Abstract
Early embryonic development is a dynamic process that relies on proper cell-cell communication to form a correctly patterned embryo. Early embryo development-related ligand-receptor pairs (eLRs) have been shown to guide cell fate decisions and morphogenesis. However, the scope of eLRs and their influence on early embryo development remain elusive. Here, we developed a computational framework named TimeTalk from integrated public time-course mouse scRNA-seq datasets to decipher the secret of eLRs. Extensive validations and analyses were performed to ensure the involvement of identified eLRs in early embryo development. Process analysis identified that eLRs could be divided into six temporal windows corresponding to sequential events in the early embryo development process. With the interpolation strategy, TimeTalk is powerful in revealing paracrine settings and studying cell-cell communication during early embryo development. Furthermore, by using TimeTalk in the blastocyst and blastoid models, we found that the blastoid models share the core communication pathways with the epiblast and primitive endoderm lineages in the blastocysts. This result suggests that TimeTalk has transferability to other bio-dynamic processes. We also curated eLRs recognized by TimeTalk, which may provide valuable clues for understanding early embryo development and relevant disorders.
Similar content being viewed by others
Introduction
Early embryo development is a multi-level regulatory process in which fertilized egg undergoes rounds of cleavage to form a self-organized, hollow sphere structure called blastocyst1. At the beginning of the early embryo development, the paternal gametes sperm and maternal gametes oocytes fused to form a one-cell embryo called the zygote in fertilization. Then, the one-cell embryo zygote experiences rounds of cell cleavage division with maternal factor decay and zygotic genome activation (ZGA), which is called maternal to zygotic transition2. After the fourth cleavage division, the embryo begins to compact into blastomeres. With blastomere formation, there are two critical differentiation events: the first cell fate decision and the second cell fate decision. In the first cell fate decision from the 8-cell stage to the 32-cell stage, the cells in the embryo segregate into two lineages: trophectoderm (TE) and inner cell mass (ICM), while TE lineage will develop into the placenta. However, in the second cell fate decisions from the early-blastocyst stage to the late-blastocyst stage, the cells in the ICM lineage differentiated into epiblast (EPI) and primitive endoderm (PE) lineage. The EPI lineage will develop into the fetus, and the PE lineage will develop into the Yolk sac1,3,4. The phenomena intrigue evolving research on the mechanisms of the two fate decision events4,5.
Cell-cell communication is a fundamental process in which, for a given cell, the membrane protein (i.e., receptor) is bound by the protein secreted by other cells or itself, called a ligand, inducing intracellular signaling responses6. Recently, many studies illustrated that cell-cell communication participates in the two fate-decision events during early embryo development. For example, during the first fate decision, LIF-JAK/STAT pathway-related pair Lif-Lifr is essential for ICM lineage maintenance7. Nevertheless, during the second fate decision, the communication between cells mediated by Fgf4-Fgfr1 and Fgf4-Fgfr2 interplay with master regulator Nanog and Gata6 promote PE lineage formation and maturation1. The progenitor cells in the ICM with high expression of Gata6 express high levels of the FGF receptor genes Fgfr1 and Fgfr2. Similarly, the progenitor cells in the ICM with high expression of Nanog express high levels of Fgf4. The binding of FGF4 secreted by Nanoghigh cells to FGFR1 or FGFR2 in Gata6high cells constructs a positive feedback loop with GATA6 to activate gene expression of primitive endoderm program1,8. Moreover, the cell-cell communication mediated by Fgf4-Fgfr1 of Nanoghigh cells promotes EPI exiting from pluripotency9. Thus, Fgf4-Fgfr1 and Fgf4-Fgfr2 regulate PE and EPI formation.
In conclusion, previous research indicates that cell-cell communication via ligand-receptor pairs has critical functions during early embryo development. We termed these early embryo development-related ligand-receptor pairs as eLRs. Studying eLRs will help us better understand cell-cell communication during early embryo development. However, the scope of eLRs and their influence on early embryo development is still lacking.
With the advancement of functional genomics research, more cell-cell communication priori knowledge resources, including ligand-receptor databases, have been accumulated in recent years6. On the other hand, the development of single-cell RNA sequencing (scRNA-seq) technology has made it possible to infer the cell-cell communication events between different cells. Consequently, integrating single-cell transcriptomic sequencing data and prior knowledge to infer cell-cell communication has emerged as a new research direction in bioinformatics. As for early embryo development studies, research performed scRNA-seq over each development stage to obtain the time-course scRNA-seq data to measure the dynamic changes in cell states and types. However, it is critical to note that definitive cell types do not emerge during early embryo development until blastocyst formation. However, commonly used tools for cell-cell communication inference, such as CellPhoneDB10 and CellChat11, are designed to study cell-cell communication between given cell types and cannot meet the requirements of early embryo development research. Moreover, the need for multiple time points of cell-cell communication analysis during early embryo development and the need to elucidate potential causal relationships between cell-cell communication and gene regulatory networks during dynamic changes of early embryo development also pose challenges to existing cell-cell communication research.
To address these issues, we developed and applied a computational framework named TimeTalk for utilizing temporal series information to study the dynamics of autocrine signaling within the embryos and to identify early embryo development-related ligand-receptor pairs (eLRs) from integrated public time-course mouse scRNA-seq datasets. Our analysis identified 430 eLRs, including previously reported Fgf4-Fgfr1 and Fgf4-Fgfr2. Additionally, we conducted thorough in silico analyses to test the involvement of identified eLRs in early embryo development. After validation, we found that the identified eLRs can be divided into six temporal windows. Furthermore, the GO analysis reveals that different temporal windows correspond with sequential early embryo development events. Moreover, we used Granger causality and network analysis to discover and check the potential regulation relationship between eLR and corresponding temporal TFs (tTFs). To broaden the application of TimeTalk to paracrine studies and other development processes, we further improved the framework, used it to investigate cell-cell communication in blastocysts, and reconstructed in-vitro embryo models named blastoids. The results indicated that the communication between EPI and PE lineages in blastoids involves shared core LR pairs and signaling pathways as in naturally developed blastocysts. In summary, the identified eLRs would be a valuable resource for better understanding early embryo development, and some would be targets to perturb some key early embryo development processes. In addition, we believed TimeTalk would be a helpful toolkit to study cell-cell communication in other development processes. TimeTalk is an R package that is available at https://github.com/ChengLiLab/TimeTalk.
Results
Curation of early-embryo development single-cell RNA-seq data sets for studying cell-cell communication
To identify and study eLRs, we collected public early embryo development scRNA-seq datasets from the mouse MII-oocyte stage to the late blastocyst stage to ensure that scRNA-seq datasets represented every stage of early embryo development. In addition, to validate the quality of scRNA-seq data, we also collected public low-input RNA-seq data (Fig. 1a).
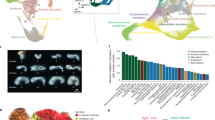
a Metadata of the collected early embryo development datasets. b Heatmap of the Pearson correlation coefficients between bulk and pseudo-bulk data. c tSNE plot of the integrated single-cell RNA-seq data. d Feature plot of the different marker genes.The unit for color gradient keys is LogNormalize, which involves dividing the feature counts for each cell by the total counts for that cell and then multiplying it by the scale.factor. This value is then transformed using the natural-log function and adding 1. e Pseudotime ordering of collected early embryo cells reveals development trajectory. The top of the figure was colored by development stages, and the bottom was colored by pseudotimes.
We first validated the quality of the scRNA-seq data. Each stage’s overall gene expression distributions were similar for the low-input RNA-seq data and the pseudo-bulk single-cell RNA-seq data (Supplementary Fig. 1a–d; for low-input RNA-seq, gene expression was quantified by RPKM; for pseudo-bulk single-cell RNA-seq data, gene expression was quantified based on the average gene expression of cells in different stages). The principal component analysis (PCA) showed that the single-cell RNA-seq and low-input RNA-seq datasets were clustered together at each stage (Supplementary Fig. 1e). By comparing the expression profile of pseudo-bulk RNA-seq data derived from scRNA-seq data and low-input RNA-seq data, we found that the pseudo-bulk and low-input RNA-seq expression profiles of the samples from each stage were highly correlated. For example, the Pearson correlation coefficient of the bulk late 2-cell expression profile with the pseudo-bulk late 2-cell expression profile was 0.91 (Fig. 1b and Supplementary Fig. 2). Consider MII-oocyte (E0) scRNA-seq data (GEO accession: GSE38495)12 and E1-E4.0 scRNA-seq datasets (GEO accession, GSE45719)13, which were obtained from two different studies. To ensure that potential batch effects could be disregarded, we performed the kBET analysis14. The results of the kBET analysis suggested that the batch effects were negligible (see Supplementary Fig. 3). Therefore, we can merge the two datasets without performing batch correction. Next, we put the scRNA-seq data into the Seurat toolkit to perform t-SNE dimension reductions (Fig. 1c) and check each stage’s marker gene expression (Fig. 1d). Upon checking the marker gene expression, and the monocle2 package15 was utilized to perform trajectory analysis and arrange the cells along the constructed trajectory (Fig. 1e).
In summary, the curated early embryo scRNA-seq datasets recapitulate the early embryo development process.
TimeTalk is a computational framework that utilizes temporal series information for eLR identification
After building the comprehensive transcriptomics datasets, we began to identify eLRs. First, we obtained the ligand-receptor list from the comprehensive CellTalkDB database16. Then, we studied the gene expression distribution of ligands and receptors across developmental stages. We found that the expression levels of ligands and receptor genes were lower (Supplementary Fig. 4a) and more dynamic (Supplementary Fig. 4b, c) in comparison with the overall trends of gene expression at each stage. Furthermore, the previously reported eLRs exhibited significant co-variation changes along the pseudotime (Supplementary Fig. 5).
According to previous studies, well-studied eLR pairs Fgf4-Fgfr1 and Fgf4-Fgfr2 interplay with transcription factors Sox2, Nanog, Oct4 (Pou5f1), Gata6 to form a complex regulatory network to guide the ICM to differentiate into EPI or PE lineage1 (Fig. 2a). Therefore, after ranking the cells according to pseudo-time inference by the monocle2 package, we investigate the expression of Fgf4, Fgfr1, Fgfr2, and related transcription factors Gata6, Pou5f1, Sox2, and Nanog (Supplementary Fig. 6). We found that the gene expression of Fgf4 and Fgfr2 were negatively correlated (Fig. 2b). Moreover, we calculate the geometry means of Fgf4 and Fgfr2 gene expression (Interaction Score, or can be noted as IS)17 to quantify the cell-cell communication activity of Fgf4-Fgfr2. We found that the IS of Fgf4-Fgfr2 was correlated with Gata6 gene expression (Fig. 2c).
a The Fgf4-Fgfr1 and Fgf4-Fgfr2 work together to regulate a core pluripotency gene network. Red lines with arrows show positive regulation, while blue lines with inhibitory marks indicate negative regulation. The black lines represent the gene expression process. This figure was created using BioRender (https://app.biorender.com/). b The ligand and receptor genes in Fgf4-Fgfr2 co-vary during development. PCC, Pearson’s correlation coefficient, SCC, Spearman’s correlation coefficient. c The dynamics of interaction score of Fgf4-Fgfr2 and transcription factor Gata6. The p-value of p_TFtoLR and p_LRtoTF were calculated by the Granger causal test, revealing the potential causal relationship between TF and LR. d The workflow of the preliminary version of TimeTalk. e Illustration of the relationship between the activated LR and TFs. Forward means LR activates TF, backward means TF regulates LR, and feedback means LR and TF mutually regulate each other. f The 229 temporal TFs (tTFs) revealed by master regulator analysis were divided into six clusters. The right side marked representative TFs of each group. g The GO analysis of six clusters of tTFs. h The dynamics of tTFs in C5 gradually decreased during development. i The line plot depicts the dynamic variations in the interaction score of Fgf4-Fgfr2 and the activity of C5 tTFs.
These results indicated that eLR gene pairs show coordinated expression patterns with core transcription factors during early embryo development. According to these observations, we developed an early-embryo ligand-receptor screening strategy, TimeTalk, to identify eLRs with the following steps. (1) Building trajectory from integrated single-cell RNA-seq datasets, ordering the single-cells along the trajectory. (2) A ligand-receptor database was used to identify the co-varying LR pairs from the pseudotime time series data (Supplementary Fig. 7, details are described in the Methods section). Meanwhile, the transcriptional regulatory network is reconstructed from scRNA-seq data to identify the temporal TFs (tTFs) during early embryo development. (3) Cluster the tTFs. (4) Inference of the regulatory relationship between the different clusters of tTF and co-varying LR pairs through the Granger causality approach by Granger causal test. (5) The LR pairs that passed the Granger causal test were considered as forward (LR regulates TF), backward (TF regulates LR), and feedback (LR and TF were mutually regulated) (Fig. 2d, e, details are described in the Methods section).
According to TimeTalk workflow, we identified 430 eLRs, and as a by-product of eLR identification, we identified 229 tTFs during early embryo development. The 229 tTFs were clustered into six distinct groups (Supplementary Data 1, Supplementary Fig. 8). Subsequent analysis revealed that the previously reported eLR Bmp4-Bmpr2 also exhibited varying Granger causality relationships with different clusters of tTFs (Supplementary Fig. 9). Cluster C1 tTFs consist of maternal factors such as Nfya18, Obox5, and Atf2 and they are enriched in GO terms like “oogenesis” and “germ cell development”. Cluster C2 contains ZGA genes like Zscan4f and is enriched with GO terms like “histone methylation” and “protein methylation”. Cluster C3 comprises TFs that play a role in establishing heterogeneity. For example, Sox21 contained in cluster C3 was reported to be involved in fate biases starting from the 4-cell stage19. Cluster C4 is composed of TFs like Ctcf involved in chromatin reorganization and enriched in GO terms like “histone methylation”, “chromatin silencing”, and “chromatin assembly or disassembly”, “DNA conformation change”. Cluster C5 is composed of tTFs involved in blastocyst development like Gata6, Tead4, Cdx2, Nanog, and Pou5f1 and enriched in GO terms like “trophectodermal cell differentiation”, “blastocyst development”, “blastocyst formation”, “blastocyst growth”. Cluster C6 is also composed of tTFs involved in blastocyst development like Sox2 and enriched GO term “embryonic organ development”, and “regulation of epithelial cell differentiation” (Fig. 2f, g, Supplementary Data 1). Additionally, we found that C5 tTFs regulate Fgf4-Fgfr2 activity (Fig. 2h, i, Supplementary Fig. 10).
In summary, the tTFs identified here provided potential targets for manipulating early embryo development at different temporal windows.
The predicted eLRs can be verified from extensive in-silico validations
Next, we validated the eLRs identified by TimeTalk from multiple in-silico strategies.
The candidate eLRs predicted by the TimeTalk workflow contain reported eLRs
We ranked the identified eLRs pairs according to their PCC values. A literature review yielded evidence that the top 30 positively correlated eLRs and the top 30 negatively correlated eLRs participate in early embryo development (Fig. 3a, Supplementary Data 1). The eLRs identified using TimeTalk also included ligands and receptors with established roles in early embryo development. For example, the candidate eLRs contain Bmp6-Bmpr2, which has been reported to regulate extra-embryonic lineage development20.
a The predicted eLRs; each dot represents an eLR. Previously reported eLR pairs were highlighted. b The enrichment ratio of essential genes in the sets of all protein-coding genes (all), ligands and receptors (LRs), and eLRs. c The enrichment ratio of housekeeping genes in all protein-coding genes (all), ligands and receptors (LRs), and eLRs. d The enrichment ratio of ZGA genes in the sets of all protein-coding genes (all), ligands and receptors (LRs), and eLRs. We used the hypergeometric test to assess the results’ significance in (b–d). e Heatmap of the interaction scores of some eLRs that were down-regulated by the ZGA inhibition, the ZGA-inhibited RNA-seq data downloaded from GSE71434.
The enrichment of essential gene in the candidate eLRs indicate that the genes composed of predicted eLRs were mainly required for early embryo development
Essential genes are required for organisms to survive and fit in with the environment21,22. A loss of function of any essential gene generally produces serious deleterious effects, including embryonic lethality23. Thus, essential genes among the eLRs identified using TimeTalk are likely required for early embryo development. We obtained the mouse essential gene list provided by the DEG 10 database24 and computed essential gene set enrichment ratios for all protein-coding genes (all), ligand and receptor genes (LRs), and eLRs. In comparison with all protein-coding genes, the set of essential genes was more significantly enriched with LRs and eLRs (Fig. 3b, Supplementary Data 1, “LR” vs. “all”, p = 1.628 × 10−111; “eLR” vs. “all” p = 3.049 × 10−85, p-values were obtained by hypergeometric test, upper tail). This observation was consistent with the principle that the proteins encoded by ligand and receptor genes required cells to respond to external stimuli. In addition, the set of essential genes was more enriched in eLRs in comparison with LRs (Fig. 3b, Supplementary Fig. 11a, Supplementary Data 1, “eLR” vs. “LR”, p = 4.344 × 10−14, p-values were obtained by hypergeometric test, upper tail). The enrichment of essential gene in the candidate eLRs indicate the genes composed of predicted eLRs were mainly required for the organism’s fitness.
The absence of housekeeping genes in the candidate eLRs suggests that the genes comprising the predicted eLRs were not required for individual cell survival
Housekeeping genes are stably expressed genes that maintain basic cellular functions regardless of cell type or developmental stage25. In contrast, the genes comprising eLRs should be dynamically expressed to coordinate early embryo development. This result led us to suspect that the identified eLR genes had been absent of housekeeping genes. Therefore, we obtained the housekeeping gene list provided by the HRT Atlas v1.0 database26 and calculated the housekeeping gene enrichment ratio for all protein-coding genes (all), ligand and receptor genes (LRs), and eLR genes (eLRs). As expected, in comparison with the set of all protein-coding genes, the set of eLR genes was more depleted of housekeeping genes (Fig. 3c, Supplementary Data 1, “LR” vs. “all”, p = 1.944 × 10-47; “eLR” vs. “all”, p = 5.986 × 10−16, p-values were obtained by hypergeometric test, lower tail). Moreover, the housekeeping gene enrichment ratio of the set of all ligand and receptor genes was not significantly different from that of the identified eLRs (Fig. 3c, Supplementary Fig. 11b, Supplementary Data 1). Together with the essential gene enrichment results, these findings suggest that the eLR genes identified using TimeTalk were more important for the organism’s fitness, rather than for the survival of an individual cell.
Many genes composed of the candidate eLRs were activated by the ZGA process, and the cell-cell communication mediated by the candidate eLRs required the ZGA process
ZGA is the embryogenesis process in which maternally provided factors are replaced by factors supplied by zygotic transcription27. Transcription of the ZGA gene is required for lineage specification and provides substrates for the initiation of gastrulation27,28. Therefore, ZGA is vital for early embryo development. Thus, we hypothesized that ZGA genes were more enriched in the identified eLR genes than the LR genes. The ZGA gene list was obtained from ref. 18, and the ZGA gene enrichment ratio was calculated for the protein-coding genes (all), ligand and receptor genes (LRs), and eLR genes (eLRs). The enrichment ratio of the ZGA gene in the protein-coding gene set was not significantly different from that of the LR gene set (Fig. 3d, Supplementary Data 1, Supplementary Fig. 11c, p = 0.09293, p-values were obtained by hypergeometric test, upper tail). However, the ZGA gene enrichment ratio of the eLRs was significantly different from that of the protein-coding gene set (Fig. 3d, Supplementary Fig. 11c, Supplementary Data 1, p = 0.001159, p-values were obtained by hypergeometric test, upper tail). In addition, the ZGA gene enrichment ratio of the eLRs was higher than that of the LR gene (Fig. 3d, Supplementary Fig. 11c, Supplementary Data 1, p = 0.00279, p-values were obtained by hypergeometric test, upper tail). These findings indicate that the set of eLRs identified using TimeTalk was enriched with ZGA genes when compared to both LR genes and protein-coding genes. This result suggests that the many genes composed of candidate eLRs were activated during the ZGA process.
It has been established that ZGA is essential for pre-implantation in mice29. Considering that eLR-mediated cell-cell communication is essential for early embryo development, we hypothesized that the signaling mediated by specific eLR pairs might require ZGA. Accordingly, we obtained ZGA inhibition RNA-seq datasets30 and calculated interaction scores17 (the calculation of interaction score is described in the Methods section) to quantify the cell-cell communication strength mediated by eLR pairs. As a result, ZGA inhibition altered the cell-cell communication strength of 56 eLR pairs (Fig. 3e, Supplementary Fig. 12). Thus, the establishment of candidate eLRs mediated cell-cell communication correlated with the ZGA process. Furthermore, compared to non-eLR pairs, eLR pairs show less impact from ZGA inhibition (Supplementary Fig. 13, Supplementary Data 1). This suggests that eLR pairs possess additional regulation mechanisms beyond ZGA.
In summary, the results above demonstrate that the eLR pairs identified by TimeTalk have several essential characteristics. First, many highly ranked eLR pairs have been reported in the literature in relevant contexts. Second, the identified eLR pairs are enriched with the essential gene and the ZGA gene but depleted of the housekeeping gene. Third, ZGA inhibition disrupted some types of cell-cell communication mediated by the identified eLR pairs. Finally, these characteristics supported the involvement of the identified eLR pairs in early embryo development.
The interplay between the tTFs and eLR orchestrates early embryo development
As it is previously reported, the interplay between the eLR Fgf4-Fgfr2 and tTF Gata6 formed a positive regulatory network to guide the separation of PE and EPI lineage during early embryo development8. However, no comprehensive prolling of the relationship between eLR and tTF has been conceived. Therefore, we hypothesized that investigating the interplay between eLR and TF will help us to understand the gene regulatory network to control mouse embryo development.
The identified eLRs can be clustered into 6 clusters associated with different groups of temporal TFs. Cluster 1 eLRs have a backward relationship with C1 tTFs. This result implies that maternal tTFs control the cluster 1 eLRs. This conclusion is supported by the observation that cluster 1 eLRs exhibit a higher ratio of the maternal gene (Supplementary Fig. 14a). Cluster 3 eLRs exhibit a higher ratio of ZGA genes (Supplementary Fig. 14b). In addition, cluster 1 eLRs have a feedback relationship with C5 tTFs. Interestingly, cluster 4 eLRs have a forward relationship with C1-C6 tTFs. Cluster 5 eLRs have feed relationships with C1, C2, C3, and C4 and backward relationships with C6. Cluster 6 eLRs have a backward relationship with C6 tTFs (Fig. 4a, Supplementary Data 2). Clusters 1, 2, 3, and 4 enriched MAPK signaling pathways (Fig. 4b, Supplementary Data 2). This pathway is involved in cell differentiation during early embryo development31,32.
a Heatmap of the interaction scores of all identified 430 eLRs can be grouped into 6 clusters. In the row annotation of the heatmap, the “cluster” column illustrates the classification of the eLR, C1, C2, C3, C4, C5, and C6 columns illustrate the relationship of each eLR to the C1, C2, C3, C4, C5, C6 regulons. In the column annotation of the heatmap, the “Stage” row illustrates the development stage of every single cell, the “Pseudotime” row illustrates the pseudotime, C1, C2, C3, C4, C5, C6 row illustrates the activity of every single cell. b The KEGG analysis of eLRs clustered for each clusters. c–e The cluster 1, cluster 5, and cluster 6 eLR to TF ratio, the signaling network provided by NicheNet can verify the ratio. f The distribution of eLR genes across the regulons of various tTFs. Each row in the figure corresponds to a specific tTF, with the horizontal bar indicating the ratio of eLR genes present in its regulon. g Fgf4-Fgfr1 and Fgf4-Fgfr2 interacted with different tTFs, forming an organized regulatory network.
Cluster 4,5,6 eLRs enriched KEGG term “Signaling pathways regulating pluripotency of stem cells”. Besides, clusters 4 and 6 enriched the “Hippo signaling pathway” related to trophectoderm lineage formation33(Fig. 4b, Supplementary Data 2). Upon analyzing the enrichment results, we discovered that cluster 4 eLR was enriched in the Hippo signaling pathway with 23 eLRs consisting of genes such as Cdh1, Wnt3a, Areg, Gdf5, Wnt5a, Wnt7a, Fgf1, Tgfbr1, Fzd2, Itgb2, Fzd5, Fzd1, Fzd4, Fzd9, Bmpr1b, Bmpr2, and Fzd3. Similarly, cluster 6 eLR was found to be enriched in the Hippo signaling pathway with six eLRs consisting of genes like Wnt3a, Wnt7b, Fzd7, Fzd5, Fzd3, and Fzd6 (Supplementary Fig. 15a). Most of these genes are known to be involved in the TGF-β and Wnt signaling pathways, which have crosstalk with Hippo signaling pathway34. The KEGG database also highlights this crosstalk between the Hippo, Wnt, and TGF-β signaling pathways (see web link from the KEGG database: https://www.genome.jp/pathway/mmu04390). An investigation was conducted to determine whether there is a connection between eLR activity and the Hippo signaling pathway. It was discovered that the C5 tTF includes Tead2 and Tead4, which are part of the core Hippo pathway gene sets35. Therefore, Tead2 and Tead4 can be indicators of Hippo signaling pathway activity. As for Tead2, the eLR Wnt3a-Fzd5 exhibited an increase in activity over pseudotime, with the Granger test indicating that Tead2 was responsible for activating Wnt3a-Fzd5 activity (Supplementary Fig. 15b). Prior research has demonstrated that both the Wnt3a and Fzd5 genes are crucial for subsequent trophectoderm lineage development36. Our findings, combined with this previous research, demonstrate that Hippo signaling pathway activation leads to eLR activation, indicating a potential role for the Hippo signaling pathway in regulating trophectoderm development by activating eLR activity. The eLR Cthrc1-Fzd3 exhibited a peak expression between the 4-cell stage and 16-cell stages with respect to Tead4 expression, suggesting that Tead4 has a feedback relationship with Cthrc1-Fzd3 activity (Supplementary Fig. 15c). Previous studies indicate that Cthrc1 selectively activates the planar cell polarity pathway of Wnt signaling by stabilizing the Wnt-receptor complex37. Additionally, Cthrc1 promotes trophoblast growth, migration, and invasion through reciprocal Wnt/β-catenin regulation38. Our findings suggest that the cell-cell communication triggered by Cthrc1 may also interact with the Hippo signaling pathway to promote trophectoderm lineage formation.
In addition, we verified regulation from eLR to tTF by the signaling network created from NicheNet. (Fig. 4c–e, Supplementary Fig. 16a–c, Supplementary Data 2). Besides, we investigate regulation relationships from tTFs to eLRs and find that C1 and C6 regulate more eLRs than the other cluster of tTFs (Fig. 4f, Supplementary Fig. 17, Supplementary Data 2). We used BioTapestry39 to illustrate well-studied eLRs Fgf4-Fgfr1, Fgf4-Fgfr2 interplay with temporal TFs like Sox2 and Nanog form a complex regulatory network (Fig. 4g).
In conclusion, the interplay between the tTFs and eLR orchestrates early embryo development.
TimeTalk workflow can be updated to decipher cell-cell communication in blastoid
Blastoids are a valuable system for studying early embryo development in vitro40. Therefore, we hypothesized it would be helpful to study the cell-cell communication between different blastoid lineages to check blastoids’ fidelity in modeling naturally developed blastocysts.
However, some obstacles exist to employing TimeTalk for investigating cell-cell communication in blastoids. The presence of different cell lineages in blastoids and blastocysts requires the investigation of paracrine signaling between distinct cell types. Moreover, the available public data on blastoid research primarily comprise droplet-based single-cell RNA-seq datasets, which can be relatively sparse. This sparsity poses challenges when building transcriptional networks from the available data. These obstacles may make it difficult to apply the TimeTalk workflow directly to the study of blastoid cell-cell communication. Therefore, we update the TimeTalk workflow with the following procedures: (1) As the monocle341 package can reconstruct complex development trajectories from scRNA-seq datasets, we used monocle3 to replace monocle2 to order cells in each cell type. (2) Next, to study the correlation of ligand and receptor expression between two cell types, we interpolated pseudotime the time series by CellAlign42 to get equal sample points for two cell types. (3) We used Spearman’s correlation coefficient to get co-varying LR pairs between two cell types. (4) We perform zero-preserved imputation by the ALRA algorithm43. (5) The impute matrix was used to build the transcriptional network and perform master regulator analysis by RTN package44. (6) Performing the Granger causal test to calculate the Granger causality between the gene expression of master TFs from the receiver cell types and interaction score of co-varying LR pairs from the interpolated pseudotime time series. (7) The LR pairs that passed the Granger causal test were considered forward (LR regulates TF), backward (TF regulates LR), and feedback (LR and TF were mutually regulated) (Fig. 5a).
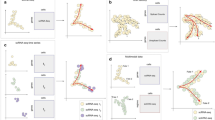
a The updated version of TimeTalk. b The re-analysis of the EPS-blastoid dataset. Trajectories were plotted in the UMAP embeddings. The color bar on the right side indicates the pseudotime. c The percentage of different cell types in naturally developed blastocysts and EPS-blastoids. d The Venn diagram shows the overlap of LR pairs in blastocyst and blastoid. There, EPI lineage as a sender cell and PE lineage as a receiver cell. e The KEGG analysis of genes consists of LR pairs mediated EPI-PE communication.
Next, we re-analyzed a public scRNA-seq data set containing EPS-blastoids and natural blastocyst45. Consistent with the original study, the EPS-blastoid captures the main lineage EPI, PE, and TE as blastocyst (Fig. 5b, Supplementary Fig. 18a–c). In addition, we found that the ICM-like lineage in EPS-blastoids also expressed 2-cell markers like Zscan4c, Zscan4d, and Zscan4f (Supplementary Fig. 18d, e). Therefore, we named these cells 2C-like cells. Besides, there are also some intermediate lineages (Supplementary Fig. 18f, g). As we annotated the cell types across different lineages, we used monocle3 to reconstruct trajectories and order cells in each lineage (Fig. 5c, Supplementary Data 3).
We are currently utilizing the updated version of TimeTalk to investigate cell-cell communication between the EPI-PE lineage in blastocysts and EPS-blastoids. As described in the previous paragraph, the interpolation and subsequent Granger causal test procedures in TimeTalk rely on three key parameters: window size of the interpolation (winsz), number of desired interpolated points (numPts), and the order of lags to include in the auxiliary regression (lag). We have set the default parameter values as winsz = 0.1, numPts = 200, and lags = 1, to identify 311 ligand-receptor pairs that mediate cell-cell communication between the epiblast and primitive endoderm in blastocyst cells. (Supplementary Data 3). TimeTalk is quite sensitive to the parameter winsz (Supplementary Fig. 19a), but relatively stable to numPts and lags parameters (Supplementary Fig. 19b, c). Moreover, it remains relatively robust when using the three critical parameters within certain ranges (Supplementary Fig. 19d). Through our analysis of TimeTalk results, we have discovered a feedback relationship between Fgf4-Fgfr2 and Gata6 with regards to cell-cell communication between the epiblast and primitive endoderm (p_LR_to_TF = 0.0184, p_TF_to_LR = 4.74 × 10-5, p-values were calculated by granger causal test Supplementary Fig. 19e). This result is consistent with previous research46, which demonstrated that Nanog in the epiblast upregulates the expression of Fgf4. The secreted Fgf4 from epiblast induced Fgf4-Fgfr2 interaction in primate endoderm cells to release Gata6 expression through ERK signaling. Gata6 then potentiates the upregulation of Fgfr2, and the feedback loop reinforces a primitive endoderm fate. Furthermore, the interpolation result can suggest that Nanog upregulates Fgf4 expression, but Fgf4 cannot regulate Nanog expression in the epiblast (SCC = 0.640, p_L_to_TF = 0.657, p_TF_to_L = 0.025, p-values were calculated by granger causal test, SCC means Spearman correlation coefficient, Supplementary Fig. 19f). Additionally, the interpolation result could also suggest the feedback relationship between Fgfr2 and Gata6 (SCC = 0.274, p_R_to_TF = 1.15×10-10, p_TF_to_LR = 9.19 × 10-5, p-values were calculated by granger causal test, Supplementary Fig 19g).
The Venn diagram shows that blastoid and blastocyst share common LR pairs identified by TimeTalk (Fig. 5d, Supplementary Data 3). These shared LR pairs include well-studied pairs Fgf4-Fgfr1 and Fgf4-Fgfr2. Moreover, the genes composed of LR pairs identified by TimeTalk enriched common signaling pathways involved in blastocyst formation. The notable examples are that both blastocysts and blastoids are enriched with “Signaling pathways regulating pluripotency of stem cells”, “TGF-beta signaling pathway”, “MAPK signaling pathway” and “Hippo signaling pathway” (Fig. 5e, Supplementary Data 3).
To the best of our knowledge, these results consistent with the previous results of blastoid’s fidelity in modeling cell-cell communication in balstocyst40, but from a different computation perspective.
Discussion
Early embryo development is a gradual and continuous process involving increasing cell heterogeneity. It is worth noting that definite cell types do not occur until blastocyst formation during this process. However, most computation toolkits for studying cell-cell communication, such as CellPhoneDB10 and CellChat11, were designed to investigate cell-cell communication in defined cell types, making it challenging to apply these tools to study early embryo development. To tackle these problems, it’s essential to create a fresh computational workflow. We have developed the TimeTalk workflow, which utilizes temporal information to infer cell-cell communication during early embryo development. TimeTalk uses trajectory analysis and the Granger causal test to examine ligand-receptor pairs with causal relationships to master transcription factors during development, thereby elucidating the potential causal relationship between ligand-receptor interaction and TF activity. To study cell-cell communication in blastoids, we introduce an interpolated pseudotime time series strategy to perform the Granger causal test of two unequal length time series to infer cell-cell communication by different types of cells, making the TimeTalk workflow applicable to paracrine signaling in various development processes. In summary, TimeTalk is suitable for studying developmental processes, particularly when scRNA-seq datasets at different time points are available.
It should be admitted that the regulation relationship between ligand-receptor interaction and transcription factors inferred by the TimeTalk should be confirmed by further wet experiments. Given the absence of high temporal resolution data, we utilized two models based on theoretical Granger causality to create simulation datasets. These datasets were then employed to examine how the interpolation strategy might impact the conclusion of Granger causality. (Supplementary Note 1). According to our simulation results, the Granger causality conclusion is affected by the choice of parameter winsz (Supplementary Figs. 20 and 21). Thus, as presented in this manuscript, we acknowledge the need for additional methods to validate the inferred potential regulatory relationship between ligand-receptor interactions and transcription factors. Regrettably, the scarcity of high temporal resolution datasets hinders our ability to address this issue. Despite this limitation, we have acknowledged that our causal analysis serves as a helpful reference, and we intend to improve our approach in the future with more accurate data. Furthermore, we advise users in the TimeTalk documentation to conduct additional data analysis and experiment with various parameter combinations to achieve more dependable results when utilizing TimeTalk for cell-cell communication analysis.
It is noteworthy that TimeTalk can be employed to investigate normal developmental processes and explore cell-cell communication during disease progression. For example, in the study of cancer development and progression, the accumulation of longitudinal scRNA-seq data in cancer research47,48 has made it possible to apply the TimeTalk framework to comprehend the cell-cell communication between cells within the tumor and its surrounding environment that promote cancer development and progression, ultimately aiding the development of effective cancer treatments.
It should be noted that the current version of the TimeTalk framework is designed for cell-cell communication mediated by ligand-receptor interactions. In contrast, other types of communication may not be accounted for. Extracellular vesicles (EVs) are an example of such communication. EVs, which are lipid bilayer structures that transport diverse biological cargo such as nucleic acids and proteins49,50, are secreted from various mammalian cell types and hypothesized to mediate long-range cell-cell communication51,52. EVs are believed to have an influential role in both normal physiological and pathological processes53, much like ligand-receptor interactions. However, previous EV profiling technologies have focused on the cargo of EVs rather than the molecules responsible for their secretion, making it difficult to decipher the function of EV-mediated cell-cell communication54,55. Recently, several studies attempted to develop single-cell profiling technology to detect EV secretion and link it to disease progression56,57. These technological advancements can help extend the TimeTalk framework to encompass EV-mediated cell-cell communication and provide insights into how it participates in development and disease.
In this study, the identified eLRs would be a valuable resource for better understanding early embryo development. Furthermore, the further study of eLRs would bring new knowledge of how different cells coordinate each other’s behavior to self-organize the embryo structure and give a new life. Moreover, the interplay between the eLRs and tTFs makes us block or enhance eLRs to manipulate tTFs activity or tune tTFs expression to regulate the downstream eLR activity to manipulate the cell-cell communication.
The process of co-evolution involves a heritable change in one entity that creates selective pressure for a change in another entity. Such entities can vary from nucleotides and amino acids to proteins, entire organisms, and potentially even ecosystems across the evolutionary time58. Co-evolution analysis was conducted on both eLR and non-eLR pairs, and the results indicate a similar distribution of co-evolution trends between the two groups (Supplementary Fig. 22). These results indicate that eLR and non-eLR are exposed to the same selection pressure.
This study assumed autocrine regulation of early embryo developmental stages during eLR screening. This hypothesis was reasonable for two reasons. First, before implantation, cells in the embryo are constrained by a glycoprotein shell called the zona pellucida1,59. Thus the embryo may be treated as a whole cell at each stage during pre-implantation. Second, previous studies illustrated that autocrine signaling is a substantial feature of the cell-to-cell communication network60,61. However, as crosstalk between the embryo and the maternal environment is essential for embryo development62, this hypothesis precludes our study from exploring maternal-embryo interactions during early embryo development. Nevertheless, the rapid growth of spatial transcriptomics applications in the maternal microenvironment60 and tissue engineering strategies63 allowing researchers to mimic maternal-embryo interactions will facilitate experiments exploring how eLRs mediate maternal-embryo crosstalk during early embryo development.
Research in Drosophila brain development recently proved that different neuron cell types were successively generated by the sequentially expressed temporal transcription factors (tTFs)64,65. This study illustrates that the interplay of eLRs and tTFs can be divided into different temporal windows. Moreover, the successive activation of tTFs and subsequent correspondent activation of cell-cell communication by eLRs indicates a wave-like mechanism to guide embryo development from a zygote (Fig. 6).
Blastoid is an ex-vivo reconstruction of blastocyst-like structure from stem cell lines to model blastocyst development40,66,67. Generating a blastoid close to the naturally developed blastocyst is an excellent way to model early embryo development68. In this study, we illustrate the fidelity of blastoids in modeling blastocysts from the cell-cell communication perspective. In addition, it should be noted that blastoid can be used to validate the function of identified eLRs.
Overall, the recognized eLR list provides valuable clues for the community to understand early embryo development and developmental diseases. Furthermore, TimeTalk would be a helpful tool for studying cell-cell communication in other developmental processes.
Methods
Ethical approval statement
After a thorough review of the sequencing studies utilized in the article, we can assure that the mouse experiments were carried out with appropriate ethical approval.
Early embryo scRNA-seq preprocessing and quality control
Publicly available pre-implantation (oocyte, zygote, early 2-cell, mid 2-cell, late 2-cell, 4-cell, 8-cell, 16-cell, early blastocyst, mid blastocyst, and late-blastocyst)12,13 scRNA-seq datasets were downloaded from the GEO database (see Availability of data and materials). These datasets were generated by the Smart-seq12 or Smart-seq269 (Oocyte Smart-seq, remains, Smart-seq2) experiment strategy and used reference genome mm9. In addition, we used the author-provided reads per kilobase per million mapped reads (RPKM) gene expression matrix for further analysis.
Custom R scripts were used for the pre-implantation scRNA-seq data to calculate the pseudo-bulk expression for an average single cell in each stage. Moreover, the PCC of pseudo-bulk and bulk RNA-seq was calculated using the function “cor” in the R stats package. Finally, PCA of pre-implantation scRNA-seq and bulk RNA-seq was performed using the function “prcomp” from the R stats package (parameters: center=TRUE, scale.=TRUE). To quantify the presence of batch effects, we conducted the kBET analysis using a χ2-based test14. The kBET analysis assessed whether the data points with different batches in the PCA space were well-mixed within random neighborhoods of a fixed size. The binary test results were averaged to generate an overall rejection rate, which is easily interpretable. A low rejection rate indicates that the batches are well-mixed and that the confounding effects can be neglected. The kBET analysis was performed by R package kBET (version: 0.99.6).
Custom R scripts generated the pseudo-bulk expression for each stage from the integrated datasets. Quantile normalization was performed with the function “normalize.quantiles” from R package preprocessCore (version 1.48.0). The ligand and receptor gene distribution analysis used the quantile normalized gene expression.
The merged expression matrix was input to Seurat (version 3.2.3)70 to perform t-SNE dimension reduction and draw marker expression by FeaturePlot.
Ligand and receptor gene expression analysis
Mouse ligand-receptor gene pairs (2034 pairs) were downloaded from CellTalkDB16 (http://tcm.zju.edu.cn/celltalkdb/download.php). The ligand and receptor genes were compared with the mm9 protein-coding genes by pseudo-bulk expression with a boxplot using the function “geom_boxplot” from ggplot2 (version 3.3.5).
We plotted each single-cell gene expression value across stages for a given ligand-receptor gene pair and fit the non-linear regression curve with the layer function “geom_path” in ggplot2 (parameters: method = “gam”, se = F)
Metric for cell-cell communication strength
Cell-cell communication strength was quantified by the interaction score proposed by ref. 17. The interaction score was defined as follows:
In which ei,j is the expression value of gene j in cell i, and nc is the number of cells of cell type c.
For single-cell autocrine, the interaction score was defined as:
TimeTalk workflow for identifying eLR
To identify the eLR from the curated early embryo scRNA-seq, we follow these steps: first, we order the cells according to their trajectories. Then, we identify co-varying eLR and temporal TF (tTFs). Finally, we obtain eLR from co-varying LR through Granger causality analysis with tTFs.
The monocle215 pipeline was used to construct the trajectory from curated scRNA-seq datasets. First, we used the “differentialGeneTest” function to get differentially expressed genes (cutoff: qval = 0.01) during early embryo development. Then, the differential expressed genes were used to perform DDRTree dimension reduction to reconstruct the development trajectory by the “reduceDimension” function. Finally, pseudotime was calculated by the “orderCells” function.
We arrange the cells by stage and pseudotime. Then, for a given LR, the ordered cells’ ligand gene and receptor gene expression values form a two time series. Then, we calculated the Pearson’s correlation coefficient (PCC) and used |PCC| = 0.1 as a cutoff to get co-varying LR. LR pairs of PCC >0.1 or PCC < −0.1 is the co-varying LR. The rationale of |PCC| = 0.1 is as follows. We first rank the positively correlated LR in ascending order by PCC, producing a concave curve. The start and endpoints of this curve are labeled as A and B, respectively. We then draw a straight line connecting AB, which is moved to obtain the tangency point C. Next, we connect AC and move the resulting straight line to obtain tangency point D. Below point D, and the curve roughly changes into a linear function. Thus, we select the vertical coordinate of point D as the cutoff value, which is 0.0987, for positive co-varying LR (Supplementary Fig. 7a). Using the same process, the cutoff for the negative co-varying LR is −0.0587 (Supplementary Fig. 7b). To establish the co-varying LR cutoff, we select the maximum value among the positive and negative co-varying LR cutoffs and round to 2 decimal places, resulting in a cutoff of 0.1.
We used the RTN package (version 2.10.1) to reconstruct the transcriptional regulatory network to get temporal TF. In brief, the log10(RPKM + 1) normalized scRNA-seq data and TF list from AnimalTFDB (version 3.0)71 to build the “tni” object from the RTN package to construct the transcriptional regulatory network. Then, the variable gene and their variance were input as a phenotype to perform master regulator analysis by function “tna.mra”. Finally, the output was considered as temporal TFs.
To reduce the computational cost, we clustered the identified tTFs by hierarchical clustering (the agglomeration method was “ward.D2”). After clustering tTFs, we calculate tTFs average expression as the activity of entire TF clusters. Then, we perform Granger causal test with the function “grangertest” from the R package lmtest (version 0.9-38). Ultimately, we test the causal relationship between tTFs activity and a given LR pairs interaction score. Because Granger causal test is an asymmetric test, we test for a given LR pair and a given cluster of tTFs whether there is a Granger-causality relationship from LR to TF or TF to LR.
As we have six clusters of tTFs, we choose the minimal p-value to represent the significance of causality from LR to tTFs or tTFs to LR. We used 0.01 as a cutoff to get candidate eLR from co-varying LR. Thus, we consider p_LRtoTF < 0.01 and p_TFtoLR < 0.01 co-varying LR as eLR.
Validation of eLRs with sets of essential genes, housekeeping genes, and ZGA genes
The essential gene set was downloaded from the DEG 10 database24 (http://origin.tubic.org/deg/public/index.php/index). The housekeeping gene set was downloaded from the HRT Atlas v1.0 database26 (http://www.housekeeping.unicamp.br/). The ZGA gene sets were obtained from Table S5 in ref. 18. The enrichment ratio of gene set A in a given gene set B was calculated with the following formula:
As shown in formula (3), the numerator indicates the number of elements that are common in both set A and set B, while the denominator represents the total number of elements in set B.
For example, the gene set enrichment ratio of essential genes in all eLR genes equals the number of essential genes among all eLRs divided by the number of genes composite of eLRs.
The p-value was calculated by the “phyper” function from R package stats (version 3.6.3). For the “phyper” function, we used lower.tail=TRUE to test whether the observed enrichment ratio was higher than the expected enrichment ratio, lower.tail=FALSE to test whether the observed enrichment ratio was lower than the expected enrichment ratio.
ZGA inhibition RNA-seq data analysis
ZGA inhibition RNA-seq data were obtained from a study by ref. 30.
The ZGA inhibition RNA-seq dataset included three conditions: early 2-cell, late 2-cell, and 2-cell treated with alpha amanitin (i.e., ZGA-inhibited condition). We treated each condition as a cell type and considered autocrine signaling within each condition.
Two rounds of hierarchical clustering identified the eLR pairs affected by ZGA inhibition. Euclidean distance and complete clustering methods were used for hierarchical clustering. The first step identified the variable eLR pairs, and the second step identified the eLR pairs affected by ZGA inhibition.
We define the delta value to measure the effect of ZGA inhibition on the cell-cell communication strength of LR pairs. The ZGA inhibition RNA-seq dataset included three conditions: early 2-cell, late 2-cell, and 2-cell treated with alpha amanitin (i.e., ZGA-inhibited condition). For a given LR pair with an interaction score IS, the delta value was defined as follows:
The formula can be reduced to:
We calculated the p-value in Supplementary Fig.13 using the “t.test” function from the R package stats (version 3.6.3). To test whether the delta value of non_eLR was lower than the delta value of eLR, we used alternative = “l” for the “t test” function.
Blastoid scRNA-seq data analysis
The scRNA-seq datasets for droplet-based EPS-generated blastoids and natural blastocysts were obtained from a study by ref. 45.
For the EPS-generated blastoid and natural blastocyst data, empty droplets were identified by the function “barcodeRanks” in the R package DropletUtils (version 1.6.1, parameters: lower = 100, fit.bounds = NULL, df = 20). After removing the empty droplets, the single-cell gene expression matrix was loaded into R by the function “CreateSeuratObject” in Seurat (parameters: min.features = 2000, min.cells = 0). Next, the natural blastocyst and EPS-generated blastoid data were integrated by CCA-based methodology in Seurat70. First, the anchors for integration were identified by the function “FindIntegrationAnchors” (parameters: dims = 1:20, k.anchor = 5, k.filter = 30). In the next step, the identified anchors were input into the function “IntegrateData” (parameters: dims = 1:20) to obtain integrated data. Finally, the integrated data were processed using the function “ScaleData” (parameters: model.use = “linear”, do.scale = TRUE, do.center = TRUE”, scale.max = 10, block.size = 1000, min.cells.to.block = 3000), “RunPCA” (parameters: npcs = 30), “RunUMAP” (parameters: reduction = “pca”, dims = 1:30, umap.method = “umap-learn”), “FindNeighbors” (parameters: dims = 1:20, reduction = “pca”), and “FindClusters” (parameters: resolution = 0.2). Cell lineages were assigned based on the following markers according to the original research of ref. 45:
Trophectoderm (TE): Cdx2, Krt8, Krt18, Ascl2, Tacstd2;
Inner cell mass or epiblast (ICM/EPI): Pou5f1, Nanog, Sox2, Esrrb, Sox15;
Primitive endoderm (PE): Gata4, Gata6, Sox17, Pdgfra, Col4a
2C like: Zscan4f, Zscan4c, Zscan4d.
We used Monocle3 to reconstruct trajectories. The integrated UMAP embedding was used to learn trajectory using the function “learn_graph” from the Monocle3 package. Finally, we choose the 2C_like cells as the root cell to calculate psedotime along the trajectory.
The extended TimeTalk workflow
The extended TimeTalk workflow including following steps: Firstly, we order cells based on the reconstructed trajectory. Next, we identify the co-varying LR from the two types of cells. Then, we identify the master regulators for each cell type. Finally, we obtain the active LR from the co-varying LR with the master regulators of a particular cell type using Granger’s causality.
As monocle341 can reconstruct complex development trajectories from scRNA-seq datasets, we used monocle3 to replace monocle2 to order cells in each cell type.
For a given LR, consider sender cells A expressed ligand(L) and receiver cell B expressed receptor (R). When examining the expression data in ordered cells of both cell type A and cell type B, two time series of unequal (or equal, if A and B are the same types of cell) length were generated. Consequently, it is necessary to calculate the correlation between these two time series. To obtain equal sample points for both cell types, we used the “interWeight” function from the cellAlign42 package to interpolate the expression data along one trajectory (window size of interpolation: winSz = 0.1, number of desired interpolated points, numPts = 200). Using this strategy, we can calculate Spearman’s correlation coefficient of interpolated ligand signals calculated from ligand gene expression in sender cells and receptor signals calculated from the receptor gene expression in receiver cells. We used SCC = 0.2 as a cutoff to identify co-varying LR.
We used the function RunALRA from the R package Seurat to perform zero-preserved imputation by the ALRA algorithm43. Each cell type’s imputed matrix was used to build the transcriptional network and perform master regulator analysis by RTN package44. The variable gene along the trajectories identified by the function graph_test from the Monocle3 package was used as a phenotype to perform master regulator analysis.
We perform the Granger test to calculate the Granger causality between the gene expression of master TFs from the receiver cell types and the interaction score of co-varying LR pairs from the interpolated pseudotime time series. Meanwhile, we calculate Spearman’s correlation coefficient to measure the correlation between the activity of co-varying LR and master TFs. We used 0.01 as a cutoff to test whether the given LR and a master TF have Granger causality. Thus, for a given LR, it will have many master TFs passed the Granger test and have many different correlation values. We choose the maximum (noted as SCC_ens) as the final correlation value to measure the correlation of LR with the TFs activity. We used SCC_ens > 0.8 or SCC_ens < −0.8 to get active LR.
Co-evolution analysis
We used the HomoloGene database (https://www.ncbi.nlm.nih.gov/homologene) to retrieve homologous ligand and receptor genes in four mammalian species Mus Musculus (mouse), Rattus norvegicus (rat), Bos Taurus (cattle), and Homo sapiens (human). Through this approach, 1119 homologous genes were obtained, which included 922 ligand-receptor pairs, of which 208 were eLR pairs, and 714 were non-eLR pairs. The ratio of nonsynonymous and synonymous substitution rates (Ka/Ks) for each gene was calculated using the following steps: amino acid sequence alignments were performed using the R package msa (version 1.28.0)72 and ClusterW algorithm73, followed by codon alignment using the codon_aln function by PAL2NAL algorithm74 in the R package orthologr (version 0.4.0)75. The KaKs ratio was then calculated using the dnastring2kaks function in the R package MSA2dist (version 1.0.0). Finally, co-evolution trends were evaluated by computing the Pearson correlation coefficient (PCC) of the Ka/Ks ratio for each ligand-receptor pair in each pairwise species comparison.
Statistics and reproducibility
All statistical analyses were performed using R (version 3.6.3). Heatmaps were generated by the R package ComplexHeatmap (version 2.9.4)76. Graphs of statistical results were generated by the R package ggplot2 (version 3.3.5). GO analysis and KEGG analysis were performed by the R package clusterProflier (version 3.14.3)77.
A collection of publicly available pre-implantation single-cell RNA-seq datasets were used to analyze the different stages of early embryo development. These stages include the oocyte (3 cells), zygote (4 cells), early 2-cell (8 cells), mid 2-cell (12 cells), late 2-cell (10 cells), 4-cell (14 cells), 8-cell (28 cells), 16-cell (50 cells), early blastocyst (43 cells), mid blastocyst (60 cells), and late blastocyst (30 cells). In total, there were 262 cells analyzed to ensure that all stages were represented in the scRNA-seq datasets. In Supplementary Note 1, we tested the impact of interpolation strategies on Granger causality conclusions by using two different time series models with theoretical causality. To determine the false positive ratio of Granger causality that resulted from interpolation, we conducted ten rounds of simulations for each model. In each round of simulation, we generated simulation data and performed the Granger test on the interpolated data 1000 times. Since each model has a theoretical Granger causality, the false positive ratio was calculated for each round.
In order to make sure that our results can be replicated, we have included the source data for the critical analysis of our manuscript as Supplementary Data. Additionally, we have uploaded our code and extensive data to Figshare (https://figshare.com/projects/TimeTalk_CB_manuscript_code_data/174498) to make it easier for others to reproduce our analyses.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The publicly available data on early embryo development used in this paper were obtained from the Gene Expression Omnibus (GEO) database (https://www.ncbi.nlm.nih.gov/geo/). The mouse oocyte scRNA-seq data were obtained from a study by ref. 12 with accession number GSE38495. The mouse scRNA-seq data covering the zygote stage to the late blastocyst stage were obtained from a study by ref. 13 with accession number GSE45719. The mouse bulk RNA-seq was obtained from studies by Wu et al. with accession numbers GSE66582. The mouse ZGA inhibition RNA-seq data were obtained from a study by ref. 30 with accession number GSE71434. The EPS-generated blastoid scRNA-seq data and natural blastocyst scRNA-seq data were obtained from a study by ref. 45 with accession number GSE135701. The processed sequencing data used in this article has been deposited in the Figshare repository (https://doi.org/10.6084/m9.figshare.23850324.v7)78. Source data underlying Figs. 2f, g, and 3a–d, and Supplementary Fig. 13 are provided in Supplementary Data 1. Source data for Fig. 4a–f and Supplementary Fig. 16a–c are provided in Supplementary Data 2. Source data for Fig. 5c–e are provided in Supplementary Data 3.
Code availability
All custom scripts required to reproduce all results reported in this manuscript have been deposited in Figshare repository (https://doi.org/10.6084/m9.figshare.23895780.v6)79. The source code for TimeTalk R package can be accessed on Github (https://github.com/ChengLiLab/TimeTalk)80. Furthermore, we have also deposited the TimeTalk R package in Zenodo (https://doi.org/10.5281/zenodo.8271645)81.
References
Zhu, M. & Zernicka-Goetz, M. Principles of self-organization of the mammalian embryo. Cell 183, 1467–1478 (2020).
Schulz, K. N. & Harrison, M. M. Mechanisms regulating zygotic genome activation. Nat. Rev. Genet. 20, 221–234 (2019).
Wang, H. & Dey, S. K. Roadmap to embryo implantation: clues from mouse models. Nat. Rev. Genet. 7, 185–199 (2006).
Rossant, J. & Tam, P. P. Blastocyst lineage formation, early embryonic asymmetries and axis patterning in the mouse. Development 136, 701–713 (2009).
Zernicka-Goetz, M., Morris, S. A. & Bruce, A. W. Making a firm decision: multifaceted regulation of cell fate in the early mouse embryo. Nat. Rev. Genet. 10, 467–477 (2009).
Armingol, E., Officer, A., Harismendy, O. & Lewis, N. E. Deciphering cell-cell interactions and communication from gene expression. Nat. Rev. Genet. 22, 71–88 (2021).
Do, D. V. et al. A genetic and developmental pathway from STAT3 to the OCT4-NANOG circuit is essential for maintenance of ICM lineages in vivo. Genes Dev. 27, 1378–1390 (2013).
Chazaud, C. & Yamanaka, Y. Lineage specification in the mouse preimplantation embryo. Development 143, 1063–1074 (2016).
Molotkov, A., Mazot, P., Brewer, J. R., Cinalli, R. M. & Soriano, P. Distinct requirements for FGFR1 and FGFR2 in primitive endoderm development and exit from pluripotency. Dev. Cell 41, 511–526.e4 (2017).
Vento-Tormo, R. et al. Single-cell reconstruction of the early maternal-fetal interface in humans. Nature 563, 347–353 (2018).
Jin, S. et al. Inference and analysis of cell-cell communication using CellChat. Nat. Commun. 12, 1088 (2021).
Ramskold, D. et al. Full-length mRNA-Seq from single-cell levels of RNA and individual circulating tumor cells. Nat. Biotechnol. 30, 777–782 (2012).
Deng, Q., Ramskold, D., Reinius, B. & Sandberg, R. Single-cell RNA-seq reveals dynamic, random monoallelic gene expression in mammalian cells. Science 343, 193–196 (2014).
Büttner, M., Miao, Z., Wolf, F. A., Teichmann, S. A. & Theis, F. J. A test metric for assessing single-cell RNA-seq batch correction. Nat. Methods 16, 43–49 (2019).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
Shao, X. et al. CellTalkDB: a manually curated database of ligand-receptor interactions in humans and mice. Brief. Bioinform. 22, bbaa269 (2021).
Kumar, M. P. et al. Analysis of single-cell RNA-Seq identifies cell-cell communication associated with tumor characteristics. Cell Rep. 25, 1458–1468 e4 (2018).
Lu, F. et al. Establishing chromatin regulatory landscape during mouse preimplantation development. Cell 165, 1375–1388 (2016).
Goolam, M. et al. Heterogeneity in Oct4 and Sox2 targets biases cell fate in 4-Cell mouse embryos. Cell 165, 61–74 (2016).
Graham, S. J. et al. BMP signalling regulates the pre-implantation development of extra-embryonic cell lineages in the mouse embryo. Nat. Commun. 5, 5667 (2014).
Rancati, G., Moffat, J., Typas, A. & Pavelka, N. Emerging and evolving concepts in gene essentiality. Nat. Rev. Genet. 19, 34–49 (2018).
Chen, H. et al. New insights on human essential genes based on integrated analysis and the construction of the HEGIAP web-based platform. Brief. Bioinform. 21, 1397–1410 (2020).
Bartha, I., di Iulio, J., Venter, J. C. & Telenti, A. Human gene essentiality. Nat. Rev. Genet. 19, 51–62 (2018).
Luo, H., Lin, Y., Gao, F., Zhang, C. T. & Zhang, R. DEG 10, an update of the database of essential genes that includes both protein-coding genes and noncoding genomic elements. Nucleic Acids Res. 42, D574–D580 (2014).
Eisenberg, E. & Levanon, E. Y. Human housekeeping genes, revisited. Trends Genet. 29, 569–574 (2013).
Hounkpe, B. W., Chenou, F., de Lima, F. & De Paula, E. V. HRT Atlas v1.0 database: redefining human and mouse housekeeping genes and candidate reference transcripts by mining massive RNA-seq datasets. Nucleic Acids Res. 49, D947–D955 (2021).
Lee, M. T., Bonneau, A. R. & Giraldez, A. J. Zygotic genome activation during the maternal-to-zygotic transition. Annu Rev. Cell Dev. Biol. 30, 581–613 (2014).
Jukam, D., Shariati, S. A. M. & Skotheim, J. M. Zygotic genome activation in vertebrates. Dev. Cell 42, 316–332 (2017).
Abe, K. I. et al. Minor zygotic gene activation is essential for mouse preimplantation development. Proc. Natl. Acad. Sci. USA 115, E6780–E6788 (2018).
Zhang, B. et al. Allelic reprogramming of the histone modification H3K4me3 in early mammalian development. Nature 537, 553–557 (2016).
Frum, T. & Ralston, A. Cell signaling and transcription factors regulating cell fate during formation of the mouse blastocyst. Trends Genet. 31, 402–410 (2015).
Pokrass, M. J. et al. Cell-cycle-dependent ERK signaling dynamics direct fate specification in the mammalian preimplantation embryo. Dev. Cell 55, 328–340.e5 (2020).
Nishioka, N. et al. The Hippo signaling pathway components Lats and Yap pattern Tead4 activity to distinguish mouse trophectoderm from inner cell mass. Dev. Cell 16, 398–410 (2009).
Attisano, L. & Wrana, J. L. Signal integration in TGF-β, WNT, and Hippo pathways. F1000Prime Rep. 5, 17 (2013).
Wang, Y. et al. Comprehensive molecular characterization of the hippo signaling pathway in cancer. Cell Rep. 25, 1304–1317.e5 (2018).
Dietrich, B., Haider, S., Meinhardt, G., Pollheimer, J. & Knöfler, M. WNT and NOTCH signaling in human trophoblast development and differentiation. Cell Mol. Life Sci. 79, 292 (2022).
Yamamoto, S. et al. Cthrc1 selectively activates the planar cell polarity pathway of Wnt signaling by stabilizing the Wnt-receptor complex. Dev. Cell 15, 23–36 (2008).
Li, Y. et al. CTHRC1 promotes growth, migration and invasion of trophoblasts via reciprocal Wnt/β-catenin regulation. J. Cell Commun. Signal. 16, 63–74 (2022).
Paquette, S. M., Leinonen, K. & Longabaugh, W. J. Bio tapestry now provides a web application and improved drawing and layout tools. F1000Res 5, 39 (2016).
Rivron, N. C. et al. Blastocyst-like structures generated solely from stem cells. Nature 557, 106–111 (2018).
Cao, J. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature 566, 496–502 (2019).
Alpert, A., Moore, L. S., Dubovik, T. & Shen-Orr, S. S. Alignment of single-cell trajectories to compare cellular expression dynamics. Nat. Methods 15, 267–270 (2018).
Linderman, G. C. et al. Zero-preserving imputation of single-cell RNA-seq data. Nat. Commun. 13, 192 (2022).
Fletcher, M. N. et al. Master regulators of FGFR2 signalling and breast cancer risk. Nat. Commun. 4, 2464 (2013).
Li, R. et al. Generation of blastocyst-like structures from mouse embryonic and adult cell cultures. Cell 179, 687–702.e18 (2019).
Schrode, N., Saiz, N., Di Talia, S. & Hadjantonakis, A. K. GATA6 levels modulate primitive endoderm cell fate choice and timing in the mouse blastocyst. Dev. Cell 29, 454–467 (2014).
Zhang, K. et al. Longitudinal single-cell RNA-seq analysis reveals stress-promoted chemoresistance in metastatic ovarian cancer. Sci. Adv. 8, eabm1831 (2022).
Davidson, S. et al. Single-Cell RNA sequencing reveals a dynamic stromal niche that supports tumor growth. Cell Rep. 31, 107628 (2020).
Valadi, H. et al. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat. Cell Biol. 9, 654–659 (2007).
Balaj, L. et al. Tumour microvesicles contain retrotransposon elements and amplified oncogene sequences. Nat. Commun. 2, 180 (2011).
Raposo, G. & Stoorvogel, W. Extracellular vesicles: exosomes, microvesicles, and friends. J. Cell Biol. 200, 373–383 (2013).
Théry, C., Zitvogel, L. & Amigorena, S. Exosomes: composition, biogenesis and function. Nat. Rev. Immunol. 2, 569–579 (2002).
Dickhout, A. & Koenen, R. R. Extracellular vesicles as biomarkers in cardiovascular disease; chances and risks. Front. Cardiovasc. Med. 5, 113 (2018).
Kalluri, R. The biology and function of exosomes in cancer. J. Clin. Investig. 126, 1208–1215 (2016).
Hessvik, N. P. & Llorente, A. Current knowledge on exosome biogenesis and release. Cell Mol. Life Sci. 75, 193–208 (2018).
Fathi, M. et al. Identifying signatures of EV secretion in metastatic breast cancer through functional single-cell profiling. iScience 26, 106482 (2023).
Schoger, E. et al. Single-cell transcriptomics reveal extracellular vesicles secretion with a cardiomyocyte proteostasis signature during pathological remodeling. Commun. Biol. 6, 79 (2023).
Fraser, H. B., Hirsh, A. E., Wall, D. P. & Eisen, M. B. Coevolution of gene expression among interacting proteins. Proc. Natl. Acad. Sci. USA 101, 9033–9038 (2004).
Plusa, B. & Piliszek, A. Common principles of early mammalian embryo self-organisation. Development 147, dev183079 (2020).
Garcia-Alonso, L. et al. Mapping the temporal and spatial dynamics of the human endometrium in vivo and in vitro. Nat. Genet. 53, 1698–1711 (2021).
Hou, R., Denisenko, E., Ong, H. T., Ramilowski, J. A. & Forrest, A. R. R. Predicting cell-to-cell communication networks using NATMI. Nat. Commun. 11, 5011 (2020).
Paria, B. C., Reese, J., Das, S. K. & Dey, S. K. Deciphering the cross-talk of implantation: advances and challenges. Science 296, 2185–2188 (2002).
Govindasamy, N. et al. 3D biomimetic platform reveals the first interactions of the embryo and the maternal blood vessels. Dev. Cell 56, 3276–3287.e8 (2021).
Holguera, I. & Desplan, C. Neuronal specification in space and time. Science 362, 176–180 (2018).
Konstantinides, N. et al. A complete temporal transcription factor series in the fly visual system. Nature 604, 316–322 (2022).
Luijkx, D., Shankar, V., van Blitterswijk, C., Giselbrecht, S. & Vrij, E. From mice to men: generation of human blastocyst-like structures in vitro. Front. Cell Dev. Biol. 10, 838356 (2022).
Zhai, J., Xiao, Z., Wang, Y. & Wang, H. Human embryonic development: from peri-implantation to gastrulation. Trends Cell Biol. 32, 18–29 (2022).
Posfai, E., Lanner, F., Mulas, C. & Leitch, H. G. All models are wrong, but some are useful: establishing standards for stem cell-based embryo models. Stem Cell Rep. 16, 1117–1141 (2021).
Picelli, S. et al. Smart-seq2 for sensitive full-length transcriptome profiling in single cells. Nat. Methods 10, 1096–1098 (2013).
Stuart, T. et al. Comprehensive integration of single-. Cell Data. Cell 177, 1888–1902.e21 (2019).
Hu, H. et al. AnimalTFDB 3.0: a comprehensive resource for annotation and prediction of animal transcription factors. Nucleic Acids Res. 47, D33–D38 (2019).
Bodenhofer, U., Bonatesta, E., Horejš-Kainrath, C. & Hochreiter, S. msa: an R package for multiple sequence alignment. Bioinformatics 31, 3997–3999 (2015).
Thompson, J. D., Higgins, D. G. & Gibson, T. J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673–4680 (1994).
Suyama, M., Torrents, D. & Bork, P. PAL2NAL: robust conversion of protein sequence alignments into the corresponding codon alignments. Nucleic Acids Res. 34, W609–W612 (2006).
Drost, H. G., Gabel, A., Grosse, I. & Quint, M. Evidence for active maintenance of phylotranscriptomic hourglass patterns in animal and plant embryogenesis. Mol. Biol. Evol. 32, 1221–1231 (2015).
Gu, Z., Eils, R. & Schlesner, M. Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32, 2847–2849 (2016).
Yu, G., Wang, L. G., Han, Y. & He, Q. Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Wang, L. The dataset needed to reproduce the results in the TimeTalk manuscript. figshare https://doi.org/10.6084/m9.figshare.23850324.v7 (2023).
Wang, L. The necessary code to generate analysis results for the TimeTalk manuscript. figshare https://doi.org/10.6084/m9.figshare.23895780.v6 (2023).
ChengLiLab. ChengLiLab/TimeTalk: v0.0.310316 (v0.0.310316). GitHub https://github.com/ChengLiLab/TimeTalk (2023).
ChengLiLab. ChengLiLab/TimeTalk: v0.0.310316 (v0.0.310316). Zenodo https://doi.org/10.5281/zenodo.8271645 (2023).
Acknowledgements
We are grateful to the researchers who produced the publicly available data used in this study. In addition, we are grateful to Prof. Jin Zhang at Zhejiang University, Prof. Dahai Zhu at Peking Union Medical College, Ming Yang of Prof. Peng Du’lab at Peking University,Yang Ding of Prof. Xiaochen Bo’s lab at Institute of Health Service and Transfusion Medicine, Molong Qu and Liang Xiong at Prof. Hongkui Dengs’ lab at Peking University,Yuting Liu, Zihan Xu, Yulin Lyu, Yaqi Wang and Qing Fang of Prof. Cheng Li’s Lab at Peking University for providing sound advice regarding this research. This work was supported by the National Natural Science Foundation of China (32288102 and 32025006 to C.L., 62173338 to H.C., 61873276 to X.B.) and the National Key Research and Development Program of China (2021YFA1100300 to C.L.), and the Beijing Nova Program of Science and Technology (20220484198 to H.C.). Part of the data analysis was performed on the High-Performance Computing Platform of the Center for Life Sciences, Peking University.
Author information
Authors and Affiliations
Contributions
L.W., H.C., H.L., X.B., and C.L. conceived and designed the study. L.W. and S.M. performed the background investigation. L.W. performed all the bioinformatics analyses. L.W. and Y.Z. created the scientific illustrations. L.W., Y.Z., Y.S., S.M., H.L. and H.C. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Sourav Chowdhury, Sagnik Sen and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors Debarka Sengupta and George Inglis. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, L., Zheng, Y., Sun, Y. et al. TimeTalk uses single-cell RNA-seq datasets to decipher cell-cell communication during early embryo development. Commun Biol 6, 901 (2023). https://doi.org/10.1038/s42003-023-05283-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-023-05283-2
- Springer Nature Limited
This article is cited by
-
The diversification of methods for studying cell–cell interactions and communication
Nature Reviews Genetics (2024)