pseudogenes.
Genome annotation and comparative genomics analysis
The replication origin (oriC) in LAC-4 genome was predicted by OriFinder76. The assembled genome sequences were annotated by using programs Glimmer 3.02 for identification of protein-coding sequences (CDSs)77 with manual validation of predicted CDSs on the basis of annotations of the BJAB7015 genome17 and the CBMAR database78, tRNAscan-SE for tRNA genes79 and RNAmmer for rRNA genes80. CDS functions were predicted using BLASTp searches81 of the NCBI non-redundant database followed by manual curation using the annotations of A. baumannii ATCC 17978 (NCBI accession no. NC_009083.1)19 and BJAB7015 (NC_021733.1)17 as references.
The virulence factor genes were predicted with the BLASTp searches against the virulence factor database (VFDB)82. The mobile genetic elements in the LAC-4 genome sequences were detected by the following online tools and/or open-access database and manual examinations: MobilomeFINDER for the tRNA/tmRNA gene-related genomic islands (GI)83, IslandViewer for the island-like regions84 and IS Finder for insertion sequence (IS) elements85. New IS names were provided by the curators of IS database85, while transposon (Tn) numbers were assigned by the Tn Number Registry86. The gene clusters carried by genomic islands (GIs) or involved in the heme utilization in the sequenced A. baumannii genomes were aligned by using the standalone package MultiGeneBlast87, an approach also known as “BLASTp searches + hit collocation”.
The genome sequence comparisons of LAC-4 with the other 20 completely sequenced A. baumannii strains were performed with the rapid multiple genome alignment tool mGenomeSubtractor88. All the annotated LAC-4 protein-coding genes (served as query) were examined by mGenomeSubtractor-facilitated BLASTn searches, using an H-value cut-off > 0.42 for conserved genes, against the other A. baumannii genomes (served as subject). The H-value (0 < = H-value < = 1.0) reflects the degree of similarity in terms of the length of match and the degree of identities at a nucleotide level between the matching gene in the subject genome and the query gene examined88.
GenBank accession number
The genome sequences of the LAC-4 chromosome and two plasmids pABLAC1 and pABLAC2 have been submitted to the GenBank under accession numbers CP007712, CP007713 and CP007714, respectively.
Antimicrobial susceptibility testing
Antimicrobial susceptibility of the clinical isolates or strains were determined using the broth microdilution protocols of Clinical Laboratory Standards Institute89 according to methods described previously5. Escherichia coli (ATCC #25922) and Pseudomonas aeruginosa (ATCC #27853) were used as quality control strains in the testing.
Molecular detection of hemO cluster genes
PCR primer pairs specific for each gene of the eight-gene hemO cluster35 were designed based on alignment of gene cluster sequences of six A. baumannii strains [strains ACICU21, SDF22, MDR-ZJ0625, TYTH-127, TCDC-AB0711526 and AB005718], available as of May, 2013. The sequence alignment was performed by using Lasergene MegAlign program from DNASTAR, Inc. (Madison, WI). Primers were chosen from regions of highly conserved sequences and are shown in Supplementary Table S5 online. The PCR amplification of the hemO gene cluster was performed for the clinical isolates using genomic DNA as a template. Individual PCR reactions involved a 50 μl reaction mixture containing 1 × 5 PRIME MasterMix, 200 nM of forward and reverse primers and 1 μl of genomic DNA template. Amplification was carried out with 5 min at 95°C; 36 cycles of 30 s at 95°C, 30 s at 55°C and 1 min at 72°C; and 10 min at 72°C. The resulting PCR products were visualized and imaged through agarose gel electrophoresis analysis.
References
Boucher, H. W. et al. Bad bugs, no drugs: no ESKAPE! An update from the Infectious Diseases Society of America. Clin Infect Dis 48, 1–12 (2009).
Azim, A. et al. Epidemiology of bacterial colonization at ICU admission with emphasis on extended-spectrum beta-lactamases (ESBL) and metallo-beta-lactamase (MBL) producing gram-negative bacteria - an Indian experience. J Med Microbiol 59, 955–60 (2010).
Karlowsky, J. A. et al. Surveillance for antimicrobial susceptibility among clinical isolates of Pseudomonas aeruginosa and Acinetobacter baumannii from hospitalized patients in the United States, 1998 to 2001. Antimicrob Agents Chemother 47, 1681–8 (2003).
Sievert, D. M. et al. Antimicrobial-resistant pathogens associated with healthcare-associated infections: summary of data reported to the National Healthcare Safety Network at the Centers for Disease Control and Prevention, 2009-2010. Infect Control Hosp Epidemiol 34, 1–14 (2013).
Valentine, S. C. et al. Phenotypic and molecular characterization of Acinetobacter baumannii clinical isolates from nosocomial outbreaks in Los Angeles County, California. J Clin Microbiol 46, 2499–507 (2008).
Huang, X. Z. et al. Molecular analysis of imipenem-resistant Acinetobacter baumannii isolated from US service members wounded in Iraq, 2003-2008. Epidemiol Infect 140, 2302–7 (2012).
Taitt, C. R. et al. Antimicrobial resistance determinants in Acinetobacter baumannii isolates taken from military treatment facilities. Antimicrob Agents Chemother 58, 767–81 (2014).
Antunes, L. C., Visca, P. & Towner, K. J. Acinetobacter baumannii: evolution of a global pathogen. Pathog Dis 71, 292–31 (2013).
Zarrilli, R., Pournaras, S., Giannouli, M. & Tsakris, A. Global evolution of multidrug-resistant Acinetobacter baumannii clonal lineages. Int J Antimicrob Agents 41, 11–9 (2013).
Poirel, L., Bonnin, R. A. & Nordmann, P. Genetic basis of antibiotic resistance in pathogenic Acinetobacter species. IUBMB Life 63, 1061–7 (2011).
Imperi, F. et al. The genomics of Acinetobacter baumannii: insights into genome plasticity, antimicrobial resistance and pathogenicity. IUBMB Life 63, 1068–74 (2011).
Mosqueda, N. et al. Globally expanding carbapenemase finally appears in Spain: nosocomial outbreak of acinetobacter baumannii producing plasmid-encoded OXA-23 in Barcelona, Spain. Antimicrob Agents Chemother 57, 5155–7 (2013).
Mugnier, P. D., Poirel, L., Naas, T. & Nordmann, P. Worldwide dissemination of the blaOXA-23 carbapenemase gene of Acinetobacter baumannii. Emerg Infect Dis 16, 35–40 (2010).
Bou, G., Oliver, A. & Martinez-Beltran, J. OXA-24, a novel class D beta-lactamase with carbapenemase activity in an Acinetobacter baumannii clinical strain. Antimicrob Agents Chemother 44, 1556–61 (2000).
Grosso, F., Quinteira, S., Poirel, L., Novais, A. & Peixe, L. Role of common blaOXA-24/OXA-40-carrying platforms and plasmids in the spread of OXA-24/OXA-40 among Acinetobacter species clinical isolates. Antimicrob Agents Chemother 56, 3969–72 (2012).
Metan, G., Sariguzel, F., Sumerkan, B., Reijden, T. & Dijkshoorn, L. Clonal diversity and high prevalence of OXA-58 among Acinetobacter baumannii isolates from blood cultures in a tertiary care centre in Turkey. Infect Genet Evol 14, 92–7 (2013).
Zhu, L. et al. Complete genome analysis of three Acinetobacter baumannii clinical isolates in China for insight into the diversification of drug resistance elements. PLoS One 8, e66584 (2013).
Adams, M. D. et al. Comparative genome sequence analysis of multidrug-resistant Acinetobacter baumannii. J Bacteriol 190, 8053–64 (2008).
Smith, M. G. et al. New insights into Acinetobacter baumannii pathogenesis revealed by high-density pyrosequencing and transposon mutagenesis. Genes Dev 21, 601–14 (2007).
Park, J. Y. et al. Complete genome sequence of multidrug-resistant Acinetobacter baumannii strain 1656-2, which forms sturdy biofilm. J Bacteriol 193, 6393–4 (2011).
Iacono, M. et al. Whole-genome pyrosequencing of an epidemic multidrug-resistant Acinetobacter baumannii strain belonging to the European clone II group. Antimicrob Agents Chemother 52, 2616–25 (2008).
Vallenet, D. et al. Comparative analysis of Acinetobacters: three genomes for three lifestyles. PLoS One 3, e1805 (2008).
Farrugia, D. N. et al. The complete genome and phenome of a community-acquired Acinetobacter baumannii. PLoS One 8, e58628 (2013).
Gao, F. et al. Genome sequence of Acinetobacter baumannii MDR-TJ. J Bacteriol 193, 2365–6 (2011).
Zhou, H. et al. Genomic analysis of the multidrug-resistant Acinetobacter baumannii strain MDR-ZJ06 widely spread in China. Antimicrob Agents Chemother 55, 4506–12 (2011).
Chen, C. C. et al. Genome sequence of a dominant, multidrug-resistant Acinetobacter baumannii strain, TCDC-AB0715. J Bacteriol 193, 2361–2 (2011).
Liou, M. L. et al. Genome sequence of Acinetobacter baumannii TYTH-1. J Bacteriol 194, 6974 (2012).
Loewen, P. C., Alsaadi, Y., Fernando, D. & Kumar, A. Genome Sequence of a Tigecycline-Resistant Clinical Isolate of Acinetobacter baumannii Strain AB031 Obtained from a Bloodstream Infection. Genome Announc 2, e01036 (2014).
Loewen, P. C., Alsaadi, Y., Fernando, D. & Kumar, A. Genome Sequence of an Extremely Drug-Resistant Clinical Isolate of Acinetobacter baumannii Strain AB030. Genome Announc 2, e01035 (2014).
Merino, M. et al. Complete Genome Sequence of the Multiresistant Acinetobacter baumannii Strain AbH12O-A2, Isolated during a Large Outbreak in Spain. Genome Announc 2, e01182 (2014).
Sahl, J. W. et al. Evolution of a pathogen: a comparative genomics analysis identifies a genetic pathway to pathogenesis in Acinetobacter. PLoS One 8, e54287 (2013).
Wright, M. S. et al. New insights into dissemination and variation of the health care-associated pathogen Acinetobacter baumannii from genomic analysis. MBio 5, e00963–13 (2014).
Harris, G. et al. A mouse model of Acinetobacter baumannii--associated pneumonia using a clinically isolated hypervirulent strain. Antimicrob Agents Chemother 57, 3601–13 (2013).
Fournier, P. E. et al. Comparative genomics of multidrug resistance in Acinetobacter baumannii. PLoS Genet 2, e7 (2006).
de Leseleuc, L., Harris, G., Kuolee, R., Xu, H. H. & Chen, W. Serum resistance, gallium nitrate tolerance and extrapulmonary dissemination are linked to heme consumption in a bacteremic strain of Acinetobacter baumannii. Int J Med Microbiol 304, 360–9 (2013).
Vinogradov, E., Maclean, L., Xu, H. H. & Chen, W. The structure of the polysaccharide isolated from Acinetobacter baumannii strain LAC-4. Carbohydr Res 390, 42–45 (2014).
Ahmed-Bentley, J. et al. Gram-negative bacteria that produce carbapenemases causing death attributed to recent foreign hospitalization. Antimicrob Agents Chemother 57, 3085–91 (2013).
Dijkshoorn, L. et al. Correlation of typing methods for Acinetobacter isolates from hospital outbreaks. J Clin Microbiol 31, 702–5 (1993).
Poirel, L., Menuteau, O., Agoli, N., Cattoen, C. & Nordmann, P. Outbreak of extended-spectrum beta-lactamase VEB-1-producing isolates of Acinetobacter baumannii in a French hospital. J Clin Microbiol 41, 3542–7 (2003).
Gogou, V. et al. Evolution of multidrug-resistant Acinetobacter baumannii clonal lineages: a 10 year study in Greece (2000-09). J Antimicrob Chemother 66, 2767–72 (2011).
Longo, B. et al. An outbreak of Acinetobacter baumannii in an intensive care unit: epidemiological and molecular findings. J Hosp Infect 64, 303–5 (2006).
Bertini, A. et al. Characterization and PCR-based replicon typing of resistance plasmids in Acinetobacter baumannii. Antimicrob Agents Chemother 54, 4168–77 (2010).
Chattoraj, D. K. Control of plasmid DNA replication by iterons: no longer paradoxical. Mol Microbiol 37, 467–76 (2000).
Segal, H. & Elisha, B. G. Characterization of the Acinetobacter plasmid, pRAY and the identification of regulatory sequences upstream of an aadB gene cassette on this plasmid. Plasmid 42, 60–6 (1999).
Hamidian, M., Nigro, S. J. & Hall, R. M. Variants of the gentamicin and tobramycin resistance plasmid pRAY are widely distributed in Acinetobacter. J Antimicrob Chemother 67, 2833–6 (2012).
Wang, S. et al. Transcriptomic response of Escherichia coli O157:H7 to oxidative stress. Appl Environ Microbiol 75, 6110–23 (2009).
Huang, H. et al. Complete genome sequence of Acinetobacter baumannii MDR-TJ and insights into its mechanism of antibiotic resistance. J Antimicrob Chemother 67, 2825–32 (2012).
Heritier, C., Poirel, L. & Nordmann, P. Cephalosporinase over-expression resulting from insertion of ISAba1 in Acinetobacter baumannii. Clin Microbiol Infect 12, 123–30 (2006).
Ramirez, M. S. & Tolmasky, M. E. Aminoglycoside modifying enzymes. Drug Resist Updat 13, 151–71 (2010).
Intorne, A. C., de Oliveira, M. V., de, M. P. L. & de Souza Filho, G. A. Essential role of the czc determinant for cadmium, cobalt and zinc resistance in Gluconacetobacter diazotrophicus PAl 5. Int Microbiol 15, 69–78 (2012).
Valencia, E. Y., Braz, V. S., Guzzo, C. & Marques, M. V. Two RND proteins involved in heavy metal efflux in Caulobacter crescentus belong to separate clusters within proteobacteria. BMC Microbiol 13, 79 (2013).
Vila, J., Marti, S. & Sanchez-Cespedes, J. Porins, efflux pumps and multidrug resistance in Acinetobacter baumannii. J Antimicrob Chemother 59, 1210–5 (2007).
Coyne, S., Courvalin, P. & Perichon, B. Efflux-mediated antibiotic resistance in Acinetobacter spp. Antimicrob Agents Chemother 55, 947–53 (2011).
Coyne, S., Rosenfeld, N., Lambert, T., Courvalin, P. & Perichon, B. Overexpression of resistance-nodulation-cell division pump AdeFGH confers multidrug resistance in Acinetobacter baumannii. Antimicrob Agents Chemother 54, 4389–93 (2010).
Hornsey, M. et al. AdeABC-mediated efflux and tigecycline MICs for epidemic clones of Acinetobacter baumannii. J Antimicrob Chemother 65, 1589–93 (2010).
Hou, P. F., Chen, X. Y., Yan, G. F., Wang, Y. P. & Ying, C. M. Study of the correlation of imipenem resistance with efflux pumps AdeABC, AdeIJK, AdeDE and AbeM in clinical isolates of Acinetobacter baumannii. Chemotherapy 58, 152–8 (2012).
Marchand, I., Damier-Piolle, L., Courvalin, P. & Lambert, T. Expression of the RND-type efflux pump AdeABC in Acinetobacter baumannii is regulated by the AdeRS two-component system. Antimicrob Agents Chemother 48, 3298–304 (2004).
Yoon, E. J., Courvalin, P. & Grillot-Courvalin, C. RND-type efflux pumps in multidrug-resistant clinical isolates of Acinetobacter baumannii: major role for AdeABC overexpression and AdeRS mutations. Antimicrob Agents Chemother 57, 2989–95 (2013).
Sun, J. R. et al. A truncated AdeS kinase protein generated by ISAba1 insertion correlates with tigecycline resistance in Acinetobacter baumannii. PLoS One 7, e49534 (2012).
Rosenfeld, N., Bouchier, C., Courvalin, P. & Perichon, B. Expression of the resistance-nodulation-cell division pump AdeIJK in Acinetobacter baumannii is regulated by AdeN, a TetR-type regulator. Antimicrob Agents Chemother 56, 2504–10 (2012).
Russo, T. A. et al. The K1 capsular polysaccharide of Acinetobacter baumannii strain 307-0294 is a major virulence factor. Infect Immun 78, 3993–4000 (2010).
Haseley, S. R. & Wilkinson, S. G. Structural studies of the putative O-specific polysaccharide of Acinetobacter baumannii O24 containing 5,7-diamino-3,5,7,9-tetradeoxy-L-glycero-D-galacto-nonulosonic acid. Eur J Biochem 250, 617–23 (1997).
Hu, D., Liu, B., Dijkshoorn, L., Wang, L. & Reeves, P. R. Diversity in the major polysaccharide antigen of Acinetobacter baumannii assessed by DNA sequencing and development of a molecular serotyping scheme. PLoS One 8, e70329 (2013).
Morrison, M. J. & Imperiali, B. The renaissance of bacillosamine and its derivatives: pathway characterization and implications in pathogenicity. Biochemistry 53, 624–38 (2014).
Post, D. M. et al. O-antigen and core carbohydrate of Vibrio fischeri lipopolysaccharide: composition and analysis of their role in Euprymna scolopes light organ colonization. J Biol Chem 287, 8515–30 (2012).
Tsvetkov, Y. E., Shashkov, A. S., Knirel, Y. A. & Zahringer, U. Synthesis and identification in bacterial lipopolysaccharides of 5,7-diacetamido-3,5,7,9-tetradeoxy-D-glycero-D-galacto- and -D-glycero-D-talo-non-2-ulosonic acids. Carbohydr Res 331, 233–7 (2001).
Haley, K. P. & Skaar, E. P. A battle for iron: host sequestration and Staphylococcus aureus acquisition. Microbes Infect 14, 217–27 (2012).
Wilks, A. & Burkhard, K. A. Heme and virulence: how bacterial pathogens regulate, transport and utilize heme. Nat Prod Rep 24, 511–22 (2007).
Hom, K. et al. Small molecule antivirulents targeting the iron-regulated heme oxygenase (HemO) of P. aeruginosa. J Med Chem 56, 2097–109 (2013).
Skaar, E. P., Gaspar, A. H. & Schneewind, O. IsdG and IsdI, heme-degrading enzymes in the cytoplasm of Staphylococcus aureus. J Biol Chem 279, 436–43 (2004).
Penwell, W. F., Arivett, B. A. & Actis, L. A. The Acinetobacter baumannii entA gene located outside the acinetobactin cluster is critical for siderophore production, iron acquisition and virulence. PLoS One 7, e36493 (2012).
Gaddy, J. A. et al. Role of Acinetobactin-mediated Iron Acquisition Functions in the Interaction of Acinetobacter baumannii ATCC 19606T with Human Lung Epithelial Cells, Galleria mellonella Caterpillars and Mice. Infect Immun 80, 1015–24 (2012).
Antunes, L. C., Imperi, F., Towner, K. J. & Visca, P. Genome-assisted identification of putative iron-utilization genes in Acinetobacter baumannii and their distribution among a genotypically diverse collection of clinical isolates. Res Microbiol 162, 279–84 (2010).
Turton, J. F., Gabriel, S. N., Valderrey, C., Kaufmann, M. E. & Pitt, T. L. Use of sequence-based typing and multiplex PCR to identify clonal lineages of outbreak strains of Acinetobacter baumannii. Clin Microbiol Infect 13, 807–15 (2007).
Diancourt, L., Passet, V., Nemec, A., Dijkshoorn, L. & Brisse, S. The population structure of Acinetobacter baumannii: expanding multiresistant clones from an ancestral susceptible genetic pool. PLoS One 5, e10034 (2010).
Gao, F. & Zhang, C. T. Ori-Finder: a web-based system for finding oriCs in unannotated bacterial genomes. BMC Bioinformatics 9, 79 (2008).
Delcher, A. L., Bratke, K. A., Powers, E. C. & Salzberg, S. L. Identifying bacterial genes and endosymbiont DNA with Glimmer. Bioinformatics 23, 673–9 (2007).
Srivastava, A., Singhal, N., Goel, M., Virdi, J. S. & Kumar, M. CBMAR: a comprehensive beta-lactamase molecular annotation resource. Database (Oxford) 2014, bau111 (2014).
Schattner, P., Brooks, A. N. & Lowe, T. M. The tRNAscan-SE, snoscan and snoGPS web servers for the detection of tRNAs and snoRNAs. Nucleic Acids Res 33, W686–9 (2005).
Lagesen, K. et al. RNAmmer: consistent and rapid annotation of ribosomal RNA genes. Nucleic Acids Res 35, 3100–8 (2007).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25, 3389–402 (1997).
Chen, L., Xiong, Z., Sun, L., Yang, J. & Jin, Q. VFDB 2012 update: toward the genetic diversity and molecular evolution of bacterial virulence factors. Nucleic Acids Res 40, D641–5 (2012).
Ou, H. Y. et al. MobilomeFINDER: web-based tools for in silico and experimental discovery of bacterial genomic islands. Nucleic Acids Res 35, W97–W104 (2007).
Dhillon, B. K., Chiu, T. A., Laird, M. R., Langille, M. G. & Brinkman, F. S. IslandViewer update: Improved genomic island discovery and visualization. Nucleic Acids Res 41, W129–32 (2013).
Siguier, P., Perochon, J., Lestrade, L., Mahillon, J. & Chandler, M. ISfinder: the reference centre for bacterial insertion sequences. Nucleic Acids Res 34, D32–6 (2006).
Roberts, A. P. et al. Revised nomenclature for transposable genetic elements. Plasmid 60, 167–73 (2008).
Medema, M. H., Takano, E. & Breitling, R. Detecting sequence homology at the gene cluster level with MultiGeneBlast. Mol Biol Evol 30, 1218–23 (2013).
Shao, Y. et al. mGenomeSubtractor: a web-based tool for parallel in silico subtractive hybridization analysis of multiple bacterial genomes. Nucleic Acids Res 38, W194–200 (2010).
CLSI. Methods for dilution antimicrobial susceptibility tests for bacteria that grow aerobically; Approved Standard - Ninth Edition. M07-A9, (Clinical and Laboratory Standards Institute, Wayne, PA, 2012).
Acknowledgements
This work was supported by grants from the Army Research Office (W911NF-12-1-0059) of U.S. Department of Defense, the 973 program, Ministry of Science and Technology, China (2015CB554202), the National Natural Science Foundation of China (31170083, 31170082 and 31130068), the Specialized Research Fund for the Doctoral Program of Higher Education, China (20130073110062), Department of Homeland Security (2009-ST-062-000018) and California State University Program for Education and Research in Biotechnology (the 2011 Entrepreneurial Joint-Venture grant). S.N.K., B.M.M. and P.J.E. were supported by scholarships as part of the grant from Army Research Office (W911NF-12-1-0059) and B.M.M. was also supported by NIH Minority Biomedical Research Support-Research Initiative for Scientific Enhancement (MBRS-RISE) Program (5R25GM061331). We thank the team of curators of the Institut Pasteur Acinetobacter MLST system for curating the data and making them publicly available at http://pubmlst.org/abaumannii/. We also thank technical assistance from S. Valentine, L. Real, D. Meas, W. Warner and M. C. Chong.
Author information
Authors and Affiliations
Contributions
H.H.X., X.H., H.Y.O. and W.C. conceived the study. X.H., S.N.K., B.M.M. and P.J.E. performed experiments. H.Y.O. and H.H.X. performed genomics and bioinformatics analyses. H.H.X., H.Y.O., W.C., X.H., Z.D. and M.O. wrote the paper. All authors read and approved the final manuscript.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Supplementary Information
Supplementary Information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Ou, HY., Kuang, S., He, X. et al. Complete genome sequence of hypervirulent and outbreak-associated Acinetobacter baumannii strain LAC-4: epidemiology, resistance genetic determinants and potential virulence factors. Sci Rep 5, 8643 (2015). https://doi.org/10.1038/srep08643
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep08643
- Springer Nature Limited
This article is cited by
-
Molecular and virulence characteristics of carbapenem-resistant Acinetobacter baumannii isolates: a prospective cohort study
Scientific Reports (2023)
-
Subtypes, resistance and virulence platforms in extended-drug resistant Acinetobacter baumannii Romanian isolates
Scientific Reports (2021)
-
A Slam-dependent hemophore contributes to heme acquisition in the bacterial pathogen Acinetobacter baumannii
Nature Communications (2021)
-
Characterization of carbapenem-resistant hypervirulent Acinetobacter baumannii strains isolated from hospitalized patients in the mid-south region of China
BMC Microbiology (2020)
-
Genetic mechanisms of antibiotic resistance and virulence in Acinetobacter baumannii: background, challenges and future prospects
Molecular Biology Reports (2020)