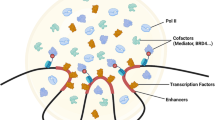
Abstract
Whether phase-separation is involved in the organization of the transcriptional machinery and if it aids or inhibits the transcriptional process is a matter of intense debate. In this Mini Review, we will cover the current knowledge regarding the role of transcriptional condensates on gene expression regulation. We will summarize the latest discoveries on the relationship between condensate formation, genome organization, and transcriptional activity, focusing on the strengths and weaknesses of the experimental approaches used to interrogate these aspects of transcription in living cells. Finally, we will discuss the challenges for future research.
Similar content being viewed by others
Introduction
The mechanisms behind gene transcription have puzzled researchers for decades. Transcription factors (TFs) are central actors in this process since they bind to regulatory elements1 triggering the recruitment of coactivators, chromatin remodelers, the Mediator complex, and RNA polymerase II (Pol II). The first textbook models of transcription were built considering the sequential, hierarchical formation of these macromolecular protein-DNA complexes through lock-and-key interactions2, but these models were insufficient to describe many aspects of transcription in cells.
One of the most perplexing observations is that the diffusion of TFs within the nucleus is not sufficient to explain their velocity in locating specific DNA targets within the entire genome3,4,5. Moreover, a wealth of evidence shows that TF-chromatin interactions are highly dynamic, with lifetimes spanning from less than a second to minutes6, in contrast to the traditional view of transcriptional complexes bound to chromatin for hours7. Puzzlingly, the stochastic and apparently inefficient TF-chromatin interactions result in sustained transcriptional bursts that can last significantly longer8 and are the consequence of many regulatory processes that involve TF binding to enhancers, enhancer looping, chromatin remodeling, and Pol II recruitment and activation through mechanism(s) not completely elucidated9,10.
Although it was widely known that liquid-liquid phase-separation (LLPS) contributes to many emergent properties of biological systems, including cellular organization11, a tsunami of reports in the last few years12 proposed that the selective phase-separation of TFs and other related molecules near target genes contributes to transcriptional regulation13. In this review, we will explore the different models explaining the nature and function of these condensates, their link with chromatin organization, and the experimental evidence both in favor and against their role in transcription.
Transcriptional hubs, condensates, or factories?
The discovery of membrane-less compartments within the nucleus dates from the nineteenth century with the foundational observations of the nucleolus14. Decades later, different electron microscopy techniques provided high-resolution views of the nuclear ultrastructure and its enormous variety of nuclear bodies with different functions (reviewed elsewhere15). In parallel, the improvement of fluorescence microscopy tools allowed the exploration of the intracellular distribution of biomolecules in live cells, leading to the observation that many actors involved in transcription do not distribute homogeneously throughout the nucleus but concentrate in distinct foci. In this section, we will discuss the different ideas aimed at explaining this heterogeneous organization. These models have granted different functions and names to these foci and still present challenges to be addressed.
A couple of decades ago, the observation of Pol II and active transcription sites concentrating as discrete foci within the nucleus of interphase cells led to the idea of transcription occurring within certain nuclear regions referred to as transcription factories (reviewed elsewhere16). This model—that contrasted with the classic idea of individual polymerases moving along an immobile DNA template17—postulates that Pol II molecules accumulate in stable clusters that physically interact with mobile chromatin loops to be transcribed18,19,20. Transcription factories may explain the transcription of some but not all genes, and consequently, they probably coexist with other modes of transcription21. In fact, the observation of Pol II molecules moving along very long and highly expressed genes22 suggests a more complex scenario than a static transcriptional platform.
Deeper scrutiny of transcription foci highlighted that they may concentrate many transcription-related molecules that interact among them weakly and with genomic loci forming networks with lifetimes below the minute time scale23. These dynamic networks are thought to favor the protein-protein and DNA-protein interactions required for transcription and are usually referred to as transcriptional hubs24. The propensity of TFs and other transcriptional players to form hubs was attributed to a common, modular structure shared by these molecules. Particularly, eukaryotic TFs present well-folded DNA-binding domains that recognize their chromatin targets and transactivation domains that usually include intrinsically disordered regions (IDRs)25 which could interact transiently with the IDRs of Pol II, cofactors, and other TFs26,27,28. Thus, both DNA-binding and transactivation domains of TFs would be responsible for the interaction network implicated in transcriptional hubs.
In the same direction, weak, multivalent IDR-IDR interactions seem to promote the phase-separation of IDR-containing proteins, including Med1, Pol II29, TBP30, BRD426, YAP31, and OCT427 in aqueous solutions. These observations led to the idea that similar droplet-like structures of transcription-related molecules with hallmarks of liquid phases could form inside cells and could play a role in transcription regulation26,32,33. According to this idea, chromatin regions act as scaffolds recruiting TFs and related molecules; this initial seed would grow by the incorporation of many other multivalent molecules with IDRs through weak interactions, finally forming a liquid transcriptional condensate34,35. We should point out that the terms ‘hubs’ and ‘liquid transcriptional condensates’ are not precisely defined in the literature yet; some authors are using the term ‘hubs’ to indicate that they are formed by a relatively lower number of molecules23 and do not specify the mechanism involved in their formation33. In contrast, it is assumed that liquid transcriptional condensates form by LLPS and include a larger number of molecules. In any case, the term ‘transcriptional condensate’ is now used to invoke any nuclear compartment concentrating biomolecules involved in transcription (Pol II, TFs, coactivators). Additionally, transcriptional repressors can also form condensates promoting transcriptional repression36, suggesting that condensation would act by favoring the specific activity of the condensed biomolecules, either activators or repressors. In this Mini Review, we will use the term ‘transcriptional condensates’ as an operational, agnostic definition, without implying that these structures necessarily form by phase-separation.
A recent report shows that the charge patterns of IDRs define the selective partitioning of biomolecules into different condensates, with possible functional consequences37, including the segregation of different transcriptional programs into specific transcriptional condensates38. Moreover, different types of transcriptional condensates seem to control several steps of gene activation, including the formation of preinitiation complexes, transcriptional pausing and elongation, super-enhancer clustering, and mRNA splicing and processing39,40. In fact, the biochemical reactions occurring during transcription may modulate the relative composition and properties of these condensates. For example, Pol II CTD phosphorylation will drive the transition from initiation to elongation, and therefore, the exit of Pol II molecules from initiation condensates to splicing condensates41. Also, newly synthesized RNA might regulate condensate formation and dissolution42. These features would work as feedback mechanisms controlling the different steps throughout mRNA biosynthesis.
However, other authors are challenging whether phase separation is involved in the formation of transcriptional condensates, claiming that alternative theoretical frameworks that also include weak and transient interactions between multivalent biomolecules could explain many of the properties assigned to liquid condensates43,44,45,46. Scrutinizing the biophysical properties of transcriptional condensates is especially challenging given their sub-diffraction size and the complex network of players involved, which make it difficult to use the standard in vitro approaches for studies in cells. There have been a few attempts to utilize single-molecule tracking in live cells as a tool to test whether distinct diffusive behaviors can be observed inside and outside condensates as predicted for distinct phases, leading to diverse and sometimes contradictory conclusions44,47,48.
Hence, the jury is still out on whether sufficient evidence has been provided for LLPS involvement in the formation of transcriptional condensates.
Chromatin organization under the phase-separation lens
Chromatin presents a hierarchical organization at different scales that span from the micron-sized territories occupied by each chromosome, sub-micron features as topologically associating domains (TADs) and loops49, to the recently characterized nano-domains50. Although it is now widely accepted that many aspects of this organization are intimately related to genome function51, the relationship is still far from being understood49. Complementary approaches52,53,54,55 focusing on the physical properties of this biopolymer and/or its biochemical features, provided some clues regarding chromatin folding in the nucleus and its relevance to its functions. Chromatin-chromatin interactions are invoked as one of the major forces driving genome self-organization, defining domains and compartments, among other features56. At a sub-micron spatial scale, active processes such as loop extrusion drive the formation of loops and TADs and, combined with the passive, local exploration of the chromatin chain by diffusion, allow the interactions between relevant transcriptional regulatory elements, including enhancers and promoters55,56.
In addition, many aspects of chromatin organization have been analyzed under the phase-separation framework52,57. For example, reports indicate that chromatin behaves as a liquid phase in vitro58 and at the sub-micrometer/sub-second scales in cells59, whereas it behaves as a solid phase at the micrometers/minutes scales and also in live cells59. Additionally, the formation of constitutive and facultative heterochromatin domains might be produced by the phase-separation of associated heterochromatin proteins60,61, exemplifying the relevance of condensation to genome function.
More broadly, given that the structure of the genome influences its related functions56, we can expect an analogous interplay between transcriptional condensates and genome organization; this means that the former might shape the latter and vice versa (Fig. 1). In this sense, chromatin could act as a scaffold recruiting multivalent transcription-related molecules and triggering the formation of transcriptional condensates (Fig. 1a). From a chemical point of view, chromatin provides an adequate surface to promote the formation of condensates even at lower concentrations of the engaging molecules than those expected for a standard phase-separation process62. Given that transcriptional condensates may involve the interactions between multiple, distant chromatin regions, the 3D genome architecture can also contribute to condensate formation providing an adequate topological framework. In this sense, it has been shown that CTCF-dependent chromatin loops recruit Pol II, BRD4, and Mediator molecules, driving transcriptional condensate formation63 (Fig. 1b). In addition, the chromatinic microenvironment of condensates seems to modulate their shape and size (Fig. 1c), as very dense environments could limit the growth and coalescence of synthetic condensates64 while low-density chromatin regions would favor their formation65. However, the functional consequences of this potential modulation remain elusive.
Illustrations of chromatin (pink) and condensates (green) schematically represent the ways they can influence each other. a Chromatin can act as a nucleation site for condensate formation. b Chromatin structure driven by CTCF-cohesin loops (cohesin complexes represented in blue) can act as a structural framework favoring condensate formation. c Chromatin microenvironment can exert mechanical forces on nuclear condensates. Local microenvironments would generate greater forces impairing condensate growth. d Condensate formation and growth can pull in two distant chromatin regions, driving chromatin looping. e Condensate growth can exert mechanical forces on surrounding chromatin, pushing it apart. f Condensate can bring multiple chromatin regions together, forming a multi-way chromatin hub.
Conversely, nuclear condensates themselves can reorganize the local chromatin architecture66. For example, it has been shown that TFs such as FOXA1 and SOX2 co-condense with DNA in vitro, exerting a tension force on the DNA strand which can bring together distal regions67,68. This force observed in vitro could contribute to chromatin looping associated with transcription regulation. Thus, chromatin-sticky condensates may bring together genomic loci through coalescence involving capillary forces65,69 (Fig. 1d), whereas, in other cases, the growth of a non-sticky condensate may exclude chromatin, pulling apart certain genomic loci65 (Fig. 1e).
Although further work needs to be done to demonstrate that many of these processes observed in vitro or with synthetic condensates in cells also occur in vivo, several studies suggest that transcriptional condensates could regulate gene expression by modulating genome structure. Specifically, the formation of phase-separated condensates of certain TFs has been linked to a 3D rearrangement of the genome and the consequent activation of target genes as observed during somatic cell reprogramming70, B-cell to macrophage trans-differentiation71 and leukemogenesis72. The use of synthetic TFs capable of phase-separate also illustrates how transcriptional condensates targeted to specific loci could increase chromatin accessibility, promote long-range contacts between genes and regulatory regions, and induce transcription of these target genes66.
Transcriptional condensates have also been postulated as responsible for the formation of multi-way hubs comprising many genes involved in similar biological functions (Fig. 1f). This is the case for housekeeping genes in mammalian cells73 wherein the contacts between their promoters are driven by the TF Ronin. Similarly, a report claims that HSF1 is responsible for the coalescence of long-range, distal genes responding to heat-shock stress in yeast74. These examples illustrate additional ways in which transcriptional condensates may contribute to chromatin architecture, thus modulating gene expression.
A functional link between transcriptional condensates and transcriptional activity: evidence and caveats
In the previous sections, we discussed some ideas on the nature of transcriptional condensates and reviewed evidence linking them with chromatin organization. The concepts presented above point to the inevitable relevant question of what the function of these condensates in transcriptional control might be. In this last section, we will present and discuss the evidence either in favor of or against the role of transcriptional condensates in transcriptional regulation. However, interpreting the complex and sometimes apparently contradictory spectrum of evidence requires first understanding the strengths and limitations of the different strategies usually employed to address this question. In Table 1, we provide a representative overview of the current literature describing the biological models, experimental approaches, and relevant results from key studies in the field.
One important experimental caveat in all these studies is how to selectively affect the ability of a transcriptional player to form condensates without altering other important activities. For example, IDRs of TFs and coactivators appear essential for condensate formation in many cases27, but they are also linked to activation functions25 and even to the DNA target searching process75,76. Therefore, it is extremely difficult to discriminate among the many functions IDRs can have on transcriptional regulation, including biomolecular condensation. This important limitation makes it relevant to understand the methodological aspects of the specific studies; Fig. 2 summarizes recent and commonly used experimental strategies designed to test whether transcriptional condensates are involved in transcriptional regulation.
a Correlative strategies generally involve using fluorescence microscopy to compare the spatial distributions of the condensates of a TF of interest (green) with those of other biomolecules with a known role in transcription, such as coactivators or Pol II (red). The merged images illustrate different degrees of colocalization (yellow): i) ‘total colocalization’, where both biomolecules colocalize in the same condensates, and ii) ‘partial colocalization’, where only a subset of condensates colocalize with the condensates formed by the other biomolecule. b Causal strategies involve performing perturbations of TF structure and analyzing their impact on certain properties in the cells. Top panels: TFs usually present a modular structure with a DNA-binding domain (DBD), and an IDR responsible for transactivation, target searching, and, often, condensate formation. TF structure can be experimentally altered by different approaches, including i) pharmacological targeting (oval), ii) deletions/truncations (scissors), and iii) point-mutations (cross). Analyzing the TF spatial distribution and the TF-driven transcriptional activity after these perturbations allows for exploring the contribution of the specific TF structural features to condensate formation and activity. Bottom panels: Synthetic TFs (green-purple) can be designed by fusing domains of the TF of interest to moieties with known properties. iv) Designed DNA binding: replacing the TF DBD with a well-studied, bacterial TF DBD, or RNA-guided dCas9 (not illustrated), and introducing a specific transcription reporter (cyan) driven by this DBD allow studying how the TF IDR contributes to transcriptional activity at these designed DNA targets. v) Designed nucleation: replacing the TF DBD with a responsive module capable of light- or chemical-controlled oligomerization helps to understand the contribution of the TF IDR to condensate formation. vi) Designed condensation: this strategy includes analyzing the effects of replacing the TF IDR with an exogenous IDR with enhanced condensation propensity, helping to understand the role of condensation on the TF activity at its natural DNA targets. These strategies can also be combined or modified to analyze more complex scenarios, according to the specific system.
Correlative studies (Fig. 2a) often include the use of different fluorescence microscopy techniques and labeling strategies to analyze the colocalization of nuclear condensates and key molecular players in transcriptional regulation, either in live or fixed cells. Noteworthy, fixation has been reported to produce artifacts and change the appearance of condensates77, therefore some colocalization experiments performed in fixed cells should be validated on live samples if possible. In the particular case of co-immunofluorescence studies, it is also imperative to perform additional experiments on live-cell systems and purified proteins to characterize a cellular object as a condensate.
Correlative reports have shown the colocalization between TFs condensates and other transcriptional players or epigenetic marks (Fig. 2a.i) such as Med1, BRD4, H3K27ac, and Pol II27,35,78,79. Even though, in many cases, the colocalization is partial (Fig. 2a.ii), at least a fraction of TF condensates appears to co-occupy, within the resolution limit of the techniques, sites of active transcription. Similarly, TF condensates were found to colocalize with target enhancers or target genes labeled by DNA-FISH26,27 and with fluorescently labeled reporter genes39,43,80,81,82,83,84. In addition, a recent study found that condensate proximity with a gene locus correlates with RNA burst enhancement85. Taken together, these results point to at least a population of transcriptional condensates being involved in transcription.
Causal studies (Fig. 2b), which means probing beyond correlation of a cause-effect link between transcriptional condensates and transcription, are intrinsically more difficult to achieve. There have been three main strategies used thus far: pharmacological/chemical treatments, deletions/truncations/mutations of key transcriptional players, and synthetic/chimera TFs coupled with reporter genes. Chemical manipulation (Fig. 2b.i) includes—but is not limited to—the use of aliphatic alcohols, which, in principle, perturb weak hydrophobic interactions, resulting in the dissolution of some liquid condensates86. Even though this behavior was traditionally considered the fingerprint of LLPS and this dissolution correlated with decreased transcriptional activity26,27,79,80,87, nowadays, this treatment is considered to have many confounding effects on cells beyond affecting condensates88,89,90. Nevertheless, it might still be useful to explore the chemical properties of the intermolecular interactions involved in condensates rather than to establish evidence of phase separation91.
While pharmacological studies are scarce, there is an interesting report wherein a compound specifically targets the androgen receptor’s IDR, disrupting both the receptor condensates and its transcriptional activity78. Targeting the IDR of the oncogenic driver TF MYC also affects its transcriptional activity92, but the effect on MYC condensates has not been evaluated. Beyond a potentially suitable tool to causally link condensates to transcriptional activity, pharmacological manipulation of condensates can potentially be used for clinical applications93. We expect that high-throughput screening strategies will amplify the pharmacological toolkit to target IDRs and transcriptional condensates in the near future.
Deletions, truncations (Fig. 2b.ii), and point-mutations (Fig. 2b.iii) in different transcriptional players are suitable strategies to test causality, with many examples shown in Table 1. However, it is worth noting that given the likely lack of true modularity in TF domains76, the complex intertwined connections between a measured phenotype and untested pleiotropic effects of mutations make the lines between correlation and causality a bit blurry. For example, and as we pointed out above, deleting an IDR of a TF might affect its mechanism of action at different levels, making it difficult to discriminate which one is responsible for the change in the phenotypic output.
Synthetic/chimeric TFs have provided the most promising experimental evidence thus far to test the hypothesis of a causal link between transcriptional condensates and transcriptional activity (Table 1). Synthetic TFs usually contain a DNA-targeting moiety (such as a catalytically dead Cas9, or the DNA-binding domain of a eukaryotic/prokaryotic TF) (Fig. 2b.iv), an activation domain of interest, and an ’optogenetic’ domain (e.g., CRY2), which allows control of oligomerization and condensate formation in a switch-on/off fashion using light39,43,66,80,81 (Fig. 2b.v). These constructs may sometimes include additional IDRs (e.g., FUS IDR) to enhance their condensation propensity (Fig. 2b.vi). Many of these approaches are frequently combined with the use of reporter genes with specific binding sequences for the DNA-binding domain of the synthetic TF and stem-loops within the reporter nascent RNA that bind to coating proteins fused to fluorescent tags, providing a spatially resolved, readable transcriptional output (Fig. 2b.iv). The intrinsic artificial nature of these constructions is a valid concern, and therefore, the extrapolation of the results obtained from synthetic TFs with reporter genes to the biology of natural TFs in a native context remains unclear.
Many studies using synthetic/chimera TF strategies pointed to a positive functional role of condensates in transcriptional activity39,66,80,81,83,87 (Table 1). Nevertheless, a more complex picture is portrayed in a study wherein DNA binding and oligomerization are independently controlled within a synthetic TF. Here, the authors analyze how artificial TF clustering correlates with the transcriptional output of a fluorescent reporter gene, reporting the occurrence of transcription with and without clustering. Interestingly, only 4% of TF clusters colocalize with nascent RNA signals, with a wide spectrum of effects ranging from inhibition to sustained activation depending on the artificial TF82. In fact, other groups have argued that transcriptional condensates are either neutral or even inhibitory for transcription23,43,84. In this sense, a clean experimental design in yeast demonstrated that the activation of a reporter gene did not depend on clustered TF recruitment84.
Both subtle differences and, in some cases, opposite conclusions among studies on the role of condensates in transcription could be due to a plethora of factors. From cell-type to gene-specific effects and all the way to protein-specific action can explain the observed differences. Moreover, as with most biological processes, transcriptional condensates may have an ideal composition range for optimal activity wherein any excess or deficiency might have inhibitory effects, as illustrated by others23,94,95, and this optimal concentration ratio could be system- and/or condition-specific.
Outlook/perspectives
Over the last few years, we witnessed striking adjustments in how we understand transcription. One of the most stunning changes is the multi-phase picture of the nucleus96, where some of the membrane-less subnuclear compartments concentrate transcriptional actors interacting through weak, multivalent bonds. Although substantial work has been done to elucidate whether these subnuclear structures play a role in the spatial and/or temporal control of transcription, we are still missing fundamental pieces to understand this complex transcriptional puzzle (Box 1).
One of the most relevant challenges to explain the relatively slow progress in this specific topic is the absence of adequate tools to study these nuclear compartments. Particularly, there is not a consensus in the scientific community on the evidence required to firmly assess LLPS involvement in the formation of a given type of condensates in cells (discussed in detail elsewhere44,97,98). Because of these technical difficulties, probing a functional relationship between biomolecular condensation and transcriptional regulation is still very challenging13. A major criticism in the field is that many observations performed in vitro or by the artificial manipulation of proteins to generate condensates in cellular systems cannot be extrapolated to native proteins in physiological conditions. Forcing local molecular concentrations higher than their physiological values may favor non-specific interactions between multivalent molecules and the formation of abnormal condensates99. Moreover, chemical cell fixation might promote the aberrant distribution of many nuclear biomolecules77,100,101, perturb chromatin organization101,102, and underestimate the transient nature of relevant interactions, overall preventing the straightforward interpretation of studies using fixed cells. In the same direction, it might be risky attempting to directly correlate live, single-cell imaging observations with those obtained from experiments based on the chromosome conformation capture procedure, a method based on crosslinking chromatin regions that provides information about the three-dimensional organization of the genome103.
Despite these limitations, growing evidence points toward a functional link between (some) transcriptional condensates and transcriptional regulation (Table 1). The state-of-the-art data suggests that this relationship is more complex than the initial view of transcriptional condensates concentrating transcription-activating molecules near gene promoters and thus promoting transcription. Indeed, the small sample of key reports compiled in this Mini Review attempts to be representative of the abundant literature in the field, ruling out a univocal relation between condensates and transcription regulation.
Our unresolved transcriptional puzzle suggests that transcriptional condensates with different compositions and functions may coexist in a single cell and be part of another temporal/spatial regulatory layer of transcription. Biomolecular condensates may be directly engaged in this regulation or could also contribute to shape chromatin structure at different scales, thus indirectly modulating gene expression.
References
Spitz, F. & Furlong, E. E. Transcription factors: from enhancer binding to developmental control. Nat. Rev. Genet. 13, 613–626 (2012).
Coulon, A., Chow, C. C., Singer, R. H. & Larson, D. R. Eukaryotic transcriptional dynamics: from single molecules to cell populations. Nat. Rev. Genet. 14, 572–584 (2013).
Mazzocca, M., Fillot, T., Loffreda, A., Gnani, D. & Mazza, D. The needle and the haystack: single molecule tracking to probe the transcription factor search in eukaryotes. Biochem. Soc. Trans. 0, 1–12 (2021).
Jana, T., Brodsky, S. & Barkai, N. Speed–specificity trade-offs in the transcription factors search for their genomic binding sites. Trends Genet. 37, 421–432 (2021).
Woringer, M. & Darzacq, X. Protein motion in the nucleus: from anomalous diffusion to weak interactions. Biochem. Soc. Trans. 46, 945–956 (2018).
Wagh, K., Stavreva, D. A., Upadhyaya, A. & Hager, G. L. Transcription factor dynamics: one molecule at a time. Annu. Rev. Cell Dev. Biol. 39, 277–305 (2023).
Perlmann, T., Eriksson, P. & Wrange, O. Quantitative analysis of the glucocorticoid receptor-DNA interaction at the mouse mammary tumor virus glucocorticoid response element. J. Biol. Chem. 265, 17222–17229 (1990).
Stavreva, D. A. et al. Transcriptional bursting and co-bursting regulation by steroid hormone release pattern and transcription factor mobility. Mol. Cell 75, 1161–1177.e11 (2019).
Rodriguez, J. & Larson, D. R. Transcription in living cells: molecular mechanisms of bursting. Annu. Rev. Biochem. 89, 1–24 (2020).
Tunnacliffe, E. & Chubb, J. R. What is a transcriptional burst? Trends Genet. 36, 288–297 (2020).
Botterbusch, S. & Baumgart, T. Interactions between phase-separated liquids and membrane surfaces. Appl. Sci. (Basel) 11, 1288 (2021).
Dolgin, E. The shape-shifting blobs that shook up cell biology. Nature 611, 24–27 (2022).
Wagh, K., Garcia, D. A. & Upadhyaya, A. Phase separation in transcription factor dynamics and chromatin organization. Curr. Opin. Struct. Biol. 71, 148–155 (2021).
Pederson, T. The nucleolus. Cold Spring Harb. Perspect. Biol. 3, a000638 (2011).
Belmont, A. S. Nuclear compartments: an incomplete primer to nuclear compartments, bodies, and genome organization relative to nuclear architecture. Cold Spring Harb. Perspect. Biol. 14, a041268 (2022).
Papantonis, A. & Cook, P. R. Transcription factories: genome organization and gene regulation. Chem. Rev. 113, 8683–8705 (2013).
Alberts, B. et al. Molecular biology of the cell. 4th edn, (Garland Science, 2002).
Iborra, F. J., Pombo, A., Jackson, D. A. & Cook, P. R. Active RNA polymerases are localized within discrete transcription “factories’ in human nuclei. J. cell Sci. 109, 1427–1436 (1996).
Cook, P. R. The organization of replication and transcription. Science 284, 1790–1795 (1999).
Eskiw, C. H., Rapp, A., Carter, D. R. & Cook, P. R. RNA polymerase II activity is located on the surface of protein-rich transcription factories. J. Cell Sci. 121, 1999–2007 (2008).
Pownall, M. E. et al. Chromatin expansion microscopy reveals nanoscale organization of transcription and chromatin. Science 381, 92–100 (2023).
Leidescher, S. et al. Spatial organization of transcribed eukaryotic genes. Nat. Cell Biol. 24, 327–339 (2022).
Chong, S. et al. Tuning levels of low-complexity domain interactions to modulate endogenous oncogenic transcription. Mol. Cell 82, 2084–2097.e2085 (2022).
Palacio, M. & Taatjes, D. J. Merging established mechanisms with new insights: condensates, hubs, and the regulation of RNA polymerase II transcription. J. Mol. Biol. 434, 167216–167216 (2022).
Ferrie, J. J., Karr, J. P., Tjian, R. & Darzacq, X. “Structure”-function relationships in eukaryotic transcription factors: the role of intrinsically disordered regions in gene regulation. Mol. Cell, 1–15. https://doi.org/10.1016/j.molcel.2022.09.021 (2022).
Sabari, B. R. et al. Coactivator condensation at super - enhancers links phase separation and gene control. Science 361, eaar3958 (2018).
Boija, A. et al. Transcription factors activate genes through the phase-separation capacity of their activation domains. Cell, 1–14 https://doi.org/10.1016/j.cell.2018.10.042 (2018).
Boehning, M. et al. RNA polymerase II clustering through carboxy-terminal domain phase separation. Nat. Struct Mol. Biol. 25 https://doi.org/10.1038/s41594-018-0112-y (2018).
Cho, W.-K. et al. Mediator and RNA polymerase II clusters associate in transcription-dependent condensates. Science 412–415, eaar4199 (2018).
Basu, S. et al. Unblending of transcriptional condensates in human repeat expansion disease. Cell 181, 1062–1079.e1030 (2020).
Cai, D. et al. Phase separation of YAP reorganizes genome topology for long-term YAP target gene expression. Nat. Cell Biol. 21, 1578–1589 (2019).
Shin, Y. et al. Spatiotemporal control of intracellular phase transitions using light-activated optoDroplets. Cell 168, 159–171.e114 (2017).
Chong, S. et al. Imaging dynamic and selective low-complexity domain interactions that control gene transcription. Science 2555, 1–16 (2018).
Shrinivas, K. et al. Enhancer features that drive formation of transcriptional condensates. Mol. Cell 75, 549–561.e547 (2019).
Stortz, M., Pecci, A., Presman, D. M. & Levi, V. Unraveling the molecular interactions involved in phase separation of glucocorticoid receptor. BMC Biol. 18, 59 (2020).
Rawat, P. et al. Stress-induced nuclear condensation of NELF drives transcriptional downregulation. Mol. Cell 81, 1013–1026.e1011 (2021).
Lyons, H. et al. Functional partitioning of transcriptional regulators by patterned charge blocks. Cell 186, 327–345.e328 (2023).
Mann, R. & Notani, D. Transcription factor condensates and signaling driven transcription. Nucleus 14, https://doi.org/10.1080/19491034.2023.2205758 (2023).
Wei, M. T. et al. Nucleated transcriptional condensates amplify gene expression. Nat. Cell Biol. https://doi.org/10.1038/s41556-020-00578-6 (2020).
Peng, L., Li, E. M. & Xu, L. Y. From start to end: phase separation and transcriptional regulation. Biochim. Biophys. Acta Gene Regu Mech. 1863, 194641–194641 (2020).
Guo, Y. E. et al. Pol II phosphorylation regulates a switch between transcriptional and splicing condensates. Nature. https://doi.org/10.1038/s41586-019-1464-0 (2019).
Henninger, J. E. et al. RNA-mediated feedback control of transcriptional condensates. Cell, 1–19. https://doi.org/10.1016/j.cell.2020.11.030 (2021).
Trojanowski, J. et al. Transcription activation is enhanced by multivalent interactions independent of phase separation. Mol. Cell 82, 1878–1893.e1810 (2022).
McSwiggen, D. T., Mir, M., Darzacq, X. & Tjian, R. Evaluating phase separation in live cells: diagnosis, caveats, and functional consequences. Genes Dev. 1–16. https://doi.org/10.1101/gad.331520.119 (2019).
Erdel, F. et al. Mouse heterochromatin adopts digital compaction states without showing hallmarks of hp1-driven liquid-liquid phase separation. Mol. Cell 1–14 https://doi.org/10.1016/j.molcel.2020.02.005 (2020).
Mittag, T. & Pappu, R. V. A conceptual framework for understanding phase separation and addressing open questions and challenges. Mol. Cell 82, 2201–2214 (2022).
Kent, S. et al. Phase-separated transcriptional condensates accelerate target-search process revealed by live-cell single-molecule imaging. Cell Rep. 33, 108248–108248 (2020).
Bo, S., Hubatsch, L., Bauermann, J., Weber, C. A. & Frank, J. Stochastic dynamics of single molecules across phase boundaries. Phys. Rev. Res. 3, 043150 (2021).
Finn, E. H. & Misteli, T. Molecular basis and biological function of variability in spatial genome organization. Science 365, eaaw9498 (2019).
Szabo, Q. et al. Regulation of single-cell genome organization into TADs and chromatin nanodomains. Nat. Genet. 52, 1151–1157 (2020).
Cremer, T. et al. The interchromatin compartment participates in the structural and functional organization of the cell nucleus. BioEssays 42, 1–18 (2020).
Conte, M. et al. Loop-extrusion and polymer phase-separation can co-exist at the single-molecule level to shape chromatin folding. Nat. Commun. 13, 4070 (2022).
Maeshima, K., Tamura, S., Hansen, J. C. & Itoh, Y. Fluid-like chromatin: toward understanding the real chromatin organization present in the cell. Curr. Opin. Cell Biol. 64, 77–89 (2020).
Bajpai, G., Amiad Pavlov, D., Lorber, D., Volk, T. & Safran, S. Mesoscale phase separation of chromatin in the nucleus. Elife 10 https://doi.org/10.7554/eLife.63976 (2021).
Mirny, L. A., Imakaev, M. & Abdennur, N. Two major mechanisms of chromosome organization. Curr. Opin. Cell Biol. 58, 142–152 (2019).
Misteli, T. The self-organizing genome: principles of genome architecture and function. Cell 183, 28–45 (2020).
Rippe, K. Liquid-liquid phase separation in chromatin. Cold Spring Harb. Perspect. Biol. 1–24. https://doi.org/10.1101/cshperspect.a040683 (2021).
Gibson, B. A. et al. Organization of chromatin by intrinsic and regulated phase separation. Cell 1–15. https://doi.org/10.1016/j.cell.2019.08.037 (2019).
Nozaki, T. et al. Condensed but liquid-like domain organization of active chromatin regions in living human cells. Sci. Adv. 9, eadf1488 (2023).
Strom, A. R. et al. Phase separation drives heterochromatin domain formation. Nature 547, 241–245 (2017).
Larson, A. G. et al. Liquid droplet formation by HP1α suggests a role for phase separation in heterochromatin. Nature 547, 236–240 (2017).
Pancholi, A. et al. RNA polymerase II clusters form in line with surface condensation on regulatory chromatin. Mol. Syst. Biol. 17, 1–26 (2021).
Lee, R. et al. CTCF-mediated chromatin looping provides a topological framework for the formation of phase-separated transcriptional condensates. Nucleic Acids Res. 1–20. https://doi.org/10.1093/nar/gkab1242 (2021).
Lee, D. S. W. et al. Size distributions of intracellular condensates reflect competition between coalescence and nucleation. Nat. Phys. https://doi.org/10.1038/s41567-022-01917-0 (2023).
Shin, Y. et al. Liquid nuclear condensates mechanically sense and restructure the genome. Cell 175, 1481–1491.e1413 (2018).
Kim, Y. J. et al. Light-activated macromolecular phase separation modulates transcription by reconfiguring chromatin interactions. Sci. Adv. 9, 1–18 (2023).
Quail, T. et al. Force generation by protein–DNA co-condensation. Nat. Phys. 17, 1007–1012 (2021).
Tuan N. et al. Chromatin sequesters pioneer transcription factor Sox2 from exerting force on DNA. bioRxiv, 13, 3988 (2022).
Strom, A. R. et al. Condensate-driven interfacial forces reposition DNA loci and measure chromatin viscoelasticity. bioRxiv https://www.biorxiv.org/content/10.1101/2023.02.27.530281v1 (2023).
Wang, J. et al. Phase separation of OCT4 controls TAD reorganization to promote cell fate transitions. Cell Stem Cell, 1–16 https://doi.org/10.1016/j.stem.2021.04.023 (2021).
Christou-Kent, M. et al. CEBPA phase separation links transcriptional activity and 3D chromatin hubs. Cell Rep. 42, 112897 (2023).
Oka, M. et al. Phase-separated nuclear bodies of nucleoporin fusions promote condensation of MLL1/CRM1 and rearrangement of 3D genome structure. Cell Rep. 42, 112884 (2023).
Dejosez, M. et al. Regulatory architecture of housekeeping genes is driven by promoter assemblies. Cell Rep. 42, 112505 (2023).
Chowdhary, S., Kainth, A. S., Paracha, S., Gross, D. S. & Pincus, D. Inducible transcriptional condensates drive 3D genome reorganization in the heat shock response. Mol. Cell 82, 4386–4399.e4387 (2022).
Brodsky, S. et al. Intrinsically disordered regions direct transcription factor in vivo binding specificity. Mol. Cell, 1–13. https://doi.org/10.1016/j.molcel.2020.05.032 (2020).
Staller, M. V. Transcription factors perform a 2-step search of the nucleus. Genetics 222, iyac111 (2022).
Shawn Irgen-Gioro, V. W. S. C. Fixation can change the appearance of phase separation in living cells. eLife, 11, e79903 (2022).
Xie, J. et al. Targeting androgen receptor phase separation to overcome antiandrogen resistance. Nat. Chem. Biol. https://doi.org/10.1038/s41589-022-01151-y (2022).
Zhang, H. et al. Reversible phase separation of HSF1 is required for an acute transcriptional response during heat shock. Nat. Cell Biol. 24 https://doi.org/10.1038/s41556-022-00846-7 (2022).
Ma, L. et al. Co-condensation between transcription factor and coactivator p300 modulates transcriptional bursting kinetics. Mol. Cell 81, 1682–1697.e1687 (2021).
Schneider, N. et al. Liquid-liquid phase separation of light-inducible transcription factors increases transcription activation in mammalian cells and mice. Sci. Adv. 7 https://doi.org/10.1126/sciadv.abd3568 (2021).
Wu, J. et al. Modulating gene regulation function by chemically controlled transcription factor clustering. Nat. Commun. 13, 2663 (2022).
Kawasaki, K. & Fukaya, T. Functional coordination between transcription factor clustering and gene activity. Mol. Cell 83, 1605–1622.e1609 (2023).
Meeussen, J. V. W. et al. Transcription factor clusters enable target search but do not contribute to target gene activation. Nucleic Acids Res. 51, 5449–5468 (2023).
Du, M. et al. Direct observation of a condensate effect on super-enhancer controlled gene bursting. Cell https://doi.org/10.1016/j.cell.2023.12.005 (2023).
Shi, M. et al. Quantifying the phase separation property of chromatin-associated proteins under physiological conditions using an anti-1,6-hexanediol index. Genome Biol. 22, 229 (2021).
Ahn, J. H. et al. Phase separation drives aberrant chromatin looping and cancer development. Nature. https://doi.org/10.1038/s41586-021-03662-5 (2021).
Itoh, Y. et al. 1,6-hexanediol rapidly immobilizes and condenses chromatin in living human cells. Life Sci. Alliance 4, e202001005 (2021).
Ulianov, S. V. et al. Suppression of liquid–liquid phase separation by 1,6-hexanediol partially compromises the 3D genome organization in living cells. Nucleic Acids Res. 49, 10524–10541 (2021).
Duster, R., Kaltheuner, I. H., Schmitz, M. & Geyer, M. 1,6-Hexanediol, commonly used to dissolve liquid-liquid phase separated condensates, directly impairs kinase and phosphatase activities. J. Biol. Chem. 296, 100260 (2021).
Muzzopappa, F. et al. Detecting and quantifying liquid-liquid phase separation in living cells by model-free calibrated half-bleaching. Nat. Commun. 13, 7787 (2022).
Boike, L. et al. Discovery of a functional covalent ligand targeting an intrinsically disordered cysteine within MYC. Cell Chem. Biol. 28, 4–13.e17 (2021).
Mitrea, D. M., Mittasch, M., Gomes, B. F., Klein, I. A. & Murcko, M. A. Modulating biomolecular condensates: a novel approach to drug discovery. Nat. Rev. Drug Discov. 21, 841–862 (2022).
Solvie, D. et al. MYC multimers shield stalled replication forks from RNA polymerase. Nature 612, 148–155 (2022).
Chen, L. et al. Hormone-induced enhancer assembly requires an optimal level of hormone receptor multivalent interactions. Mol. Cell 83, 3438-3456.e12 (2023).
Feric, M. & Misteli, T. Phase separation in genome organization across evolution. Trends Cell Biol. 31, 671–685 (2021).
Musacchio, A. On the role of phase separation in the biogenesis of membraneless compartments. EMBO J. 41, e109952 (2022).
Alberti, S., Gladfelter, A. & Mittag, T. Considerations and challenges in studying liquid-liquid phase separation and biomolecular condensates. Cell 176, 419–434 (2019).
Darzacq, X. & Tjian, R. Weak multivalent biomolecular interactions: a strength versus numbers tug of war with implications for phase partitioning. RNA 28, 48–51 (2022).
Schnell, U., Dijk, F., Sjollema, K. A. & Giepmans, B. N. G. Immunolabeling artifacts and the need for live-cell imaging. Nat. Methods 9, 152–158 (2012).
Teves, S. S. et al. A dynamic mode of mitotic bookmarking by transcription factors. eLife 5, 066464 (2016).
Hoffman, E. A., Frey, B. L., Smith, L. M. & Auble, D. T. Formaldehyde crosslinking: a tool for the study of chromatin complexes. J. Biol. Chem. 290, 26404–26411 (2015).
Gavrilov, A., Razin, S. V. & Cavalli, G. In vivo formaldehyde cross-linking: it is time for black box analysis. Brief. Funct. Genomics, 1–3. https://doi.org/10.1093/bfgp/elu037 (2014).
Zhang, Y. et al. Nuclear condensates of p300 formed though the structured catalytic core can act as a storage pool of p300 with reduced HAT activity. Nat. Commun. 12, 4618 (2021).
Song, L. et al. Hotspot mutations in the structured ENL YEATS domain link aberrant transcriptional condensates and cancer. Mol. Cell, 1–19. https://doi.org/10.1016/j.molcel.2022.09.034 (2022).
Mazzocca, M. et al. Chromatin organization drives the search mechanism of nuclear factors. Nat. Commun. 14, 6433 (2023).
Zhu, K. et al. An intrinsically disordered region controlling condensation of a circadian clock component and rhythmic transcription in the liver. Mol. Cell 83, 3457–3469.e3457 (2023).
Yoshida, S. R., Maity, B. K. & Chong, S. Visualizing protein localizations in fixed cells: caveats and the underlying mechanisms. J. Phys. Chemit. B https://doi.org/10.1021/acs.jpcb.3c01658 (2023).
Acknowledgements
This work was supported by grants from ANPCyT (PICT 2020-00818 and PICT-2018-1921 to V.L.; PICT 2019-0397 to D.M.P.). Universidad de Buenos Aires (UBACyT 20020190100101BA to V.L.) and CONICET (PIP 2022-11220210100212CO to V.L.). The funding agencies had no role in writing the manuscript. The authors would like to apologize to all researchers whose work has not been cited; this field is growing exponentially, and it would be impossible to reference all the related work.
Author information
Authors and Affiliations
Contributions
M.S., D.M.P., and V.L. wrote the manuscript with equal contributions.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Primary Handling Editors: Manuel Breuer and David Favero.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Stortz, M., Presman, D.M. & Levi, V. Transcriptional condensates: a blessing or a curse for gene regulation?. Commun Biol 7, 187 (2024). https://doi.org/10.1038/s42003-024-05892-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-024-05892-5
- Springer Nature Limited
This article is cited by
-
RNA glues it all
Nature Cell Biology (2024)
-
Chromatin structure and dynamics: one nucleosome at a time
Histochemistry and Cell Biology (2024)