Abstract
Transposable elements (TEs) are a major source of genetic polymorphisms and play a role in chromatin architecture, gene regulatory networks, and genomic evolution. However, their functional role in pigs and contributions to complex traits are largely unknown. We created a catalog of TEs (n = 3,087,929) in pigs and found that young SINEs were predominantly silenced by histone modifications, DNA methylation, and decreased accessibility. However, some transcripts from active young SINEs showed high tissue-specificity, as confirmed by analyzing 3570 RNA-seq samples. We also detected 211,067 dimorphic SINEs in 374 individuals, including 340 population-specific ones associated with local adaptation. Mapping these dimorphic SINEs to genome-wide associations of 97 complex traits in pigs, we found 54 candidate genes (e.g., ANK2 and VRTN) that might be mediated by TEs. Our findings highlight the important roles of young SINEs and provide a supplement for genotype-to-phenotype associations and modern breeding in pigs.
Similar content being viewed by others
Introduction
TEs and common repeats are ubiquitous sequences that can copy and insert themselves throughout the eukaryotic and prokaryotic genomes1,2,3,4,5,6,7. The movement of TEs is often accompanied by an increase in their abundance, comprising a large fraction of genomic sequences8,9. According to the mechanism of transposition, TEs can be generally classified into (1) RNA-mediated class I elements (retrotransposons), including long terminal repeats (LTRs), long interspersed nuclear elements (LINEs), and short interspersed nuclear elements (SINEs); and (2) RNA-independent class II elements (DNA transposons)10. TE classes could be further divided into distinct families or subfamilies based on their age (active period) and DNA sequence characteristics.
At the predominant view of the 1960s–1990s, TEs were described as selfish or junk DNA11. Thanks to the availability of whole-genome sequence of various species and the ongoing development of bioinformatics tools12,13,14,15, our knowledge of TEs has progressed at a fast pace. TEs are known to play an essential role in shaping genomic sequences and contributing to the diversity in genome size and chromosome structure1,16,17. Most TEs, in fact, are fixed, inactive, and not randomly distributed in the genome18,19. However, several TE families are still actively transposing and serving as a major source of genetic polymorphisms between individuals, such as the Alu, L1, and SVA TE families in the human genome20.
It is evident in many species (e.g., humans and rice) that the impacts of active TEs on genome evolution are wide-ranging, including admixture, adaptation, footprints of selection, and population structure21,22,23,24. For example, the polymorphic TEs detected in the 1000 Genomes Project, consisting of 16,192 loci in 2504 individuals across 26 human populations, successfully recapitulated human evolution and captured the signal for positive selection on recent human TE insertions20,25,26.
In addition to their direct influence on DNA sequence, there is also emerging evidence that TEs have important functional contributions to gene regulatory networks and epigenome variation. For instance, TEs can directly affect gene transcriptional structure by provoking various forms of alternative splicing, including exonization, exon skipping, and intron retention (3′ and 5′), to generate novel protein-coding sequences or premature ends27,28,29,30. TEs can disrupt the existing cis-regulatory elements, such as promoters, enhancers, and insulators, or provide novel ones31,32,33,34. They can also serve as a rich source of non-coding RNAs, including lncRNAs, circRNA, small RNAs, and microRNA targets35,36,37,38. Moreover, the silencing of TEs has a close connection with epigenetic regulatory mechanisms, such as DNA methylation, piRNA, histone modifications, and RNA interference18,39,40,41,42. Importantly, it has been reported that the complex interactions between TEs and epigenetic elements could allow for rapid phenotypic adaptation to environmental changes40,42,43.
Pig (Sus scrofa), one of the earliest domesticated animals, is estimated to have been domesticated approximately 10,000 years ago in Asia and Europe independently44. It serves as an indispensable source of animal protein and an important biomedical model for humans45,46. Currently, a total of 22 pig assemblies are publicly available on NCBI47,48,49,50,51, accompanied by the availability of massive high-throughput whole-genome sequences. These provide researchers with ideal materials to advance the current development of genomic research in pigs. However, the study of TEs in the pig genome is still in its infancy. A few previous studies have mainly focused on their diversity and distribution47,48,49,50,51, yet the functional and evolutionary importance of TEs in pigs has largely been overlooked. In our recent study52, we identified novel introgressions in Eurasian boars from Asian and European pig populations using SINE (PRE-1 subfamily) polymorphisms, suggesting that a portion of TEs are still active in the current pig genome. However, these studies are far from sufficient to comprehensively understand the important roles of TEs in pigs.
In this study, we created a comprehensive and high-quality atlas of TEs so far in pigs and classified SINE families into four categories based on their ages using the newly built pipelines. We then systematically explored the genomic bias of these SINE categories by combining large-scale multi-omics data from 21 tissues, including three-dimensional chromatin architecture, chromatin accessibility, histone modifications, transcription factor binding sites (TFBS), and DNA methylation. We estimated the contribution of active SINEs to tissue-specific gene expression by cross-examining 3570 published RNA-seq samples from 52 tissues and 27 cell types. Furthermore, we created an atlas of SINEs using 374 whole-genome sequence data to study the roles of young SINEs in pig population admixture and local adaptation. The TE-mediated adaptation has been found in functional regions, such as the almost fixed dimorphic SINEs observed in laboratory-inbred Bama Xiang pigs at the upstream region of the LEP gene. Finally, by mapping these dimorphic SINEs to 4072 loci associated with 97 complex traits in pigs, we propose 54 candidate genes that might regulate complex traits through TEs.
Results
Composition of young SINE families in the pig genome
To thoroughly detect TEs, we developed the Pig TE Detection and Classification (PigTEDC) pipeline (Fig. 1a) and applied it to the pig genome (Sus scrofa 11.1). This pipeline used a combination of similarity-, structure-, and de novo-based methods. We also classified all potential pig TEs into classes/superfamilies and families and derived their consensus sequences based on existing TE repositories (RepBase update and Dfam 2.0 databases).
a A schematic of the Pig Transposable Element Detection and Classification (PigTEDC) pipeline. It is composed of three TE detection approaches, which use similarity-, structure-, and de novo-based algorithms. b The proportion of TEs from different superfamilies in the count. c The proportion of TEs from different superfamilies in length. d Classification of pig TE superfamilies and families (≥3000 copies in each family). e Sequence divergence distribution for TE superfamilies (upper panel) and families (bottom panel) in the pig genome. Sequence divergence distributions are plotted in bins of 0.01 increments. f Phylogenetic tree and sequence divergence distribution for SINE families in the pig genome. On the right panel, the x-axis represents the divergence, and the y-axis represents the counts of the SINE families. g Boxplots display the proportion of genomic SVs formed by different SINE families. h Boxplots display the proportion of genomic SVs formed by different SINE subfamilies. See Supplementary Data 1 for their definitions. i Classification of pig SINE families based on their ages. The line inside each boxplot represents the median.
Excluding nested TEs, we found 3,087,929 TEs occupying 37.9% (947 MB) of the pig genome. Two-thirds of TE copies (insert in the genome) were assigned to a specific family, with retrotransposons being the most common type (~90%). Similar to previous studies47,48,49,50,51, LTR (9.25%), LINE (27.57%), and SINE (54.95%) were the most common retrotransposons, whereas DNA transposons only accounted for 8.12% of TEs (Fig. 1b). SINE being the most abundant in count but covering less genome size than LINE (Fig. 1c).
Out of 532 TE families, 65 (with over 3000 TE copies each) make up 84.6% of all classified TEs (Fig. 1d). These include PRE1f in SINE/tRNA (170,511 copies), MIR in SINE/MIR (45,927 copies), L1B-SSc in LINE/L1 (35,819 copies), and MLT1D in LTR/ERVL-MaLR (7866 copies). The stacking plots in Fig. 1e show the divergence distribution for superfamilies or families. Our analysis of pig TEs revealed two bursts at 10% and 30%, estimated to have occurred 20 and 60 million years ago, which is similar to the divergence distribution of TEs in the human genome53. Obviously, most TE families amplified around 70–50 Mya (divergence at 30 ± 5%). This was during the Paleocene Epoch (65–54 Mya) which created new ecological niches for surviving mammals, birds, reptiles, and marine animals54. The most recent burst of TEs was mainly related to the SINE/tRNA, LINE/L1, and LTR/ERV1 families. Among these, SINE/tRNA remains the most active in the modern pig genome55,56,57. Further exploring the ages of highly homologous subfamilies in SINE classes (Fig. 1f with an average divergence of 4%, labeled in purple), we found that 3 out of 26 SINE families (PRE1-SS, PRE0-SS, and PRE1a) were recently most prolific and shown to be polymorphic within pig breeds in a previous study52, and thus considered as young SINE families.
We next analyzed young SINE families (PRE1-SS, PRE0-SS, and PRE1a) to classify them into subfamilies with high resolution. We examined all the full-length young SINEs from 14 publicly available pig genomes, identified 978,506 non-redundant young SINEs, and created their consensus sequence through multiple sequence alignment (Supplementary Figs. 1, 2). Subsequently, using a minimum spanning tree analysis, we recategorized the 90 subfamilies of SINE into 17 large (size >10,000), 12 medium (size ≥3000 and ≤10,000), and 61 small (size <3000) subfamilies (Supplementary Figs. 3, 4, Supplementary Data 1).
When comparing the locations of young SINEs with medium-length (range from 200 to 300 bp) structure variations (SVs) in 14 assemblies to the pig reference, we found that most polymorphic SINEs belonged to the PRE1-SS, PRE1a, and PRE0-SS families, accounting for an average of 90.75% of the medium-length SVs (Fig. 1g). Especially for the L13 subfamily accounted for an average of 36.15% of the medium-length SVs (Fig. 1h). Our findings indicate that only a certain group of recently active SINE subfamilies played a major role in causing SVs (about 250 bp long) in various pig breeds during recent evolution. To simplify our analysis, we classified all SINEs into four categories: youngest (L13 subfamilies), younger (SINE families other than L13 subfamilies and non-L13), older (non-young PRE families), and oldest (non-PRE families; ancient families) (Fig. 1i, Supplementary Fig. 5).
Widespread roles of young SINEs in gene regulatory networks
Previous studies have proposed that TEs may be co-opted into regulatory sequences of genes through diverse epigenetic mechanisms58,59,60,61. To test this, we examined how SINE subfamilies affect genome features such as 3D chromatin architecture62, chromatin accessibility, histone modifications, TFBS, and DNA methylation after only mapping unique reads (Fig. 2a).
a The five types of genomic features used in this study included 3D chromatin architecture, chromatin accessibility, histone modifications, DNA methylation, and TFBS. b The distribution of SINE families between 3D chromatin architectures (Compartments A vs. B) and near topologically associating domains is examined. c The read density distributions of chromatin accessibility and histone modifications near transcripts were analyzed across four different SINE groups. d Boxplots display the enrichment of four SINE groups in 15 distinct chromatin states across 14 tissues. The line inside each boxplot represents the median.
We observed a highly enrichment of all SINEs in the A compartments (active), while there was a depletion in the B compartments (inactive) (Wilcoxon test, P-values < 10−16, Fig. 2b, Supplementary Fig. 6). After dividing A/B compartments into topologically associating domains, it was observed that CTCF binding sites were enriched in the boundary regions of these domains. SINEs also exhibited a similar but slightly weaker trend of enrichment, while young SINEs showed higher but more variable enrichment compared to old SINEs (Fig. 2b, Supplementary Fig. 7). We next explored the distribution of SINE families on the chromatin accessibility and nucleosome positioning near transcripts using the published chip-seq (14 tissues) and MNase-seq (five tissues) datasets, respectively63,64. TE enrichment in chromatin showed age-specific patterns, with youngest and younger SINE families depleted from open chromatin but enriched near the nucleosome (Fig. 2c). Older SINE families were relatively highly enriched for open chromatin, especially in the stomach, adipose, and cerebellum. We further studied the relationship of SINE families with four active epigenetic marks (H3K4me1—primed enhancers, H3K4me3—enriched in transcriptionally active promoters, H3K27ac—which distinguishes active enhancers from poised enhancers, and H3K36me3—actively transcribed gene bodies) and two repressive marks (H3K9me3—constitutively repressed genes and H3K27me3—facultatively repressed genes). In Fig. 2c, we observed that most SINE families were underrepresented in all four active marks, consistent across tissues. However, young SINEs were highly enriched for H3K9me3, which indicates permanent repression, but not for H3K27me3, which has development-dependent repressed characteristics65. In general, compared to old SINE groups, young SINE groups had a higher over-/under-representation for all histone modification types, especially the youngest group.
In addition, we investigated the enrichment of SINE families in 15 chromatin states across 14 tissues66. We observed that young SINE groups had lower enrichment in most chromatin states compared to old SINE groups. The degree of SINE enrichment in chromatin states was similar across all 14 tissues (Fig. 2d, Supplementary Data 2). The enrichment characteristics of different SINE groups can be divided into four distinct patterns based on their degree of enrichment (Fig. 3a). The oldest SINE group was highly enriched in most cases (80%, 168 out of 210 combinations of 14 tissues and 15 states), while the young SINE group showed three enrichment patterns in the remaining combinations (enlarged inset). In two of these patterns, only the youngest SINE group was highly enriched in TssAHet (flanking active TSS without ATAC) and EnhAWk (weakly active enhancer). Overall, SINE groups were depleted from active promoters and enhancers, except for weak TSS and enhancers. Young SINE groups were more depleted than old ones. This suggests that new SINEs may be silenced by histone modifications and DNA methylation, while older ones may be tolerated by the pig genome.
a Hierarchical clustering of enrichment patterns in 15 chromatin states for four SINE groups across 14 tissues (left panel). The heatmap shows three distinct enrichment patterns for high enrichment in the young SINE group (right panel). b A heatmap for the enrichment of transcription factor binding motifs in SINE families, chromatin accessibility, and histone modifications. c The signal density of MeDIP-seq and CpG islands within different SINE families. The shading represents the 95% confidence intervals, shown as error bars above and below the mean column. d Boxplots display the DNA methylation levels on different SINE families. “L” and “R” represent the upstream and downstream directions of the SINE body. For example, “L250” represents the 0 to 250 bp window upstream of SINE, and “L500” represents the 250 to 500 bp window upstream of SINE. The line inside each boxplot represents the median.
TEs carrying TFBS may contribute to the regulation of genes67,68. We performed motif enrichment analysis of SINEs to explore their possible contributions (Fig. 3b). In total, 31 TFBS were predicted to have binding motifs in at least one SINE family, mostly in old ones (96.8%). 83.9% of the TFBS related to open chromatin, indicating that young SINEs were rarely exapted into regulatory regions69 and were repressed by less chromatin accessibility. The youngest SINE-specific TFBS related to the ZNF148 gene has been shown to promote the development of a muscle phenotype70. Besides, three members of the RFX transcription factor family were amplified in both young and older SINEs and involved in immune, reproductive, and nervous pathways71. For instance, RFX1 and RFX3 were found to be major histocompatibility complex (MHC) class II promoter binding proteins that functioned as trans-activators of the hepatitis B virus enhancer72,73.
Given that TEs play major roles in gene expression regulation by shaping the epigenetic modifications74, we analyzed the epigenetic states of SINE families by examining DNA methylation (MeDIP), density of CG (CpG) sequence contexts, and AT:GC content (CpG islands) (Fig. 3c). The results showed that almost all SINE families exhibited a depletion in genomic regions with CpG islands, and the young SINE families were more highly methylated than the old ones. Similarly, we found that CG methylation levels in SINE bodies, particularly in young SINEs, were higher than in their flanking regions across 10 different tissues. This corresponds to the enrichment of SINE families in H3K9me3 (Fig. 3d).
A previous study revealed that piRNAs play a major role in TE silencing via the ping-pong cycle in pig germline75. We further distinguished small non-coding RNAs into three classes to investigate the relationship between piRNA density and SINE families (Supplementary Fig. 8). Unlike siRNA and miRNA, piRNA was highly enriched for SINE-related sequences or sequence flanks, and there was a negative correlation between piRNA density and the age of the SINE subfamily. Our result agrees with findings in humans that young SINE families are more prone to piRNA targeting76, which may be the main reason for their high methylation levels, as we observed above.
Young SINE-associated transcriptome profiling in pigs
TEs can modify host gene transcription directly by remodeling alternative splice events or providing cis-regulatory sequences77,78,79. To test this, we analyzed PacBio long-read isoform sequences from 38 pig tissues to detect transcripts of SINE-associated exonization and alternative splice sites80,81. We estimated the contribution of TEs to gene expression across 52 tissues and 27 cell types by analyzing 3570 published RNA-seq samples from the EBI database (Fig. 4a, Supplementary Data 3).
a Overview of RNA-seq libraries in 3570 samples across 52 tissues and 27 types of cells. b The bar plot indicates the proportion of functional regions affected by SINE across four different categories of SINE-associated transcripts. The x-axis groups indicate the relationship between the SINE-associated transcript and annotated genes, while the colors represent the position of the SINE within the SINE-associated transcript. c Boxplots display the CG methylation levels on young SINE families. Young-T group represents the SINE families that derived the young SINE-associated transcripts. The Younger and Youngest groups represent all the younger and youngest SINEs in the entire genome, respectively. d The boxplots display the read density of chromatin accessibility and histone modifications in the Young-T, Younger, and Youngest groups. The line inside each boxplot represents the median.
After processing the raw data using LoRDEC82, we obtained 30,331,870 error-corrected Iso-seq reads with an average length of 2797 bp. Of these, 7.48% (2,267,973) were TE-related transcripts (the transcripts containing TE sequences)83. Notably, 68.81% (1,560,568) of the TE-derived transcripts were recognized as SINE-associated transcripts inserted by nearly full-length SINE (average coverage of 87.76%). This suggests that TE-derived transcripts, particularly SINEs, are abundant in the pig transcriptome. We next classified 337,746 young (younger and youngest) SINE-associated transcripts into four categories based on their genomic location compared to known transcripts in the pig genome annotations81 (Supplementary Fig. 9). Out of these, 1028 perfectly matched with 517 genes and 47 lncRNAs (Supplementary Data 4), while 62,304 potentially offered novel alternative splice events for 8103 genes and 405 lncRNAs (overlapping with at least one splice junction of a known transcript). The remaining young SINE-associated transcripts were classified as either 150,469 exon-covered transcripts (Overlapping exons without splice junction on same or opposite strands) or 130,180 intronic transcripts (Located in an intron), which had no complete structural similarity with the available transcript annotation.
When comparing the locations of young SINEs in their derived transcripts, we noticed a higher proportion of them in UTRs (Fig. 4b and Supplementary Data 5). Especially, for SINE-associated annotated transcripts that perfectly matched, 81.52% were found to have SINE in their 3′-UTRs. This may affect gene expression by increasing the length of the UTRs84 or by directly inserting into the regulatory region via a mechanism similar to Staufen-mediated decay (SMD)85. For example, we found a full-length PRE0-SS was inserted in the 3′-UTR of the pig PDK1 gene, which is consistent with a previous report that Alu and B1 regulate both human and mouse orthologs of PDK1 by SMD86. Besides, we found that young SINEs that produced transcripts (Young-T) had lower average CG methylation levels than all young SINEs in most tissues (Fig. 4c). Similarly, these young SINEs were more enriched in open chromatin and histone modifications, particularly H3K4me3, suggesting that they were more likely active across multiple tissues (Fig. 4d).
To study young SINE-associated transcripts, we analyzed transcriptome data from 52 tissues and 27 cell types using Salmon tools87 to measure gene and SINE-associated transcript abundance (RNA-seq counts). Normalized gene expression by DESeq288 allowed us to create t-SNE plots that is mostly consistent with tissue types (Fig. 5a). We also conducted a co-expression network analysis of 14,403 genes using the WGCNA R package89 to investigate the expression characteristics of young SINE-associated transcripts across a wide range of tissues and cell types (Supplementary Fig. 10).
a The t-SNE plots display the expression differentiation among different tissues and cells. b Top 20 results of functional enrichment analysis for young SINE-associated genes. c The bar plot indicates the enrichment of 248 young-T SINEs in chromatin states and histone modifications across different tissues.
As a result, 13,872 genes were grouped into 40 modules (number of genes >30), with most modules showing high tissue specificity and playing key roles in particular organ systems in pigs (Supplementary Figs. 11, 13, Supplementary Data 6). The results were supported by gene-to-gene networks of topological clustering using the Markov clustering (MCL) algorithm90 (Supplementary Fig. 14). Importantly, 17.9% of co-expressed genes (2744) were related to young SINE-associated transcripts and were mainly found in specific tissue modules (trachea, adipose, fetal thymus, and alveolar macrophages) (Supplementary Fig. 15).
Young SINE-related genes, as shown in Fig. 5b, were found to be highly enriched in neural development, cellular metabolism, muscle development, and immune response, which may have played a role in natural selection and domestication of modern pigs91,92,93. For instance, 186 genes in module ME2 had high expression in brain tissues and were significantly enriched in chemical synaptic transmission (GO:0007268), brain development (GO:0007420), and neuron projection morphogenesis (GO:0048812) (Supplementary Fig. 16). Correspondingly, there were 248 young SINEs that were associated with these 186 genes. These SINEs were more enriched in active epigenetic marks (H3K4me1, H3K4me3, and H3K27ac) and depleted from the repressive mark (H3K27me3) at the nervous system (cerebellum, cortex, and hypothalamus) than other tissues, indicating that young SINEs exhibited strong and concordant tissue specificity in both transcript expression and epigenetic regulation (Fig. 5c).
The roles of young SINEs in population admixture and local adaptation in pigs
Since the majority of TE polymorphisms in the pig genome are young SINEs47,55,57,94, we created a thorough map of dimorphic SINEs using whole-genome sequencing data from over 300 pigs, which represent most Eurasian pig breeds. This map allows us to investigate the roles of young SINEs and their associated genes in pig population admixture and local adaptation.
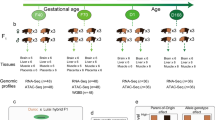
To investigate the effect of sequencing depth and dimorphic SINE detection tools on the detection of dimorphic SINEs, we benchmarked polymorphic TE detection tools that previously performed well in human projects under different sequencing depths13,95 (Supplementary Figs. 17–25, Supplementary methods) and customized the Pig TE Polymorphism (PigTEP) pipeline to improve its performance in pig whole-genome sequence datasets (Fig. 6a, Supplementary methods). We then used this pipeline to identify dimorphic SINEs in 374 individuals from 25 diverse populations (N ≥ 5) using a uniform sequencing depth of 10× (Supplementary Fig. 26, Supplementary Data 7, average mapped bases: 27.18 GB and average mapping rates: 99.44%). These individuals were assigned to 10 major groups: PYGMY, ISEA, CHD, KOD, AWB, TWB, EUD, EWB, MINI, and COM (Fig. 6b).
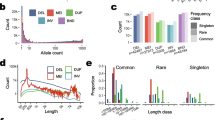
a The Pig TE polymorphism pipeline. The pipeline was constructed to identify both dimorphic SINEs and SNPs for each individual simultaneously. Ref+ refers to the insertion of a TE into the reference genome, whereas Ref- refers to the deletion of a TE from the reference genome. b Overviews of whole-genome re-sequencings in 374 individuals. c Venn diagram represents the distribution of dimorphic SINEs among different populations. d PCA plot displays the genetic relationship based on dimorphic SINEs among 374 individuals. e PCA plot displays the genetic relationship based on dimorphic SINEs among 364 individuals from modern pigs. f Phylogenetic tree based on dimorphic SINEs for 374 individuals. g Population structure based on dimorphic SINEs for 374 individuals when K was 3 and 10.
We identified a total of 211,067 dimorphic SINEs in the pig genome, with almost half located in non-intergenic regions. Out of these, 189,966 Ref+ refer to the insertion of a TE into the reference genome, while 21,101 Ref- refer to the deletion of a TE from the reference genome. Most of these SINEs (64.89%) were rare, with minor allele frequencies of less than 5% in the entire pig population (Supplementary Fig. 27), and they showed variable frequency distribution among groups (Supplementary Fig. 28). While over 85% of the dimorphic SINEs were shared among groups, there were still 30,441 dimorphic SINEs (PYGMY and ISEA accounted for 60.83% and 26.01%, respectively) that were unique to a single group (Fig. 6c).
Principal component analysis (PCA) analysis of dimorphic SINE genotypes distinguished four species of the Suidae (Fig. 6d). PC1 separated Porcula slavania from Sus species, while PC2 and PC3 (19.41% and 13.67%, respectively) showed genetic separation between Asian and Western breeds (Fig. 6e). Korean domestic pigs (KOD) showed closer genetic similarity to Western breeds than to Chinese breeds, likely due to gene flow and introgression mediated by humans. Our results were confirmed by the TE-based phylogenetic tree and genetic admixture (Fig. 6f, g; Supplementary Fig. 29), consistent with previous studies on SNP-based genotypes44,96. The comparison of Chinese and European domestic pigs confirmed our previous findings on TE-based introgression between Northern Chinese domestic pigs and European domestic pigs55. Korean wild boars clustered with other Asian wild boars instead of European pigs, unlike Korean domestic pigs.
To identify dimorphic SINEs linked to local adaptation, we chose the 10 ancestral components with the lowest coefficient of variation (CV = 0.229; Fig. 6g). For each dimorphic SINE between cluster i and the other clusters, we computed pairwise Fsti values and alpha coefficients (using Bayescan)97 to measure their divergence in allele frequencies at specific loci. Loci with higher Fst values and positive alpha values indicate positive selection. We found 337 dimorphic SINEs with high Fsti and positive alpha coefficient in the gene functional regions, including exon, splice, UTR5, UTR3, and upstream regions, of 330 genes (Supplementary Data 8). 77.94% of the genes were found in both PYGMY (n = 223) and ISEA (n = 42), while the remaining 75 were linked to breed-specific traits of domestic pigs (Fig. 7a). For example, the PRE1 insertion in the promoter of the IGFBP7 gene, which is associated with tumor suppression98, was more common in Chinese indigenous breeds than in commercial breeds99. Furthermore, the upstream of FRZB (high signals in H3K4me1 and H3K27ac, Fig. 7b), which is associated with pig growth traits100, was inserted by a population-specific dimorphic SINE from Southern Chinese domestic pigs.
a The scatter diagram displays the 75 genes that may be associated with local adaptation. The x-axis represents the Fst, and the y-axis represents population frequency. b Chromatin accessibility and histone modifications for FRZB (chr15:88332856–88377275) and its dimorphic SINE (chr15:88377793). c Bar plot displays the population frequency of the dimorphic SINEs in the first exon of RUNX3. d Overviews of nine candidate genes under local adaptation. Bar charts indicate the population frequency of candidate dimorphic SINEs and the average TPM values of their corresponding candidate genes across tissues. e The scatter diagram displays the linkage disequilibrium between T-SNPs and dimorphic SINEs. The x-axis represents the chromosome, and the y-axis represents the r2 values.
A fixed dimorphic SINE was found in the first exon of the RUNX3 gene of Goettingen miniature pigs, MiniLEWE, and Southern Chinese domestic pigs, particularly Luchuan pigs (Fig. 7c). RUNX3 gene is known as a tumor suppressor gene in human T-cell malignancy101 and plays a key role in the TGF-β induced signaling pathway102. In addition, 30 genes in long-term laboratory-inbred Bama Xiang pigs103 showed extreme Fst and were significantly enriched in the AMPK signaling pathway (Corrected P-values = 0.00295, Supplementary Data 9), with SLC3A2 (upstream) and SIRT1 (UTR3) genes having the dimorphic SINEs with a perfectly fixed frequency.
Out of 75 candidate genes, nine were mapped to swine quantitative trait-associated (QTX) data associated with phenotypic traits104 (Fig. 7d). Four of these genes, related to laboratory-inbred Bama Xiang pigs, were associated with fat content and body weight, consistent with selective breeding103. The LEP gene, highly expressed in adipose tissue, produces leptin, a hormone regulating appetite, fat storage, and body weight105. These findings demonstrate that dimorphic SINEs are a valuable source for studying genomic ancestry and local adaptive evolution in pigs.
Mapping young SINEs to the genetic associations of complex traits
To investigate the relationship between dimorphic SINEs and complex traits, we collected 4072 trait-associated SNPs (T-SNPs) from 79 published GWAS studies of 97 complex traits in pigs, including reproduction, production, meat and carcass, health, and exterior traits (Supplementary Fig. 30). As shown in Fig. 7e, 127 dimorphic SINEs associated with traits and in linkage disequilibrium (LD, r2 > 0.3) with T-SNPs were identified using 296 domestic pigs (109 Asian and 187 European). Specially, it was found that these dimorphic SINEs were more prevalent in the TxFlnkWk (Weak transcribed at gene), indicating their potential for gene regulation (Fig. 8a).
a Bar plot displays the enrichment of dimorphic SINEs in different chromatin states. b Heatmap displays the frequency of T-dimorphic SINEs among different pig populations. The darker red color represents a higher population frequency for dimorphic SINEs. c The population frequency of dimorphic SINEs in ELOVL3 gene among different pig populations. d The population frequency of dimorphic SINEs in ANK2 gene among different pig populations. e The expression of the ANK2 gene in the top 10 tissues sorted by gene expression. f The population frequency of dimorphic SINEs in VRTN gene among different pig populations. g The expression of the VRTN gene at the top 10 tissues sorted by gene expression. h The VRTN gene (chr7:97614707–97624273) structure and the neighboring SINE-associated transcript (chr7:97613985–97615896). i Chromatin accessibility and histone modifications for the upstream of VRTN gene. H3K27me3 signals for the upstream of VRTN gene.
54 genes influenced by dimorphic SINEs were linked to intramuscular fat composition and teat number. Many of these genes were specifically expressed in certain tissues (Z-score >2), such as the nervous system (plasmacytoid dendritic cells, choroid plexus, hypothalamus, and brain), reproductive system (testis, oviduct, and oocyte), and muscle satellite cells (Supplementary Fig. 31, Supplementary Data 10). Importantly, most of the intronic T-dimorphic SINEs exhibited breed-specific MAF between Chinese and Western pigs, which is consistent with their differences in fatty acid content and teat number (Fig. 8b).
We found a 320 kb dimorphic SINE hotspot (chr14:112,965,840–113,285,513; r2 > 0.3) linked to intramuscular fat composition, containing six dimorphic SINEs and eight genes. Two genes, C14H10orf76 (ARMH3, r2 = 0.89) and GBF1 (r2 = 0.86), are essential for Golgi maintenance and secretion106. The ELOVL3 gene is a strong candidate gene for fatty acid composition107,108. A low-frequency dimorphic SINE was found in its intron region, while multiple T-dimorphic SINEs were found within its upstream region of 15 to 50 kb. The dimorphic SINE (chr14:113,199,425) near 27 kb upstream was found at high frequency in Chinese domestic pigs, especially Southern Chinese domestic pigs (Fig. 8c).
Furthermore, pairwise LD (r2 = 0.88) was observed between the dimorphic SINE (chr8:109,447,835) located in the ANK2 intron and the T-SNP linked to C14:0, C16:0, and C16:1n7 fatty acid content in backfat109. Ankyrin-B (AnkB), an alternatively spliced variant of ANK2, is linked to obesity susceptibility in humans110. We found that the insertion of the T-dimorphic SINE was almost fixed in Western domestic pig populations (Fig. 8d) and located in an LD block of 15 kb (r2 > 0.5, chr8:109,439,023–109,454,866, Supplementary Fig. 32). We observed that ANK2 is a gene that is ubiquitously expressed in pigs and highly enriched in the nervous system (Fig. 8e). Two SINE-associated transcripts overlapped with ANK2 exons and were significantly correlated with ANK2 expression (Supplementary Data 11). ANK2 expression was significantly upregulated in cultivars with high-fat deposition such as Songliao black pigs compared with those with low-fat deposition like Landrace pigs111 and fast-growing chickens compared with slow-growing chickens112.
In addition, we found a high LD (r² = 0.75) between a T-SNP (chr7:97,606,621) and a T-dimorphic SINE (chr7:97,615,896) in the first intron of the VRTN gene—the gene suggested to be associated with teat number and the most promising candidate gene to increase the number of thoracic vertebrae (ribs) in pigs113. We observed a clear difference in frequency of the dimorphic SINE between Chinese indigenous breeds and commercial breeds (Fig. 8f). In addition, VRTN gene is highly expressed in embryonic stem cells, embryos, and ovaries, suggesting its role in early pig development (Fig. 8g). Especially, a novel transcript (2191 bp in length) derived from the bimorphic SINE covers the first exon of VRTN and is significantly associated with VRTN expression (Fig. 8h, P-values = 3.03 × 10−201, Supplementary Data 11). This transcript was supported by the RNA-seq exon coverage in NCBI annotation (Supplementary Fig. 33). This region exhibited the open chromatin and enhancer signals (H3K4me1) while was facultatively repressed in most tissues (H3K27me3) (Fig. 8i). An obvious decline was observed in the repressed states of stem cells and embryo-related tissues, corresponding to the tissue-specific expression in VRTN, suggesting that this region was crucial for VRTN, and this dimorphic SINE was more likely to affect its expression.
Discussion
In this study, we built a pig genome atlas of TEs using the PigTEDC pipeline, combining the similarity-, structure-, and de novo-based methods. Our findings show that almost a third (947 MB) of the pig genome is made up of TEs, mainly non-LTR retrotransposons (SINE and LINE). SINE is shorter and more complete than LINE. Similar to our previous findings52, the PRE1-SS, PRE0-SS, and PRE1a families in SINE/tRNA are the most recent and have the most polymorphic insertions. These polymorphic SINEs contribute nearly 90% of medium-length SVs across different assemblies, especially the L13 subfamily, with 36.15% of the youngest SINEs in the pig genome.
Gene regulatory network is influenced by genomic components, chromatin accessibility, histone modifications, DNA methylation, and cis-regulatory elements such as TFBS, promoters, and enhancers. TEs linked to specific chromosome features can impact gene regulatory networks in multiple ways as listed above. To our knowledge, this is the first time that large-scale multi-omics data were used to fully explore the relationships between TEs and chromosome features in the pig genome. Our findings showed that SINEs were highly enriched in the A compartment, and that the enrichment of SINEs in chromatin was associated with their ages. For instance, young SINE families were frequently enriched in close chromatin-like nucleosomes but highly depleted from open chromatin.
As expected, SINEs were highly depleted from all active chromatin tags, and more signals of constitutive heterochromatin tags (H3K9me3 peaks) were observed on SINEs. The exception was H3K27me3, which was associated with facultative suppressor genes and cannot permanently silence SINEs. Most histone modifications in SINE decreased as the TE’s age increased, which was in line with the distribution of DNA methylation on SINE and its contribution to TFBS. However, young SINEs, especially the youngest SINE family, were highly enriched in weakly active enhancer regions of hypothalamus tissue (Fold >1.5). We speculate that the relationship between SINE and its host genome is a combination of both arms race and co-evolution, depending on how the symbiosis turned out.
In the former case of parasitism, the young SINEs were more likely to be treated as new invaders that were constitutively silenced by histone modifications and DNA methylation of the host genome (e.g., PIWI-piRNA pathway during the TE in testis), while the old SINEs mutated and gained new regulatory potential, and thus were tolerated or even co-opted by the pig genome. In the latter case of mutualism, there might be rare cases where the SINEs were positively selected by nature, thereby helping the host genome better adapt to the local environment in the long run.
The use of long-read isoform sequencing provided us a more complete characterization of full-length transcripts, which made it possible to identify the young SINE-associated transcripts. Meanwhile, the Iso-seq reads we used here were collected from ~40 pig tissues, which ensured the investigation of the abundance and tissue specificity of young SINE-associated transcripts.
Our findings showed that the vast majority of young SINE-associated transcripts were non-coding RNAs that covered exons or fell within introns. A total of 3112 genes were found to be associated with young SINE-associated transcripts, and nearly 88% of them were enriched in co-expressed modules with high tissue specificity. The transcripts derived from young SINEs exhibited lower levels of CG methylation and were more enriched in open chromatin and histone modifications than whole young SINEs. Specifically, some young SINEs exhibited strong and consistent tissue specificity in both transcript expression and epigenetic regulation. This is consistent with previous findings in other species114,115,116, suggesting that SINE insertions may be a crucial component of genes and regulate tissue-specific expression of their target genes.
Dimorphic SINEs belong to SVs that are more sensitive to sequencing depth than SNPs. In this study, we benchmarked four dimorphic SINE detection tools under different sequencing depths to ensure the unbiased detection of dimorphic SINEs. Our findings showed that MELT (version 2.2.2) had robust performance in both Ref+ and Ref- detection (Supplementary Figs. 17–18). As expected, we found that the number of detected dimorphic SINEs increased with sequencing depth, especially from 5× to 10×, which nearly doubled the average number of dimorphic SINEs (Supplementary Fig. 19). Considering the sequencing depth in the current 838 publicly available whole-genome sequence datasets in pigs, we retained 374 individuals whose sequencing depth was greater than 10× and down-sampled their sequencing depth to ~10× through a strategy of randomly removing reads (average mapped bases: 27.17 GB and average mapping rates: 99.43%). Finally, the PigTEP pipeline was developed to identify both dimorphic SINEs and SNPs in individuals simultaneously. This pipeline will help other researchers explore the role of dimorphic SINEs in pig genomic study and breeding.
The contribution of TEs was underestimated in pig genomic research, despite the active role of SINEs under selective pressure. However, our understanding of SINEs in pig population genetics is limited without a comprehensive map of dimorphic SINEs based on large-scale re-sequencing data.
We genotyped and analyzed 211,067 dimorphic SINE loci in 374 individuals from 25 pig populations. These loci showed high variability in allele frequencies among populations. Based on these SINEs, we identified ten major clusters that corresponded to geographic differentiation. These SINEs with high pairwise Fst value can help us understand local adaptation in domestic pigs and identify candidate genes for economically important traits. Our findings confirm previous studies (e.g., IGFBP7) and identify new candidate genes.
GWAS studies have discovered thousands of QTLs for important pig traits based on SNPs, but most of these loci remain functionally uncharacterized. One possible reason is that phenotypic changes may be affected by SVs (TEs) in linkage disequilibrium with SNPs. In this study, 127 dimorphic SINEs were found to be in linkage disequilibrium with significant GWAS SNPs of complex traits, and nearly a third of them showed high tissue specificity in expression. Some of these dimorphic SINEs can generate novel transcripts (in H3K4me1 and H3K27me3), as exon-covered transcripts were found upstream of the VRTN gene. Future research is needed to validate how these dimorphic SINEs regulate target genes in specific tissues (e.g., VRTN gene in embryonic stem cells) and affect complex traits.
Methods
Overview of the PigTEDC pipeline
The PigTEDC pipeline was composed of three TE detection approaches using different algorithms to achieve its overall efficiency:
The similarity-based method was represented by the widely known RepeatMasker (V4.0.6) (http://www.repeatmasker.org), which search sequence using the TE consensus sequence from RepBase Update (https://www.girinst.org/repbase/) and Dfam 2.0 databases117 against homologous regions of the pig genome. The method was used to detect all known TEs, including DNA transposons, LTR, SINE, and LINE.
The structure-based method was used to capture the particular TE families based on their known sequence structure and motifs, which further enhanced the results of the similarity-based method to enhance the power of detecting known TEs. In our pipeline, HelitronScanner (V1.1)118 was used to improve DNA transposons detection, both LTRharvest (V2.0)119 and LTR-Finder (V1.07)120 were used to enrich the LTR results, and SINE-Finder (V1.1)121 was used to increase SINE detection.
The de novo-based method was used to identify the missing pig TEs from the known TE database, which employed the clusters of the repetitive sequences in the genome based on various methods. There were a total of three tools in this category. RepeatModeler (V1.0.11) was used to automate the runs of RECON (Multiple alignment clustering) and RepeatScout (Consensus seed clustering) for the pig genome (http://www.repeatmasker.org/RepeatModeler/). Red122 and P-Clouds123 were used to capture all potential repetitive sequences that included TEs and simple repeats in the pig genome using machine learning and oligonucleotide clustering methods, respectively.
The classification of pig TEs detected in the PigTEDC pipeline was further performed by the RepeatMasker through the following three parts:
Order and family classification of known TEs. The known TEs generated from both similarity-based and structure-based methods can be directly classified into four orders by referring to the RepBase update database, including DNA transposons, LTR, SINE, and LINE. The results of RepeatModeler from the de novo-based method also provided parts of known TEs as above. Considering the complex structure of nested TEs, we removed these TEs from RepeatMasker using parseRM_GetNesting.pl. After merging known TEs with each order, we removed the redundant TEs detected by different tools by using bedtools. For example, some SINEs can be identified by both RepeatMasker and SINE-Finder. At last, the non-redundant known TEs from each order were aligned to the consensus sequences of various families belonging to that order to confirm their specific families.
Match of known TEs with their family consensus sequences. Considering the different TEs have different coverage towards their family consensus sequences, we located the coverage area for each TE family using cross_match software (http://www.phrap.org/consed/). The parameters of cross_match was set as “-gap_init −25 -gap_ext −5 -minscore 10 -minmatch 6 -alignments -bandwidth 50 -word_raw”. Parts of TEs failed to pass the paired match with their family consensus sequences, thus were regarded as overly fragmented TEs. The passed TEs consisting of full-length and fragmented TEs, had clear and specific family classifications.
Family classification of unknown or novel TEs. In the step of classifying unknown Pig TEs, we first merged all repetitive sequences from both Red and P-Clouds tools, and kept parts of them with at least three copies using hs-blastn. We then filtered these multicopy sequences to keep potential TE sequences by aligning them with various non-TEs repetitive sequences, including tandem repeat, gene, tRNA, and rRNA. Meanwhile, these potential TE sequences were additionally filtered as unknown TEs by excluding known TE sequences from the RepBase update database. In addition, merging all unknown TEs from RepeatMasker, RepeatModeler, and Red & P-Clouds, we further reduced the presence of redundant sequences by CD-HIT (parameters: -c 0.90 and –n 8) and picked the representative one of each cluster as the consensus sequence for each unknown TE family. Finally, to obtain the genomic locations of unknown TEs, we performed the genome-wide identification for each unknown TE family using RepeatMasker with the custom-build library.
Divergence distribution of pig TEs
Divergences of TEs from consensus sequences were obtained using the Kimura two-parameter model from RepeatMasker124. The divergence levels were corrected for CpG content by DCpG = D/(1 + 9FCpG)125. Histograms with a bin size of 0.01 were plotted to show the distribution of divergence levels. Activity periods were estimated assuming a substitution rate of 5 × 10−9 substitutions/site per year126,127. Distribution histograms for sequence divergence of TE were plotted using a 0.01 bin size.
Processing of young SINEs
Annotation of young SINEs on 14 genome assemblies. For each pig genome assembly, we employed RepeatMasker to search for young SINEs in our customized repeat library (PRE1-SS, PRE0-SS, and PRE1a). And then “parseRM_GetNesting.pl” was used to remove the nesting and nested young SINEs, followed by a length filtering to remove elements with length <200 bp and >300 bp using a shell script.
Clustering and filtering of young SINEs. To ensure that the young SINEs used for subfamily classification were highly homologous, we utilized a clustering-based approach to keep all non-nested young SINEs from 14 genomes with a sequence identity of >90%, implemented in CD-HIT-EST128 with the parameter that “-T 0 -c 0.9 -M 0 -n 5 -p 0”. we finally retained 52 of 17,294 SINE clusters (n > 500) involving 1,157,133 young SINEs.
Removing the duplicated sequences of young SINEs. Due to the existence of shared SINEs from different pig genomes, we removed the completely duplicated sequences from our young SINE datasets using seqkit (https://github.com/shenwei356/seqkit).
Construction of consensus sequence. Because the running time and memory usage for multiple sequence alignment (MSA) of large-scale genomic sequences can be enormous, we randomly extracted 100,000 young SINEs to conduct the MSA. Using MEME suite programs (http://web.mit.edu/meme_v4.11.4/share/doc/motif-consensus.html), we scanned each column in a letter’s (A, T, C, G) frequency matrix of MSA using the “50% rule” that any letters with frequency less 50% of the maximum were discarded. Finally, a consensus sequence with a length of 261 bp was created for the subfamily classification.
Phylogenetic tree for SINEs
26 SINE families were used to construct the phylogenetic tree. Multiple sequence alignments of their consensus sequences were performed with mafft129 (V7.407) at the default setting. Then, IQ-TREE130 was used to create maximum likelihood (ML) trees for SINE (with 100 fast bootstrap replicates). Finally, EVOLVIEW131 was used to visualize the phylogenetic tree for SINE families.
Subfamily classification of young SINEs
We conducted subfamily classification for young SINEs using the COSEG pipeline (http://www.repeatmasker.org/COSEGDownload.html). First, sequence homology analysis was done using cross_match software (https://www.phrap.com/) with the parameters (-gap_init −25 -gap_ext −5 -minscore 100 -minmatch 6 -alignments -bandwidth 50 -word_raw). Then, “preprocessAlignments.pl” was used to determine the consensus range and create input files for COSEG programs with the following parameters: the minimum distance between sites (-maxEdgeGap) was 10; the consensus sequence ranged from 1 to 251 bp (-minConsRange 1 -maxConsRange 251; refer to Supplementary Fig. 1). In the end, a minimum spanning tree of young SINEs was constructed to define their subfamilies using COSEG programs with the following parameters: the minimum subfamily size (-m) was 100; Alkes Price’s p-value method (-k) was used132; 2 bp (-t) co-segregating mutations were used when developing subfamilies.
Identification of dimorphic SINEs among the 14 genome assemblies
-
(1)
Detection of SVs among the 14 genome assemblies. We employed Minimap2 (V2.17)133 to identify SVs in the assemblies compared to the pig reference assembly. First, we performed a cross-breed full-genome alignment (PAF file) for each genome assembly using Minimap2 with the following parameters: “-c -x asm20 –cs”. Then, we used paftools.js (a JavaScript script within Minimap2) to identify the confident/callable regions and call the variants from the asm-to-ref alignment with the following parameters: “call -L1000”.
-
(2)
Identification of dimorphic SINEs from SVs. We first retained the SVs that had a similar size (>200 bp and <300 bp) with overlapping SINE using a shell script. Then, the inserted sequences of the resulting SVs were then aligned against the consensus sequences of SINE families and young SINE subfamilies using BLASTn version 2.2.31+ with default parameters. Finally, the SINE families or subfamilies of these SVs were defined by their best high-quality match between consensus sequences of SINE families or subfamilies.
3D chromatin architecture and chromatin accessibility
The 3D chromatin architecture and ATAC-seq used in this study, including A/B compartments, Hi-C, and TADs, were downloaded from the FRAGENCODE project62. MNase-seq reads were filtered with Trim_galore tools to obtain and retain high-quality reads with the following parameters: “-q 20 --phred 33 --stringency 3 -e 0.1”. The filtered reads were then aligned to the Duroc reference genome (Sus scrofa 11.1) using the BWA134. DANPOS2135 (V2.26) was then used to call nucleosome binding peaks and to generate nucleosome occupancy profiles with the following parameters: “--span 1 --smooth_with 20 --wideth 40”, which were further normalized with the mean score of the whole genome.
Histone modifications
The histone modifications used in this study include four active epigenetic marks (H3K4me1, H3K4me3, H3K27ac, and H3K36me3) and two repressive marks (H3K9me3 and H3K27me3). Histone modification data were quality-filtered using Trim_galore tools with the following parameters: “-q 20 --phred33 --stringency 3 -e 0.1”. ChIP-seq data were aligned to the Duroc reference genome using BWA. Peaks were called using MACS2136 (V2.1.1) with the following parameters: “-q 0.05”. The computeMatrix module from deepTools137 (V.3.5.0) was used to transform and compute the corresponding data matrix from the modification signal over a set of SINE regions. The plotProfile module was used to turn the compressed matrix into summary plots.
Definition of chromatin status
As described in a previous study66, we defined 15 distinct chromatin states across 14 tissues and grouped them into the following seven categories: (1) promoters included TssA (Strongly active promoters/transcripts), TssAHet (Flanking active TSS without ATAC), and TssBiv (Transcribed at gene); (2) TSS-proximal transcribed regions included TxFlnk (Transcribed at gene), TxFlnkWk (Weak transcribed at gene), and TxFlnkHet (Transcribed region without ATAC); (3) enhancers included EnhA (Strong active enhancer), EnhAMe (Medium enhancer with ATAC), EnhAWk (Weak active enhancer), EnhAHet (Active enhancer no ATAC), and EnhPois (Poised enhancer); (4) repressed regions included Repr (Repressed polycomb) and ReprWk (Weak repressed polycomb); (5) quiescent regions; (6) ATAC_Is (ATAC island); (7) TssBiv (Bivalent/poised TSS).
Transcription factor binding sites
To identify TFBS genome-wide, we used the FIMO software138 from the MEME Suite139 to look for their occurrences in the Duroc reference genome for the 746 Transcription Factors (TFs) cataloged in the Vertebrate 2020 JASPAR database140. We set the parameters of fimo software as “p-value = 1e-4”, “--max-stored-scores 100000000” and “--alpha=1”. Among the predicted TFBS, those obtained from the 31 known TF motif were used to profile the resulting density of different SINE families.
DNA methylation
MeDIP-Seq data were quality-filtered using Fastp tools141 with the following parameters: “-q 20 -u 30 -l 30 -w 16”. MeDIP-Seq data were aligned to the Duroc reference genome using Hisat2142. Duplicate reads were removed from the bam files using Sambamba tools143. The filtered bam files were used to identify the MeDIP-enriched regions based on a clustering approach using SICER2144.
The WGBS data were aligned to the Duroc reference genome using Bismark145 (V0.23.0) with the parameters that “--ambig-bam”. The statistical analysis and visualization of DNA methylation levels on TEs and the whole genome were performed using the methPlot script from BatMeth2146, and the R language utilizing the “data.table” and ggplot2 packages.
Identification of small non-coding RNAs
To study non-coding RNAs in young SINE families, we did the following steps: (1) We merged all raw data and removed adapters with sRNAseqAdapterRemover from TBtools147. (2) We used the “collapse_reads_md.pl” script of Mirdeep2148 (V0.1.2) to remove repetitive sequences. (3) We downloaded known pig piRNAs from piRBase149 (V3.0) (http://bigdata.ibp.ac.cn/piRBase/) and used Bowtie to map the collapsed reads to them, saving the unaligned reads. (4) We used Mirdeep2’s mapper and miRDeep2 to identify candidate microRNAs and their genomic locations based on these unaligned reads. (5) We mapped the remaining reads that did not align to piRNAs or miRNAs to the Duroc reference genome using Mirdeep2’s mapper, filtering for a length of 21 nt to identify candidate siRNAs. We excluded any small non-coding RNAs that had multiple hits in the reference genome.
Identification of SINE-associated transcripts and gene quantification
Using LoRDEC82 software with parameters “-k 21 -s 3”, we processed the raw data by their corresponding RNA-seq data. After merging the error-corrected Iso-seq reads, we aligned the consensus sequence of young SINE families to it using BLAST software with parameters “-evalue 1e−5 -max_target_seqs 1”. We identified 2,267,973 candidate TE-derived transcripts containing young TE insertions (transcripts contain more than 80% of the sequence of SINE). As previously studied81, we categorized young SINE-associated transcripts into four groups by comparing their genomic location to known transcripts in the currently available pig genome annotations. These four groups, as shown in Supplementary Fig. 9, include: (1) perfect match with known annotation; (2) at least one splice junction in common with a known transcript; (3) exonic overlap without a matched splice junction on the same or opposite strands; (4) located in a reference intron.
To study the cis-functionality of young SINE-associated transcripts, we used Salmon tools87 to measure the abundance (RNA-seq counts) of genes and SINE-associated transcripts. The RNA-seq counts were transformed to log2-counts per million (log CPM, suitable for linear modeling150).
Construction of co-expression network
We used 14,403 genes from 3570 samples to explore young SINE-associated transcripts across different tissues and cell types. We analyzed gene expression using DESeq288 and built a co-expression network with WGCNA R package89. The network was constructed with the following parameters: “corType = pearson, maxBlockSize = 20,000, power = 4, minModuleSize = 30, mergeCutHeight = 0.3”. The power was set to 4 (Supplementary Fig. 10). The blockwiseModules function was used to create a correlation network, a cluster tree, and modules. The modules were merged and analyzed using the plotDendroAndColors function.
KEGG enrichment and Gene ontology analysis
We analyzed 2744 genes associated with young SINE-associated transcripts using Metascape151. We used human gene IDs for enrichment of KEGG pathway and Gene Ontology terms, with the entire human gene list as the gene background after converting all pig gene IDs.
Benchmarking detection tools and sequencing depth for dimorphic SINE detection
We benchmarked four types of dimorphic SINEs detection tools that have been reported to have superior performance in the previous projects13,95,152, including STEAK153 (http://github.com/applevir/STEAK), TranSurVeyor154 (https://github.com/Mesh89/TranSurVeyor), MELT155 (https://melt.igs.umaryland.edu/), and RetroSeq156 (https://github.com/tk2/RetroSeq). Their performance was evaluated using Sniffles157 (https://github.com/fritzsedlazeck/Sniffles) with 20× PacBio sequencing from the same individual. We compared the counts and verification rates (the proportion of dimorphic SINEs supported by Sniffles) of identified dimorphic SINEs across a range of sequencing depths (5x, 10x, 15x, 20x, 30x, and 50x) to assess their detection powers (Supplementary Figs. 16–17). The results showed that MELT showed the robust performance on both Ref+ and Ref- dimorphic SINEs than the other three dimorphic SINEs detection tools. Therefore, the MELT was selected in our PigTEP pipeline. Using the MELT program, we further performed the dimorphic SINEs detection using NGS data of Meishan and Duroc breeds across a range of sequencing depths (5x, 10x, 15x, 20x, 30x, and 50x). We found that the number of dimorphic SINEs increased with the sequencing depth, especially from 5x to 10x that increased the average number of dimorphic SINEs by nearly one-fold (Supplementary Fig. 18). After evaluating the sequencing depth of current 838 publicly available pig next-generation sequence datasets (Downloaded from the NCBI SRA database https://www.ncbi.nlm.nih.gov/sra), we retained 374 individuals whose sequencing depth was greater than 10x for dimorphic SINEs identification (average mapped bases: 27.17 GB and average mapping rates: 99.43%).
Identification of SNPs and dimorphic SINEs from re-sequencing data
Preprocessing of next-generation sequencing. The 374 individuals were selected to identify the SNP and dimorphic SINEs, after their sequencing depth was standardized around 10x. We first performed the preprocessing as following steps: Step 1: Quality control. Quality control was conducted for each raw re-sequencing data using the fastp (https://github.com/OpenGene/fastp) with the following parameters: The quality value that a base was qualified (-q) was 20; The percent of bases were allowed to be unqualified (-u) was 30; The required length of kept reads (-l) was 50; Step 2: Read alignment. Clean reads from all individuals were aligned to the Duroc reference genome (Sus scrofa 11.1) using BWA134 (https://github.com/lh3/bwa) with the BWA-MEM algorithm; Step 3: Processing the alignment files. The mapped reads were subsequently processed for format conversion (view), position sorting (sort), merging (merge), and statistics (stats) using Samtools152 (http://samtools.sourceforge.net/). Then, the MarkDuplicates method from the Picard package (https://sourceforge.net/projects/picard/) was used to remove the PCR duplicates that were introduced during library construction.
Genome-wide SNP detection. After preprocessing of the alignments, the DNAseq mode of Sentieon (https://github.com/Sentieon/sentieon-dnaseq) was next used to identify genome-wide SNP in the following steps: Step 4: Indel realignment and Base quality score recalibration (BQSR). The processed alignment bam files were subjected to realign and recalibration with the parameters of “Realigner” and “QualCal”; Step 5: SNP genotype calling. Genotype calling of SNP for each individual was performed using the “Haplotyper” algorithm. Step 6: Merge of VCF files. All 374 g-vcf files were joined together using the “GVCFtyper” algorithm. Then, the newly merged VCF were filtered to retain high-quality SNPs using the vcftools with the following parameters: “--max-missing 0.8 --maf 0.05 --min-alleles 2 --max-alleles 2 --recode”.
Genome-wide dimorphic SINEs detection. The MELT tool (https://melt.igs.umaryland.edu/) was used to discover the dimorphic SINEs (Ref+ and Ref-) in the following steps: Step 7: Discovery of deletion. With the processed bam files from Step 3, we used the Deletion-Genotype module of MELT to identify the “Ref-” dimorphic SINEs based on our previously identified TE sets (bed file, n = 3,087,929); Step 8: Building retrotransposon. Referring to the results of Step 7, we build three customized reference files for PRE1-SS, PRE0-SS, and PRE1a families using the “BuildTransposonZIP” module; Step 9: Discovery of insertion. For each bam file, we carried out the identification of “Ref+” dimorphic SINEs using four modules of MELT step by step, including “IndivAnalysis”, “GroupAnalysis”, “Genotype”, and “MakeVCF”.
All detected dimorphic SINEs and SNPs were processed using gene-based annotations in ANNOVAR158. The pig annotation files were downloaded from the NCBI database for the Duroc reference genome (Sus scrofa 11.1). Dimorphic SINEs and SNPs were classified into eight categories based on their genome locations, including exonic regions, splicing sites, intronic regions, 5′ and 3′ untranslated regions (UTRs), upstream and downstream regions, and intergenic regions.
Principal component analysis
We filtered identified dimorphic SINEs (Ref+ and Ref-) from 374 pigs using PLINK159 v1.9 with following parameters: “--maf 0.01 --mind 0.8 --geno 0.8”. Next, we performed principal component analysis on the filtered dimorphic SINEs using PLINK v1.9 with the parameters: “--pca”. We presented the eigenvectors and eigenvalues for each individual in the PCA biplot using R packages.
TE-based phylogenetic tree
To establish evolutionary relationships between individuals, we transformed the dimorphic SINEs dataset (Ref+ and Ref-) into a pseudo-SNPs dataset (A/T/C/G) by randomly replacing data under the condition that individual polymorphism was not changed. The transformed dataset was reduced to 44,192 dimorphic SINEs using PLINK v1.9 with the following parameters: “--maf 0.1 --indep --pairwise 50 10 0.2”, based on a linkage disequilibrium threshold of 0.2 and a minor allele frequency threshold of 0.1. The phylogenetic tree was constructed with 1000 bootstrap replicates using the maximum-likelihood approach implemented in SNPhylo160 (V1.10.2).
Analysis of genetic admixture
We used the program ADMIXTURE161 (V1.3.0) to analyze population structure in our study. This estimated genetic admixture among different pig breeds using all identified dimorphic SINEs. We tested 14 cases (ranging from K = 2 to 15) to identify genetic clusters for 374 pigs, using default parameters. Results were visualized using StructureSelector162.
Identifying the dimorphic SINEs associated with local adaptation
As shown in Fig. 6g, the optimal number of ancestral components was inferred to be K = 10 with the lowest CV (0.229), resulting in the eleven distinct genetic clusters of G01: PYGMY, G02: ISEA, G03: Southern Chinese domestic pigs (WUZ, LUC, and XIA), G04: laboratory-inbred Bama Xiang pigs, G05: Eastern Chinese domestic pigs (CHDO, WANB, EHS, and MEI), G06: Asian wild boars (KWB, CWB, and TWB) plus Northern Chinese domestic pigs (LAI, HET, and MIN), G07: Goettingen miniature pigs and MiniLEWE163, G08: Korean domestic pigs (JEJ and PEN) plus European domestic pigs, G09: European wild boars plus IBE and Yucatán miniature pigs, G10: Landrace/Yorkshire crossbreeds (YL)164, and G11: Duroc pigs.
We calculated the pairwise Fsti value and estimated alpha coefficient97 of dimorphic SINEs between cluster i and the remaining clusters to measure their locus-specific divergence in allele frequencies. Pairwise Fsti values were calculated for each dimorphic SINEs using the vcftools165 (V0.1.16) with the default parameters. BayeScan program97 (v2.1) was used to identify putative adaptive dimorphic SINEs. based on different allele frequencies among populations, and we performed it using default settings (prior odds to 10, iterations to 5000, and burn-in to 50,000).
Linkage disequilibrium analysis
We collected a total of 4072 trait-associated SNPs (T-SNPs) from 79 published GWAS studies of 97 complex traits. These included 18 reproduction, 22 production, 36 meat and carcass, six health, and two exterior traits. We combined the dimorphic SINEs and 4072 T-SNPs into a new variant dataset from 296 domestic pigs (109 Asian and 187 European domestic pigs). We analyzed the linkage disequilibrium between T-SNPs and dimorphic SINEs using PLINK v1.9 with the parameter “--ld”. We only considered dimorphic SINEs with a relative T-SNP distance less than 10 kb and R2 greater than 0.3 as candidate dimorphic SINEs. Linkage disequilibrium analysis for ANK2 gene (chr8:109,439,023–109,454,866) was performed using Haploview software166. LDBlockShow167 (V1.40) tool was used for the linkage disequilibrium analysis of large regions like 320 kb dimorphic SINEs hotspots (chr14:112,965,840–113,285,513).
Statistics and reproducibility
The Z-score of each chromatin state for each TE group was calculated using permutation tests in regioneR168, with the number of permutations set to 1000. For the association analysis of young SINE-associated transcripts and genes, we used R code and generalized linear models (GLM) to model transcript expression levels of each gene. We considered the expression level of young SINE-associated transcripts (within their gene body) and the type of tissues or cells as explanatory variables. We used a Bonferroni significance threshold of 1.42 × 10−6 (0.05/35,135) as the standard threshold.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All public datasets of pigs used in our analysis had sufficient ethical approval. Five pig MNase-seq samples were downloaded from NCBI (BioProject ID PRJNA417273) representing brain, heart, kidney, liver, and muscle tissues. H3K4me1, H3K4me3, H3K27ac, and H3K27me3 in 14 tissues were downloaded from NCBI under BioProject ID PRJNA762083. In addition, we downloaded one H3K9me3 sample (BioProject ID PRJNA152995), and four H3K36me3 (BioProject ID PRJNA529704) and H3K27me3 (BioProject ID PRJEB31243) samples each. Chromatin states of pigs were analyzed in 14 different tissues, which were downloaded from (http://genome.ucsc.edu/s/zhypan/susScr11_15_state_14_tissues_new). This study used MeDIP-Seq data from 80 samples, which were downloaded from NCBI under BioProject ID PRJNA143661. Whole Genome Bisulfite Sequencing data from 246 samples across 10 tissues were downloaded from 16 studies (Supplementary Data 12). Data from 177 samples across 20 pig small non-coding RNA studies were collected (Supplementary Data 13). Two PacBio long-read isoform sequencing datasets were downloaded from NCBI: BioProject ID PRJCA000349 (pooled 38 samples) and PRJNA351265. Transcriptome data from 52 tissues and 27 types of cells can be found in Supplementary Data 3. The source data for the main figures is available in the online resource of figshare (https://doi.org/10.6084/m9.figshare.23856213.v4)169.
References
Ricci, M., Peona, V., Guichard, E., Taccioli, C. & Boattini, A. Transposable elements activity is positively related to rate of speciation in mammals. J. Mol. Evol. 86, 303–310 (2018).
Platt, R. N. 2nd, Blanco-Berdugo, L. & Ray, D. A. Accurate transposable element annotation is vital when analyzing new genome assemblies. Genome Biol. Evol. 8, 403–410 (2016).
Sahebi, M. et al. Contribution of transposable elements in the plant’s genome. Gene 665, 155–166 (2018).
Cordaux, R. & Batzer, M. A. The impact of retrotransposons on human genome evolution. Nat. Rev. Genet. 10, 691–703 (2009).
Levin, H. L. & Moran, J. V. Dynamic interactions between transposable elements and their hosts. Nat. Rev. Genet. 12, 615–627 (2011).
Streva, V. A., Faber, Z. J. & Deininger, P. L. LINE-1 and Alu retrotransposition exhibit clonal variation. Mob. DNA 4, 16 (2013).
Flynn, J. M. et al. RepeatModeler2 for automated genomic discovery of transposable element families. Proc. Natl. Acad. Sci. USA 117, 9451–9457 (2020).
Beck, C. R. et al. LINE-1 retrotransposition activity in human genomes. Cell 141, 1159–1170 (2010).
Ade, C., Roy-Engel, A. M. & Deininger, P. L. Alu elements: an intrinsic source of human genome instability. Curr. Opin. Virol. 3, 639–645 (2013).
Wicker, T. et al. A unified classification system for eukaryotic transposable elements. Nat. Rev. Genet. 8, 973–982 (2007).
Biemont, C. A brief history of the status of transposable elements: from junk DNA to major players in evolution. Genetics 186, 1085–1093 (2010).
Arkhipova, I. R. Using bioinformatic and phylogenetic approaches to classify transposable elements and understand their complex evolutionary histories. Mob. DNA 8, 19 (2017).
Hoen, D. R. et al. A call for benchmarking transposable element annotation methods. Mob. DNA 6, 13 (2015).
Goerner-Potvin, P. & Bourque, G. Computational tools to unmask transposable elements. Nat. Rev. Genet. 19, 688–704 (2018).
Xing, J., Witherspoon, D. J. & Jorde, L. B. Mobile element biology: new possibilities with high-throughput sequencing. Trends Genet. 29, 280–289 (2013).
Sotero-Caio, C. G., Platt, R. N. 2nd, Suh, A. & Ray, D. A. Evolution and diversity of transposable elements in vertebrate genomes. Genome Biol. Evol. 9, 161–177 (2017).
Chenais, B., Caruso, A., Hiard, S. & Casse, N. The impact of transposable elements on eukaryotic genomes: from genome size increase to genetic adaptation to stressful environments. Gene 509, 7–15 (2012).
Zeh, D. W., Zeh, J. A. & Ishida, Y. Transposable elements and an epigenetic basis for punctuated equilibria. Bioessays 31, 715–726 (2009).
Bourque, G. et al. Ten things you should know about transposable elements. Genome Biol. 19, 199 (2018).
Stewart, C. et al. A comprehensive map of mobile element insertion polymorphisms in humans. PLoS Genet. 7, e1002236 (2011).
Manthey, J. D., Moyle, R. G. & Boissinot, S. Multiple and independent phases of transposable element amplification in the genomes of piciformes (woodpeckers and allies). Genome Biol. Evol. 10, 1445–1456 (2018).
Schmitz, J. SINEs as driving forces in genome evolution. Genome Dyn. 7, 92–107 (2012).
Carpentier, M. C. et al. Retrotranspositional landscape of Asian rice revealed by 3000 genomes. Nat. Commun. 10, 24 (2019).
Marmoset Genome, S. & Analysis, C. The common marmoset genome provides insight into primate biology and evolution. Nat. Genet 46, 850–857 (2014).
Rishishwar, L., Tellez Villa, C. E. & Jordan, I. K. Transposable element polymorphisms recapitulate human evolution. Mob. DNA 6, 21 (2015).
Rishishwar, L. et al. Evidence for positive selection on recent human transposable element insertions. Gene 675, 69–79 (2018).
Van’t Hof, A. E. et al. The industrial melanism mutation in British peppered moths is a transposable element. Nature 534, 102–105 (2016).
Mao, H. et al. A transposable element in a NAC gene is associated with drought tolerance in maize seedlings. Nat. Commun. 6, 8326 (2015).
Feschotte, C. Transposable elements and the evolution of regulatory networks. Nat. Rev. Genet. 9, 397–405 (2008).
Attig, J. et al. Splicing repression allows the gradual emergence of new Alu-exons in primate evolution. Elife 5, e19545 (2016).
Hirsch, C. D. & Springer, N. M. Transposable element influences on gene expression in plants. Biochim. Biophys. Acta Gene Regul. Mech. 1860, 157–165 (2017).
Trizzino, M. et al. Transposable elements are the primary source of novelty in primate gene regulation. Genome Res. 27, 1623–1633 (2017).
Wang, J. et al. MIR retrotransposon sequences provide insulators to the human genome. Proc. Natl. Acad. Sci. USA 112, E4428–E4437 (2015).
Sundaram, V. et al. Widespread contribution of transposable elements to the innovation of gene regulatory networks. Genome Res. 24, 1963–1976 (2014).
Funkhouser, S. A. et al. Evidence for transcriptome-wide RNA editing among Sus scrofa PRE-1 SINE elements. BMC Genom. 18, 360 (2017).
Jeck, W. R. et al. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA 19, 141–157 (2013).
Roberts, J. T. et al. Continuing analysis of microRNA origins: formation from transposable element insertions and noncoding RNA mutations. Mob. Genet. Elem. 3, e27755 (2013).
Chishima, T., Iwakiri, J. & Hamada, M. Identification of transposable elements contributing to tissue-specific expression of long non-coding RNAs. Genes 9, 23 (2018).
Fedoroff, N. V. Presidential address. Transposable elements, epigenetics, and genome evolution. Science 338, 758–767 (2012).
Rey, O., Danchin, E., Mirouze, M., Loot, C. & Blanchet, S. Adaptation to global change: a transposable element-epigenetics perspective. Trends Ecol. Evol. 31, 514–526 (2016).
Lanciano, S. & Mirouze, M. Transposable elements: all mobile, all different, some stress responsive, some adaptive? Curr. Opin. Genet. Dev. 49, 106–114 (2018).
Horvath, V., Merenciano, M. & Gonzalez, J. Revisiting the relationship between transposable elements and the eukaryotic stress response. Trends Genet. 33, 832–841 (2017).
Lerat, E., Casacuberta, J., Chaparro, C. & Vieira, C. On the importance to acknowledge transposable elements in epigenomic analyses. Genes 10, 258 (2019).
Groenen, M. A. et al. Analyses of pig genomes provide insight into porcine demography and evolution. Nature 491, 393–398 (2012).
Cooper, D. K. et al. The pathobiology of pig-to-primate xenotransplantation: a historical review. Xenotransplantation 23, 83–105 (2016).
Ekser, B., Markmann, J. F. & Tector, A. J. Current status of pig liver xenotransplantation. Int. J. Surg. 23, 240–246 (2015).
Fang, X. et al. The sequence and analysis of a Chinese pig genome. Gigascience 1, 16 (2012).
Li, M. et al. Genomic analyses identify distinct patterns of selection in domesticated pigs and Tibetan wild boars. Nat. Genet. 45, 1431–1438 (2013).
Li, M. et al. Comprehensive variation discovery and recovery of missing sequence in the pig genome using multiple de novo assemblies. Genome Res. 27, 865–874 (2017).
Groenen, M. A. M. et al. Analyses of pig genomes provide insight into porcine demography and evolution. Nature 491, 393–398 (2012).
Vamathevan, J. J. et al. Minipig and beagle animal model genomes aid species selection in pharmaceutical discovery and development. Toxicol. Appl. Pharm. 270, 149–157 (2013).
Zhao, P. et al. PRE-1 revealed previous unknown introgression events in Eurasian boars during the middle pleistocene. Genome Biol. Evol. 12, 493 (2020).
Chalopin, D., Naville, M., Plard, F., Galiana, D. & Volff, J. N. Comparative analysis of transposable elements highlights mobilome diversity and evolution in vertebrates. Genome Biol. Evol. 7, 567–580 (2015).
Gingerich, P. D. Environment and evolution through the Paleocene-Eocene thermal maximum. Trends Ecol. Evol. 21, 246–253 (2006).
Zhao, P. et al. PRE-1 revealed previous unknown introgression events in Eurasian boars during the middle pleistocene. Genome Biol. Evol. 12, 1751–1764 (2020).
Zhou, R. et al. Developmental stage-specific A-to-I editing pattern in the postnatal pineal gland of pigs (Sus scrofa). J. Anim. Sci. Biotechnol. 11, 90 (2020).
Chen, C. et al. SINE jumping contributes to large-scale polymorphisms in the pig genomes. Mob. DNA 12, 17 (2021).
Jordan, K. W., He, F., de Soto, M. F., Akhunova, A. & Akhunov, E. Differential chromatin accessibility landscape reveals structural and functional features of the allopolyploid wheat chromosomes. Genome Biol. 21, 176 (2020).
Vrljicak, P. et al. Analysis of chromatin accessibility in decidualizing human endometrial stromal cells. FASEB J. 32, 2467–2477 (2018).
Benoit, M. et al. Environmental and epigenetic regulation of Rider retrotransposons in tomato. PLoS Genet. 15, e1008370 (2019).
Cournac, A., Koszul, R. & Mozziconacci, J. The 3D folding of metazoan genomes correlates with the association of similar repetitive elements. Nucleic Acids Res. 44, 245–255 (2016).
Foissac, S., Djebali, S., Munyard, K., Vialaneix, N. & Giuffra, E. Multi-species annotation of transcriptome and chromatin structure in domesticated animals. BMC Biol. 17, 108 (2019).
Yumei, L. et al. Human exonization through differential nucleosome occupancy. Proc. Natl. Acad. Sci. USA 115, 8817–8822 (2018).
Kern, C. et al. Functional annotations of three domestic animal genomes provide vital resources for comparative and agricultural research. Nat. Commun. 12, 1821 (2021).
Saksouk, N., Simboeck, E. & Dejardin, J. Constitutive heterochromatin formation and transcription in mammals. Epigenetics Chromatin 8, 3 (2015).
Pan, Z. et al. Pig genome functional annotation enhances the biological interpretation of complex traits and human disease. Nat. Commun. 12, 5848 (2021).
Luis, V. -C. J., Vivien, H., Laura, A. & Josefa, G. Diverse families of transposable elements affect the transcriptional regulation of stress-response genes in Drosophila melanogaster. Nucleic Acids Res. 47, 6842–6857 (2019).
Jordi, M., Fatima, M., Jordi, P. & Casacuberta, J. M. Plant lineage-specific amplification of transcription factor binding motifs by miniature inverted-repeat transposable elements (MITEs). Genome Biol. Evol. 10, 1210–1220 (2018).
Simonti, C. N., Pavlicev, M. & Capra, J. A. Transposable element exaptation into regulatory regions is rare, influenced by evolutionary age, and subject to pleiotropic constraints. Mol. Biol. Evol. 34, 2856–2869 (2017).
Bakke, J. et al. Transcription factor ZNF148 is a negative regulator of human muscle differentiation. Sci. Rep. 7, 8138 (2017).
Piasecki, B. P., Burghoorn, J. & Swoboda, P. Regulatory Factor X (RFX)-mediated transcriptional rewiring of ciliary genes in animals. Proc. Natl. Acad. Sci. USA 107, 12969–12974 (2010).
Reith, W. et al. Cloning of the major histocompatibility complex class II promoter binding protein affected in a hereditary defect in class II gene regulation. Proc. Natl. Acad. Sci. USA 86, 4200–4204 (1989).
Siegrist, C. A. et al. RFX1 is identical to enhancer factor C and functions as a transactivator of the hepatitis B virus enhancer. Mol. Cell. Biol. 13, 6375–6384 (1993).
Noshay, J. M., Anderson, S. N., Zhou, P., Ji, L. & Springer, N. M. Monitoring the interplay between transposable element families and DNA methylation in maize. PLOS Genet. 15, e1008291 (2019).
Gebert, D., Ketting, R. F., Zischler, H. & Rosenkranz, D. piRNAs from pig testis provide evidence for a conserved role of the piwi pathway in post-transcriptional gene regulation in mammals. PLoS ONE 10, e0124860 (2015).
Ha, H., Song, J., Wang, S., Kapusta, A. & Feschotte, C. A comprehensive analysis of piRNAs from adult human testis and their relationship with genes and mobile elements. Bmc Genom. 15, 545 (2014).
Jang, H. S. et al. Transposable elements drive widespread expression of oncogenes in human cancers. Nat. Genet 51, 611–617 (2019).
Moschetti, R., Palazzo, A., Lorusso, P., Viggiano, L. & Marsano, R. M. “What you need, baby, I got it”: transposable elements as suppliers of cis-operating sequences in Drosophila. Biol. Basel 9, 25 (2020).
Sundaram, V. & Wysocka, J. Transposable elements as a potent source of diverse cis-regulatory sequences in mammalian genomes. Philos. Trans. R. Soc. Lond. B Biol. Sci. 375, 20190347 (2020).
Li, Y. et al. A survey of transcriptome complexity in Sus scrofa using single-molecule long-read sequencing. DNA Res. 25, 421–437 (2018).
Beiki, H. et al. Improved annotation of the domestic pig genome through integration of Iso-Seq and RNA-seq data. BMC Genom. 20, 344 (2019).
Salmela, L. & Rivals, E. LoRDEC: accurate and efficient long read error correction. Bioinformatics 30, 3506–3514 (2014).
Chung, N. et al. Transcriptome analyses of tumor-adjacent somatic tissues reveal genes co-expressed with transposable elements. Mob. DNA 10, 39 (2019).
Goubert, C., Zevallos, N. A. & Feschotte, C. Contribution of unfixed transposable element insertions to human regulatory variation. Philos. Trans. R. Soc. Lond. B Biol. Sci. 375, 20190331 (2020).
Brogna, S. & Wen, J. Nonsense-mediated mRNA decay (NMD) mechanisms. Nat. Struct. Mol. Biol. 16, 107–113 (2009).
Lucas, B. A. et al. Evidence for convergent evolution of SINE-directed Staufen-mediated mRNA decay. Proc. Natl Acad. Sci. USA 115, 968–973 (2018).
Patro, R., Duggal, G., Love, M. I., Irizarry, R. A. & Kingsford, C. Salmon provides fast and bias-aware quantification of transcript expression. Nat. Methods 14, 417–419 (2017).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinform. 9, 559 (2008).
van Dongen, S. & Abreu-Goodger, C. Using MCL to extract clusters from networks. Methods Mol. Biol. 804, 281–295 (2012).
Zhu, Y. et al. Signatures of selection and interspecies introgression in the genome of Chinese domestic pigs. Genome Biol. Evol. 9, 2592–2603 (2017).
Bosse, M. et al. Artificial selection on introduced Asian haplotypes shaped the genetic architecture in European commercial pigs. Proc. Biol. Sci. 282, 20152019 (2015).
Chen, M. et al. Genome-wide detection of selection signatures in Chinese indigenous Laiwu pigs revealed candidate genes regulating fat deposition in muscle. BMC Genet. 19, 31 (2018).
Richardson, S. R. et al. The influence of LINE-1 and SINE retrotransposons on mammalian genomes. Microbiol Spectr. 3, MDNA3-0061–MDNA3-0062014 (2015).
Wang, L. & Jordan, I. K. Transposable element activity, genome regulation and human health. Curr. Opin. Genet. Dev. 49, 25–33 (2018).
Ai, H. et al. Adaptation and possible ancient interspecies introgression in pigs identified by whole-genome sequencing. Nat. Genet. 47, 217–225 (2015).
Foll, M. & Gaggiotti, O. A genome-scan method to identify selected loci appropriate for both dominant and codominant markers: a Bayesian perspective. Genetics 180, 977–993 (2008).
Godina, C. et al. Prognostic impact of tumor-specific insulin-like growth factor binding protein 7 (IGFBP7) levels in breast cancer: a prospective cohort study. Carcinogenesis 42, 1314–1325 (2021).
Wu, Q. et al. Linkage disequilibrium and functional analysis of PRE1 insertion together with SNPs in the promoter region of IGFBP7 gene in different pig breeds. J. Appl. Genet. 59, 231–241 (2018).
Wang, Z. et al. Single nucleotide polymorphism scanning and expression of the FRZB gene in pig populations. Gene 543, 198–203 (2014).
Zhou, W. N. et al. RUNX3 plays a tumor suppressor role by inhibiting cell migration, invasion and angiogenesis in oral squamous cell carcinoma. Oncol. Rep. 38, 2378–2386 (2017).
Fainaru, O. et al. Runx3 regulates mouse TGF-beta-mediated dendritic cell function and its absence results in airway inflammation. EMBO J. 23, 969–979 (2004).
Zhang, L. et al. Development and genome sequencing of a laboratory-inbred miniature pig facilitates study of human diabetic disease. iScience 19, 162–176 (2019).
Fu, Y. et al. A gene prioritization method based on a swine multi-omics knowledgebase and a deep learning model. Commun. Biol. 3, 502 (2020).
Barb, C. R., Hausman, G. J. & Czaja, K. Leptin: a metabolic signal affecting central regulation of reproduction in the pig. Domest. Anim. Endocrinol. 29, 186–192 (2005).
Chan, C. J. et al. BioID performed on Golgi enriched fractions identify C10orf76 as a GBF1 binding protein essential for Golgi maintenance and secretion. Mol. Cell Proteom. 18, 2285–2297 (2019).
Viterbo, V. S. et al. Genome wide association study of fatty acid composition in Duroc swine. Asian-Australas. J. Anim. Sci. 31, 1127–1133 (2018).
Westerberg, R. et al. ELOVL3 is an important component for early onset of lipid recruitment in brown adipose tissue. J. Biol. Chem. 281, 4958–4968 (2006).
Crespo-Piazuelo, D. et al. Identification of strong candidate genes for backfat and intramuscular fatty acid composition in three crosses based on the Iberian pig. Sci. Rep. 10, 13962 (2020).
Lorenzo, D. N. & Bennett, V. Cell-autonomous adiposity through increased cell surface GLUT4 due to ankyrin-B deficiency. Proc. Natl. Acad. Sci. USA 114, 12743–12748 (2017).
Xing, K. et al. Comparative adipose transcriptome analysis digs out genes related to fat deposition in two pig breeds. Sci. Rep. 9, 12925 (2019).
Claire D’Andre, H. et al. Identification and characterization of genes that control fat deposition in chickens. J. Anim. Sci. Biotechnol. 4, 43 (2013).
van Son, M. et al. A QTL for number of teats shows breed specific effects on number of vertebrae in pigs: bridging the gap between molecular and quantitative genetics. Front. Genet. 10, 272 (2019).
Miao, B. et al. Tissue-specific usage of transposable element-derived promoters in mouse development. Genome Biol. 21, 255 (2020).
Smith, C. E. L. et al. A tissue-specific promoter derived from a SINE retrotransposon drives biallelic expression of PLAGL1 in human lymphocytes. PLoS ONE 12, e0185678 (2017).
Jjingo, D. et al. Mammalian-wide interspersed repeat (MIR)-derived enhancers and the regulation of human gene expression. Mob. DNA 5, 14 (2014).
Hubley, R. et al. The Dfam database of repetitive DNA families. Nucleic Acids Res. 44, D81–D89 (2016).
Xiong, W., He, L., Lai, J., Dooner, H. K. & Du, C. HelitronScanner uncovers a large overlooked cache of Helitron transposons in many plant genomes. Proc. Natl. Acad. Sci. USA 111, 10263–10268 (2014).
Ellinghaus, D., Kurtz, S. & Willhoeft, U. LTRharvest, an efficient and flexible software for de novo detection of LTR retrotransposons. BMC Bioinform. 9, 18 (2008).
Xu, Z. & Wang, H. LTR_FINDER: an efficient tool for the prediction of full-length LTR retrotransposons. Nucleic Acids Res. 35, W265–W268 (2007).
Wenke, T. et al. Targeted identification of short interspersed nuclear element families shows their widespread existence and extreme heterogeneity in plant genomes. Plant Cell 23, 3117–3128 (2011).
Girgis, H. Z. Red: an intelligent, rapid, accurate tool for detecting repeats de-novo on the genomic scale. BMC Bioinform. 16, 227 (2015).
Gu, W., Castoe, T. A., Hedges, D. J., Batzer, M. A. & Pollock, D. D. Identification of repeat structure in large genomes using repeat probability clouds. Anal. Biochem. 380, 77–83 (2008).
Tarailo-Graovac, M. & Chen, N. Using RepeatMasker to identify repetitive elements in genomic sequences. Curr. Protoc. Bioinform. 4, 4 10 11–14 10 14 (2009).
Liu, G. E., Alkan, C., Jiang, L., Zhao, S. Y. & Eichler, E. E. Comparative analysis of Alu repeats in primate genomes. Genome Res. 19, 876–885 (2009).
Groenen, M. A. M. A decade of pig genome sequencing: a window on pig domestication and evolution. Genet Sel. Evol. 48, 23 (2016).
Tajaddod, M. et al. Transcriptome-wide effects of inverted SINEs on gene expression and their impact on RNA polymerase II activity. Genome Biol. 17, 220 (2016).
Li, W. & Godzik, A. Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22, 1658–1659 (2006).
Katoh, K. & Standley, D. M. A simple method to control over-alignment in the MAFFT multiple sequence alignment program. Bioinformatics 32, 1933–1942 (2016).
Minh, B. Q. et al. IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol. Biol. Evol. 37, 1530–1534 (2020).
Subramanian, B., Gao, S., Lercher, M. J., Hu, S. & Chen, W. H. Evolview v3: a webserver for visualization, annotation, and management of phylogenetic trees. Nucleic Acids Res. 47, W270–W275 (2019).
Price, A. L., Eskin, E. & Pevzner, P. A. Whole-genome analysis of Alu repeat elements reveals complex evolutionary history. Genome Res. 14, 2245–2252 (2004).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
Chen, K. et al. DANPOS: dynamic analysis of nucleosome position and occupancy by sequencing. Genome Res. 23, 341–351 (2013).
Liu, T. Use model-based Analysis of ChIP-Seq (MACS) to analyze short reads generated by sequencing protein-DNA interactions in embryonic stem cells. Methods Mol. Biol. 1150, 81–95 (2014).
Ramirez, F., Dundar, F., Diehl, S., Gruning, B. A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187–W191 (2014).
Grant, C. E., Bailey, T. L. & Noble, W. S. FIMO: scanning for occurrences of a given motif. Bioinformatics 27, 1017–1018 (2011).
Bailey, T. L., Johnson, J., Grant, C. E. & Noble, W. S. The MEME suite. Nucleic Acids Res. 43, W39–W49 (2015).
Mathelier, A. et al. JASPAR 2016: a major expansion and update of the open-access database of transcription factor binding profiles. Nucleic Acids Res. 44, D110–D115 (2016).
Chen, S., Zhou, Y., Chen, Y. & Gu, J. fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34, i884–i890 (2018).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Tarasov, A., Vilella, A. J., Cuppen, E., Nijman, I. J. & Prins, P. Sambamba: fast processing of NGS alignment formats. Bioinformatics 31, 2032–2034 (2015).
Zang, C. et al. A clustering approach for identification of enriched domains from histone modification ChIP-Seq data. Bioinformatics 25, 1952–1958 (2009).
Krueger, F. & Andrews, S. R. Bismark: a flexible aligner and methylation caller for Bisulfite-Seq applications. Bioinformatics 27, 1571–1572 (2011).
Zhou, Q., Lim, J. Q., Sung, W. K. & Li, G. An integrated package for bisulfite DNA methylation data analysis with Indel-sensitive mapping. BMC Bioinform. 20, 47 (2019).
Chen, C. et al. TBtools: an integrative toolkit developed for interactive analyses of big biological data. Mol. Plant 13, 1194–1202 (2020).
Friedlander, M. R., Mackowiak, S. D., Li, N., Chen, W. & Rajewsky, N. miRDeep2 accurately identifies known and hundreds of novel microRNA genes in seven animal clades. Nucleic Acids Res. 40, 37–52 (2012).
Wang, J. et al. piRBase: a comprehensive database of piRNA sequences. Nucleic Acids Res. 47, D175–D180 (2019).
Law, C. W., Chen, Y., Shi, W. & Smyth, G. K. voom: Precision weights unlock linear model analysis tools for RNA-seq read counts. Genome Biol. 15, R29 (2014).
Karnovsky, A. et al. Metscape 2 bioinformatics tool for the analysis and visualization of metabolomics and gene expression data. Bioinformatics 28, 373–380 (2012).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Santander, C. G. et al. STEAK: a specific tool for transposable elements and retrovirus detection in high-throughput sequencing data. Virus Evol. 3, vex023 (2017).
Rajaby, R. & Sung, W. K. TranSurVeyor: an improved database-free algorithm for finding non-reference transpositions in high-throughput sequencing data. Nucleic Acids Res. 46, e122 (2018).
Gardner, E. J. et al. The Mobile Element Locator Tool (MELT): population-scale mobile element discovery and biology. Genome Res. 27, 1916–1929 (2017).
Keane, T. M., Wong, K. & Adams, D. J. RetroSeq: transposable element discovery from next-generation sequencing data. Bioinformatics 29, 389–390 (2013).
Sedlazeck, F. J. et al. Accurate detection of complex structural variations using single-molecule sequencing. Nat. Methods 15, 461–46 (2018).
Wang, K., Li, M. Y. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Lee, T. H., Guo, H., Wang, X., Kim, C. & Paterson, A. H. SNPhylo: a pipeline to construct a phylogenetic tree from huge SNP data. BMC Genom. 15, 162 (2014).
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664 (2009).
Li, Y. L. & Liu, J. X. StructureSelector: a web-based software to select and visualize the optimal number of clusters using multiple methods. Mol. Ecol. Resour. 18, 176–177 (2018).
Reimer, C. et al. Assessing breed integrity of Gottingen Minipigs. BMC Genom. 21, 308 (2020).
Keel, B. N., Nonneman, D. J., Lindholm-Perry, A. K., Oliver, W. T. & Rohrer, G. A. Porcine single nucleotide polymorphisms and their functional effect: an update. BMC Res. Notes 11, 860 (2018).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Barrett, J. C., Fry, B., Maller, J. & Daly, M. J. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21, 263–265 (2005).
Dong, S. S. et al. LDBlockShow: a fast and convenient tool for visualizing linkage disequilibrium and haplotype blocks based on variant call format files. Brief. Bioinformatics 22, bbaa227 (2021).
Gel, B. et al. regioneR: an R/Bioconductor package for the association analysis of genomic regions based on permutation tests. Bioinformatics 32, 289–291 (2016).
Zhao, P. Source data for young SINEs in pig genomes impact gene regulation, genetic diversity, and complex traits. Figshare https://doi.org/10.6084/m9.figshare.23856213.v4 (2023).
Acknowledgements
This work is financially supported by the Project of Sanya Yazhou Bay Science and Technology City (SKJC-2020-02-007), the National Natural Science Foundation of China (31941007), the Science and Technology special fund of Hainan Province (ZDYF2022XDNY237), the AFRI grant numbers 2019-67015-29321 and 2021-67015-33409 from the USDA National Institute of Food and Agriculture (NIFA), and the High-performance Computing Platform of YZBSTCACC. We thank Sylvain Foissac from Université de Toulouse and other members associated with the FRAGENCODE project for their contributions to data of chromatin accessibility used in this manuscript.
Author information
Authors and Affiliations
Contributions
P.Z., Z.W., L.F., and G.L. conceived and designed the experiments. P.Z. designed analytical strategy and performed all analysis process. Y.G. assisted the DNA methylation analysis. Z.P. assisted the Histone modifications analysis. L.L. and X.L. assisted the bioinformatics analysis process. L.G., H.Z., X.H., and L.Q. collected and prepared for sequencing data. P.Z., G.L., L.F., Z.W., H.Z., and D.Y. wrote and revised the paper. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Clément Goubert, Miguel Branco, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: George Inglis. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhao, P., Gu, L., Gao, Y. et al. Young SINEs in pig genomes impact gene regulation, genetic diversity, and complex traits. Commun Biol 6, 894 (2023). https://doi.org/10.1038/s42003-023-05234-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-023-05234-x
- Springer Nature Limited