Abstract
Plant-soil feedbacks (PSFs) are suggested to be major drivers of plant species coexistence and exotic invasions in natural plant communities, where species with more positive PSFs are thought to be more abundant in communities. Most evidence for this comes from mesocosm experiments with single species, but whether the results are transposable to diverse plant communities is mostly not verified and remains debated. We performed a combined monoculture and community experiment to test whether PSFs in monocultures predict PSFs in communities, and to infer the role of PSFs in invasive plant success. We found that (1) PSFs from monocultures were poor predictors for PSFs in plant communities, (2) competitive strength of invasive species did not consistently depend on PSF, and (3) dominant species experienced a significantly stronger negative PSFs than non-dominant species when grown in community. Hence, PSFs of plant species in monocultures seem less predictive for their abundance in plant communities or for invasibility than previously assumed. Nevertheless, PSF—and particularly negative PSF—seems indeed a major driver of plant species coexistence, with a strong species-specific pathogenic effect on dominant plants facilitating the persistence of rare species.
Similar content being viewed by others
Introduction
Plant-soil feedbacks (PSF) have been suggested to be an important regulating mechanism for plant species dynamics in grassland communities1,2. The accumulation of mutualistic mycorrhizal fungi in the rhizosphere induces positive PSFs around conspecific plants3, while soil-borne pathogens can induce negative PSFs by hampering their growth or that of their offspring4,5,6. These species-specific pathogens are one of the major drivers of species richness-productivity relationships7,8. PSF has also been proposed to influence community structure by benefitting the establishment of invasive species through positive feedbacks9,10 and species coexistence through negative or positive feedbacks11,12,13. The enemy release hypothesis (ERH)14 postulates that non-native plants can become invasive because species-specific pathogens are absent in the introduced range15,16. It has indeed been shown that invasive plants often encounter more positive or less negative PSFs than natives17,18, though some studies showed the opposite19,20,21.
In what manner PSF influences plant species dominance is debated22. Some studies showed that dominant plant species have net positive PSFs or weaker negative PSFs than co-occurring rare species17,23,24, whereas other studies found that PSF has negative effects on most plants12, with dominant plant species accumulating more species-specific pathogens3, maintaining plant diversity through negative frequency dependency2,12,13. Although it has been theoretically predicted that negative PSFs can promote plant species coexistence25,26,27,28,29, supporting experimental evidence is lacking.
Most experimental setups so far measured PSF in monoculture mesocosm experiments, assigning fixed PSF values to plant species17,30,31,32, though some works have suggested that interspecific competition in communities between plants may alter individual PSFs33,34,35,36. For example, more recent experiments have shown that PSFs from monocultures are not correlated to these found in the field37,38. Hence, there is a need for community experiments12,39,40, where the interaction between competition and PSF can be measured. Some earlier experiments indeed did investigate PSF in artificial communities21,34,41,42,43, though these only included pairs of two to three species or did not focus on the effect of communities on PSF.
The aim of this research was to investigate if monoculture mesocosm experiments are good predictors for PSFs in plant communities. We therefore combined monoculture and community PSF experiments for 10 co-occurring native grassland species and three invasive species. The communities consisted of these 10 native species with or without one of the three invasive species. We either used a full microbial inoculum (PSFtot) or an extracted pathogen/saprobe filtrate (PSFpath) from the soil of the conditioning phase of the experiments to determine PSF for each species in both monoculture and community in the response phase (see Extended Data Fig. 1). A sterilized inoculum was used as control. If PSFs of species in monocultures are good predictors for PSF in communities, then this species-specific PSF might indeed influence plant dominance17. Species with more positive PSFs in monoculture will thrive more in unsterilized communities than in sterilized ones, leading to increased size or dominance, unlike species with more negative PSFs in monoculture. On the other hand, if monoculture experiments appear to be poor predictors of PSFs in community, then what factors contribute to PSFs in community that disrupt the correlation with monoculture PSFs? We hypothesize that in this alternative case, PSF in communities is affected by plant dominance44, where large, dominant species will accumulate more pathogens and thus receive a more negative PSF3. The PSF a species receives then depends on that species’ dominance within the community26,27,29. Lastly, we tested whether the enemy release hypothesis applied to the three invasive species of our experiment (Solidago gigantea, Avena sterilis and Lupinus polyphyllus). We hypothesized that these species are stronger competitors and represent higher proportions of the community biomass in unsterilized communities than in sterilized communities.
Results and discussion
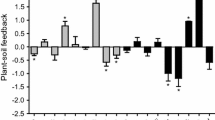
For most species both PSFtot and PSFpath were negative regardless of being grown in monoculture or community (Fig. 1, Extended Data Figs. 2–4). However, for nine out of ten species PSFpath was less negative in community than in monoculture, showing that the impeding effect of pathogens on plant growth was reduced when grown in community (Fig. 1b). These results are in line with earlier meta-analyses, where monocultures grown in the greenhouse showed more negative feedbacks than field experiments38,40. This could result from pathogens often being species-specific3,17: while in monocultures the entire pathogen fraction will be represented by the same species-specific pathogens, in more biodiverse communities pathogens specific for each plant species will occur in lower proportions. Since most mutualistic arbuscular mycorrhizal fungi are generalists45 this negative effect of higher plant species diversity is probably less relevant for positive feedbacks. Due to this asymmetry in specificity, PSF might become less negative with rising biodiversity of the plant community.
a Shows results for standardized PSFtot from both mutualists and pathogens. b Shows results for standardized PSFpath from the pathogen fraction alone. Grey areas show when PSF is more positive when grown alone than in community, for white areas PSF is more positive in communities. Significance of regression was evaluated using a linear model (see ‘Methods’). Error bars indicate standard errors, n = 5 for calculation PSF in community per species per soil treatment, only communities without addition of invasives were used; n = 6 for monocultures. Species numberings: 1. Achillea millefolium, 2. Agrostis capillaris, 3. Anthoxanthum odoratum, 4. Centaurea jacea, 5. Holcus lanatus, 6. Leucanthemum vulgare, 7. Lotus corniculatus, 8. Plantago lanceolata, 9. Rumex acetosa, 10. Trifolium pratense.
We found that PSFs measured in monocultures were not correlated to these in communities (Fig. 1a, b, resp. R² = 0.29 & 0.12, Ext. Data Fig. 2a, b, resp. R² = 0.06 & 0.03). Our results confirm that PSF measured in monoculture experiments are thus not representative for community, and consequently neither for field conditions37,38. Since most earlier works on PSF mechanisms concentrated on monoculture experiments, we see a clear need for individual PSFs from different plant species measured in a community setting, in order to fully explain how PSF and community structure influence one another. As far as we know, only one earlier study was able to show a significant correlation between PSF in monocultures and plant abundance in the field17.
We here show that larger, more dominant species (in terms of biomass) in our experimental community were exposed to significantly more negative feedbacks than smaller species (Fig. 2a, b for above-ground biomass, Extended Data Figs. 5, 6 for below-ground biomass and total biomass, for both PSFtot and PSFpath, Table 1). We attribute this dominance-driven negative PSF to pathogen fractioning, where pathogens specific for low-abundant or smaller species represent low proportions of the soil community, in contrast to pathogens specific for more dominant or larger plant species. Since this mechanism is not applicable in monocultures, PSF was here not dependent on species biomass (Extended Data Fig. 7a, b, Table 1). Some of the smaller species in the community, such as Lotus corniculatus and Centaurea jacea even showed a significantly positive PSFpath in community whereas they had a negative or insignificant PSFpath in monoculture (Fig. 1 and Extended Data Figs. 2–4). This seems to be an indirect effect of the larger species being stronger affected in communities by negative PSF, enabling a better performance of the smaller or rare species in the community in the presence of soil microbes through a reduced competition from these larger species. The PSFs in community, especially for smaller, less competitive plants, are thus a combination of the direct feedback induced by its own species-specific soil biota, and of the indirect effect from altered competition with other plant species and their PSFs. Though PSFpath can never be positive for a certain species in monoculture (since mutualists are absent), we here show that it can become positive for these small species in communities, demonstrating that the positive effects from the reduced competition can overwhelm the negative effect of their own species-specific pathogens. PSF can thus be a strong driver of community composition and species coexistence, in contrast to conclusions of some earlier works which found that PSF in community seemed to be of minor importance due to their effects being overwhelmed by other interactions37,40,43. Our results support that negative PSFs in community can promote plant coexistence and thus biodiversity through negative frequency dependency12,27. Just as PSF alters with e.g. water availability46 or nutrient availability47,48—it also changes depending on the community in which the plant occurs and its dominance within that community. This negative frequency dependency seems most significant when calculated for root biomass (Extended Data Fig. 5a, b, Table 1). This might be explained by soil pathogens being associated with the rhizosphere of plants, making root biomass the main determinant for pathogen load, thus also PSF. The effect is also significant when calculated for above-ground and total biomass, though only for PSFpath (Fig. 2b, Extended Data Fig. 6b, Table 1). The negative frequency dependency also seems to be present for the PSFtot (Fig. 2a), though not significant (P = 0.14). The presence of mutualists may obscure the directional effect of pathogens. This effect could perhaps be explained by different plant species benefitting to varying degrees of mutualistic mycorrhizal fungi49. Plant communities more dependent on mutualistic mycorrhizae, or where mutualists are more abundant, might show a reduced negative frequency dependency.
a Shows results for standardized PSFtot from both mutualists and pathogens. b Shows results for standardized PSFpath from the pathogen fraction alone. Significance of regression was evaluated using a mixed linear model (see ‘Methods’). R² indicated on the graph is the marginal R², conditional R² = 0.31. Trendline was made with the function nls, y = 0.59/x0.36 – 1, all parameters highly significant, CI = 99.5%.
Furthermore, grasses showed significantly more negative PSF than legumes or non-leguminous forbs in communities (Fig. 2a, b and Extended Data Figs. 6a, b, P < 0.001 for both PSFtot and PSFpath, Table 1), though this effect was not present in monocultures (Extended Data Fig. 7a, b). These results are in line with the meta-analysis by Kulmatiski et al.12, who attributed this to high root to shoot ratios and related to that a higher vulnerability to pathogens. Our results support these results that grasses seem to accumulate more pathogens rather than that they benefit less from mutualists, as this trend was significant for both PSFtot as for PSFpath. Other than simply accumulating more pathogens, grass-specific pathogens might be less species-specific, where pathogens can affect different closely related plant species24,50, though we are not aware of any studies showing this specifically for grasses. Lastly, though PSF is significantly negatively impacted by species size in community for all functional groups, this negative dependency on size seems to be stronger for legumes (significant interaction term between species biomass and functional group, see Table 1).
Assuming that the enemy release hypothesis applies to the three investigated invasive species of our study, we expected these species to be less negatively impacted by PSF than native species, making them stronger competitors17,18. However, the competition effect of invasive species on native species was independent of soil treatment in our study (interaction term not significant in Table 2, Fig. 3a, Extended Data Fig. 8a). Along with this, two out of three invasive species did not significantly represent a higher proportion of the total community biomass depending on soil treatment (Fig. 3b and Extended Data Fig. 8b; Table 2). The only invasive species that did represent a significant higher proportion of the community when PSF was present was Lupinus polyphyllus, which was also the only species to receive a positive PSFtot in monoculture (Extended Data Fig. 3). Hence, our results do not generally support the ERH, and are in line with other studies emphasizing that invasion processes are complex, and cannot fully be explained by the ERH alone but also depends on other factors19.
a Above-ground community biomass of 10 native plant species in response to soil treatments and presence/absence of one invasive species. Sterile = sterilized soil treatment, PSF = unsterilized soil treatment, Pathogen = sterilized soil treatment + addition of pathogen/saprobe filtrate. b Biomass of the invasive plants (% of total above-ground community biomass) in relation to soil treatments. Letters indicate significant differences depending on a the presence of an invasive species or b soil treatment, calculated with Tukey’s HSD test, α = 0.05, n = 10. Error bars indicate standard error. A. sterilis Avena sterilis, L. polyphyllus Lupinus polyphyllus, S. gigantea Solidago gigantea.
As discussed above, PSF seems to become less negative with increasing biodiversity of the plant community due to species-specific pathogens representing lower proportions of the total pathogen fraction. The validity of the ERH diminishes as the biodiversity of the invaded community increases, causing invasive species to lose their competitive edge. Invasiveness due to enemy release might thus mostly apply in communities of low biodiversity. It has already been shown theoretically and experimentally that more biodiverse communities are more resistant to invasion than communities of low diversity51,52,53. In these studies, low invasibility was predicted to result from the presence of species of many different functional guilds52 or the low levels of resources51 that co-occur with diverse communities. The reduction in negative PSF for natives and accompanying larger competition experienced by invaders due to the higher native biomass production might be an additional explanation, as it has been shown that biodiversity increases productivity54 due to a reduced species-specific pathogen load7,8. As plant-pathogen relationships are often species-specific, we predict that the biodiversity effect on plant invasions through pathogens does not saturate with increasing biodiversity due to functional similarity of species55, but is more in line with the singular hypothesis of biodiversity where each plant-pathogen interaction is unique56,57. Rather than actively driving the decrease in diversity and nutrient enrichment58,59,60,61, the invasion of some alien species may be a consequence of these disturbances43,62,63, e.g. in communities that lost diversity through nutrient enrichment, where these invasive species could again benefit from the ERH.
Our results do confirm that on a community scale PSF has a significant net negative effect on biomass production12 (Fig. 3a for above-ground biomass, Extended Data Fig. 8a for total biomass; Table 2). Besides this, the addition of an invasive plant species may also significantly reduce total native biomass (Fig. 3a and Table 2), though this was only significant for one out of three species in our experiment (A. sterilis). The other two invasives also reduced native biomass, though not significantly.
Future experimental PSF research should include community mesocosm experiments. Though results from monocultures can disentangle fundamental processes of PSF, community experiments clearly give very different results (Fig. 4a, b) and seem more representative for field conditions. Especially measurements of PSF in communities of varying biodiversity and functional composition might prove promising. These community experiments are needed to disentangle the mechanisms of how species interactions and community processes influence PSF and subsequently the ERH and invasion success. We propose future experiments should investigate the effect of PSF on invasion of non-native species in communities of differing diversity. Earlier works where a significant effect of PSF on competition of invasive plants was found, often worked with low biodiverse systems21 (often one-on-one competition), opposed to our results from a more biodiverse system. We expect that as PSF becomes less negative in more biodiverse communities, most invasive species will benefit less from the ERH and will show lower competitiveness and subsequently lower invasion success. Though labour intensive measurements, it might also prove valuable to include root mass additionally to above-ground biomass in PSF experiments as we have shown here that root mass might be more determining PSF than above-ground or total weight in community.
a Shows results for ΔPSFtot from both mutualists and pathogens. R2 indicated on the graph is the marginal R², conditional R2 = 0.87. Trendline was made with the function nls, y = 0.72/x0.28 – 1, all parameters significant, CI = 95%. b Shows results for ΔPSFpath from the pathogen fraction alone. Conditional R² = 0.43, y = 1.11/x0.25 – 1, all parameters significant, CI = 95%. Significance of regression was evaluated using a mixed linear model (see ‘Methods’). n = 10 for communities, only communities without invasives were used; n = 6 for monocultures. Error bars indicate standard errors. Species numberings: 1. Achillea millefolium, 2. Agrostis capillaris, 3. Anthoxanthum odoratum, 4. Centaurea jacea, 5. Holcus lanatus, 6. Leucanthemum vulgare, 7. Lotus corniculatus, 8. Plantago lanceolata, 9. Rumex acetosa, 10. Trifolium pratense.
Methods
Monoculture experiment
In order to measure the PSFs that plant species experience when grown alone, a monoculture experiment was set up with 10 native species, typically co-occurring in moist mesotrophic European grasslands (Achillea millefolium, Centaurea jacea, Leucanthemum vulgare (Asteraceae), Lotus corniculatus, Trifolium pratense (Fabaceae), Plantago lanceolata (Plantaginaceae), Agrostis capillaris, Anthoxanthum odoratum, Holcus lanatus (Poaceae), Rumex acetosa (Polygonaceae)) from 3 different functional groups (grasses (3 species), leguminous (2) and non-leguminous species (5)). In addition, 3 exotic invasive species in mesotrophic grasslands in Europe (Avena sterilis (Poaceae), Lupinus polyphyllus (Fabaceae), Solidago gigantea (Asteraceae)) were included in the study, one from each of the functional groups mentioned above. Seeds of natives and of L. polyphyllus were obtained from Cruydt-Hoeck (The Netherlands), seeds of A. sterilis from B&T World Seeds (France), and of S. gigantea were collected in the region of Brussels during the autumn of 2021.
The experiment was split into two phases: the conditioning phase, where plants are subjugated to soil microbial communities and when (species-specific) mutualists and pathogens could accumulate, and the response phase, where plants of the same species were grown in the conditioned soil under different soil treatments (Extended Data Fig. 1, see further for the different soil treatments). During the conditioning phase, seeds were germinated on universal potting soil (pH = 6.0, Viano, Belgium) for one to three weeks depending on species, after which they were transplanted into 0.5 l pots (10 × 8 cm, Soparco, France). Plants were grown separately in a sand/inoculum mixture (95-5 vol%) during eight weeks (March and April 2022) in a greenhouse in a randomized setup. The sand fraction did not contain any organic matter, was composed 100% of dried quartz sand, and concentrations of N and P were below detection limits (Type M31, Sibelco NV, Belgium). The inoculum was collected from a moist mesotrophic grassland in a Belgian nature reserve (Doode Bemde; 50.815509, 4.644803). Soil samples (0–10 cm deep) were taken randomly from 10 different locations within this grassland and thoroughly mixed. This soil was stored for 3 days at 5 °C before usage. Plants received a half-strength Hoagland solution17 per week, i.e. each plant individual received 2.5 ml per week in the first two weeks, 5 ml per week in the next four weeks and 7.5 ml per week in the last two weeks. At the end of the conditioning phase each individual received a total of 30.0 mg N, 2.0 mg P, 222.0 mg K, 55.3 mg Ca, 31.5 mg Mg, 4.9 mg Fe, 0.03 mg Cu, 0.40 mg B, 0.24 mg Mn, 0.12 mg Zn and 0.07 mg Mo64. Pots were watered upon demand two to three times per week into the saucers to avoid leaching out of pathogens and nutrients. At the end of the conditioning phase plants were harvested and disposed. The conditioned soil was used for the response phase.
Three different treatments were applied to the soil: (i) Sterilized soil, by steam sterilization for 1 h at 121 °C in a VAPOUR-line autoclave of VWR. Due to the use of a small soil inoculum in a background of mineral soil, the differences in effect of soil sterilization on nutrient release are negligible65. (ii) Untreated soil, where the conditioned soil was directly used to grow plants of the response phase. (iii) pathogen/saprobe filtrate, where soil from the conditioning phase was first filtered through multiple analytical Retsch sieves (mesh sizes of 1000, 500, 250, 180, 125 and 63 µm) with 0.5 l water to retain sand, root particles and other coarse materials. This solution was then filtered through an analytical 20 µm Retsch test sieve—retaining arbuscular mycorrhizal fungi and their spores—after which the filtrate, containing pathogens and non-mycorrhizal microbes that could pass the 20 µm pore-size filter, was collected17. After sterilizing the soil (as above), this filtrate was added back. Though nitrogen-fixing bacteria from the genus Rhizobium and closely related taxa could also pass through these sieves, visual checks of the roots of all leguminous plants showed the absence of root nodules in all treatments. In this second phase, plants were grown in soil conditioned by the same species during eight weeks, receiving equivalent nutrient dosages as in the first phase, after which they were harvested. The plant material was dried for 72 h at 70 °C prior to weighing above and below-ground dry biomass. For each of the 13 species and three soil treatments, six repetitions were performed for a total of 234 plants in each phase (13 species × 3 soil treatments × 6 repetitions).
Community experiment
To compare PSF from monocultures with PSF in plant communities, the same 10 native species were grown randomly together in 5 l pots (23 cm diameter, Göttinger, Germany), with one individual per species per pot. The same procedure was followed as for the monoculture experiments unless mentioned otherwise. In the conditioning phase, soil was conditioned by the whole community instead of a single species. Every community received 10 times more nutrient solution as in the monoculture experiment in order to receive the same amount of nutrients per plant. For the response phase, the same three soil treatments as for the monoculture experiment were applied, though an extra factor was included into this design. Some plant communities were assigned an invasive species (one out of three aforementioned invasive species), placed in the middle of the community in a full factorial design for a total of 1200 native plants in each phase (10 species/community × 3 soil treatments × 4 addition possibilities × 10 repetitions) and 90 invasive plants in the second phase (30 plants of each invasive species, 3 soil treatments × 10 repetitions). Invasive plants were included into the plant communities in order to investigate whether they were stronger competitors when PSF was present, and thus if the ERH applies to these invasive species. Above-ground dry biomass was measured for all 1290 individual plants, and below-ground dry biomass for half of all repetitions, thus for 645 plants in total, after careful separation of roots of different species.
PSF calculations
In this work we distinguish two types of plant-soil feedback: PSFtot is the feedback plants receive from all soil biota, both pathogens and mutualists. It was calculated for each individual plant separately—grown in unsterilized conditioned soil (see above, soil treatment ii)—as the difference between that individuals biomass and the average biomass of the same species when grown in sterilized soil (soil treatment i). Since this difference depends not only on the effect size of feedback, but also on how large a certain plant species becomes, we standardized this term by dividing by the average biomass of the same species when grown in sterilized soil (see Eq. 1). PSFpath is the feedback plants receive from only pathogens (and saprobes) and is calculated as PSFtot but with soil treatment ii replaced by treatment iii, i.e. soil treated with the pathogen/saprobe filtrate. Standardized PSF effect sizes were thus measured as follows65:
All PSFs are calculated using above-ground, below-ground or total biomass, as indicated in captions.
Statistics
In order to investigate whether PSF measured in monocultures was representative for communities we performed a standardised major axis regression (package smatr66) between PSF (PSFtot & PSFpath) in monocultures and in communities (Fig. 1a, b and Extended Data Fig. 2a, b), after checking for the model assumptions (visual plots and function shapiro.test).
Since PSF in communities was not correlated to PSF in monocultures, we examined whether PSF in community depends on the dominance of the species itself, measured as dry weight biomass. We therefore made mixed linear regressions (function lmer, R package lme467) assessing the relations between PSF and species biomass in communities (Fig. 2a, b and Extended Data Figs. 5, 6, Table 1). Both species biomass (as average dry weight in grams of the sterilized treatment), functional group and their interaction were inserted as fixed factors. As species biomass was inversely transformed (1/x) for improved model fit, a constant of 1 was added to the response variable (standardized effect size of PSF), since the function 1/x cannot exist as a negative. The minimum value for standardized effect size is −1, when an individual would be extremely small when growing in conditioned soil (see Eq. 1). By increasing the standardized effect size in the model by 1 we ensured that the new minimum value of the response variable is 0, the asymptote of the function 1/x. The community to which each individual belonged (as repetition number) and the presence of an invasive species in that community (four levels: no invasive added, Avena sterilis added, Lupinus polyphyllus added or Solidago gigantea added) were inserted as random factors in order to account for differences between communities and the possible impact invasives could have on the reduction in species-specific pathogens. As shown by the negligible amounts of variance explained by these random factors compared to the residual variance in all models and the negligible rise in R² when these variables are added (= difference between conditional R² and marginal R²), these random factors had no significant impact on regressions. Assumptions were visually checked, except for multicollinearity between different fixed values, which was analysed using VIF values with a cut-off of <3. For the relation between the difference in PSF between communities and monocultures and species biomass (Extended Data Fig. 5a, b, Table 1) the same mixed linear regressions were used, but with species biomass (transformed as above) as fixed factor and functional group as random factor.
In order to show that this relation between species biomass and PSF was only present in communities and not in monocultures, we made multiple linear regressions relating PSF and species biomass from the monoculture experiment (Extended Data Fig. 7a, b, Table 1). Explanatory variables were species biomass, functional group and their interaction as above, but without random factors and no transformation of the response variable since this did not improve model fit. Assumptions were checked as for the linear regressions above and with VIF values.
Significant effect sizes of PSF in both monocultures and communities (Extended data Figs. 3, 4) were calculated with one-sample t-tests. PSFs were statistically significant when they were significantly different from 0. Assumptions were checked with the Shapiro-Wilk test. When these were not met, significances were calculated with the Wilcoxon one-sample signed rank test.
The influence of both soil treatment (sterilized, sterilized + pathogen/saprobe filtrate, no treatment) and the presence of an invasive species were analysed using two-way ANOVA’s with interaction term (Fig. 3a, Extended Data Fig. 8a and Table 2) and one-way ANOVA’s (Fig. 3b, Extended Data Fig. 8b and Table 2). An interaction term was added in order to assess whether the effect of invasive species altered depending on the soil treatment. Assumptions were checked as above. Post hoc tests were also performed as above.
Data acquisition was done in Microsoft Excel version 2210 and analyses were performed in R version 4.2.168. All figures were made with ggplot, and edited in Inkscape 1.2.1.
Limitations of our methods
Pathogen/saprobe-filtrate preparation through a 20 µm pore-size filter is a standardized work method in the field of PSF8,17,69,70,71,72, but it has its limitations. Sieving a suspension through a 20 µm filter retains micro-arthropods, nematodes, AMF, enchytraeids and collembola, but still contains pathogenic and saprobic soil bacteria and fungi73,74,75. We thus assumed that both saprobic bacteria and fungi had no significant effect on plant growth within these 8 weeks as we assigned the effects observed in PSFpath to (species-specific) pathogens. Besides this, some nematode eggs can pass certain smaller pore-sizes76, possibly slightly altering the PSF measured, though it has to be verified whether these eggs can have a significant impact on plant growth within 8 weeks. Furthermore, some soil organisms are better at recolonizing the soil after filtration and others are more sensitive to the filtration process73, creating a bias in microbe abundance and thus in the PSF measured. Lastly, total native biomass seems lower—though not significantly lower—when grown in the unsterilized soil treatment (= PSFtot) than when grown in the sterilized soil treatment with the addition of the pathogen/saprobe filtrate (= PSFpath) (Fig. 3a, b and Extended Data Fig. 8a, b). This could be attributed to pathogens having to recolonize the soil after addition to the sterilized soil, reducing their negative effect on plant growth. Another, not mutually exclusive, explanation is that some of the filtered pathogens in the soil were more sensitive to the soil processing.
Data availability
The data that support the findings of this study are available in figshare with the identifier https://doi.org/10.6084/m9.figshare.23295482.
References
Bever, J. D., Mangan, S. A. & Alexander, H. M. Maintenance of plant species diversity by pathogens. Annu. Rev. Ecol. Evol. Syst. 46, 305–325 (2015).
Lekberg, Y. et al. Relative importance of competition and plant–soil feedback, their synergy, context dependency and implications for coexistence. Ecol. Lett. 21, 1268–1281 (2018).
Bever, J. D., Platt, T. G. & Morton, E. R. Microbial population and community dynamics on plant roots and their feedbacks on plant communities. Annu. Rev. Microbiol. 66, 265–283 (2012).
Janzen, D. H. Herbivores and the number of tree species in tropical forests. Am. Nat. 104, 501–528 (1970).
Connell, J. H. Dynamics of Populations (Center for Agricultural Publication and Documentation, 1971).
Liu, Y., Yu, S., Xie, Z.-P. & Staehelin, C. Analysis of a negative plant-soil feedback in a subtropical monsoon forest: Recruitment of tree juveniles. J. Ecol. 100, 1019–1028 (2012).
Maron, J. L., Marler, M., Klironomos, J. N. & Cleveland, C. C. Soil fungal pathogens and the relationship between plant diversity and productivity: soil pathogens, productivity and invasibility. Ecol. Lett. 14, 36–41 (2011).
Schnitzer, S. A. et al. Soil microbes drive the classic plant diversity–productivity pattern. Ecology 92, 296–303 (2011).
Knevel, I. C., Lans, T., Menting, F. B. J., Hertling, U. M. & van der Putten, W. H. Release from native root herbivores and biotic resistance by soil pathogens in a new habitat both affect the alien Ammophila arenaria in South Africa. Oecologia 141, 502–510 (2004).
Reinhart, K. O. & Callaway, R. M. Soil biota and invasive plants. New Phytol. 170, 445–457 (2006).
Molofsky, J. & Bever, J. D. A novel theory to explain species diversity in landscapes: positive frequency dependence and habitat suitability. Proc. R. Soc. Lond. B Biol. Sci. 269, 2389–2393 (2002).
Kulmatiski, A., Beard, K. H., Stevens, J. R. & Cobbold, S. M. Plant-soil feedbacks: a meta-analytical review. Ecol. Lett. 11, 980–992 (2008).
Crawford, K. M. et al. When and where plant‐soil feedback may promote plant coexistence: a meta‐analysis. Ecol. Lett. 22, 1274–1284 (2019).
Keane, R. Exotic plant invasions and the enemy release hypothesis. Trends Ecol. Evol. 17, 164–170 (2002).
Wolfe, L. M. Why alien invaders succeed: support for the escape‐from‐enemy hypothesis. Am. Nat. 160, 705–711 (2002).
Vilà, M., Maron, J. L. & Marco, L. Evidence for the enemy release hypothesis in Hypericum perforatum. Oecologia 142, 474–479 (2005).
Klironomos, J. N. Feedback with soil biota contributes to plant rarity and invasiveness in communities. Nature 417, 67–70 (2002).
Callaway, R. M., Thelen, G. C., Rodriguez, A. & Holben, W. E. Soil biota and exotic plant invasion. Nature 427, 731–733 (2004).
Colautti, R. I., Ricciardi, A., Grigorovich, I. A. & MacIsaac, H. J. Is invasion success explained by the enemy release hypothesis? Ecol. Lett. 7, 721–733 (2004).
Shannon, S., Flory, S. L. & Reynolds, H. Competitive context alters plant–soil feedback in an experimental woodland community. Oecologia 169, 235–243 (2012).
Fahey, C. & Flory, S. L. Soil microbes alter competition between native and invasive plants. J. Ecol. 110, 404–414 (2022).
Schroeder, J. W., Dobson, A., Mangan, S. A., Petticord, D. F. & Herre, E. A. Mutualist and pathogen traits interact to affect plant community structure in a spatially explicit model. Nat. Commun. 11, 2204 (2020).
Mangan, S. A. et al. Negative plant–soil feedback predicts tree-species relative abundance in a tropical forest. Nature 466, 752–755 (2010).
Anacker, B. L., Klironomos, J. N., Maherali, H., Reinhart, K. O. & Strauss, S. Y. Phylogenetic conservatism in plant-soil feedback and its implications for plant abundance. Ecol. Lett. 17, 1613–1621 (2014).
Chesson, P. Mechanisms of maintenance of species diversity. Annu. Rev. Ecol. Syst. 31, 343–366 (2000).
Bever, J. D. Soil community feedback and the coexistence of competitors: conceptual frameworks and empirical tests. New Phytol 157, 465–473 (2003).
Bonanomi, G., Giannino, F. & Mazzoleni, S. Negative plant-soil feedback and species coexistence. Oikos 111, 311–321 (2005).
Kulmatiski, A., Heavilin, J. & Beard, K. H. Testing predictions of a three-species plant-soil feedback model: testing a plant-soil feedback model. J. Ecol. https://doi.org/10.1111/j.1365-2745.2010.01784.x (2011).
Revilla, T. A., Veen, G. F., Eppinga, M. B. & Weissing, F. J. Plant–soil feedbacks and the coexistence of competing plants. Theor. Ecol. 6, 99–113 (2013).
Engelkes, T. et al. Successful range-expanding plants experience less above-ground and below-ground enemy impact. Nature 456, 946–948 (2008).
Teste, F. P. et al. Plant-soil feedback and the maintenance of diversity in Mediterranean-climate shrublands. Science 355, 173–176 (2017).
Wilschut, R. A. et al. Root traits and belowground herbivores relate to plant–soil feedback variation among congeners. Nat. Commun. 10, 1564 (2019).
Abbott, K. C. et al. Spatial heterogeneity in soil microbes alters outcomes of plant competition. PLoS ONE 10, e0125788 (2015).
Casper, B. B. & Castelli, J. P. Evaluating plant-soil feedback together with competition in a serpentine grassland. Ecol. Lett. 10, 394–400 (2007).
Ke, P.-J. & Wan, J. Effects of soil microbes on plant competition: a perspective from modern coexistence theory. Ecol. Monogr. 90, e01391 (2020).
Beals, K. K. et al. Predicting plant-soil feedback in the field: meta-analysis reveals that competition and environmental stress differentially influence PSF. Front. Ecol. Evol. 8, 191 (2020).
Schittko, C., Runge, C., Strupp, M., Wolff, S. & Wurst, S. No evidence that plant-soil feedback effects of native and invasive plant species under glasshouse conditions are reflected in the field. J. Ecol. 104, 1243–1249 (2016).
Forero, L. E., Grenzer, J., Heinze, J., Schittko, C. & Kulmatiski, A. Greenhouse- and field-measured plant-soil feedbacks are not correlated. Front. Environ. Sci. 7, 184 (2019).
van der Putten, W. H. et al. Plant-soil feedbacks: the past, the present and future challenges. J. Ecol. 101, 265–276 (2013).
Heinze, J., Sitte, M., Schindhelm, A., Wright, J. & Joshi, J. Plant-soil feedbacks: a comparative study on the relative importance of soil feedbacks in the greenhouse versus the field. Oecologia 181, 559–569 (2016).
Kardol, P., Cornips, N. J., van Kempen, M. M. L., Bakx-Schotman, J. M. T. & van der Putten, W. H. Microbe-mediated plant–soil feedback causes historical contingency effects in plant community assembly. Ecol. Monogr. 77, 147–162 (2007).
Hendriks, M. et al. Spatial heterogeneity of plant–soil feedback affects root interactions and interspecific competition. New Phytol. 207, 830–840 (2015).
Crawford, K. M. & Knight, T. M. Competition overwhelms the positive plant–soil feedback generated by an invasive plant. Oecologia 183, 211–220 (2017).
Bennett, J. A. & Klironomos, J. Mechanisms of plant–soil feedback: interactions among biotic and abiotic drivers. New Phytol. 222, 91–96 (2019).
Moora, M. et al. Alien plants associate with widespread generalist arbuscular mycorrhizal fungal taxa: evidence from a continental-scale study using massively parallel 454 sequencing. J. Biogeogr. 38, 1305–1317 (2011).
Dudenhöffer, J.-H., Luecke, N. C. & Crawford, K. M. Changes in precipitation patterns can destabilize plant species coexistence via changes in plant–soil feedback. Nat. Ecol. Evol. 6, 546–554 (2022).
Manning, P., Morrison, S. A., Bonkowski, M. & Bardgett, R. D. Nitrogen enrichment modifies plant community structure via changes to plant–soil feedback. Oecologia 157, 661–673 (2008).
in ’t Zandt, D., van den Brink, A., de Kroon, H. & Visser, E. J. W. Plant-soil feedback is shut down when nutrients come to town. Plant Soil 439, 541–551 (2019).
Moora, M. Mycorrhizal traits and plant communities: perspectives for integration. J. Veg. Sci. 25, 1126–1132 (2014).
Münzbergová, Z. & Šurinová, M. The importance of species phylogenetic relationships and species traits for the intensity of plant-soil feedback. Ecosphere 6, art234 (2015).
Tilman, D. Niche tradeoffs, neutrality, and community structure: a stochastic theory of resource competition, invasion, and community assembly. Proc. Natl. Acad. Sci. USA 101, 10854–10861 (2004).
Fargione, J., Brown, C. S. & Tilman, D. Community assembly and invasion: an experimental test of neutral versus niche processes. Proc. Natl. Acad. Sci. USA 100, 8916–8920 (2003).
Ruijven, J., De Deyn, G. B. & Berendse, F. Diversity reduces invasibility in experimental plant communities: the role of plant species. Ecol. Lett. 6, 910–918 (2003).
Grace, J. B. et al. Integrative modelling reveals mechanisms linking productivity and plant species richness. Nature 529, 390–393 (2016).
Walker, B. H. Biodiversity and ecological redundancy. Conserv. Biol. 6, 18–23 (1992).
Eisenhauer, N., Hines, J., Maestre, F. T. & Rillig, M. C. Reconsidering functional redundancy in biodiversity research. NPJ Biodivers. 2, 1–4 (2023).
Eisenhauer, N. et al. Plant diversity effects on soil microorganisms support the singular hypothesis. Ecology 91, 485–496 (2010).
Vanderhoeven, S., Dassonville, N. & Meerts, P. Increased topsoil mineral nutrient concentrations under exotic invasive plants in Belgium. Plant Soil 275, 169–179 (2005).
Hawkes, C. V., Wren, I. F., Herman, D. J. & Firestone, M. K. Plant invasion alters nitrogen cycling by modifying the soil nitrifying community. Ecol. Lett. 8, 976–985 (2005).
Dassonville, N., Vanderhoeven, S., Gruber, W. & Meerts, P. Invasion by Fallopia japonica increases topsoil mineral nutrient concentrations. Ecoscience 14, 230–240 (2007).
Peltzer, D. A. et al. Punching above their weight: low-biomass non-native plant species alter soil properties during primary succession. Oikos 118, 1001–1014 (2009).
Hansen, M. J. & Clevenger, A. P. The influence of disturbance and habitat on the presence of non-native plant species along transport corridors. Biol. Conserv. 125, 249–259 (2005).
Bauer, J. T. Invasive species: “back-seat drivers” of ecosystem change? Biol. Invasions 14, 1295–1304 (2012).
Minden, V., Schaller, J. & Olde Venterink, H. Plants increase silicon content as a response to nitrogen or phosphorus limitation: a case study with Holcus lanatus. Plant Soil 462, 95–108 (2021).
Pernilla Brinkman, E., Van der Putten, W. H., Bakker, E.-J. & Verhoeven, K. J. F. Plant-soil feedback: experimental approaches, statistical analyses and ecological interpretations: design and analysis of feedback experiments. J. Ecol. 98, 1063–1073 (2010).
Warton, D. I., Duursma, R. A., Falster, D. S. & Taskinen, S. smatr 3—an R package for estimation and inference about allometric lines: The smatr 3 - an R package. Methods Ecol. Evol. 3, 257–259 (2012).
Bates, D., Machler, M., Bolker, B. & Walker, S. Fitting Linear Mixed-Effects Models Using (2015).
R Core Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/. (2022).
McCarthy-Neumann, S. & Kobe, R. K. Conspecific plant–soil feedbacks reduce survivorship and growth of tropical tree seedlings. J. Ecol. 98, 396–407 (2010).
Hilbig, B. E. & Allen, E. B. Plant-soil feedbacks and competitive interactions between invasive Bromus diandrus and native forb species. Plant Soil 392, 191–203 (2015).
Pizano, C., Kitajima, K., Graham, J. H. & Mangan, S. A. Negative plant–soil feedbacks are stronger in agricultural habitats than in forest fragments in the tropical Andes. Ecology 100, e02850 (2019).
Lankau, R. A., Wheeler, E., Bennett, A. E. & Strauss, S. Y. Plant–soil feedbacks contribute to an intransitive competitive network that promotes both genetic and species diversity. J. Ecol. 99, 176–185 (2011).
van de Voorde, T. F. J., van der Putten, W. H. & Bezemer, T. M. Soil inoculation method determines the strength of plant–soil interactions. Soil Biol. Biochem. 55, 1–6 (2012).
Wang, M. et al. Removal of soil biota alters soil feedback effects on plant growth and defense chemistry. New Phytol. 221, 1478–1491 (2019).
Bardgett, R. The Biology of Soil: A Community and Ecosystem Approach (OUP Oxford, 2005).
Wurst, S., Allema, B., Duyts, H. & Van Der Putten, W. H. Earthworms counterbalance the negative effect of microorganisms on plant diversity and enhance the tolerance of grasses to nematodes. Oikos 117, 711–718 (2008).
Acknowledgements
We thank Lisa Partoens, Joren M. Snoeks, Loïc den Godt op Aardt and Jani Sleutel for their help with seedling transplantations in the greenhouse. Furthermore, we acknowledge Piet De Becker and the organizations VHM and Natuurpunt for their admittance to take soil samples in the nature reserve Doode Bemde.
Author information
Authors and Affiliations
Contributions
E.P.G. and H.O.V. conceived the ideas and designed methodology; E.P.G. and F.V.P. collected the data; E.P.G. analysed the data; E.P.G. led the writing of the manuscript; E.P.G., V.M., F.V.P. and H.O.V. contributed critically to the drafts and all authors gave final approval for publication.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Goossens, E.P., Minden, V., Van Poucke, F. et al. Negative plant-soil feedbacks disproportionally affect dominant plants, facilitating coexistence in plant communities. npj biodivers 2, 27 (2023). https://doi.org/10.1038/s44185-023-00032-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s44185-023-00032-4
- Springer Nature Limited