Abstract
In host-symbiont systems, interspecific transmissions create opportunities for host switches, potentially leading to cophylogenetic incongruence. In contrast, conspecific transmissions often result in high host specificity and congruent cophylogenies. In most bird-feather mite systems, conspecific transmission is considered dominant, while interspecific transmission is supposedly rare. However, while mites typically maintain high host specificity, incongruent cophylogenies are common. To explain this conundrum, we quantify the magnitude of conspecific vs. interspecific transmission in the brood parasitic shiny cowbird (Molothrus bonariensis). M. bonariensis lacks parental care, allowing the assessment of the role of horizontal transmission alone in maintaining host specificity. We found that despite frequent interspecific interactions via foster parental care, mite species dispersing via conspecific horizontal contacts are three times more likely to colonize M. bonariensis than mites transmitted vertically via foster parents. The results highlight the previously underappreciated rate of transmission via horizontal contacts in maintaining host specificity on a microevolutionary scale. On a macroevolutionary scale, however, host switches were estimated to have occurred as frequently as codivergences. This suggests that macroevolutionary patterns resulting from rare events cannot be easily generalized from short-term evolutionary trends.
Similar content being viewed by others
Introduction
With the advance of cophylogenetic analytical and methodological frameworks, many host-symbiont systems have been assessed for the relative contribution of codiversification versus other types of events that shape coevolutionary histories of hosts and symbionts. The majority of studies have suggested that strict codiversification between hosts and symbionts (i.e., temporal and topological congruence of host and parasite phylogenetic branching pattern) is rare1,2,3,4,5,6,7,8,9,10,11,12,13,14, on average, being only 7% as common as other coevolutionary events15. Generally, cophylogenetic incongruence (i.e., the disagreement between host and symbiont phylogenetic branching patterns at the macroevolutionary scale) may be caused by several evolutionary events, such as duplication (speciation of a symbiont within a single-host species), sorting (extinction and missing the boat), failure of the symbiont to speciate, and host switching (or host shift)16. Among these events, host switching is typically the most frequent event1,4,9,16,17. At the microevolutionary scale, host switching is also a biologically intriguing event leading to the evolution of multihost symbionts, especially when it occurs between phylogenetically distant hosts1,18,19. Most host switches occur via interspecific horizontal transfers (Fig. 1: qhi), promoting incongruence in host and symbiont phylogenies20,21,22. In contrast, conspecific vertical transmission, i.e., from biological parents to offspring (Fig. 1: qvc), is expected to maintain single-host symbionts (i.e., high host specificity) and produce congruent host and symbiont phylogenies (strict codiversification)22,23. Yet, despite the perceived dominance of vertical transmission and low horizontal transmission rates24,25,26,27,28, some host-symbiont systems may simultaneously display both incongruent cophylogenetic patterns and high host specificity3,5,16,29,30. This conundrum challenges the role of vertical conspecific transmission in promoting codiversification and maintaining host specificity.
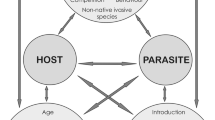
A Conspecific (vertical/horizontal) vs interspecific (vertical/horizontal) transmissions in different bird systems (brood parasites and non-brood parasites); B two hypotheses on how different symbiont transmission types may affect the mite host specificity and cophylogenetic congruence with their avian hosts. Our null hypothesis (H0) proposes that a lower conspecific transmission rate (in comparison to the interspecific rate) results in lower host specificity, lower number of codivergence events, and a higher number of host switches; whereas a higher rate of conspecific transmission (H1) is expected to result in higher host specificity, higher number of codivergence events, and lower number of host switches. Artwork by L.G.A.P.
A major and well-studied host-symbiont system is that of birds and feather mites (Acariformes: Analgoidea and Pterolichoidea), where mites have high levels of dependence and specificity with their avian hosts31,32,33,34. These symbiotic organisms spend their entire life cycle on their host (full-time, obligate symbionts). With a few exceptions (e.g., some skin mites), they do not have a specialized dispersal stage and seem to lack any other adaptations for long-range dispersal between hosts33,35,36. Therefore, the most important dispersal mode of feather mites across host individuals should be via parental care, i.e., vertical conspecific transmission from host parents to offspring (Fig. 1A: qvc)16,25,37,38,39. Consequently, the diversification of these symbionts is expected to be driven largely by host evolution. However, multiple cophylogenetic studies have shown that host switches are relatively common in feather mites3,5,6,32,40, suggesting that host switches are in fact one of the main drivers of feather mite diversification3. Thus, the current biological expectations are in conflict: one suggests that vertical transmission should be prevalent, leading to congruence between host and symbiont phylogenies, while observations show widespread phylogenetic incongruence among mites, despite their high host specificity. Therefore, it is crucial to understand the relative contribution of two types of conspecific transmission: vertical and horizontal (qvc and qhc), promoting cophylogenetic concordance and high host specificity vs interspecific transmission (qvi and qhi), that can generate cophylogenetic discordance and low host specificity (Fig. 1). In feather mites, vertical conspecific transmission (qvc) occurring from parents to chicks during the nesting period can be accurately measured25,39. However, quantifying conspecific horizontal transmission (qhc), which occurs in the form of social transmission (physical contact between hosts), is particularly difficult as it takes place outside of the nesting period. As a result, vertical transmission (qvc) has been overemphasized while conspecific horizontal transmission (qhc) has been largely overlooked in the literature. This has contributed to the uncertainty regarding the role of conspecific horizontal transmission in shaping both host specificity and cophylogenetic congruence.
The brood parasitic shiny cowbird, Molothrus bonariensis (Passeriformes: Icteridae), provides an excellent model to investigate the relationship between conspecific and interspecific transmission and host specificity in feather mites. Like all obligate brood parasites, this bird neither builds nests nor displays parental care, effectively preventing vertical conspecific transmission from biological parents to chicks (qvc = 0). This is a generalist bird parasitizing more than 90 different passerine species in 17 families in South America and beyond41,42,43. Given that M. bonariensis is a brood parasite, one would expect that this bird might have (i) specific, single-host mite species that co-diverged with their hosts over a long evolutionary time and (ii) foster parent mite species whose coevolutionary history has been mostly driven by host shifts rather than codivergence events. Molothrus-specific mites (i.e., single-host mites consistently found only on M. bonariensis) can be transferred only by horizontal contact between conspecific hosts, i.e., other M. bonariensis (Fig. 1A: qhc). In contrast, Molothrus-alien mites (i.e., mites found both on M. bonariensis and one or more species of its foster parents) would be transferred mostly via vertical interspecific care (Fig. 1A: qvi), and, potentially to a much lesser extent, by horizontal interspecific interaction (Fig. 1A: qhi; see justification in the section “Molothrus-specific mites have higher species richness …”). At the macroevolutionary timescale, the constant transmission of mites from foster parents likely has provided more opportunities for host switches than what would have been expected in non-brood parasite bird-mite systems. The M. bonariensis system, therefore, can be used to evaluate the magnitude of horizontal conspecific transmission (qhc) vs. interspecific transmissions (i.e., vertical qvi, and horizontal qhi) and their influence in long-term co-evolution in host-symbiont systems.
Here we accomplish the above goal by quantifying rates of different types of short-term transmission that reflect host specificity and comparing coevolutionary patterns among multiple mite lineages (e.g., Proctophyllodes, Amerodectes, Trouessartia) that offer independent biological replicates in the Molothrus system. Specifically, we evaluate the effective horizontal versus vertical mite transmission rates assuming that each mite species on each M. bonariensis specimen resulted from at least a single successful host switch event. The distribution of mites in each host specificity category, therefore, should reflect the relative conspecific and interspecific transmission rates. Then we used event-based cophylogenetic reconciliation analyses to estimate the number of four coevolutionary events (codivergence, duplication, host switch, extinction) that occurred in this system on the macroevolutionary scale. We discuss the implications of our results to explain the observed macroevolutionary and microevolutionary patterns in this system.
Our null hypothesis is that if horizontal conspecific transmission (qhc) is lower than interspecific transmission (rates qvi and qhi) in the Molothrus system, then both host specificity and cophylogenetic congruence should be low due to a high frequency of interspecific transmission from foster parents (qvi) and the absence of conspecific vertical transmission (qvc) (Fig. 1B: H0). Otherwise, if conspecific transmission is higher than interspecific (H1: qhc > qvi + qhi), host specificity should be high, while the level of cophylogenetic congruence cannot be predicted because many host switch opportunities may arise over a long evolutionary time. Hypothesis H1 (Fig. 1B: H1) favors a greater role of horizontal transmission in maintaining host specificity than it is assumed in the literature, e.g.,25,39. Furthermore, these two hypotheses (H1 and H0, respectively) can also answer the question of whether or not host-specific symbionts can persist via the horizontal transmission route alone.
Results
Molothrus-specific mites have higher species richness, prevalence, and abundance than Molothrus-alien mites
To identify Molothrus host-specific mites and mites primarily associated with Molothrus foster parents, we conducted an extensive mite survey focusing on M. bonariensis (144 specimens). To aid in identifying and classifying mites into the Molothrus-specific vs alien categories, we also sampled mites from its common foster parents (27 species, 69 specimens) (Supplementary Data 1 and 2) and considered the literature data. Out of the 144 M. bonariensis specimens inspected for feather mites, 5 individuals entirely lacked mites, and 139 had mites. Mites were identified and assigned to three categories, Molothrus-specific, Molothrus-alien, and quill-and-skin mites QSM; with the former two categories representing feather vane mites lacking a vector transmission, while the latter category representing rare ecological groupings, some of which can be transmitted by a vector (see Methods). Molothrus-specific mites (5 species) were exclusively and consistently found on 130 M. bonariensis specimens with moderate to high prevalence (13–60%, Table 1: Proctophyllodes molothrus, Amerodectes molothrus, Xolalgoides sp.1, Mesalgoides sp.1, and Trouessartia sp.6)44,45. Molothrus-alien mites (19 species, Table 1) were recorded on few host individuals (prevalence 0.69–7.5%). Twenty-six out of the 69 Molothrus-alien records could be attributed to known M. bonariensis foster parents based on 8 mite species identified with morphological and molecular evidence (Supplementary Note 5, Supplementary Data 1). Lastly, QSM mites (5 species, Table 1) were similar to the Molothrus-alien category as these mites were also present at low prevalence (1.4–5.6%) (Table 1). Cumulative mite abundance (the total number of all mite specimens in each category) was also higher in Molothrus-specific mites in comparison with the two other categories (Table 1).
The co-occurrence pattern of Molothrus-specific and Molothrus-alien or QSM mites was also evidenced by our quantitative data: (i) in 52.5% of cases, only Molothrus-specific mites were found on a particular host individual; while (ii) Molothrus-specific mites were more prevalent (34.5%) than either Molothrus-alien (16.4%) or QSM (6%) in co-occurring mite category records (Table 2); and (iii) only in 2.5% of cases, mites originating solely from the foster parents (qvi+qhi) were found (Table 2). These data provide evidence for the low rate of horizontal interspecific transmission (Fig. 1: qhi).
The rate of horizontal social symbiont transmission is high
Based on the prevalence and co-occurrence pattern of mites, we inferred that the effective conspecific transmission rate is around 3.9 times the interspecific transmission rate (74.8% vs. 18.9%, Table 2). Since Molothrus-specific mites can only disperse between M. bonariensis via horizontal conspecific transmission (qhc) and Molothrus-alien mites predominantly disperse via vertical interspecific transmission from foster parents (qvi), it can be inferred that cowbird-to-cowbird transmission is at least 3.9 times more frequent than foster parent transmission in this system. Because the QSM group can disperse either from foster parents to M. bonariensis or horizontally (by host social contacts or via phoresy), we cannot determine if they belong to conspecific or interspecific transmissions. Assuming that QSM mites disperse either only via (foster) parental care or only via host horizontal contacts, we estimate the overall ratio between Molothrus-to-Molothrus and foster parent transmission in the M. bonariensis system is within the interval of 3.0–4.3 (74.8/25.2–81.1/18.9) (Table 2), where the lower value (3.0) is a conservative estimate of horizontal conspecific vs. vertical interspecific transmission rate. Thus one can conclude that Molothrus-to-Molothrus transmission (qhc) is at least three times greater than foster parent transmission (qvi) and horizontal interspecific transmission (qhi), i.e., qhc > (qvi + qhi) = 3:1.
Molothrus-associated mites are sister taxa to mites associated with other Molothrus species or foster parents phylogenetically
In order to understand macroevolutionary patterns between mites and Molothrus, we obtained molecular data from 118 specimens of field-collected mites associated with Molothrus and/or putative foster parents (29 bird species from 21 genera and 10 families, Genbank accessions CO1: MW814590–MW814707 and HSP70: MW829221–MW829276; see Table 1, Supplementary Data 2, Supplementary Note 3). We reconstructed mite phylogenies using 6 loci by supplementing our dataset with 153 mite sequences and 127 hosts from previous publications5,6 (Fig. 2). The sampling of mite phylogeny encompasses ca. 48.6% of the total named mite species for the families Proctophyllodidae and Trouessartiidae (n = 253), and ca. 18.8% of the total host species (n = 425) (considering New World Passeriformes records only and unique mite species haplotypes)46. Out of the 29 M. bonariensis-associated mite species, we provided phylogenetic information for two Molothrus-specific and one Molothrus-alien species (highlighted by orange in Fig. 2) and identified their sister taxa. They belong to three families/subfamilies of mites—Pterodectinae, Proctophyllodinae, and Trouessartiidae, and therefore represent three independent host-symbiont coevolutionary histories (Fig. 2).
Maximum Likelihood phylogeny of feather mites based on six genes (5 nuclear, 1 mitochondrial). Nodal support higher than 75% (estimated by ultrafast bootstrap with 1000 replicates) is shown by thicker lines. The major mite lineages, Trouessartiidae, Proctophyllodinae and Pterodectinae, are highlighted. Mites from cowbirds (Molothrus bonariensis and M. ater) and their foster parent birds: finches (Sicalis flaveola and S. luteola) and other Icteridae are highlighted. Portions of this phylogeny, exemplifying important host shifts to Molothrus are given for each mite lineage on insets (A–D); these cases are further considered in detail in Fig. 3. Artwork by L.G.A.P.
The sister taxa of two of the three mite species, Amerodectes molothrus and Proctophyllodes molothrus, are mites from M. ater (M. bonariensis’ sister species) from Mexico (A, B in Figs. 2 and 3). A species delimitation analysis based on genetic distances within and between the sister pairs, Proctophyllodes sp. ex. M. ater - Proctophyllodes molothrus (CO1 K2P = 5.5%, intraspecific distance for P. molothrus 0.2–0.3%) and Amerodectes tretiakae - Amerodectes molothrus (CO1 K2P = 9.5%, intraspecific distance for A. molothrus = 0.5%) (Fig. 3A, B), unambiguously placed these four genetically distinct OTUs as separate species (Assemble Species by Automatic Partitioning (ASAP) algorithm score = 5.50). The third species, Trouessartia sicaliae, shows similarity to mites from putative foster parents, Sicalis flaveola and S. luteola (Thraupidae) (CO1 K2P = 3.6–6.5%) (Fig. 3C). Another M. ater-associated mite species, Proctophyllodes egglestoni, displays a similar pattern in that it is closely related to mites from Agelaius phoeniceus (CO1 K2P = 0.2%), a common foster parent of M. ater43 (Fig. 3D).
A–D Mite molecular phylogenetic trees, cophylogenetic reconciliation analysis, and schematic explanations of cophylogenetic scenarios (for complete phylogenies, see Fig. 2; and Supplementary Figs. 1–5). Nodal support is given for each branch (ultrafast bootstrap, 1000 replicates). Molecular CO1 K2P genetic distances are given as percentages. See the Result section for a detailed explanation of cophylogenetic scenarios. Artwork by L.G.A.P.
Host switches and codivergences occur at nearly the same frequencies
For the three mite lineages, A, B+C, and D (Fig. 3), we performed separate cophylogenetic analyses using dated host and parasite phylogenies (Fig. 3, Supplementary Note 4, Supplementary Figs. 1–5) to identify either ongoing (with gene flow) or historical host switching (no gene flow). For all these lineages, a significant phylogenetic congruence was detected, p < 0.01 (see Methods, Supplementary Note 4, Supplementary Figs. 1–5). Our time estimates provide a temporal context for mite-bird colonization events (Fig. 3) and suggest that host switches were temporally possible with respect to host diversification. For the three Molothrus-associated mite lineages, we recovered 2 codivergences and 3 host switch events. Two codivergence events represent speciation that occurred within the Molothrus genus, between Amerodectes molothrus—A. tretiakae (Fig. 3A) and Proctophyllodes molothrus—P. sp. (Fig. 3B). However, host switches occurred at their ancestral nodes: in Amerodectes, there was a host switch from ancestral Molothrus to ancestral Sicalis (Fig. 3A); in Proctophyllodes, there was 1 host switch from an unknown host to ancestral Molothrus (Fig. 3B); and in Trouessartia, T. sicaliae switched its host from Sicalis flaveola to M. bonariensis (Fig. 3D). Proctophyllodes egglestoni, on the other hand, has experienced ongoing transmission from foster parents to M. ater, possibly through both vertical and horizontal routes as some foster parent species may also form mixed flocks with M. ater47 (Fig. 3C). Below we present a detailed description of topology- and time-informed coevolutionary scenarios in the cowbird system (Fig. 3A–D).
Amerodectes molothrus complex
A putative host switch onto ancestral Molothrus, followed by host switch of Amerodectes mites from ancestral Molothrus to their foster parents (ancestral Sicalis) (Fig. 3A: τ2), which resulted in their codivergence on Molothrus and possibly on Sicalis (Fig. 3A: τ2–τ0). This scenario is evidenced in all our cophylogenetic reconciliations and by the fact that mites from Sicalis and Molothrus are monophyletic (Fig. 2A). Our host-based calibration time estimates for this host switch make it temporally possible (7.9 Mya), contrarily to our fossil mite calibration (15.5 Mya), given that it occurred before the origin of both Molothrus and Sicalis (Supplementary Figs. 1 and 2). Our divergence time estimates of mites from Molothrus ater and M. bonariensis (4.4/3.7 Mya, host/fossil calibration) predate the divergence time estimates of their hosts (i.e., ~1 Mya), but are close to the extreme value (3.8 Mya) from the literature48,49,50. In contrast, Amerodectes mites associated with Sicalis flaveola/luteola originated after their hosts split into species (0.6/1.2 Mya vs 4.8–5.6 Mya) (Supplementary Fig. 1, 2). Taken together, our cophylogenetic evidence and time estimates suggest that in the Amerodectes molothrus complex, a host switch occurred from the ancestral Molothrus to ancestral Sicalis.
Proctophyllodes molothrus complex
A putative host switch onto ancestral Molothrus, followed by codivergence on Molothrus (Fig. 3B: τ2–τ0). Because this group is sister to the mite lineage having hosts from many families (Cardinalidae, Icteriidae, Passerellidae, and Icteridae) (Fig. 2B), its ancestral host cannot be identified with certainty. Our time estimates for this host switch (1.9–5.0/3.7–9.6 Mya host/fossil calibration, respectively) are plausible, given that they are completely (former) or partially (latter) within the timing of origin of Molothrus.
Proctophyllodes egglestoni
Ongoing mite transmission from foster parents and possibly by social contact (Fig. 3C: τ0). Mites from Molothrus ater clustered with those of the red-winged blackbird, Agelaius phoeniceus, and their CO1 genetic distances were nearly identical (0.2%) (Figs. 2C and 3C). The high sequence identity is indicative of an ongoing mite transmission from foster parent to Molothrus and/or via social contacts in mixed flocks. This is a multihost mite, mostly associated with icterids47.
Trouessartia sicaliae complex
Likely host switch to M. bonariensis from foster parents followed by on-host divergence (Fig. 3D: τ1–τ0). One DNA sequence of Trouessartia from M. bonariensis was recovered as nested within the Trouessartia sicaliae morphospecies associated with thraupids Sicalis spp. (Figs. 2D and 3D), which are known foster parents of M. bonariensis. However, the Trouessartia lineage from M. bonariensis had substantial CO1 genetic distance (3.6%), indicating that this Sicalis to Molothrus switch occurred sometime in the past, or is a result of a transfer from an unsampled host. Ongoing gene flow via conspecific social horizontal transmission occurring at low rate cannot be excluded.
Discussion
To investigate the magnitude of host-specific symbiont transmission via horizontal conspecific contact (qhc) vs interspecific transmissions (qvi and qhi), we studied a generalist brood parasitic passerine, the shiny cowbird M. bonariensis, and its obligatory symbiotic feather mites. Despite the lack of parental care in the host and the absence of regular parent-to-offspring mite vertical transmission (qvc = 0), five mite species (Molothrus-specific) were able to colonize new host generations horizontally via host social conspecific contact (qhc). In contrast, mites originating from the foster parents (Molothrus-alien) and/or dispersing through other modes (QSM) had a higher species richness (24 species, 11 genera), despite their overall low prevalence. This is an expected outcome because M. bonariensis has a broad range of foster parent hosts (n = 97)43, which contributes to the great diversity of mites in the Molothrus-alien category.
Based on the known biology of M. bonariensis, social transmission of Molothrus-specific mites is only possible via horizontal conspecific contact between bird individuals (Fig. 1: qhc), which can occur when their fledglings leave their foster parent’s nest for the formation of foraging and roosting flocks, during courtship, copulation, and other conspecific interactions51,52,53,54,55,56. Molothrus-specific mites do not occur on any other bird species as shown by our extensive survey and previous research44,45,57. Thus, there is no evidence that these mites are transmitted via foster parental care or that M. bonariensis foster parents can serve as vectors of Molothrus-specific mites. Furthermore, horizontal interspecific transmission (qhi) should be minimal in all cases as alien-only mites were found in only 2.5% of M. bonariensis individuals vs specific-only in 40.3% (Table 2); and even this small figure (2.5%) can be fully or partially attributed to foster parent transmission. Our estimation qhc > (qvi+qhi) = 3:1 (see above) supports our alternative hypothesis (Fig. 1: H1), suggesting that conspecific horizontal transmission of host-specific mites (qhc) is higher than vertical interspecific transmission and horizontal interspecific transmission (qvi+qhi and qvi > >qhi). It is also possible that mites transmitted from foster parents are replaced by Molothrus-specific mites through competitive exclusion. Unfortunately, here we cannot provide direct evidence for this hypothesis because samples of immature bird stages were not available for study. Under an alternative scenario, when horizontal interspecific transmission of Molothrus-alien mites is equal to or greater than horizontal conspecific transmission of Molothrus-specific mites (i.e., qhi ≥ qhc), hosts harboring alien-only mites would be expected to be found at the same or greater frequency as hosts harboring only Molothrus-specific mites. However, this is not the case in this system.
In the cowbird system, all Molothrus-specific mites were transmitted horizontally via conspecific contacts, which likely occurred gradually with host age as the birds experienced an increasing rate of social contact. This has been shown in feather mites associated with the Australian bushturkeys (Galliformes: Megapodiidae), a host that has minimal parent-to-offspring contact (eggs are buried, young birds are fully fledged on hatching)58. In M. bonariensis, horizontal conspecific transmission has maintained host specificity of its mite symbionts and also the continuity of the mites’ generations, resulting in 90.3% prevalence of Molothrus-specific mites among hosts carrying any mites. On a macroevolutionary scale, the coevolutionary patterns of Molothrus-associated mites, however, exhibit a range of possible events, from strict codivergence to incomplete and complete host switches, providing snapshots of how a low rate of interspecific transmission on a macroevolutionary timescale could result in potential host switch events. Our coevolutionary analyses revealed a pronounced pattern of host switching and codiversification. As an example, the two most common and abundant mite species, Amerodectes molothrus and Proctophyllodes molothrus, were involved in historical host switches (Fig. 3A, B). Subsequently, these two newly established mite lineages have become genetically isolated and specific to their new hosts. For multihost mites, failure to speciate with ongoing gene flow might be more common. For example, an exceptional multihost generalist mite, Proctophyllodes egglestoni, can probably be transferred from M. ater to its foster parents (Agelaius phoeniceus) and back through aggressive (e.g., nest defense) and gregarious (e.g., mixed-flocks) behaviors59. These ongoing transmissions are supported by the shallow genetic distances between the mites from the two hosts, COX1 K2P = 0.2% (Fig. 3D). However, mites using this strategy were not found in the Molothrus bonariensis system, thus allowing us to accurately quantify the rate of foster parent transmission in M. bonariensis.
Previous studies have suggested the presence of horizontal conspecific and vertical interspecific transmission in cowbirds and other brood parasitic birds by the presence of host-specific or parent-specific symbionts, albeit with no quantitative data or phylogenetic analyses57,60,61,62,63. Furthermore, even in non-brood parasitic systems, horizontal conspecific transmission has also been observed as the main transmission route of symbionts, such as feather mites from red-billed choughs64, barn swallows65, and in feather lice from bee-eaters64,65,66. Thus, our data on the high rate of social (horizontal) transmission in the Molothrus system potentially represent a more general pattern in feather mites that needs to be further investigated in other bird systems as well.
In host-symbiont systems, the general expectation is that parent-offspring vertical transmission results in both cophylogenetic congruence and high host specificity7,14,16,23,67. However, in the feather mite-bird symbiotic system, low cophylogenetic congruence and relatively high host specificity have been simultaneously detected in the same systems3,5,6,32. If we assume that most feather mites disperse vertically25,39,68, only the high host specificity can be explained; the low cophylogenetic congruence cannot be explained in terms of a larger contribution of predominant vertical transmission. Our work may reconcile these opposing observations by (i) estimating relative rates of ongoing horizontal and foster parent vertical transmission (microevolutionary scale) and (ii) quantifying codivergence vs non-codivergence coevolutionary events (macroevolutionary scale). The high host specificity observed in the Molothrus-specific mites can be therefore explained solely by the high rates of horizontal conspecific transmission, despite the lack of vertical conspecific transmission. On the macroevolutionary scale, although brood parasitic birds have qhc larger than qvi, the frequency of host switches cannot be extrapolated from these rates directly (Fig. 1B: H1). Here we observe an equal number of codivergence vs host-switch events. We therefore suggest that on a microevolutionary scale, both vertical (qvc, qvi) and horizontal transmissions (qhc, qhi) can affect host specificity of feather mites, while these transmissions may not substantially influence their coevolutionary scenarios on the macroevolutionary scale. For example, the probability of a successful host switch, an event affecting host-symbiont coevolution, may depend on multiple factors other than transmission rates, such as the competitive abilities of a particular symbiont species, common resources shared by unrelated hosts, and niche availability16,69,70.
In summary, our work highlights that symbiont horizontal transmission via conspecific social contacts is an important dispersal mode of Molothrus-specific mites onto new host individuals in the Molothrus-feather mite system. This horizontal transmission alone (without parent-offspring vertical transmission) can maintain highly abundant, dominant, and host-specific species of obligate mite symbionts on their hosts. Here, we identified five independently evolved lineages of mites specific to M. bonariensis that colonize new generations of hosts exclusively through horizontal route of transmission. These symbionts persist on M. bonariensis at high prevalence and abundance despite the constant influx of new diverse mite colonists transmitted vertically from over 90 species of foster parents. On average, mite species dispersing via conspecific horizontal contacts are three times more likely to colonize M. bonariensis than mites transmitted vertically via foster parents. Our data therefore provide evidence challenging the traditional view of the importance of vertical transmission as the main force generating host specificity on a microevolutionary scale23. On a macroevolutionary scale, we show that horizontal transmission maintained these Molothrus-specific mites on their hosts over a long evolutionary time, at least 1.38 Mya since the split of M. bonariensis and M. ater. There were both codivergence and host switch events, occurring nearly at the same frequencies. This suggests that macroevolutionary patterns, which are based on rare coevolutionary events, cannot be easily generalized from short-term evolutionary trends, such as transmission mode and rates.
Methods
Taxon sampling
We sampled 144 M. bonariensis specimens for feather mites in Brazil, 22 captured in the wild (by plucking infested feathers), 10 roadkill donated bodies (by washing), and 112 museum dry skins (by feather-ruffling) (see Supplementary Notes 1 and 2 for sampling details, and Supplementary Data 1 for shiny cowbird mite list). We have complied with all relevant ethical regulations for animal use (permits MMA 57944-3, CEUA 12/2017). For museum samples, to exclude potential cross-contamination, confidence scores were applied as detailed in Supplementary Note 2. This dataset was used to estimate the mite transmission rates. To identify foster parent mite species and classify mites into Molothrus-specific and Molothrus-alien categories, we also sampled mites from (i) putative Molothrus bonariensis foster parent species:43 68 specimens (27 species, 19 genera, 10 families) captured with mist nets from different regions in Brazil, and (ii) 1 specimen of Molothrus ater, which is sister to M. bonariensis (see supplementary file P21.MCC.tre in ref. 71), from Mexico (Supplementary Data 2: column “Data description”). At least one representative of each mite morphospecies of the two common subfamilies Proctophyllodinae, and Pterodectinae, and the family Trouessartiidae, was chosen per bird species and per region for DNA sequencing (Supplementary Data 2). A total of 118 mite specimens were sequenced for two genes (174 new sequences); in addition, 153 previously deposited GenBank sequences of 153 mite species from 127 bird species (90 genera, 36 families) were included in the analyses as well (Supplementary Data 2).
Definition of host specificity categories
Using morphological data (from museum skins, dead birds, and field-captured mites, 365 mite records: 29 mite morphospecies, 12 genera, 8 families from 144 M. bonariensis; Table 1; Supplementary Data 1; Supplementary Note 1 and 2), we identified mite species and assigned them in three categories (Molothrus-specific, Molothrus-alien and Quill-and-skin QSM mites) based on the following criteria: exclusivity of an association with M. bonariensis (i.e., a single-host mite species occurring only on M. bonariensis)72, phylogenetic relationships and genetic distances as compared to mites from foster parents or one of its sister brood parasitic species, Molothrus ater71, and transmission types:
-
(i)
Molothrus-specific—mite species found exclusively on Molothrus bonariensis; in molecular trees they should be distantly related to mites known from foster parent passerines and be closely related to mites from one of its sister host species, Molothrus ater.
-
(ii)
Molothrus-alien (foster parent mites)—found principally on M. bonariensis foster parent birds and also they can be found on M. bonariensis (likely as a result of transmission from foster parents); in phylogenetic trees, they should be closely related (i.e., have zero length or shallow branches) to mites specific to a particular foster parent passerine species.
-
(iii)
Quill-and-skin mites (QSM)—inhabit the host skin (e.g., Microlichus, Metamicrolichus, Strelkoviacarus) or live inside feather quills (Dermoglyphus). These mites are grouped together because of their restricted contact transmission in comparison to that of plumage feather mites (groups i and ii). For example, when two bird individuals come into contact, the likelihood of sharing skin or quill mites is likely to be much lower than that of sharing plumage mites. Furthermore, skin mites are naturally rare and have low prevalence and abundance73 in comparison with plumage mites74. Some skin-inhabiting mites also have the ability to be horizontally transferred across interspecific and intraspecific hosts via phoresy on louse flies (Diptera: Hippoboscidae)36,75,76. As a result, in terms of their transmission patterns, these mites cannot be directly compared to plumage-inhabiting feather mites (groups i and ii).
DNA Amplification and Sequencing
We sequenced the mitochondrial cytochrome c oxidase subunit I (CO1) gene (1026 nt), a marker that is useful to identify species and phylogenies below the genus level6,21,77. To reconstruct deeper mite divergences we considered 5 candidate loci (EF1-α, SRP54, HSP70, 28 S, and 18 S) used in a recent phylogenetic study5, and selected the nuclear heat shock protein cognate 5 Hsc70-5 (HSP70) gene (1674 nt) based on its highest Internode Certainty index (IC) value in RaxML 8.2.1078,79, and amplified this gene for all mites having a unique CO1 haplotype.
We added our sequence data to a large dataset (144 terminals and six genes: 18 S, 28 S, EF1-α, SRP54, HSP70, and CO1; 8546 bp total) generated previously;5 9 additional proctophyllodid terminals (CO1-only)6,80 were also included (Supplementary Data 2). Gabucinia (Pterolichoidea: Gabuciniidae) was used as a distant outgroup5,6. Our final aligned matrix had 271 terminals, of which 118 were newly sequenced (Supplementary Data 2: GenBank accession numbers, CO1: MW814590–MW814707; HSP70: MW829221–MW829276). See Supplementary Note 3 for details on DNA isolation, amplification, and sequencing.
Phylogenetic inference
We inferred a Maximum Likelihood (ML) phylogeny in IQ-Tree 1.6.1081 under a codon model for the protein-coding genes. Branch support values were estimated by Ultrafast bootstrap with 1000 replicates for the consensus tree82 (Fig. 2). The best model for each gene partition was estimated in IQ-Tree prior to analyses using the ModelFinder algorithm and corrected Akaike Information Criterion (AICc):83 18 S + 28 S (GTR + F + I + G4), EF1-α (KOSI07 + F3X4 + G4), SRP54 (MGK + F1X4 + G4), HSP70 (MG + F3X4 + G4) and CO1 (MG + F3X4 + G4). All these analyses were run using a single command: iqtree -s data.phy -st CODON5 -alrt 1000 -bb 1000 -nt 7 -p model. The consensus tree was visualized and edited in FigTree 1.4.484. The tree was rooted using the outgroup genus Gabucinia (Pterolichoidea) after the analysis.
Species delimitation
For molecular species delimitation, we used the Assemble Species by Automatic Partitioning (ASAP) algorithm85 and the CO1 locus;77,86 see87 for limitations of CO1-only species delimitations. We also evaluate whether putative species form monophyletic lineages by inferring CO1 phylogenies in RaxML 8.2.10 using the GTR + G + I model and 100 bootstrap replicates79. We calculated K2P distances in the R package ‘ape’ v.5.388.
Divergence time estimation
We performed a Bayesian divergence time estimation in BEAST v2.6.189 using a Relaxed Clock Log Normal and the Birth and Death prior model, expecting multiple extinctions in feather mites5. Partition schemes and substitution models were found in PartitionFinder v 2.1.1 (GTR + I + G for all partitions, except for the CO1 position 3: GTR + G). For additional details, see Supplementary Note 4. We compared our divergence time estimates with the following known time divergence estimates for the host birds: Molothrus (originated 7.4, diversified 4.3 Mya); Molothrus ater/bonariensis split 1.0-2.2 Mya; Sicalis (originated 9.1 Mya, diversified 7.2 Mya), Sicalis flaveola/luteola split 4.8-5.6 Mya71. Other published estimates of the Molothrus ater/bonariensis split range from 0.8–1.2 to 2.8-3.8 Mya48,49,50,90.
Cophylogenetic analysis
To quantify the number of different coevolutionary events in the Molothrus system, we performed separate parsimony-based reconciliation cophylogenetic analyses in eMPRess91 for each phylogenetically independent mite lineage (Fig. 2): Pterodectinae (Amerodectes A), Proctophyllodinae (Proctophyllodes B + C), and Trouessartiidae (Trouessartia D). Default event cost values were used as the software automatically selects the average best fitting reconciliations. Symbiont trees were inferred in IQ-Tree 1.6.10 as above (see the ‘Phylogenetic Inference’ section). Bird phylogenies were obtained from BirdTree71. TreeAnnotator v2.6.1 was used to summarize the 1000 host trees into a maximum credibility tree with node heights calculated as median heights. We found phylogenetic information for all bird species except Polioptila dumicola. A uniform OTU naming scheme was used for both molecular and morphological species. Ongoing gene flow was considered to be present between two mite populations from different hosts if their CO1 K2P distances were equal or lower than 5%77,87.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
DNA sequences were deposited into GenBank (accession MW814590–MW814707 and MW829221–MW829276). Supplementary Information provide the mite data and host information (Supplementary Data 1); GenBank accession numbers, host and mite collection (Supplementary Data 2); mite collection data and likely foster parents for the two host specificity categories, Molothrus-alien and QSM (Supplementary Data 3); Divergence time estimates using well-known host codivergence events (Supplementary Fig. 1) or mite outgroup fossil information (Supplementary Fig. 2); Maximum parsimony cophylogenetic reconciliations for the mite (sub)families: Trouessartiidae, Proctophyllodinae, Pterodectinae (Supplementary Figs. 3–5).
References
Araujo, S. B. L. et al. Undestanding host-switching by ecological fitting. PLoS One 10, 1–17 (2015).
Clayton, D. H., Bush, S. E. & Johnson, K. P. Ecology of congruence: past meets present. Syst. Biol. 53, 165–173 (2004).
Doña, J. et al. Cophylogenetic analyses reveal extensive host-shift speciation in a highly specialized and host-specific symbiont system. Mol. Phylogenet. Evol. 115, 190–196 (2017).
Fecchio, A. et al. Diversification by host switching and dispersal shaped the diversity and distribution of avian malaria parasites in Amazonia. Oikos (2018) https://doi.org/10.1111/oik.05115.
Klimov, P. B., Mironov, S. V. & OConnor, B. M. Detecting ancient codispersals and host shifts by double dating of host and parasite phylogenies: Application in proctophyllodid feather mites associated with passerine birds. Evolution 71, 2381–2397 (2017).
Matthews, A. E. et al. Cophylogenetic assessment of New World warblers (Parulidae) and their symbiotic feather mites (Proctophyllodidae). J. Avian Biol. 49, 1–17 (2018).
Page, R. D. M. Introduction. In: Tangled Trees: phylogeny, cospeciation and coevolution. (Chicago University, 2003).
Weckstein, J. D. Biogeography explains cophylogenetic patterns in toucan chewing lice. Syst. Biol. 53, 154–164 (2004).
Trivelloni, V. & Panassiti, B. A field synopsis, systematic review, and meta-analyses of cophylogenetic studies: what is affecting congruence between phylogenies? MANTER J. Parasite Biodivers. (2022) https://doi.org/10.32873/unl.dc.manter24.
Hoberg, E. P. & Brooks, D. R. A macroevolutionary mosaic: episodic host-switching, geographical colonization and diversification in complex host-parasite systems. J. Biogeogr. 35, 1533–1550 (2008).
Brooks, D. R., Hoberg, E. P. & Boeger, W. A. In the eye of the cyclops: the classic case of cospeciation and why paradigms are important. Comp. Parasitol. 82, 1–8 (2015).
Da Graça, R. J. et al. Topological congruence between phylogenies of Anacanthorus spp. (Monogenea: Dactylogyridae) and their Characiformes (Actinopterygii) hosts: a case of host-parasite cospeciation. PLoS One 13, 1–14 (2018).
Míguez-Lozano, R., Rodríguez-González, A. & Balbuena, J. A. A quantitative evaluation of host-parasite coevolutionary events in three genera of monopisthocotylean monogeneans. Vie Milieu 67, 103–119 (2017).
Mironov, S. V., Klimov, P. B., Block, N. L. & OConnor, B. M. Feather mites of the new genus Bernierinyssus gen. n. (Acariformes: Pteronyssidae) from endemic Malagasy warblers (Passeriformes: Bernieridae)—a lineage showing symbiotic cospeciation with their avian hosts. Syst. Appl. Acarol. 25, 1765–1802 (2020).
De Vienne, D. M. et al. Cospeciation vs host-shift speciation: Methods for testing, evidence from natural associations and relation to coevolution. N. Phytol. 198, 347–385 (2013).
Clayton, D. H., Bush, S. E. & Johnson, K. P. Coevolution of Life on Hosts. (University of Chicago Press, 2015). https://doi.org/10.7208/chicago/9780226302300.001.0001.
Doña, J. & Johnson, K. P. Assessing symbiont extinction risk using cophylogenetic data. Biol. Conserv. 250, 108705 (2020) https://doi.org/10.1016/j.biocon.2020.108705.
Agosta, S. J., Janz, N. & Brooks, D. R. How specialists can be generalists: resolving the and ‘parasite paradox’ and implications for emerging infectious disease. Zoologia 27, 151–162 (2010).
Agosta, S. J. & Klemens, J. A. Ecological fitting by phenotypically flexible genotypes: Implications for species associations, community assembly and evolution. Ecol. Lett. 11, 1123–1134 (2008).
Banks, J. C. & Paterson, A. M. Multi-host parasite species in cophylogenetic studies. Int. J. Parasitol. 35, 741–746 (2005).
Doña, J. et al. Persistence of single species of symbionts across multiple closely-related host species. Sci. Rep. 9, 4–5 (2019).
Antonovics, J. et al. The evolution of transmission mode. Philos. Trans. R. Soc. B Biol. Sci. 372, 7–11 (2017).
Hayward, A., Poulin, R. & Nakagawa, S. A broadscale analysis of host‐symbiont cophylogeny reveals the drivers of phylogenetic congruence. Ecol. Lett. 24, 1681–1696 (2021).
Doña, J., Serrano, D., Mironov, S., Montesinos-Navarro, A. & Jovani, R. Unexpected bird–feather mite associations revealed by DNA metabarcoding uncovers a dynamic ecoevolutionary scenario. Mol. Ecol. 28, 379–390 (2019).
Doña, J. et al. Vertical transmission in feather mites: insights into its adaptive value. Ecol. Entomol. 42, 492–499 (2017).
Clayton, D. H. & Johnson, K. P. What’s bugging brood parasites? Trends Ecol. Evol. 16, 9–10 (2001).
Clayton, D. H. & Tompkins, D. M. Ectoparasite virulence is linked to mode of transmission. Proc. R. Soc. B Biol. Sci. 256, 211–217 (1994).
Harbison, C. W., Bush, S. E., Malenke, J. R. & Clayton, D. H. Comparative transmission dynamics of competing parasite species. Ecology 89, 3186–3194 (2008).
Matthews, A. E. et al. Dispersal-limited symbionts exhibit unexpectedly wide variation in host specificity. Syst. Biol. c, 1–10 (2023).
Solter, L. F. Transmission as a predictor of ecological host specificity with a focus on vertical transmission of microsporidia. J. Invertebr. Pathol. 92, 132–140 (2006).
Dabert, J. Feather mites (Astigmata; Pterolichoidea, Analgoidea) and birds as models for cophylogenetic studies. Phytophaga 14, 409–424 (2004).
Doña, J., Proctor, H., Mironov, S., Serrano, D. & Jovani, R. Host specificity, infrequent major host switching and the diversification of highly host-specific symbionts: the case of vane-dwelling feather mites. Glob. Ecol. Biogeogr. 27, 188–198 (2018).
Gaud, J. & Atyeo, W. T. Feather mites of the world (Acarina, Astigmata): the supraspecific taxa. Ann. du Musée R. l’Afrique Cent. Sci. Zool. 277, (Pt.1 1–193 (text) & Pt. 2 1–436 (illustrations)) (1996).
Proctor, H. C. Feather mites (Acari: Astigmata): ecology, behavior, and evolution. Annu. Rev. Entomol. 48, 185–209 (2003).
OConnor, B. M. Life-history modifications in astigmatid mites. In: Mites: Ecological and Evolutionary Analyses of Life-history Patterns. (ed. Houck, M. A.) 136–159 (Chapman & Hall, 1994).
Fain, A. A review of the family Epidermoptidae Trouessart parasitic on the skin of birds (Acarina: Sarcoptiformes). Verh. van K. Vlaam. Acad. voor Wet. Lett. en schone kunsten van België, Klasse der Wet. 84, 1–176 (I);-1–144 (II) (1965).
Brooke, M. D. L. Vertical transmission of feather lice between adult blackbirds Turdus merula and their nestlings: a lousy perspective. J. Parasitol. 96, 1076–1080 (2010).
Johnson, K. P., Weckstein, J. D., Bush, S. E. & Clayton, D. H. The evolution of host specificity in dove body lice. Parasitology 138, 1730–1736 (2011).
Mironov, S. V. & Malyshev, L. L. Dynamics of infection the Chaffinch nestlings Fringilla coelebs with feather mites (Acari: Analgoidea). Parazitologiia 36, 356–374 (2002). [In Russian with English summary].
Ehrnsberger, R., Mironov, S. V. & Dabert, J. A preliminary analysis of phylogenetic relationships of the feather mite family Freyanidae Dubnin,1953 (Acari: Astigmata). Biol. Bull. 38, 181–201 (2001).
Post, W., Cruz, A. & McNair, D. B. The North American invasion pattern of the shiny cowbird. J. F. Ornithol. 64, 32–41 (1993).
Sykes Jr, P. W. & Post, W. First specimen and evidence of breeding by the shiny cowbird in Georgia. Oriole 66, 45–51 (2001).
Lowther, P. E. Lists of victims and hosts of the parasitic cowbirds (Molothrus). Field Museum of Natural History, Chicago, Illinois, USA https://www.fieldmuseum.org/blog/brood-parasitism-host-lists (2019).
Mironov, S. V., Literak, I. & Čapek, M. New feather mites of the subfamily Pterodectinae (Acari: Astigmata: Proctophyllodidae) from passerines (Aves: Passeriformes) in Mato Grosso do Sul, Brazil. Zootaxa 38, 1–38 (2008).
Pedroso, L. G. A. & Hernandes, F. A. Two new feather mites of the genus Proctophyllodes Robin (Acariformes: Proctophyllodinae) from passerines in Brazil. Syst. Appl. Acarol. 26, 1081–1096 (2021).
Doña, J., Proctor, H., Mironov, S., Serrano, D. & Jovani, R. Global associations between birds and vane-dwelling feather mites. Ecology 97, 3242–3242 (2016).
Atyeo, W. T. & Braasch, N. L. The feather mite genus Proctophyllodes (Sarcoptiformes: Proctophyllodidae). Bull. Univ. Nebrasca State Mus. 5, 1–354 (1966).
Barker, K., Burns, K. J., Klicka, J., Lanyon, S. M. & Lovette, I. J. New insights into New World biogeography: an integrated view from the phylogeny of blackbirds, cardinals, sparrows, tanagers, warblers, and allies. Auk 132, 333–348 (2015).
Remsen, J. V., Powell, A. F. L. A., Schodde, R., Barker, F. K. & Lanyon, S. M. A revised classification of the Icteridae (Aves) based on DNA sequence data. Zootaxa 4093, 285–292 (2016).
Gómez, R. O. & Lois-Milevicich, J. Phylogenetic signal in the skull of cowbirds (Icteridae) assessed by multivariate and cladistic approaches. Zool. Anz. 286, 52–57 (2020).
Linz, G. M., Michael L. Avery & Dolbeer, R. A. Ecology and management of blackbirds (Icteridae) in North America. 252 pages, CRC Press (2017).
Louder, M. I. M., Ward, M. P., Schelsky, W. M., Hauber, M. E. & Hoover, J. P. Out on their own: a test of adult-assisted dispersal in fledgling brood parasites reveals solitary departures from hosts. Anim. Behav. 110, 29–37 (2015).
Ortega, C. R., Cruz, A. & Mermoz, M. E. Issues and controversies of Cowbird (Molothrus spp.) management. Ornithol. Monogr. 57, 6–15 (2005).
Sick, H. Ornitologia Brasileira. (Nova Fronteira, 1997).
Soler, M. Avian brood parasitism: behaviour, ecology, evolution and coevolution. (Springer International Publishing, 2017).
Labrador, M. D. M., Doña, J., Serrano, D. & Jovani, R. Feather mites at night: an exploration of their feeding, reproduction, and spatial ecology. Ecology 103, 1–5 (2022).
Mena, M. et al. Parasites of the shiny cowbird, Molothrus bonariensis, and the austral blackbird, Curaeus curaeus, (Passeriformes: Icteridae) in Chile. Braz. J. Vet. Parasitol. 29, 1–10 (2020).
Proctor, H. C. & Jones, D. N. Geographical structuring of feather mite assemblages from the Australian Brush-Turkey (Aves: Megapodiidae). J. Parasitol. 90, 60–66 (2004).
Peer, B. D., Rothstein, S. I., Kuehn, M. J. & Fleischer, R. C. Host defenses against cowbird (Molothrus spp.) Parasitism: Implications for Cowbird Management. Ornithol. Monogr. 57, 84–97 (2005).
Atyeo, W. T. & Gaud, J. Feather mites of obligate brood parasites. J. Parasitol. 69, 455–458 (1983).
Hahn, D. C., Price, R. D. & Osenton, P. C. Use of lice to identify cowbird hosts. Auk 117, 943–951 (2000).
Hilario-Pérez, A. & Dowling, A. P. G. Nasal mites from specimens of the brown-headed cowbird (Icteridae: Molothrus ater) from Texas and Arkansas, U.S.A. Acarologia 58, 296–301 (2018).
Lindholm, A. K., Venter, G. J. & Ueckermann, E. A. Persistence of passerine ectoparasites on the diederik cuckoo Chrysococcyx caprius. J. Zool. 244, 145–153 (1998).
Blanco, G., Tella, J. L. & Potti, J. Feather mites on group-living Red-billed Choughs: a non-parasitic interaction? J. Avian Biol. 28, 197–206 (1997).
Blanco, G. & Frías, O. Symbiotic feather mites synchronize dispersal and population growth with host sociality and migratory disposition. Ecography 24, 113–120 (2001).
Darolova, A., Hoi, H., Kristofik, J. & Hoi, C. Horizontal and vertical ectoparasite transmission of three species of Malophaga, and individual variation in European Bee-Eaters (Merops apiaster). J. Parasitol. 87, 256–262 (2001).
Dick, C. W., Esbérard, C. E. L., Graciolli, G., Bergallo, H. G. & Gettinger, D. Assessing host specificity of obligate ectoparasites in the absence of dispersal barriers. Parasitol. Res. 105, 1345–1349 (2009).
Dabert, J. & Mironov, S. V. Origin and evolution of feather mites. Exp. Appl. Acarol. 23, 437–454 (1999).
Bush, S. E. & Clayton, D. H. The role of body size in host specificity: reciprocal transfer experiments with feather lice. Evolution 60, 2158 (2006).
Mestre, A., Poulin, R. & Hortal, J. A niche perspective on the range expansion of symbionts. Biol. Rev. 95, 491–516 (2020).
Jetz, W., Thomas, G. H., Joy, J. B., Hartmann, K. & Mooers, A. O. The global diversity of birds in space and time. Nature 491, 444–448 (2012).
Wells, K. & Clark, N. J. Host specificity in variable environments. Trends Parasitol. 35, 452–465 (2019).
Hill, D. S., Wilson, N. & Corbet, G. B. Mites associated with british species of Ornithomya (Diptera: Hippoboscidae). J. Med. Entomol. 4, 102–122 (1967).
Matthews, A. E. et al. Feather mite abundance varies but symbiotic nature of mite-host relationship does not differ between two ecologically dissimilar warblers. Ecol. Evol. 8, 1227–1238 (2018).
Fain, A. & Grootaert, P. Obsevations sur des Acarines (Acari: Epidermoptidae), parasites d’Ornithomyia avicularia (L.) (Diptera: Hippoboscidae) de Belgique. Bull. Annls Soc. r. Belg. Entomol. 132, 183–186 (1996).
Mironov, S. V., Bochkov, A. V. & Fain, A. Phylogeny and evolution of parasitism in feather mites of the families Epidermoptidae and Dermationidae (Acari: Analgoidea). Zool. Anz. 243, 155–179 (2005).
Doña, J. et al. DNA barcoding and minibarcoding as a powerful tool for feather mite studies. Mol. Ecol. Resour. 15, 1216–1225 (2015).
Salichos, L., Stamatakis, A. & Rokas, A. Novel information theory-based measures for quantifying incongruence among phylogenetic trees. Mol. Biol. Evol. 31, 1261–1271 (2014).
Stamatakis, A. RAxML version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Hernandes, F. A., Matthews, A. E. & Boves, T. J. Four new feather mite species of the genus Amerodectes Valim & Hernandes (Acariformes: Proctophyllodidae) from New World warblers (Passeriformes: Parulidae) in the USA. Syst. Appl. Acarol. 23, 946–968 (2018).
Nguyen, L. T., Schmidt, H. A., Von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Minh, B. Q., Nguyen, M. A. T. & Von Haeseler, A. Ultrafast approximation for phylogenetic bootstrap. Mol. Biol. Evol. 30, 1188–1195 (2013).
Kalyaanamoorthy, S., Minh, B. Q., Wong, T. K. F., Von Haeseler, A. & Jermiin, L. S. ModelFinder: fast model selection for accurate phylogenetic estimates. Nat. Methods 14, 587–589 (2017).
Rambaut, A. FigTree FigTree v1.4.4. Institute of Evolutionary Biology, University of Edinburgh, Edinburgh. http://tree.bio.ed.ac.uk/software/figtree/ (2018).
Puillandre, N., Brouillet, S. & Achaz, G. ASAP: assemble species by automatic partitioning. Mol. Ecol. Resour. 21, 609–620 (2021).
Hebert, P. D. N., Cywinska, A., Ball, S. L. & DeWaard, J. R. Biological identifications through DNA barcodes. Proc. R. Soc. B Biol. Sci. 270, 313–321 (2003).
Klimov, P. B., Skoracki, M. & Bochkov, A. V. Cox1 barcoding versus multilocus species delimitation: validation of two mite species with contrasting effective population sizes. Parasites Vectors 12, 1–15 (2019).
Paradis, E. & Schliep, K. Ape 5.0: an environment for modern phylogenetics and evolutionary analyses in R. Bioinformatics 35, 526–528 (2019).
Bouckaert, R. et al. BEAST 2: a software platform for bayesian evolutionary analysis. PLoS Comput. Biol. 10, 1–6 (2014).
Rothstein, S. I., Pattern, M. A. & Fleicher, R. C. Phylogeny, specialization, and brood parasite-host coevolution: some possible pitfalls of parsimony. Behav. Ecol. 13, 1–10 (2002).
Santichaivekin, S. et al. EMPRess: a systematic cophylogeny reconciliation tool. Bioinformatics 37, 2481–2482 (2021).
Acknowledgements
We thank the following museum curators for providing access to host skins for our sampling (see Supplemenary Note 2 for museum abbreviations): Dr. Glayson Ariel Bencke (MCN); Dr. Carla Suertegaray Fontana from (MCT); Antenor Silva Junior and Ma. Patrícia Weckerlin e Silva (MHNCI); Dr. Luis Fábio Silveira (MZUSP); Dr. Alexandre Aleixo (MPEG). We thank the following people who helped us organize field trips—Dr. Gerturd Müller (UFPel), Dr. Fabiana F. Bernardon (UFPel), Dr. Lilian Manica (UFPR), Dr. Luiz dos Anjos (UEL), Dr. Edson Guilherme da Silva (UFAC), Dr. Mauro Pichorim (UFRN), Dr. Luciano Naka (UFPE), Dr. Roberto Cavalcanti (UNB), Dr. Sérgio Posso (UFMS), and Dr. Marco A. Pizo (UNESP-RC). Dr. Augusto F. Batisteli donated salvaged shiny cowbirds. Molecular work and analyses were done by L.G.A.P. and P.B.K. at the University of Michigan Museum of Zoology. L.G.A.P. was funded by São Paulo Research Foundation (FAPESP)—2018/21504-0 and 2016/11671-1. P.B.K. was supported by a grant from the Ministry of Science and Higher Education of the Russian Federation within the framework of the Federal Scientific and Technical Program for the Development of Genetic Technologies for 2019-2027 (agreement No. 075-15-2021-1345, unique identifier RF 193021×0012); S.V.M. was supported by the Russian Foundation for Basic Research (RFBR), grant No. 20-04-00500; F.A.H. was supported by National Council for Scientific and Technological Development, CNPq-Brazil Researcher (304479/2019-5).
Author information
Authors and Affiliations
Contributions
L.G.A.P. and P.B.K. performed molecular laboratory work and contributed to writing the manuscript. L.G.A.P. was responsible for the figures’ art schemes. F.A.H., Q.H., K.P.J., H.R.B., S.V.M., A.R.P. and B.M.OC. contributed with discussion and critical revision.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Kasun Bodawatta and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Tobias Goris. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pedroso, L.G.A., Klimov, P.B., Mironov, S.V. et al. Horizontal transmission maintains host specificity and codiversification of symbionts in a brood parasitic host. Commun Biol 6, 1171 (2023). https://doi.org/10.1038/s42003-023-05535-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-023-05535-1
- Springer Nature Limited