Abstract
In human, Lymphoblastoid cell lines (LCLs) from the CEPH/CEU (Centre d'Etude du Polymorphisme Humain – Utah) family resource have been extensively used for examining the genetics of gene expression levels. However, we noted that CEU/CEPH cell lines were collected and transformed approximately thirty years ago, much earlier than the other cell lines from the pertaining individuals, which we suspected could potentially affect gene expression, data analysis and results interpretation. In this study, by analyzing RNA sequencing data of CEU and the other three European populations as well as an African population, we systematically examined and evaluated the potential confounding effect of LCL age on gene expression levels and patterns. Our results indicated that gene expression profiles of CEU samples have been biased by the older age of CEU cell lines. Interestingly, most of CEU-specific expressions are associated with functions related to cell proliferation, which are more likely due to older age of cell lines than intrinsic characters of the population. We suggested the results be carefully explained when CEU LCLs are used for transcriptomic data analysis in future studies.
Similar content being viewed by others
Introduction
As a spontaneous replicating source of normal cells or DNA from a single individual, Lymphoblastoid cell lines (LCLs) have been widely used and substantially accelerated the process of biological investigations. In human genetics and genomics, LCLs provide a constant supply of DNA material for variety of assays and studies. For example, LCLs were applied as tools in vitro for evaluating drug targets and pathways1, also in vitro cell model for pharmacogenomic studies exploring genetic variation by drug dosage or cytotoxicity2. Besides, LCLs have provided unlimited genetic materials for human genetic studies3 and for human population genetic studies to characterize genetic variation of different individuals from multiple populations4,5,6. Especially, LCL derived DNA from the Coriell Cell Repositories (http://ccr.coriell.org/) were used for whole-genome genotyping and sequencing in the International HapMap Project and the 1000 Genome Project, the two largest international collaborative efforts in human genomics field since the Human Genome Project. Apart from genomics studies, LCL derived RNA was also used recently for gene expression studies. For example, Epstein–Barr virus-immortalized lymphoblastoid cell lines from the CEPH/CEU (Centre d'Etude du Polymorphisme Humain – Utah) family resource have been used for examining the genetics of gene expression levels5,7. Especially, several recent papers reported differential gene expression between CEU and YRI (Yoruba in Ibadan, Nigeria)8,9,10.
However, CEU/CEPH cell lines were collected and transformed much earlier than the other cell lines from the pertaining individuals11,12, which we suspected could potentially affect gene expression. Indeed, some previous studies reported that the older age of CEU cell lines compared to those more recently established cell lines could bias gene expression heterogeneity between populations9. In this study, taking advantage of the availability of RNA sequencing (RNA-Seq) data which allow for relatively unbiased measurements of expression levels across the entire length of transcripts7, we systematically examined and evaluated the potential confounding effect of LCL age on gene expression levels and patterns. This dataset is ideal to address the question we asked in this study. On the one hand, RNA-Seq data of three European populations (GBR, TSI and FIN), which are genetically close to CEU but the LCLs of these three population samples were established very recently, can be used as very good controls to examine whether and to what extent the gene expression profile of CEU deviated from normal level. On the other hand, analysis can be done for verification of the results reported by previous studies based on microarray data. Especially the differential gene expression between CEU and YRI identified by previous studies can be re-examined in the new RNA-Seq data (see Materials and Methods).
Results
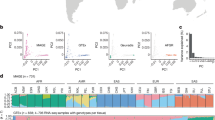
We analyzed both RNA-Seq data and DNA sequence data obtained from an identical set of samples (462 individuals, Table 1) representing the four European populations (EUR), i.e. Utah residents with Northern and Western European ancestry from the CEPH collection (CEU, n = 91), Tuscans from Italy (TSI, n = 93), Finnish in Finland (FIN, n = 95) and British in England and Scotland (GBR, n = 94) and one African population, i.e. Yoruba in Ibadan, Nigeria (YRI, n = 89). When we compared gene expression level between populations, we found that CEU-YRI pair showed larger differences (measured by VST based on RNA-Seq data) than those non-CEU-YRI pairs. Similarly, CEU-EUR pairs showed larger VST compared to other EUR pairs while genetic difference (measured by FST based on DNA variation data) between any European population pairs was very small. These results indicated CEU could have a different expression profile compared with the other European populations. In addition, we observed a strong positive correlation between expression differentiation (VST) and genetic differentiation (FST) (Figure 1A, r2 = 0.8, p < 0.01). This correlation between expression differentiation and genetic differentiation was even much stronger when CEU samples were excluded from the analysis (r2 = 0.98, p < 0.01, Figure 1B). Correspondingly, we did observe that population pairs with CEU involved showed apparent deviation from the correlation relationship (Figure 1A). Therefore, it seemed that the gene expression profile of CEU could be different from those of the other European populations, despite the overall transcriptomic profile of CEU is unexpected to be significantly different from those of the other European populations given the small genetic difference among European populations (mean FST = 0.005, Figure 1A). Indeed, our further analysis did reveal a significant deviation of gene expression profile of CEU from those of the other three European populations (TSI, FIN and GBR) (Figure 1C, t test p < 2.2e-16).
(A) Distribution of genetic differentiation (FST) and expression differentiation (VST) between each pair of the five populations (CEU/TSI/FIN/GBR/YRI). The asterisks represent population pairs CEU involved and the triangles represent non-CEU involved pairs. The four red markers on the upper panel show the population pairs between YRI and European populations, the blue markers on the bottom right panel show the population pairs between CEU and non-CEU European populations and the gray markers on the bottom left panel show the population pairs between non-CEU European populations. The gray dashed line represents the regression line between FST and VST for the 10 population pairs. The correlation between the mean VST values and mean FST values is shown above the plot. (B) Distribution of genetic differentiation (FST) and expression differentiation (VST) between each pair of the four non-CEU populations (TSI/FIN/GBR/YRI). The three red markers on the upper panel show the population pairs between YRI and European populations and the three gray markers on the bottom panel show the population pairs between non-CEU European populations. The gray dashed line represents the regression line between FST and VST for the 6 population pairs. The correlation between the mean VST values and mean FST values is shown above the plot. (C) Distribution of VST between each pair of European populations. The red solid, dashed and dotted lines represent population pairs between CEU and three other non-CEU Europeans (TSI/FIN/GBR). The blue solid, dashed and dotted lines represent population pairs between three non-CEU Europeans (TSI/FIN/GBR). (D) Venn diagram of DE genes. The yellow circle represents the number of DE genes between CEU and other non-CEU European populations (TSI/FIN/GBR) and the purple circle represents the number of DE genes between any two non-CEU European populations (TSI/FIN/GBR).
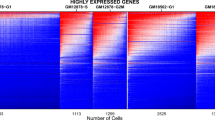
The above results suggested that some special factors contributed to the unique expression profile of CEU because the gene expression difference could not be explained by the small genetic difference between CEU and the other European populations (mean FST = 0.005). Spirited by this understanding and sense, we identified 2,420 genes showing significant expression differentiation between CEU and the other three European populations but no significant expression differentiation among the non-CEU European populations (TSI/FIN/GBR) (Figure 1D). We further performed functional annotation and enrichment analysis of these 2,420 differentially expressed (DE) genes between CEU and the other European populations (Figure 1D), taking all the 14,178 expressed genes as a background control. Notably, we identified a series of cell specific GO (Gene Ontology) functions including endomembrane system, Golgi vesicle transport, intracellular organelle part, cell, cytoplasmic part and cellular response to topologically incorrect protein, etc. (Bonferroni-corrected p < 0.01, see Materials and Methods, Table 2). Since these DE genes are enriched to functions related to cell secretion and cell proliferation, which played an important role in cell subculture, it is intuitive and reasonable to attribute CEU-specific gene expressions to the relative older age of CEU cell lines.
Further, we performed a comparative analysis of RNA sequencing data between African (YRI) and European populations. In this way, YRI was used as an outgroup to reduce the potential background noise in comparing gene expression data in closely related populations. Interestingly, the DE genes between CEU and YRI were over-represented in the 2,420 genes showing CEU-specific gene expressions compared with DE genes between non-CEU European (TSI/FIN/GBR) and YRI (one side Fisher exact test p = 0, Figure 2). These results again indicated that a substantial proportion of DE genes between CEU and YRI (around 24%, Figure 2) were unlikely due to genetic difference between the two populations, but were instead likely to be resulted from old age of cell lines. Finally, we compared the DE genes identified between CEU and YRI based on RNA-Seq data with the previously reported DE genes between CEU and YRI based on microarray platform8. As a result, we found that 31 DE genes reported by previous studies, as representative signatures of differential gene expression between African and European populations, could be false positive results due to older age of CEU cell lines, i.e. TRIP4, PITPNB, TAOK3, SAMD8, GOLGA7, PTPN12, ACOT9, FTSJ3, C19orf12, UTP14A, CLEC2D, RNF170, ALG11, UTP14C, HOOK3, YES1, PPP3CC, SERPINB9, PPHLN1, SYNCRIP, TM9SF3,GBP1, SFMBT1, FMR1, PAPLN, SNTB1, OXER1, TNFRSF13B, FAM91A1, KIAA1033, SLC39A8. Therefore, we suggested caution be paid to these genes on expression analysis involving LCLs from the CEPH/CEU family resource in future work.
Venn diagram of DE genes.
The big and red circles represent the number of DE genes between CEU and YRI. The green circles represent the 2420 genes described in Figure 1D. The blue circles represent the number of DE genes between YRI and the other non-CEU European populations: TSI, FIN and GBR respectively in (A)–(C).
Discussion
This study was initially motivated by an observation in our data analysis of apparent deviation of gene expression profile of CEU samples from those of the other populations including several European populations which are genetically very closely related to CEU. Lymphoblastoid cell lines (LCLs) are a resource that provides investigators with the nearly unique opportunity to perform in-depth studies of molecular and complex phenotypes using the same collection of samples. However, results based on LCLs in human genetics can be sometimes controversial. The transformation that immortalized the LCLs, through the infection of primary B cells with EBV, was known to result in certain artifacts13. Cell lines that often carry chromosomal abnormalities14 might have pronounced batch effects related to preparation and/or growth rates13 and the Epstein–Barr virus (EBV) transformation itself could alter the methylation status15 and expression levels of a subset of genes16,17. Most importantly, during the long-term subculture, genotypic errors were incorporated mostly in late-passage, but not in early-passage LCLs18. We noted that CEU/CEPH cell lines were collected and transformed approximately thirty years ago, much earlier than the other cell lines from the pertaining individuals, which we suspected could potentially affect gene expression. Indeed, it has been reported that the greater number of the validated non-germline mutations in the CEU cell line perhaps reflected the greater age of the CEU cell culture4. Since gene expression could be treated as an important heritable trait5,19,20, it was expected to detect a profound difference in the gene expression profiles between newly established and mature LCLs21, also the older age of CEU cell lines compared to those more recently established cell lines could bias gene expression heterogeneity between populations13,22. Although this unwanted bias caused by the age of the cell lines has been noted before in the literature, few studies have systematically evaluated the influence of this effect on gene expression patterns. To this end, we took advantage of recently available RNA-Seq data and methods that allowed us to address this question. We found apparent deviation of gene expression profile of CEU samples from those of the other populations including several European populations which were genetically very closely related to CEU. Therefore, it was reasonable to infer that gene expression level and pattern of CEU cell lines have been biased by the older age of CEU cell lines, which would spark concern about CEPH cell lines. However, the CEU-specific expression could also be the result of environment effects or gene by environment interactions, as suggested previously23. So, our analysis did not rule out the possibility that other factors might also affect gene expression levels. We emphasize here, however, that our current study is not a comprehensive one to access the relationship between cell line age and gene expression, but rather provide some warning messages for interpretation of the results of previous studies based on CEU cell lines and useful information for future study design.
In addition, we identified 2,420 genes whose expression levels are highly differentiated between CEU and the other European populations and probably associated with old age of CEU cell lines. Notably, these 2,420 genes showing CEU-specific expression were enriched in the 3,259 genes reported as eQTL in European populations (CEU/GBR/TSI/FIN) in the previous study7 (one side fisher exact test p = 0.047), compared to 501 eQTL genes in YRI population. These results could be best explained by some special characters in CEU cell lines which were very likely due to the earlier established time of CEU cell lines. Since there is no easy way to correct the biased gene expression in CEU empirically or statistically, what we could do is avoiding the possible false-positive results by referring genes as we have already listed in this paper and do not over-interpret the differential gene expression if they are found in the comparisons between CEU and other populations. In brief, we suggested these CEU-specific gene expression be explained with caution and this issue of cell line age be carefully considered in the analysis of CEU gene expression data, especially when CEU LCLs were used for transcriptomic data analysis in future studies.
Methods
RNA-Seq data and gene expression quantification
We downloaded an RNA-Sequencing dataset from ArrayExpress, which spanned the whole genome expression data in transformed lymphoblastoid cell lines (LCLs) obtained from 5 populations (in total 462 samples, Table 1) with different ancestries: Utah residents with Northern and Western European ancestry (CEU, 91 samples), British in England and Scotland (GBR, 94 samples), Tuscans in Italy (TSI, 93 samples), Finnish in Finland (FIN, 95 samples) and Yoruba in Ibadan, Nigeria (YRI, 89 samples), respectively7. The sample sizes for two sexes are nearly equal (Table 1). Reads mapping and quality control were fulfilled by the original study7, which resulted in 57,195 Ensembl genes in total. Then, we quantified reads for the whole transcripts and each quantification was filtered to exclude those with missing data for >10% of the individuals in all of the five populations. This resulted in 14,178 Ensembl genes and 111,120 transcripts on 22 autosomes and X chromosome. The transcript read counts were subsequently normalized by per kilobase per million reads (RPKM) measure.
Replication of gene expression differences between CEU and YRI on a microarray platform
We checked whether the RNA-Seq dataset could replicate gene expression differences between CEU and YRI in a previous study8, the gene expression data of which were generated with the Affymetrix GeneChip Human Exon 1.0 ST array and it reported 383 differential transcript clusters between the CEU and YRI samples, of which 306 were located in protein coding region. We could replicate 266 (87%) of them based on Benjamini-Hochberg corrected t test (p < 0.05).
Genotype data and estimation of genetic differentiation between populations
Single nucleotide polymorphisms (SNPs) data of the same 462 samples was obtained from the 1000 Genomes Project Phase I dataset24. In each population, only the polymorphic SNPs on 22 autosomes and X chromosome were included. For each SNP, genetic difference between populations was measured with the commonly used FST according to Wright's approximate formula25. FST value was calculated based on allele frequencies estimated from unrelated individuals in each population.
Quantification of expression difference between populations
To quantify population differentiation with respect to expression levels, we calculated VST for each of the transcript between any two of the four populations. VST is a measure of the proportion of variance on expression level explained by between-population differences and is analogous to the commonly used population genetics parameter FST, which measures allele frequency differences between populations. For a single transcript compared between two populations, VST is calculated as: (VT – VS)/VT, where VT is the total variance across all individuals of the pair of populations and VS is the average within-population variance weighted by each population sample size. VS = (V1*n1 + V2*n2)/(n1 + n2), where V1 is the within-population variance of population 1, V2 is the within population variance of population 2 and n1 and n2 are the numbers of individuals sampled from population 1 and 2, respectively. VST values range from 0 to 1, with values near 1 signifying that the majority of gene expression variance for a transcript segregates between populations rather than within populations.
Statistical test for differential expression between populations
For each transcript, we applied Shapiro-Wilk test of expression levels' normality, with the cutoff of Bonferroni-corrected p < 0.01. We found the majority of transcripts followed normal distribution in gene expression levels for all the 5 populations (92%, 96%, 95%, 94% and 94% transcripts for CEU, GBR, FIN, TSI and YRI, respectively). Therefore, T-test was applied to test whether the transcript expressions were significantly different between any two populations, with the cutoff of Bonferroni-corrected p < 0.01.
Functional annotation and enrichment analysis of differential expression
Due to the unique properties of the RNA-Seq data, the differential expression of longer transcripts is more likely to be identified than that of shorter transcripts with the same effect size26. This transcript length bias complicates the downstream GO analysis27. To correct this confounding effect, we applied “GOseq”27 to perform functional annotation and enrichment analysis of these 2,420 differentially expressed (DE) genes between CEU and the other European populations (Figure 1D), using all the 14,178 expressed genes as a comparison background.
References
Morag, A., Kirchheiner, J., Rehavi, M. & Gurwitz, D. Human lymphoblastoid cell line panels: novel tools for assessing shared drug pathways. Pharmacogenomics 11, 327–340, 10.2217/pgs.10.27 (2010).
Li, L. et al. Gemcitabine and cytosine arabinoside cytotoxicity: association with lymphoblastoid cell expression. Cancer Res 68, 7050–7058, 10.1158/0008-5472.CAN-08-0405 (2008).
Kasowski, M. et al. Variation in transcription factor binding among humans. Science 328, 232–235, 10.1126/science.1183621 (2010).
Abecasis, G. R. et al. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073, 10.1038/nature09534 (2010).
Spielman, R. S. et al. Common genetic variants account for differences in gene expression among ethnic groups. Nat Genet 39, 226–231, 10.1038/ng1955 (2007).
Stranger, B. E. et al. Relative impact of nucleotide and copy number variation on gene expression phenotypes. Science 315, 848–853, 10.1126/science.1136678 (2007).
Lappalainen, T. et al. Transcriptome and genome sequencing uncovers functional variation in humans. Nature 501, 506–511, 10.1038/nature12531 (2013).
Zhang, W. et al. Evaluation of genetic variation contributing to differences in gene expression between populations. Am J Hum Genet 82, 631–640, 10.1016/j.ajhg.2007.12.015 (2008).
Stranger, B. E. et al. Population genomics of human gene expression. Nat Genet 39, 1217–1224, 10.1038/ng2142 (2007).
Storey, J. D. et al. Gene-expression variation within and among human populations. Am J Hum Genet 80, 502–509, 10.1086/512017 (2007).
Dausset, J. et al. Centre d'etude du polymorphisme humain (CEPH): collaborative genetic mapping of the human genome. Genomics 6, 575–577 (1990).
Integrating ethics and science in the International HapMap Project. Nature reviews. Genetics 5, 467–475, 10.1038/nrg1351 (2004).
Akey, J. M., Biswas, S., Leek, J. T. & Storey, J. D. On the design and analysis of gene expression studies in human populations. Nat Genet 39, 807–808; author reply 808-809 10.1038/ng0707-807 (2007).
Redon, R. et al. Global variation in copy number in the human genome. Nature 444, 444–454, 10.1038/nature05329 (2006).
Hannula, K. et al. Maternal and paternal chromosomes 7 show differential methylation of many genes in lymphoblast DNA. Genomics 73, 1–9, 10.1006/geno.2001.6502 (2001).
Carter, K. L., Cahir-McFarland, E. & Kieff, E. Epstein-barr virus-induced changes in B-lymphocyte gene expression. J Virol 76, 10427–10436 (2002).
Caliskan, M., Cusanovich, D. A., Ober, C. & Gilad, Y. The effects of EBV transformation on gene expression levels and methylation profiles. Hum Mol Genet 20, 1643–1652, 10.1093/hmg/ddr041 (2011).
Oh, J. H. et al. Genotype instability during long-term subculture of lymphoblastoid cell lines. J Hum Genet 58, 16–20, 10.1038/jhg.2012.123 (2013).
Emilsson, V. et al. Genetics of gene expression and its effect on disease. Nature 452, 423–428, 10.1038/nature06758 (2008).
Oleksiak, M. F., Churchill, G. A. & Crawford, D. L. Variation in gene expression within and among natural populations. Nat Genet 32, 261–266, 10.1038/Ng983 (2002).
Caliskan, M., Pritchard, J. K., Ober, C. & Gilad, Y. The effect of freeze-thaw cycles on gene expression levels in lymphoblastoid cell lines. PLoS One 9, e107166, 10.1371/journal.pone.0107166 (2014).
Stranger, B. E. et al. Population genomics of human gene expression. Nat Genet 39, 1217–1224, 10.1038/ng2142 (2007).
Idaghdour, Y., Storey, J. D., Jadallah, S. J. & Gibson, G. A genome-wide gene expression signature of environmental geography in leukocytes of Moroccan Amazighs. PLoS Genet 4, e1000052, 10.1371/journal.pgen.1000052 (2008).
Abecasis, G. R. et al. An integrated map of genetic variation from 1,092 human genomes. Nature 491, 56–65, 10.1038/nature11632 (2012).
Weir, B. S. & Cockerham, C. C. Estimating F-Statistics for the Analysis of Population-Structure. Evolution 38, 1358–1370 (1984).
Oshlack, A. & Wakefield, M. J. Transcript length bias in RNA-seq data confounds systems biology. Biol Direct 4, 14, 10.1186/1745-6150-4-14 (2009).
Young, M. D., Wakefield, M. J., Smyth, G. K. & Oshlack, A. Gene ontology analysis for RNA-seq: accounting for selection bias. Genome Biol 11, R14, 10.1186/gb-2010-11-2-r14 (2010).
Acknowledgements
These studies were supported by the Strategic Priority Research Program of the Chinese Academy of Sciences (CAS) (XDB13040100), by the National Science Foundation of China (NSFC) grants (91331204; 31171218; 31301083). S.X. is Max-Planck Independent Research Group Leader and member of CAS Youth Innovation Promotion Association. S.X. also gratefully acknowledges the support of the National Program for Top-notch Young Innovative Talents of The “Wanren jihua”. All funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
S.X. conceived and designed the study; Y.Y., L.T. and D.L. analyzed the data; S.X. and Y.Y. wrote the paper.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Yuan, Y., Tian, L., Lu, D. et al. Analysis of Genome-Wide RNA-Sequencing Data Suggests Age of the CEPH/Utah (CEU) Lymphoblastoid Cell Lines Systematically Biases Gene Expression Profiles. Sci Rep 5, 7960 (2015). https://doi.org/10.1038/srep07960
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep07960
- Springer Nature Limited