Abstract
Chemotaxis–the movement of cells along chemical gradients—leads to collective behaviors when cells coordinate their movements. Here, using Escherichia coli as a model, we demonstrate a distinct type of bacterial collective response in acidic environments containing organic acids. Bacterial populations immersed in such environments collectively condensed into millimeter-sized focal points. Furthermore, this bacterial condensation fostered the formation of small, tightly packed cell aggregates, resembling non-surface-attached biofilms. These cell aggregates were physically displaced by the free-swimming condensing cells, leading to the segregation of the two cell populations. Bacterial condensation relied on feedback between the tendency of these bacteria to neutralize the pH and their chemotactic repulsion from low pH. Sustained cell condensation occurred when the bacteria occupied only part of the acidic environment, either dynamically or due to physical constraints. Such condensed bacterial populations can mitigate acid stress more efficiently, a principle that may be applicable to other stress conditions.
Similar content being viewed by others
Introduction
The ability of bacteria to tolerate pH stress influences their survival and behavior in a variety of environments1,2. In particular, weak organic acids, such as lactate and acetate, present an acute challenge to neutrophilic bacteria by entering the cell in their protonated state and directly acidifying the cytoplasmic space3,4,5,6,7,8. These organic acids are commonly found in natural environments, including the human gut, tumors, soil, and plants9,10,11,12,13,14. Bacteria have evolved a variety of mechanisms to cope with such stress, including adaptive expression of systems that actively export protons out of the cell, neutralize protons within the cytoplasm, or repair the resulting molecular damage1,15,16,17,18,19. In addition, the bacterial cytoplasm has an inherent buffering capacity that can neutralize excess protons 5,20. By neutralizing the effect of organic acid intake, cells can also reduce the amount of free protons in the environment20, and thus, in principle, cooperatively cope with the acid stress.
Chemotaxis also enables bacteria to alleviate pH stress by allowing them to migrate away from acidic environments (pH taxis) and thereby avoid the acidic conditions altogether21,22. The neutrophilic bacterium Escherichia coli (E. coli) is a well-known model for studying molecular and behavioral aspects of bacterial chemotaxis23,24,25. Here we used the non-pathogenic MG1655(IS1) strain, with robust chemotaxis behavior26,27,28. E. coli chemotaxis relies on dedicated methyl-accepting chemotaxis proteins that sense temporal changes in chemoeffector concentrations along the swimming trajectory of the cells, and, through phosphotransferase signaling, bias their swimming toward beneficial ‘attractant’ effectors and away from harmful ‘repellent’ effectors29,30. In particular, migration away from acidic environments is primarily mediated by the Tsr receptors21,22. Interestingly, cells lacking the Tsr receptors actively migrate towards acidic environments, down to a pH below 5, a behavior mediated by the Tar receptor21. The combined effect of Tsr and Tar was suggested to allow bacteria to navigate toward neutral pH22,31. Overall, the ability of bacteria to track chemical gradients in their environment influences many aspects of their ecology32.
Although each bacterial cell can independently follow external gradients, collective behaviors can emerge when bacteria modulate their local environment in a way that drives coordinated responses. Collective propagation of bacteria in dilute agar plates (swim plates) or in capillary tubes emerges due to consumption of amino acids or oxygen by the bacteria, leading to self-induced chemoeffector gradients23,28,33,34. Moreover, addition of certain carbon sources, such as succinate, fumarate, or malate, to the growth medium in these plates leads to spontaneous condensation of the bacteria into discrete cell accumulations35,36,37. Formation of these cell accumulations is triggered by the secretion of secondary metabolites, such as aspartate or glycine, which, in turn, act as attractants. A similar, but weak and short-lived, bacterial accumulation was also observed independent of cell proliferation38. In contrast, the secretion of the chemoattractant autoinducer-2 (AI-2) did not lead to similar large-scale bacterial condensation, but rather, enhanced the formation of compact micro-scale cell aggregates39,40.
Notably, when bacterial suspensions are embedded in a larger uniform acidic environment (Fig. 1a), both pH neutralization and pH-taxis are ineffective: the cell-free acidic environment effectively acts as a reservoir that limits the ability of the embedded cell population to modify the external pH, and the chemically uniform environment renders bacterial chemotaxis ineffective. Here, we show that under these challenging conditions, bacteria collectively condense into distinct focal points, independent of cell proliferation. Cells accumulated in these millimeter-sized focal points reach a very high cell density, yet remain suspended and motile. This long-lived condensation readily and considerably reduces the acid stress experienced by the cells. Furthermore, bacterial condensation also fosters auto-aggregation of cells and formation of tightly-packed bacterial clumps with a typical size of 10–20 µm. Interestingly, however, these cell aggregates are physically displaced by the condensing free-swimming (‘planktonic’) cells, leading to the segregation of the two cell populations. Under proper conditions, these cell aggregates may develop into mature biofilms41,42, including an extra-cellular matrix and additional factors that, in turn, promote a broad stress resistance43. Overall, bacterial condensation results in more efficient pH neutralization and may enable bacteria to further activate pH-specific adaptive responses under milder stress conditions.
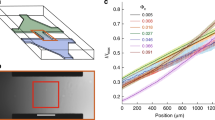
a Illustration contrasting ‘isolated’ bacterial suspensions (left) with ‘embedded’ populations immersed in a larger uniform environment (right). b Bacterial cells suspension (1 ml, OD600 = 1) in motility medium with pH 5.2 and 0.1 mM KPi applied onto an agarose hydrogel plate with identical medium (1% agarose, ~8 mm thick, 90 mm diameter). Cells were grown at 33.5 °C and expressed GFP (see Methods). Shown on the right is a typical fluorescence image of the plate taken 30 min after applying the cells. Below is a zoom-in image. c Similar cell suspensions in motility medium with pH 5.5 and OD600 = 0.4 applied to the ‘thin layer’ setup (illustrated; see Methods). Here, cells were restricted to swim within a disk-like thin layer by a hydrogel mesh in identical medium that occupied the rest of the chamber (3% agarose, pH 5.5). Initially, the entire chamber, including both the bacterial suspension and the hydrogel, formed a uniform acidic environment (pH 5.5) with low buffering capacity (0.1 mM KPi). The panel below shows top-view fluorescence images of the cell suspension (d = 5 mm and L = 10.7 mm) at various times after placing the cells in the setup; note, the grayscale was 0-1000 in the first two images and 0-5000 later (see also SI Video 1). The corresponding cell-density distributions across the bacterial layer are shown in top-right plot; here, the intensity was radially-averaged and presented symmetrically (see Methods). Experiments were done at 30 °C. (d) Similar experiments, as in (c), were conducted with a larger cell layer (d = 10 mm and L = 18 mm). Fluorescence image of the entire cell layer is shown at 40 min (left) along with the time course demonstrating the coalescence of two focal points (see also SI Videos 2-3).
Results
Bacterial condensation in low pH environments
To study the behavior of chemotactic bacterial populations in acidic environments, we used GFP-expressing E. coli cells suspended in a commonly used motility medium containing lactic acid and titrated to pH 5.2. This medium supports cell motility but does not support cell proliferation (see Methods). Surprisingly, we found that applying a thin layer of such cell suspension on top of an agarose-hydrogel layer containing identical (acidic) motility medium, led, within 30 minutes, to condensation of the bacterial population into discrete focal points (Fig. 1b). Moreover, these bacterial accumulations spontaneously formed an irregular lattice-like structure with a typical spacing of approximately 1.8 mm. Notably, the bacterial layer was thin enough (approximately 0.5 mm) to allow effective diffusion across the layer and, hence, efficient exchange of chemicals between the bacterial layer and the cell-free region underneath. Thus, under these conditions, the bacterial layer is effectively ‘chemically coupled’ to the hydrogel layer underneath (Fig. 1b).
To study bacterial condensation in a more controlled setting, we constructed a thin layer of bacterial suspension (~120 µm) embedded within a larger uniform environment at pH 5.5 (Fig. 1c). Bacteria were confined to a thin disk-like layer imprinted in an agarose hydrogel mesh, which restricted them from swimming out of the layer (see Methods). The aspect ratio of the bacterial layer was such that the distance of the bacteria from the cell-free region underneath was typically much smaller than the distance to the lateral edge of the bacterial suspension (Fig. 1c). Thus, the local environment within the bacterial suspension was tightly coupled to the cell-free environment through diffusion perpendicular to the bacterial layer. In this setting, hereafter, referred to as the ‘thin layer’ setup, the initially uniform bacterial population clearly and robustly condensed into a single focal point (Fig. 1c, lower panel). The spatial distribution of the bacteria around the focal point developed over time and had a typical diameter of approximately 1 mm (Fig. 1c, upper-right plot). In contrast, isolated populations (Fig. 1a) suspended in the same acidic medium did not condense and remained uniformly distributed (Supplementary Fig. S1).
Interestingly, the spatial pattern of bacterial accumulations depended on the diameter of the bacterial layer for a diameter smaller than 1 mm, no condensation could be observed (Supplementary Fig. S2); for a diameter of approximately 5 mm, cells typically condensed into a single focal point (Fig. 1c); and for a diameter of approximately 10 mm, cells condensed into several focal points typically arranged in a ring (Fig. 1d). Despite the high cell density at the center of the bacterial accumulations (OD600 ~ 6), the cells remained both suspended and motile, making the bacterial accumulations effectively dynamic. In particular, bacterial accumulations less than approximately 3 mm apart effectively interacted and ultimately coalesced (Fig. 1d). Notably, when two bacterial accumulations coalesced, the local cell density should have doubled. However, cells tended to avoid such high density and instead formed a ring of cells, most likely due to oxygen considerations (see Discussion). The tendency of cell accumulations to coalesce at short distances may also account for the fact that only a single focal point was stable in suspensions with a smaller diameter (d = 5 mm).
In contrast to the pattern of the bacterial focal points observed on agarose plates (Fig. 1b), the condensation observed in Figs. 1c, d had a clear radial symmetry. This symmetry most likely arises from the residual coupling of the cell suspension to the cell-free environment through the lateral edge of the cell layer. Such coupling may cause a very weak cell-density perturbation that can lead to the observed symmetry. In fact, expanding the bacterial suspension to occupy the entire upper layer of the chamber, and thereby eliminating the coupling at the edges, resulted in the bacteria still spontaneously condensing into focal points, but the radial symmetry of their arrangement was absent (Supplementary Fig. S3). Yet, for much larger bacterial layers (Fig. 1b) the location of focal points becomes more orderly, most likely due to effective correlations between them.
Bacterial condensation is mediated by chemotactic response to self-induced pH gradients
To identify which chemoreceptors are involved in the observed bacterial condensation, we repeated the ‘thin layer’ experiments, as in Fig. 1c, using cells lacking either the Tsr or Tar chemoreceptors. Notably, these receptors respond to pH gradients in opposite ways: Tsr mediates repulsion from low pH and Tar mediates attraction to low pH21,22. Since high cell density can lead to oxygen deprivation, we also tested cells lacking the Aer receptor, which mediates redox (oxygen) taxis, known as aerotaxis44. The condensation time-course of the different strains was characterized by monitoring the cell density at the peak (Fig. 2). In all cases, the cell accumulation tended to reach saturation after ~2 h, albeit at different levels. As expected from pH taxis away from acidic environments, cells lacking Tsr did not condense and maintained a uniform distribution, whereas cells lacking Tar exhibited enhanced condensation compared to wild-type cells. Additionally, cells lacking the Aer receptor exhibited similar accumulation to that of wild-type cells (Supplementary Fig. S4), suggesting that oxygen deprivation is not a dominant force in the accumulation of wild-type cells. However, deletion of the Aer receptor in a Tar− host clearly enhanced bacterial accumulation (Fig. 2), indicating that aerotaxis limits the accumulation of the Tar− cells. Thus, condensation behaviors align with the roles of Tar and Tsr in pH taxis, while oxygen limitation restricted the bacterial condensation primarily when the cell density exceeds that of wild-type cells.
Experiments were done using the ‘thin layer’ setup (see Fig. 1c) and the cell accumulation is plotted as function of time for wild-type cells (all the chemoreceptors present; blue symbols) or cells missing either Tsr (red symbols), Tar (green symbols), or both Tar and Aer (open gray symbols). Cell accumulation was defined as the difference between the cell density at the center of the focal point and the baseline density away from it (right-top illustration). Data is shown for at least three repetitions with each strain.
Given the initially uniform pH in the chamber, we hypothesized that condensation was driven by pH-taxis triggered by self-induced pH gradients. To test the ability of the bacteria to neutralize the pH in their environment, we conducted experiments with large-volume (20 ml) and well-mixed cell suspensions in the same low-pH medium used in the bacterial condensation experiments. Indeed, with either lactate (Fig. 3a, black-filled circles) or acetate (Fig. 3a, green-open circles) in the medium, cells could clearly neutralize the pH. Moreover, the ability of the cells to change the pH depended on their density (shown for lactate) and occurred over a timescale comparable to that of the bacterial condensation shown in Fig. 2. As expected, the pH modulation diminished by adding a strong pH buffer to the medium (10 mM KPi; Fig. 3a, blue-open circles). On the other hand, no significant accumulation of chemoeffectors, other than low pH, could be detected in these bacterial suspensions (Supplementary Fig. S5). Furthermore, by repeating the ‘thin-layer’ experiments (as in Fig. 1c) with a pH-sensitive fluorescein dye added to the bacterial suspension (SI Methods), a significant increase in pH could be directly detected in the vicinity of the bacterial accumulation (Fig. 3b).
a The pH of cell suspensions, in motility medium (20 ml, 0.1 mM KPi), was followed over time under continuous stirring at 30 °C. The pH time trace is shown for experiments with cell density (OD600) of 0 (no cells; light-gray symbols), 0.1 (darker gray symbols), or 0.4 (black symbols). Experiments were also done with cells (OD600 0.4) immersed in a similar medium but with either stronger buffer (10 mM KPi, blue symbols) or with acetate (1 mM) replacing the lactate (open green circles). Symbols represent the average over three repetitions, and the shaded area represents the standard deviation around the mean. b Cell suspension in motility medium was applied to the ‘thin-layer’ setup, as in Fig. 1c, except that a pH-sensitive dye (40 μM fluorescein) was added uniformly to the cell suspension (see Methods). Cells did not express GFP here. The fluorescence intensity profile across the bacterial accumulation (normalized to its value at the edges) is shown for experiments with wild-type, Tar−Aer−, or CheY−CheZ− cells, as labeled. Note that the elevated fluorescence intensity indicates elevated pH (see SI Methods). c The saturated cell accumulation in the ‘thin-layer’ setup (Fig. 1c) when using a medium with different carbon sources (1 mM) and either weaker (0.1 mM KPi) or stronger (10 mM KPi) buffering capacity, as labeled. d The saturated cell accumulation in the ‘thin-layer’ setup when cells were added at varying initial densities. Error bars represent the standard deviation across multiple repetitions (N ≥ 3). e Experiments similar to those described in Fig. 1c, using the ‘thin-layer’ setup, were conducted with the medium set to alkaline pH (9.6) and with glycerol serving as the sole energy source. The time course of cell accumulation is shown on the left for experiments with either Tar− (green) or Tsr− (red) cells, along with a top-view fluorescence image at t = 1 h (inset; Tsr− cells) and the corresponding radially-averaged cell-density distribution (right plot, presented symmetrically). f Numerical solutions of the model described in the text (Eq. 1–3), demonstrating the expected cell-density distribution at different times (left plot, initial cell density 0.4) and the saturated cell accumulation at varying initial cell densities (right plot). Throughout the figure, cell densities and cell accumulation were measured in OD600 (omitted for clarity).
To further understand the conditions required for cell condensation, we repeated the ‘thin-layer’ experiments (as in Fig. 1c) with different carbon sources in the motility medium and with either a weak (0.1 mM KPi) or strong (10 mM KPi) buffer. The saturated cell accumulation observed in these experiments is shown in Fig. 3c. Lactate and acetate clearly supported bacterial condensation in a weak buffer, but no condensation could be observed with glycerol or gluconate (Fig. 3c). Consistently, lactate and acetate supported cell-mediated pH elevation, but glycerol and gluconate did not (Fig. 3a and Supplementary Fig. S6). Importantly, we found that the addition of a strong buffer (10 mM KPi) to the medium, which limits the ability of the cells to modulate the external pH, also inhibited the bacterial condensation in all cases (Fig. 3c).
Overall, these results suggest that bacterial condensation is a cooperative chemotactic response triggered by positive feedback between the tendency of the cells to neutralize the pH in their local environment and their repulsion from low pH. As might be expected from such a cooperative response, bacterial condensation depended on the initial cell density in a highly non-linear manner (Fig. 3d, discussed in the Model section below).
Condensation in alkaline pH: the roles of Tar and Tsr
Given the mechanism suggested above for cell condensation, we expected similar condensation in alkaline pH, provided that cells could lower the pH. To test this hypothesis, we repeated the ‘thin-layer’ experiments, as in Fig. 1c, with the overall pH set to 9.6 and with glycerol replacing the organic acids in the medium. We verified that cells indeed tended to neutralize (lower) the pH under these conditions (Supplementary Fig. S6). We found that in this case as well, cells condensed over a time scale comparable to that observed in the acidic pH medium (Fig. 3e). Moreover, the bacterial accumulation exhibited a similar spatial distribution as in the acidic case (~1 mm wide), suggesting a comparable condensing force (Fig. 3e). Note that glycerol could not support bacterial condensation at low pH (Fig. 3c). Importantly, the roles of Tar and Tsr reversed in alkaline pH: while Tsr mediated condensation in acidic environments (Fig. 2), Tar mediated condensation in alkaline environments. Thus, we conclude that, depending on the environmental conditions, both Tar and Tsr can drive bacterial condensation, consistent with their roles in pH taxis.
A model for self-induced bacterial condensation in acidic environments
To test if feedback between the tendency of cells to neutralize the pH in their environment and their repulsion from low pH is sufficient to account for the observed bacterial condensation (Fig. 1c), we modeled the dynamics of cell density, \(n(\overrightarrow{r},\,t)\), in the presence of a degradable repellent \(c(\overrightarrow{r},t)\) (here protons) based on the Keller-Segel equations45 (see SI Model for details):
E1. \(\frac{\partial n(\overrightarrow{r},\,t)}{\partial t}={D}_{cell}\cdot {\nabla }^{2}n-\overrightarrow{\nabla }(n\cdot {\overrightarrow{v}}_{d})\) (cell-density dynamics)
E2. \(\frac{\partial c(\overrightarrow{r},t)}{\partial t}={D}_{H}\cdot {\nabla }^{2}c-{r}_{H}\cdot n\) (proton-density dynamics)
where \({D}_{{cell}}\) is the effective cell diffusion coefficient, due to active random swimming; \({\vec{v}}_{d}\) is the chemotaxis-induced drift velocity; \({D}_{H}\) is the proton diffusion coefficient; and \({r}_{H}\) is the proton neutralization rate, approximated by \({r}_{H}=\gamma {c}^{3/2}\), based on the data shown in Fig. 3 and S7 (see also SI Model). The chemotactic drift velocity was estimated based on ref. 46.
E3. \({\overrightarrow{v}}_{d}=\beta \overrightarrow{\nabla }f;f=\,{{\mathrm{ln}}}\left(\frac{c+{K}_{off}}{c+{K}_{on}}\right)\) (chemotaxis)
with \(\beta\) being the chemotaxis coefficient, and \({K}_{{on}}\) and \({K}_{{off}}\) are the effective dissociation constants characterizing the sensory response to protons21,46,47. The rest of the model parameters were estimated using available data21 (summarized in SI Model Table M1).
The numerical solution of this model under conditions that mimic the ‘thin-layer’ experiments can indeed demonstrate bacterial condensation similar to that observed experimentally, with similar spatial and time scales (Fig. 3f and SI Model Fig. M4). Moreover, this model also captures the highly non-linear dependence of cell condensation on the initial uniform cell density (Fig. 3d and f), except that the simulated condensation kept increasing with cell density rather than reaching saturation. This discrepancy may be due to aerotaxis, which becomes effective at high cell density but was not included in the model. Overall, the model confirms that self-induced pH-taxis can indeed lead to the observed bacterial condensation.
Coupling to the environment is essential for sustained bacterial condensation
A dominant property of the setups tested here (Fig. 1b, c) is the effective chemical coupling between the bacterial layer and the larger cell-free environment underneath. To test the role of this coupling, we introduced a PDMS layer at the bottom of the bacterial layer (Fig. 4a). This layer is permeable to oxygen but disrupts the free diffusion of protons from the reservoir to the bacterial layer, leading to a more ‘isolated’ cell population, with only the lateral edges of the bacterial layer in contact with the larger environment. Evidently, bacterial condensation was robust in the ‘coupled’ setup, but diminished in the partly ‘isolated’ setup (Fig. 4a; blue and orange symbols, respectively). At an ambient pH of 5.5, no significant condensation was observed in the partly ‘isolated’ setup (Fig. 4a, left), and at a lower ambient pH of 5.2, only reduced and transient condensation was observed (Fig. 4a, right). Furthermore, fully isolated populations did not exhibit condensation and remained uniform even at pH 5.2 (Supplementary Fig. S1). Thus, chemical coupling between the bacterial suspension and a cell-free environment is essential for maintaining sustained cell accumulations.
a In addition to the ‘thin-layer’ setup (upper illustration, Fig. 1c), we also used here a modified setup where an oxygen-permeable PDMS layer was added at the bottom of the bacterial layer (lower illustration). The added PDMS layer partly decouples the cell suspension from the rest of the chamber (except at the lateral edges) and thus, leads to a partly isolated cell population. Time traces of the cell accumulation are shown for experiments conducted either with (orange symbols) or without (blue symbols) the PDMS layer, and with an initial pH of 5.2, 5.5 or 5.8 (as labeled). Error bars represent standard deviation in, at least, three repetitions. b Using the model (Eqs. 1-3), we plotted the pH field (lower-left plot), 10 min after triggering a static cell accumulation: \(n(r)={n}_{0}\left(\right.1+(1/2)\cdot \exp [-{2(r/w)}^{2}]\)), where \(n\) is the cell density, \({n}_{0}\) set to OD600 0.4, and w = 1 mm (top-left plot). The temporal evolution of the corresponding pH baseline and maximal gradient, as well as the chemotactically-induced cell drift are shown on the right (top to bottom, respectively) for the two realizations, with (orange lines) or without (blue lines) the PDMS layer. See SI Model for details and parameters values. In particular, cell drift velocity corresponds to \({{{\rm{V}}}}_{D} \sim \nabla {{\rm{pH}}}/(1+{10}^{{pH}-p{K}_{{on}}})\), where \(p{K}_{{on}}=-\log ({K}_{{on}})\). To avoid edge effects, the setup was assumed to be much larger than the cell-density perturbation.
To explore the potential role of coupling, we used the model to evaluate how a static Gaussian cell-density perturbation modulates the proton (pH) distribution in either a ‘coupled’ or ‘isolated’ setup (Fig. 4b). The corresponding modulation in the proton density was estimated numerically by considering proton diffusion that is either three-dimensional, mimicking the ‘coupled’ setup (Fig. 4b, blue lines), or two-dimensional, mimicking the ‘isolated’ setup (Fig. 4b, orange lines). See SI Model for details. As expected, in the ‘isolated’ setup, the baseline pH increased more rapidly and the self-induced gradients were stronger (Fig. 4b). On the other hand, since the pH modulation tends to saturate close to neutral pH (Fig. 3a and Supplementary Fig. S7), the pH gradient eventually decreased as the pH baseline increased (Fig. 4b). However, the bacterial chemotactic response depends not only on the local gradient but also on the ability of the cells to sense the gradient, which is restricted to a certain dynamic range set by the sensory properties of the receptors21, \({K}_{{on}}\) and \({K}_{{off}}\). Evidently, no condensation was observed when the initial ambient pH was closer to pH 5.8 (Fig. 4a, right-lower plot). Thus, for a short time, the chemotactic cell drift is larger in the ‘isolated’ setup (Fig. 4b), but then declines as the baseline pH approaches the upper limit of the bacterial sensory dynamic range. These properties are consistent with the observation that ‘isolated’ cell populations failed to form sustained condensations (Fig. 4a). Thus, coupling to a large reservoir, while limiting the self-induced pH gradients, is essential for maintaining the overall baseline pH within the dynamic range of the bacterial chemotactic responses.
Dynamically-formed ‘embedded’ cell populations
A bacterial population can be restricted to occupy only part of a larger environment not only by physical constraints, as described so far (Fig. 1), but also dynamically, by additional external chemical gradients. An example is shown in Fig. 5, where an initially uniform cell population faces a local source of conflicting effectors; a source that was both set to acidic pH (repellent) and contained MeAsp (attractant). As previously demonstrated21, when the buffering capacity of the medium is strong, bacteria tend to form a narrow band (~1 mm wide) at a distance from the source, balancing the opposing forces (Fig. 5; ‘strong buffer’). Notably, this band of cells is chemically coupled to the rest of the channel. Interestingly, repeating these experiments now using a medium with lower buffering capacity, we observed a clear condensation of the bacterial band into a single focal point (Fig. 5, ‘weak buffer’). Condensation was not observed in the absence of the source (Supplementary Fig. S8).
A uniform cell suspension (OD600 0.1 in motility medium, pH 7) was placed in a long channel (see illustration), and a reservoir on the left was filled with motility medium set to pH 4.5 (repellent), but also contained MeAsp (10 mM; attractant), see Methods for details. Experiments were conducted with the medium containing either a ‘strong buffer’ (upper panel; 1 mM KPi) or ‘weak buffer’ (lower panel; no phosphate buffer). Fluorescence images were taken at the indicated times (in minutes). White bar: 1 mm. In both cases, the combination of attractant and repellent gradients led to strong accumulation of the cells in a narrow band, effectively ‘coupled’ to the rest of the channel (top-right illustration), however, with the weaker buffer, self-induced pH gradients (blue arrows) cause the cells to further condense into a focal point (lower-right illustration).
Bacterial condensation fosters formation of cell aggregates
The experiments described thus far were conducted with cells pre-grown at 33.5 °C. We next sought to test whether bacterial condensation may occur with cells pre-grown at 37 °C, which could be more relevant in the context of human environments. Evidently, bacterial condensation could also be observed in experiments with cells pre-grown at 37 °C (Fig. 6a and Supplementary Fig. S9). However, in this case, following their condensation into a millimeter-sized focal point, part of the bacteria tended to aggregate and form free-floating clumps comprising hundreds of cells, with a typical diameter of 10–20 μm (Fig. 6a and SI Videos 4-7). Interestingly, such aggregates were clearly observed with cells pre-grown at 37 °C, independent of the temperature at which the experiment was conducted, whether 24 °C, 30 °C, or 37 °C (Supplementary Fig. S9). In contrast, cell aggregates were not observed with cells pre-grown at 33.5 °C, regardless of the temperature at which the experiments were conducted (Supplementary Fig. S9). Notably, large-scale bacterial condensation was observed in all cases. Thus, we concluded that growing the cells at 37 °C was essential for the formation of aggregates.
a Experiments using the ‘thin-layer’ setup (as in Fig. 1c) were repeated with cells pre-grown at 37 °C (see “Methods”). A log-scale fluorescence intensity image, at t ~ 3 h, is shown (on the right) demonstrating both the main condensation of the free-swimming single cells (planktonic cells) at the center and the cell aggregates at the periphery. White bar: 200 μm. Additional high-magnification images of typical cell aggregates are shown on the left (white bar: 20 μm; see also SI Videos 4-7). The corresponding dynamics of cell aggregation is plotted below (blue symbols), evaluated by their integrated intensity (see Methods). Similar dynamics is also shown for experiments with a medium containing a stronger buffer that inhibits cell condensation (10 mM KPi; open triangles), or with Δag43 cells that still exhibit clear condensation (open circles). See also SI Video 8 and Supplementary Fig. S11. Note, aggregates with a diameter smaller than approximately 13 μm were not considered in this analysis. Also shown is the fit to the corresponding dynamics of the primary cell condensation (dashed gray line). b The spatial distribution of the cell aggregates at 3 h (radially averaged, see Methods) compared with the spatial distribution of the condensed free-swimming single cells (dashed gray line). c Experiments were conducted using the ‘thin-layer’ setup (Fig. 1c) with cells pre-grown at 33.5 °C and with fluorescent spheres (16 μm diameter) added to the cell suspension. Sample images of the spheres before (3 min) and after (50 min) the bacterial condensation are shown (white bar: 300 μm; SI Video 9). The corresponding spatial distribution of the spheres at 50 min is shown on the left (gray bars; radially averaged) along with the single-cell distribution (gray line). The distribution of spheres was fitted (thick black line) using a model for bacterial-condensation-induced force, which postulates that the drift velocity of the spheres is proportional to the cell-density gradient of the condensing cells (see SI Model).
To quantify cell aggregation, images like the one shown in Fig. 6a (right image) were analyzed to separately evaluate the total intensity of the cell aggregates and the total intensity of the condensed free-swimming planktonic cells (SI Methods). The time course of these intensities is shown in Fig. 6a, demonstrating that the formation of cell aggregates lags behind the condensation of the planktonic cells and occurs only when the cell density was already considerably elevated. On the other hand, preventing the bacterial condensation by adding a strong buffer to the medium (10 mM KPi) also inhibited cell aggregation (Fig. 6a, open triangles), indicating that cell aggregation is not merely triggered by the acidic environment, but rather by bacterial condensation. Given that density-dependent cell aggregation could also be observed with uniform cell populations (Supplementary Fig. S10), bacterial auto-aggregation can be triggered by the primary condensation due to the local elevation in cell density.
Antigen 43 (Ag43) is a known bacterial appendage that mediates physical contacts between bacterial cells grown at 37 °C40,48,49,50. To test if the bacterial aggregates observed here also rely on the same mechanism, we constructed a strain missing the Ag43 gene. We found that cells lacking Ag43, despite exhibiting large-scale condensation similar to that of Ag43+ cells, failed to form bacterial aggregates (Fig. 6a, open circles, Supplementary Fig. S11 and SI Video 8). Therefore, we concluded that cell aggregates are formed by Ag43-mediated contacts promoted by the local increase in cell density during bacterial condensation.
Interestingly, despite being triggered by cell condensation, cell aggregates were mostly found at the periphery of the planktonic-cell accumulation rather than its center (Fig. 6a, b). Given that cell aggregates are large and most likely non-chemotactic objects, we hypothesized that the accumulation of chemotactic planktonic cells exerts a physical force that expels these aggregates. To determine whether the accumulating bacteria can exert such physical forces, we conducted experiments with cells pre-grown at 33.5 °C and introduced fluorescent spheres into the initial cell suspension (see Methods). Note that cells grown at this temperature exhibit condensation but not cell aggregation. These spheres, approximately 16 µm in diameter, were comparable in size to the bacterial aggregates. We found that the cell accumulation indeed generates significant forces that can displace the spheres from its center (Fig. 6c, see also SI Video 9). We propose that the same physical forces also displace the free-floating bacterial aggregates (see also SI Model).
Discussion
Organic weak acids can pose an acute challenge to bacteria, mediated by an influx of protons that directly acidify the intracellular environment. Here we show that when immersed in a uniform, low-buffering-capacity acidic environment containing organic acids, E. coli populations respond by condensing into discrete focal points (Fig. 1). This condensation is triggered by an inherent instability resulting from feedback between the chemotactic tendency of these bacteria to avoid low pH environments and their ability to neutralize the pH, illustrated in Fig. 7a (Figs. 2–3). Similar condensation was found also in alkaline pH, provided that cells could neutralize the pH (Fig. 3e and Supplementary Fig. S6). The chemoreceptors Tsr and Tar drive this behavior in acidic and alkaline environments, respectively (Figs. 2 and 3e). Importantly, sustained bacterial condensation also requires an effective exchange of chemicals (coupling) between the bacterial suspension and a larger cell-free environment (Fig. 4, see Fig. 1a), which restricts the self-induced pH modulation to the range that can elicit effective chemotactic responses (Fig. 4). Such coupling can be realized either by physical constraints (e.g., Fig. 1c) or dynamically, by additional chemotactic cues (Fig. 5).
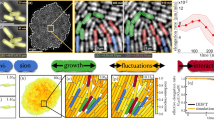
a Schematic summary of the observations made here. By neutralizing the protons associated with organic-acids, cells create self-induced pH gradients that, in turn, are sensed by the Tsr receptors and trigger collective condensation. The elevated local cell-density leads to further cell aggregations. Finally, these large and non-chemotactic cell aggregates are actively segregated from the free-swimming planktonic cells. b Benefits of bacterial condensation. The expected population-averaged pH (filled-blue symbols) and oxygen concentration (filled-green symbols) for bacteria forming static gaussian distributions with varying width (w) (see SI Model). These quantities are plotted at 3 h, corresponding to the quasi-static cell accumulation (Fig. 2). The shaded areas represent uncertainty in the rate of oxygen and proton consumption. We also estimated the population-averaged pH (open-blue symbols) or oxygen concentration (open-green symbols) experienced by cells that dynamically condense (highlighted by the black arrow; see SI Model). Notably, as the bacteria condense (smaller width) cells experience less pH stress but reduced level of oxygen.
To quantitatively assess the potential benefits of bacterial condensation, we first considered cell populations with a spatially static Gaussian distribution with varying width \(w\) and constant number of cells \({N}_{0}\): \(n\left(r\right)={N}_{0}\cdot (2/\pi {w}^{2})\cdot \exp [-{2\left(r/w\right)}^{2}]\). Using Eq. 2, we evaluated the population-averaged pH and oxygen concentrations encountered by the cells, shown in Fig. 7b, as blue and green symbols, respectively. As the population becomes more condensed (\(w\) decreases), cells experience a more neutral pH but also a lower oxygen concentration. Both trends stem from the generic non-linear dependence of diffusion time on distance, which leads to ineffective recovery of chemical depletion over larger distances. The corresponding pH and oxygen concentrations were also evaluated for dynamically condensing cell populations (Eqs. 1–3, SI Model) and are shown in Fig. 7b (open symbols), demonstrating a similar tradeoff between the low pH and oxygen depletion. Interestingly, despite this tradeoff, cells clearly favor condensation that alleviates, at least partially, the pH stress. Only when the bacterial accumulation becomes too strong does Aer-mediated aerotaxis tends to limit the cell condensation. Such oxygen-limited condensation is clear in the case of Tar− cells (Fig. 2), which tended to condense more strongly. Additionally, when two bacterial accumulations coalesce (e.g., Fig. 1d), a ring forms instead of an abrupt increase in cell density. Finally, it was previously suggested that E. coli cells aspire to moderate oxygen levels of approximately 50 µM51, although this may vary between E. coli strains27. Thus, the tradeoff between pH tolerance and oxygen limitation is dynamic and can be context dependent.
Overall, the collective condensation of bacteria observed here provides a spontaneous adaptive response to mitigate acid stress and enables bacteria to activate more specialized adaptive transcription programs under milder stress conditions. Moreover, the condensed cells remain motile, and thus, as demonstrated by their coalescence (Fig. 1d), the accumulated bacteria can move in response to gradients as a cohesive multicellular object. Interestingly, while the coalescence behavior presumably involves pH taxis, other environmental cues may also trigger collective movement of the condensing cells. Furthermore, once the acid stress has been diminished, the motile cells can readily disperse, except those that were aggregated. Such dispersion was evident under partially isolated conditions (Fig. 4a, right plot, orange symbols). Similar cell dispersion can be expected also under the coupled conditions for much longer time, such that the diffusion length exceeds the overall size of the environment.
Although the neutrophilic bacterium E. coli was used here as a model, similar behavior could be expected from other chemotactic bacteria as well. Different bacterial species have different pH preferences, such that a shift in pH from its optimal value generically leads to stress and growth inhibition19,52,53. Thus, if each cell can contribute to modifying the pH in the preferred direction, cooperative cell condensation similar to that demonstrated here could also be expected. Moreover, similar condensation can also be expected under different stress conditions, provided they are mediated by factors that: first, serve as chemorepellents, and second, can be mitigated by individual cells. Finally, once the bacteria condense, additional effectors may further influence the shape of the bacterial accumulation and its future development. A prominent candidate might be the quorum sensing molecules AI-232.
By fostering cell aggregation and potentially biofilm formation41,42 (Fig. 6), pH-induced cell condensation may more broadly enhance the bacterial tolerance to diverse stress factors. The formation of these cell aggregates was strictly dependent on Ag43 (Fig. 6 and Supplementary Fig. S11), a well-known component that holds cells together in biofilms. Thus, cell condensation can foster cell aggregation simply by elevating the local cell density (Supplementary Fig. S10). However, the secretion of AI-2 may also contribute to the formation of micro-scale aggregates by forming local gradients on the scale of the inter-bacterial distance39,54. In principle, similar micro-scale pH gradients may also form by local bacterial-induced neutralization. Interestingly, cell aggregates were displaced by the condensed planktonic cells (Figs. 6 and 7a), most likely due to an effective outward force induced by the chemotaxis-driven bacterial accumulation (SI Model). This segregation may reduce competition between the two sub-populations over nutrients and, under proper conditions, may also mediate inter-species niche segregation.
Materials and methods
Strains and Plasmids
We used the E. coli MG1655 (VF6, refs. 27,28) and its derivatives harboring deletion of tar (NL1), tsr (MK20), aer (AVE3), tar aer (MK21), cheY cheZ (MK3), or ag43 (also known as flu, MK29). Cells expressed also GFP from pTrc99a vector induced with 75 µM IPTG (pSA11).
Cell growth and motility medium
Unless otherwise specifically mentioned, cells (E. coli MG1655) were grown over night in Tryptone Broth (TB, tryptone extract 10 g/l and 5 g/l NaCl) supplemented with Amp, diluted 100-fold into fresh medium with IPTG (75 µM), and allowed to grow at 33.5 °C with shaking. Cells were harvested at an OD600 of 0.5 by centrifugation and gently washed twice in motility medium and resuspended in motility medium with final cell concentration of OD600 0.4. Unless otherwise mentioned, ‘motility medium’ included: 0.1 mM potassium phosphate (KPi), 0.94 mM lactic acid, 85.5 mM sodium chloride, 0.1 mM EDTA, and 1 µM methionine, titrated to the desired pH with NaOH. Notably, this medium supports long-term bacterial motility but not cell proliferation.
The ‘thin layer’ setup
Schematically illustrated in Fig. 1c. An agarose gel (3%, in motility medium) was casted in a cylindrical chamber made from grade-2 titanium (height: 6 mm) and sealed at the top with a mold with a thin cylindrical protrusion, which will form the bacterial layer (~120 μm). The diameter of bacterial layers (the molds) varied between 0.5 and 10 mm and were fabricated using a resin printer (Elegoo Mars 3). The diameter of the titanium chamber was mostly 10.7 mm, except in the experiments shown in Fig. 1d, where the chamber had a diameter of 18 mm. After the agarose solidified, the mold and the bottom cover were removed and the agarose was dialyzed overnight in motility buffer and a second time, for an additional 4 h, in a fresh motility buffer just before the experiment (final dialysis ratio of 104:1). Bacterial cells were then introduced into the shallow well, and the bottom and top of the chamber were covered with glass coverslips and secured in position. The entire setup was then incubated inside a humidity chamber at 30 °C, except for the aggregation experiments (Fig. 6 and Supplementary Fig. S10–11), which were performed at room temperature (24 °C). See also Fig. S9. The redistribution of the cells was followed by tracking changes in the fluorescence distribution. In the experiments described in Fig. 4 (‘isolated’), a thin oxygen-permeable PDMS (Sylgard 184, Dow Corning) slab was placed at the bottom of the agarose well, separating the swimming cells in the well from the larger chamber. Note that even in this setup, the lateral edges of the cell suspension remained in contact with the gel.
Agarose plate assay
Standard 90 mm plates were filled with agarose hydrogel (1%, in motility medium, pH 5.2). As in the ‘thin layer’ setup, once the agarose solidified, the plates were dialyzed overnight in a motility buffer and a second time, for an additional 4 hours, in a fresh motility buffer just before the experiment (final dialysis ratio of 102:1). GFP-expressing bacteria were then applied by pipetting 1 ml of the bacterial suspension on top the hydrogel and allowed to expand and form a thin layer. Plates were then kept at room temperature (24 °C). To observe the fluorescence distribution, plates were illuminated using uniform LED illumination at 470 nm (runVIEW Mini Blue, Cleaver Scientific), and viewed through a dedicated filter screen using a DLSR camera (Canon EOS2000D). The image in Fig. 1b presents the green channel of the RGB image.
The ‘channel’ setup
This assay has been fully described in ref.21, and is schematically illustrated in Fig. 5. A long channel (44 mm long, 5 mm wide, and 0.2 mm high; ibidi, μ-Slide) was filled with cell suspension (OD600 0.1) in motility medium (pH 7, 9.4 mM lactic acid). One end of the channel was sealed, and the other was filled with a hydrogel (1.5% agarose) containing the same motility medium that was titrated to pH 4.5 (repellent) and supplemented with α-methyl-aspartate (10 mM; attractant; Sigma-Aldrich). The channel was then incubated at 30 °C inside a humidity chamber. The redistribution of the cells was followed by tracking changes in the fluorescence distribution.
Fluorescence microscopy
The bacterial distribution was followed using a Nikon Ti microscope equipped with a 20 × (0.5 NA) or 4 × (0.2 NA) objective, a decoded controlled stage and a ‘perfect-focus’ system. Large images were obtained by stitching smaller images (approximately 2 mm × 2 mm each) with 20% overlap using a built-in function in NIS Elements software (Nikon). The bacterial distribution inside the well was extracted from the images by removing the measured background intensity and normalizing to the average intensity within each well. Radial averaging was done by measuring the fluorescence intensity around the center of focal points (Radial Profile Angle plugin for ImageJ). The fluorescence distribution of the fluorescein dye shown in Fig. 3b was obtained using a 60x (0.7 NA) objective to minimize background fluorescence due to the diffusion of fluorescein into the agarose hydrogel (see also SI Methods).
Measurements of pH neutralization by bacteria
In the experiments described in Fig. 3a, following their growth, cells were resuspended in 20 ml of motility medium modified as specified. The cell suspension was placed in a 50 ml conical beaker at 30 °C with stirring. The pH of the medium was measured continuously (Mettler Toledo, InLab Expert Pro-ISM). We confirmed that the presence of cells did not influence the pH measurements by occasionally comparing the measured pH with that of a sample from which the cells were removed by centrifugation and filtration (0.2 μm).
Testing for chemo-effectors in the neutralized bacterial suspensions
The experiments for measuring pH neutralization by the bacteria were performed as described above with the standard motility medium (containing lactate). When the pH of the bacterial suspension approached neutral pH (t = 2 h), the cells were cleared out by centrifugation and filtration (0.2 µm filter), and phosphate buffer was added to a final concentration of 10 mM (at pH 7). The pH-neutral supernatant was then applied to one side of the ‘channel’ chemotaxis assay (see above), and the response of the bacteria in the channel was followed. The results are presented in Supplementary Fig. S5.
Quantifying cell aggregates intensity
The intensity of the cell auto-aggregates within the entire bacterial layers (d = 5 mm) was separated from the planktonic cells based on their distinguished properties: auto-aggregates are larger, brighter, and move slower compared with single cells (see SI Methods and SI Videos 4–7). Briefly, a low-pass filter was used to separate the aggregates from the cell accumulation (see SI Video 10). Then, aggregates were detected, and their intensity and position were captured. In this analysis, only ‘mature’ aggregates, whose area was larger than that of a sphere with a diameter of 14 μm (10 pixels), were considered. The results of this analysis were cross-verified by eye with the original images to ensure accurate aggregate detection and to rule out data artefacts. To improve the identification of aggregates, we eliminated the non-specific attachment of bacteria to the glass coverslips by passivating them with 1% BSA for 30 minutes, after which they were thoroughly washed with double-distilled water (DDW) and then dialyzed in DDW for 3 h. BSA treatment of the glass coverslips did not, by itself, lead to auto-aggregation of cells (Supplementary Fig. S9).
Fluorescent spheres
In the experiments presented in Fig. 6c, we used red fluorescent spheres with a 16 μm diameter (FluoSpheres, ThermoFisher Scientific). Before use, spheres were thoroughly washed in motility medium to which Tween-20 (0.01%) was added. The spheres were then added to a cell suspension, which also contained Tween-20 (0.01%). The added Tween-20 prevented self-aggregation of the fluorescent spheres but still allowed bacterial condensation.
Model
Full details can be found in SI Model. Briefly, the Keller-Segel equations (Eq. 1–2) were numerically solved assuming cylindrical symmetry, calculating spatial derivatives, and using a 4th order Runge-Kutta method for the time integration. Proton uptake in the bacterial layer was accounted for by imposing absorbing boundary conditions for the protons at the bacterial layer interface (Robin boundary conditions). The parameters used in the simulations were estimated from refs. 21,55. The error bars in the calculated values shown in Fig. 7 were estimated by incorporating the uncertainty in these parameters (see SI Model for further details).
Statistics and reproducibility
To ensure reproducibility, at least several biological replicates (n ≥ 3) were done for each experiment. In the vast majority of cases, experiments were repeated also on several days with new cultures each time. The findings reported here were very robust and appeared in each experiment, although with small variations in the magnitude of the cell condensation (e.g., see the amplitude variations observed in Fig. 2). Averages and standard deviations shown throughout were calculated after ensuring the data are randomly distributed around the mean; the number of replicates used to calculate these values is indicated in the figure captions.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data are available upon request.
References
Schwarz, J., Schumacher, K., Brameyer, S. & Jung, K. Bacterial battle against acidity. FEMS Microb. Rev. 46, fuac037 (2022).
Cotter, P. D. & Hill, C. Surviving the acid test: responses of gram-positive bacteria to low pH. Microbiol. Mol. Biol. Rev. 67, 429–453 (2003). table of contents.
Slonczewski, J. L., Fujisawa, M., Dopson, M. & Krulwich, T. A. Cytoplasmic pH measurement and homeostasis in bacteria and archaea. Adv. Microb. Physiol. 55, 1–79, 317 (2009).
Axe, D. D. & Bailey, J. E. Transport of lactate and acetate through the energized cytoplasmic membrane of Escherichia coli. Biotechnol. Bioeng. 47, 8–19 (1995).
Slonczewski, J. L., Macnab, R. M., Alger, J. R. & Castle, A. M. Effects of pH and repellent tactic stimuli on protein methylation levels in Escherichia coli. J. Bacteriol. 152, 384–399 (1982).
Repaske, D. R. & Adler, J. Change in intracellular pH of Escherichia coli mediates the chemotactic response to certain attractants and repellents. J. Bacteriol. 145, 1196–1208 (1981).
Lund, P. A. et al. Understanding how microorganisms respond to acid pH is central to their control and successful exploitation. Front. Microbiol. 11, 556140 (2020).
Branson, S. R., Broadbent, J. R. & Carpenter, C. E. Internal pH and acid anion accumulation in Listeria monocytogenes and Escherichia coli exposed to lactic or acetic acids at mildly acidic pH. Front. Microbiol. 12, 803271 (2021).
Koh, A., De Vadder, F., Kovatcheva-Datchary, P. & Bäckhed, F. From dietary fiber to host physiology: short-chain fatty acids as key bacterial metabolites. Cell 165, 1332–1345 (2016).
Wolfe, A. J. The acetate switch. Microbiol. Mol. Biol. Rev. 69, 12–50 (2005).
Wang, S. P. et al. Pivotal roles for pH, lactate, and lactate-Utilizing bacteria in the stability of a human colonic microbial ecosystem. mSystems 5, e00645 (2020).
Macias-Benitez, S. et al. Rhizospheric organic acids as biostimulants: monitoring feedbacks on soil microorganisms and biochemical properties. Front. Plant Sci. 11, https://doi.org/10.3389/fpls.2020.00633 (2020).
Roussin-Léveillée, C., Mackey, D., Ekanayake, G., Gohmann, R. & Moffett, P. Extracellular niche establishment by plant pathogens. Nat. Rev. Microbiol. 6, 360–377 (2024).
Corbet, C. & Feron, O. Tumour acidosis: from the passenger to the driver’s seat. Nat. Rev. Cancer 17, 577–593 (2017).
Foster, J. W. Escherichia coli acid resistance: tales of an amateur acidophile. Nat. Rev. Microbiol. 2, 898–907 (2004).
Kanjee, U. & Houry, W. A. Mechanisms of acid resistance in Escherichia coli. Annu. Rev. Microbiol. 67, 65–81 (2013).
Guan, N. & Liu, L. Microbial response to acid stress: mechanisms and applications. Appl. Microbiol. Biotechnol. 104, 51–65 (2020).
Brameyer, S., Schumacher, K., Kuppermann, S. & Jung, K. Division of labor and collective functionality in Escherichia coli under acid stress. Commun. Biol. 5, 327 (2022).
Krulwich, T. A., Sachs, G. & Padan, E. Molecular aspects of bacterial pH sensing and homeostasis. Nat. Rev. Microbiol. 9, 330–343 (2011).
Zychlinsky, E. & Matin, A. Cytoplasmic pH homeostasis in an acidophilic bacterium, Thiobacillus acidophilus. J. Bacteriol. 156, 1352–1355 (1983).
Livne, N. & Vaknin, A. Collective responses of bacteria to a local source of conflicting effectors. Sci. Rep. 12, 4928 (2022).
Yang, Y. & Sourjik, V. Opposite responses by different chemoreceptors set a tunable preference point in Escherichia coli pH taxis. Mol. Microbiol. 86, 1482–1489 (2012).
Adler, J. Chemotaxis in bacteria. Science 153, 708–716 (1966).
Berg, H. C. E. coli in Motion (Springer, 2004).
Wuichet, K., Cantwell, B. J. & Zhulin, I. B. Evolution and phyletic distribution of two-component signal transduction systems. Curr. Opin. Microbiol. 13, 219–225 (2010).
Barker, C. S., Prüß, B. M. & Matsumura, P. Increased motility of Escherichia coli by insertion sequence element integration into the regulatory region of the flhD operon. J. Bacteriol. 186, 7529–7537 (2004).
Tamar, E., Koler, M. & Vaknin, A. The role of motility and chemotaxis in the bacterial colonization of protected surfaces. Sci. Rep. 6, 19616 (2016).
Frank, V., Piñas, G. E., Cohen, H., Parkinson, J. S. & Vaknin, A. Networked chemoreceptors benefit bacterial chemotaxis performance. mBio 7, e01824–16 (2016).
Parkinson, J. S., Hazelbauer, G. L. & Falke, J. J. Signaling and sensory adaptation in Escherichia coli chemoreceptors: 2015 update. Trends Microbiol 23, 257–266 (2015).
Bi, S. & Sourjik, V. Stimulus sensing and signal processing in bacterial chemotaxis. Curr. Opin. Microbiol. 45, 22–29 (2018).
Hu, B. & Tu, Y. Precision sensing by two Opposing gradient sensors: How does Escherichia coli find its preferred pH level? Biophys. J. 105, 276–285 (2013).
Keegstra, J. M., Carrara, F. & Stocker, R. The ecological roles of bacterial chemotaxis. Nat. Rev. Microbiol. 20, 491–504 (2022).
Fu, X. et al. Spatial self-organization resolves conflicts between individuality and collective migration. Nat. Comm. 9, 2177 (2018).
Cremer, J. et al. Chemotaxis as a navigation strategy to boost range expansion. Nature 575, 658–663 (2019).
Budrene, E. O. & Berg, H. C. Complex patterns formed by motile cells of Escherichia coli. Nature 349, 630–633 (1991).
Budrene, E. O. & Berg, H. C. Dynamics of formation of symmetrical patterns by chemotactic bacteria. Nature 376, 49–53 (1995).
Blat, Y. & Eisenbach, M. Tar-dependent and -independent pattern formation by Salmonella typhimurium. J. Bacteriol. 177, 1683–1691 (1995).
Mittal, N., Budrene, E. O., Brenner, M. P. & Van Oudenaarden, A. Motility of Escherichia coli cells in clusters formed by chemotactic aggregation. Proc. Natl. Acad. Sci. 100, 13259–13263 (2003).
Laganenka, L. et al. Chemotaxis and autoinducer-2 signalling mediate colonization and contribute to co-existence of Escherichia coli strains in the murine gut. Nat. Microbiol. 8, 204–217 (2023).
Laganenka, L., Colin, R. & Sourjik, V. Chemotaxis towards autoinducer 2 mediates autoaggregation in Escherichia coli. Nat. Commun. 7, 12984 (2016).
Sauer, K. et al. The biofilm life cycle: expanding the conceptual model of biofilm formation. Nat. Rev. Microbiol. 20, 608–620 (2022).
Cai, Y. M. Non-surface attached bacterial aggregates: a ubiquitous third lifestyle. Front. Microbiol. 11, 557035 (2020).
Hall-Stoodley, L., Costerton, J. W. & Stoodley, P. Bacterial biofilms: from the natural environment to infectious diseases. Nat. Rev. Microbiol. 2, 95–108 (2004).
Rebbapragada, A. et al. The Aer protein and the serine chemoreceptor Tsr independently sense intracellular energy levels and transduce oxygen, redox, and energy signals for Escherichia coli behavior. Proc. Natl. Acad. Sci. USA 94, 10541–10546 (1997).
Keller, E. F. & Segel, L. A. Traveling bands of chemotactic bacteria: a theoretical analysis. J. Theor. Biol. 30, 235–248 (1971).
Si, G., Wu, T., Ouyang, Q. & Tu, Y. Pathway-based mean-field model for Escherichia coli chemotaxis. Phys. Rev. Lett. 109, 048101 (2012).
Hu, B. & Tu, Y. Behaviors and strategies of bacterial navigation in chemical and nonchemical gradients. PLoS Comp. Biol. 10, e1003672 (2014).
Heras, B. et al. The antigen 43 structure reveals a molecular Velcro-like mechanism of autotransporter-mediated bacterial clumping. Proc. Natl. Acad. Sci. USA 111, 457–462 (2014).
Danese, P. N., Pratt, L. A., Dove, S. L. & Kolter, R. The outer membrane protein, antigen 43, mediates cell-to-cell interactions within Escherichia coli biofilms. Mol. Microbiol. 37, 424–432 (2000).
Vo, J. L. et al. Variation of Antigen 43 self-association modulates bacterial compacting within aggregates and biofilms. npj. Biofilmses Microbiomes 8, 20 (2022).
Taylor, B. L., Watts, K. J. & Johnson, M. S. Oxygen and redox sensing by two‐component systems that regulate behavioral responses: behavioral assays and structural studies of Aer using in vivo disulfide cross‐linking. In Methds. Enzymol. (eds. Brian R. Crane Melvin I. Simon & Crane Alexandrine) 422, 190–232 (Academic Press, 2007).
Rothschild, L. J. & Mancinelli, R. L. Life in extreme environments. Nature 409, 1092–1101 (2001).
Fernández-Calviño, D. & Bååth, E. Growth response of the bacterial community to pH in soils differing in pH. FEMS Microbiol. Ecol. 73, 149–156 (2010).
Laganenka, L. & Sourjik, V. Autoinducer 2-dependent Escherichia coli biofilm formation is enhanced in a dual-species coculture. Appl. Environ. Microbiol. 84, e02638 (2018).
Andersen, K. B. & von Meyenburg, K. Are growth rates of Escherichia coli in batch cultures limited by respiration? J. Bacteriol. 144, 114–123 (1980).
Acknowledgements
We thank Alan J. Wolfe (Loyola University, Chicago) and Oded Agam (Hebrew University) for their helpful conversations. We thank the Milner Foundation for partly funding NL’s fellowship. This work was supported by the Israeli Science Foundation and the Minerva Center for Bio-Hybrid Complex Systems.
Author information
Authors and Affiliations
Contributions
N.L. and A.V. designed the experiments; N.L. performed the experiments and numerical calculations. M.K. prepared strains. N.L. and A.V. analyzed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Dario Mizrachi and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Tobias Goris. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Livne, N., Koler, M. & Vaknin, A. Collective condensation and auto-aggregation of Escherichia coli in uniform acidic environments. Commun Biol 7, 1028 (2024). https://doi.org/10.1038/s42003-024-06698-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-024-06698-1
- Springer Nature Limited