Abstract
The widespread use of rodents in neuroscience has prompted the development of optimized viral variants for transduction of brain cells, in vivo. However, many of the viruses developed are less efficient in other model organisms, with birds being among the most resistant to transduction by current viral tools. Resultantly, the use of genetically-encoded tools and methods in avian species is markedly lower than in rodents; likely holding the field back. We sought to bridge this gap by developing custom viruses towards the transduction of brain cells of the Japanese quail. We first develop a protocol for culturing primary neurons and glia from quail embryos, followed by characterization of cultures via immunostaining, single cell mRNA sequencing, patch clamp electrophysiology and calcium imaging. We then leveraged the cultures for the rapid screening of various viruses, only to find that all yielded poor to no infection of cells in vitro. However, few infected neurons were obtained by AAV1 and AAV2. Scrutiny of the sequence of the AAV receptor found in quails led us to rationally design a custom-made AAV variant (AAV1-T593K; AAV1*) that exhibits improved transduction efficiencies in vitro and in vivo (14- and five-fold, respectively). Together, we present unique culturing method, transcriptomic profiles of quail’s brain cells and a custom-tailored AAV1 for transduction of quail neurons in vitro and in vivo.
Similar content being viewed by others
Introduction
Avian species are a powerful experimental organism in neuroscience (e.g., refs. 1,2,3,4,5,6,7). Birds display sophisticated cognitive capabilities and specialized behaviors, such as long-distance navigation8, imprinting, homing, food-caching, song learning, etc.9,10. Interestingly, these complex cognitive capabilities are presented by birds despite their diverging neuroarchitecture and, at times, suggested lower neuronal densities in comparison to mammals4,11. These thereby provide unique opportunities for comparative studies of cellular mechanisms leading to behavior3. Indeed, these capabilities and behaviors are behind the increased interest in avian neuroscience in recent years (e.g.,1,2,7,9,12,13,14,15,16,17,18,19,20).
Of the various avian species commonly employed today, we focused our attention on the domestic Japanese quail (Coturnix japonica). Quails are relatively small animals and therefore require smaller housing and animal facilities, reach sexual maturity quickly (significantly shorter than mice and chicken) and female quails lay approximately one egg per day; ideal for routine experimentations15. A particular benefit of this model over other birds is their ground-dwelling nature, which may simplify various behavioral studies (such as spatial navigation20) by reducing dimensionalities (e.g., ref. 21). Despite the latter, (and despite their extensive use in developmental biology15,22,23), quails are rarely used in neuroscience6,20. In part, we deem this to result from a lack of tools and methods for monitoring and examining neurons in quails in vivo21. More precisely, whereas mammalian neuroscience heavily relies on viral tools for the delivery of genetic optical probes for interrogating the brain (e.g., GCaMP and Channelrhodopsin; ChR)24,25, the use of viruses in quails has been primarily, if not exclusively, employed for transgenesis (and almost exclusively by use retroviruses, Lentivirus and MoMLV)6,15,26,28,29,30,31,31. Thus, whether transduction of other cell types at different developmental stages (including the adult animal) can be obtained by lentivirus or other viruses (e.g., adeno-associated virus; AAV) remains unknown15. Notably, the shortage of viral tools is not limited to the quail model; rather is a recurring theme in the avian field (refs. 32,34,35,35, but see below).
We sought to bridge this gap by screening for suitable viral vectors for the transduction of neurons of Japanese quails. To screen for numerous viral candidates, we develop a protocol for culturing primary neuronal cultures from quail embryos. Systematic characterization of cells in cultures demonstrates that the cultures are viable and contain over 11 different cellular populations, including two populations of mature neurons. We use the cultures for screening of multiple viruses commonly used in rodents and other avian species, although find that none yield efficient transduction, especially not as obtained in rodents. We then revert to rational engineering of viral AAV-capsids, which leads to the development of a single infectious AAV1 variant (AAV1-T593K; AAV1*) that exhibits significantly improved infection efficiency in vitro (eightfold). This variant also proved suitable for the transduction of primary chicken cultures. Lastly, and importantly, we found that AAV1* yields an approximately fivefold improvement of transduction of quail neurons in vivo.
Results
AAV1 poorly infects quail neurons in vivo
It is well appreciated in the avian field that viral transduction of neurons in vivo is challenging32. However, a handful of recent reports show partial success following the use of common recombinant adeno-associated viruses, for instance, AAV1 in pigeons32, barn owls14 and zebra finches36,37 (and other serotypes in canaries7, finches38, and more39,40). Thus, we first examined whether AAV1 would also be suitable for the transduction of neurons of Japanese quails in vivo. We produced and injected YFP-expressing AAV1 (see viral titers and injection procedures in Methods) into the Wulst41 of two months old quails, followed by cryosectioning and fluorescence imaging of brain slices (without immunostaining for YFP, see Methods) over the course of three to 8 weeks. Unfortunately, we found weak expression in one animal (and only at 7 weeks post viral injection) (Supplementary Fig. 1). The poor infection efficiency and extreme variability between the different animals could not be explained by factors such as different virus batches or virus viability, as each virus (whether made in-house or purchased) was validated on cultured cells (including primary rodent neurons). Of note, although it is plausible that detection of YFP-fluorescence could have been enhanced by immunostaining for YFP, we were not inclined to do so as our intentions were to obtain sufficient expression that would also be suitable for in vivo imaging. Thus, these results deterred us from continuing our explorations with added viral serotypes in the same manner.
Development and characterization of primary cultures from quail embryos
To rapidly screen multiple different viruses, we sought to examine in vitro transduction of cultured cells. To produce primary neuronal cultures, we explored common culturing protocols from rodents and chickens (we found no mentions for quails)33,42,43. Briefly, we dissected forebrains from seven to nine days in ovo (DIO) embryos or post-hatched chicks, mechanically dissociated and enzymatically digested the tissue, followed by plating of cells onto poly-D-lysine (PDL) covered glass coverslips; grown in standard growth media and incubator conditions (see Methods and Supplementary Table 1). Our rationale behind this staging was based on reports suggesting that neurogenesis should be completed by seven DIO44,45, thereby yielding neuronal cultures with low amounts of mature glial cells46. During the first hours after plating, we could distinguish cell bodies, some showing extending neurite-like processes (Supplementary Fig. 2a, arrowheads), but only in cultures produced from embryos. Cultures from post-hatched chicks did not yield viable cultures. Nevertheless, embryonic cultures progressively waned, and most cells died past ~four days in vitro (DIV) (Supplementary Fig. 2b, c). We next examined growing the cultures directly on the tissue culture plate itself, as well as substituted the MEM-based growth medium (frequently employed in rodent and chicken culturing protocols33,46) to an enriched Neurobasal-medium (as employed for rodent organotypic slices47) (Supplementary Table 1). These collective modifications allowed the cultures to thrive, particularly cultures produced from 9 DIO embryos, and these remained viable past three weeks (Fig. 1a, b). Cultures produced from an entire hemisphere (plated onto 60 mm culture plates) yielded a rich network of highly interconnected and branched cells, most of which (>90%) showed positive NeuN48- and NeuroTrace-staining49,50 (Fig. 1c and Supplementary Fig. 2d), although a much smaller fraction of cells displayed mature neuronal morphology (Fig. 1b, arrowheads). MAP2-staining indicated a lower estimation (~18%) of mature neurons in the culture (Fig. 1d) (discussed below).
a Micrographs of cultures produced from 7, 8, or 9-day-old embryos (days in OVO, DIO) (rows), grown for 7, 14, and 21 days in vitro (DIV) (column). b Cultures remained viable for up to 30 days (arrowheads represent examples of viable cells). c Seven DIV cultures (produced from 9 DIO embryos) co-stained with DAPI (yellow) and NeuroTrace (pink) suggest that the majority (~90%) of cells (inset: pink arrowheads) represent neurons, with a smaller fraction (10%) of non-neuronal cells, summarized in the right most panel. d MAP2-staining of cultures suggests that ~18% of cultured cells are neurons (summarized in the pie chart).
To assess the culture’s viability and to confirm the presence of neurons in the culture, we patched cells with neuronal morphology (e.g., Fig. 1d, inset) and, indeed, these exhibited robust action potential firing (under current-clamp) and, under voltage-clamp conditions, displayed prototypical barrages of excitatory postsynaptic currents (EPSCs) (Fig. 2a). We found that several electrophysiological features of quail neurons are highly similar to those of age-matched cultured neurons produced from mouse embryos (Fig. 2b and Supplementary Fig. 3a), albeit exhibit significantly lower intrinsic excitability (Fig. 2c, d and Supplementary Fig. 3b, c). Thus, our culturing protocol yields viable cultures with various cell types; however—and importantly—the various staining techniques suggest that the cultures include ~20% mature neurons.
a Representative current-clamp (top) and voltage-clamp (bottom) recordings from cultured neurons (left inset—micrograph of the patched cell). The current-clamp recording shows the resting membrane potential of the neuron (−80 mV), excitatory postsynaptic potential (EPSPs), and action potential firing (e.g., dashed rectangle and inset). During voltage-clamp, the cell was clamped at −70 mV, during which barrages of excitatory postsynaptic currents (EPSCs) could be distinguished. b Summary of electrical properties of quail neurons in vitro (n = 12). c Assessment of intrinsic neuronal excitability. Representative voltage traces (top) from a single neuron in response to current injection (bottom protocol) from which we could deduce the maximal firing rates, summarized in d. Data are presented as mean ± SEM. Color-coded traces correspond to colored data points on the plot (For the complete dataset, see Supplementary Fig. 3c).
Single-cell mRNA sequencing of embryonic quail cultures
To examine the different cell types obtained in culture, we employed single-cell mRNA sequencing (sc.mRNA seq.51) of cultured cells produced from 9 DIO embryos (grown for 7 DIV; Methods52,53,54). Unsupervised clustering of the data (based on the significant top ten or top 200 differentially expressed-genes; DEGs) acquired from ~10,000 cultured cells revealed 15 noticeable clusters of cell types (clusters #0–14); all clusters exhibiting a similar (and high) number of unique molecular identifiers (UMI or nCount) and gene counts (nFeature) (Supplementary Fig. 4a–d and Fig. 3a, b). Importantly, all cells exhibited an acceptable mitochondrial genes ratio (10–20%)55,56, demonstrating the viability of cells in cultures, and these were on par with sequencing results obtained from the brains of chicken embryos45 (Supplementary Fig. 4a, b).
a UMAP plot of 9561 cells, taken from cultures produced from an embryo’s entire hemisphere, classified based on the top ten most differentially expressed genes (DEGs), as shown in the heatmap in (b). The number of cells within each cluster are noted at the left corner of UMAP and clusters’ colors correspond to column colors in b. Red annotated genes (in b) were manually curated (i.e., identity was not automatically found in reference genomes). c Cell type identification dotplot shows mean expression levels (blue to red) and percent of cells (size of data point) in each cluster and for select markers (bottom axis). Red annotated genes were manually curated. d Feature plot showing expression of SFRP1. Mean expression (Log2 fold change) and percent of cells for each cluster are depicted below the plot.
Before proceeding to cell annotations, many transcripts had to be manually curated (e.g., Fig. 3b, red annotations), although dozens of other genes could not be identified despite multiple reference genomes (e.g., ensembl.org), including the recently published (though partially-sequenced) quail genome57. Identification and classification of cell type was done by querying multiple cell-type specific or enriched gene markers, as previously described58,59,60 (along online atlases, e.g., proteinatlas.org). The largest group of cells (2400 cells) displayed a broad and dispersive expression of most top DEGs (Fig. 3a, b—light brown; subclusters are shown in Supplementary Fig. 4c–e). This cluster (Cluster #0, Supplementary Fig. 4d) showed a combination of neuronal (STMN2, CDH2) and glial markers (FABP7, PROM1, and SLC1A3), but also moderate expression of NES (neuroepithelial marker) and VIM (radial glia marker), representing very early Radial glial cells expressing early neuronal markers; likely a type of neural committed progenitor cells that precede neurons, such as pro-neuronal radial glia (Fig. 3c, see summary in Supplementary Fig. Table 2)61,62,63. This observation is supported by our NeuroTrace and NeuN-staining results suggesting the very high abundance of seemingly neuronal cells (see Fig. 1 and Supplementary Fig. 2)64, and by the presence of two additional (and different) bona fide radial glia populations in the cultures that are distinguished from this cluster (Fig. 3c). Cluster #1 could be best classified as apical radial glia owing to PROM1 expression65 (along other glial markers, namely FABP7 and SLC1A3), whereas cluster #2 shows the expression of NES and dispersive expression of added markers (reminiscent of cluster #0), suggesting a slightly earlier differentiation state of non-apical radial glia65. We could easily distinguish two mature neuronal populations, both expressing specific pan-neuronal markers STMN2, SYT1 (Fig. 3c, clusters 6 and 7), as well as MAP2 and NeuN (RBFOX3) (Supplementary Fig. 4f). The inhibitory neuronal population exclusively expressed GAD1 and -2, DLX1 and -2, and moderate levels of SOX11, whereas excitatory neurons showed strong expression of GABBR2, GRIN (1 and 2B), and EPN2 and, uniquely, SLC17A6. Interestingly, NEUROD6 (a glutamatergic neuron marker66,67) was enriched in this cluster, but was also found at lower levels in inhibitory neurons (Fig. 3c). This transcriptome signature is less common in mammals (e.g., refs. 59,68,69). We did not detect other neuron types in this brain region. We further found three distinct microglia populations (clusters #5, 8, and 13, Supplementary Fig. 4d, e); one cluster (cluster #13, n = 130 cells) showing added proliferation markers, explicitly HMGB2, CENPF, TOP2A, and CCNB270,71. These imply that cluster #13 represents dividing microglia (Supplementary Fig. 4e). These are consistent with reports showing the complex heterogeneity in microglia populations, especially during early developmental stages compared with the adult72,73. The smallest cluster of cells (#14, n = 125 cells) is distinguished as oligodendrocyte precursor cells owing to the expression of PTPRZ1, PMP2, and PDGFRA (Fig. 3c, PDGFRA; manually annotated). Cluster #10 lies at the interface between clusters 0, 4, and 14, with neuronal progenitor cells (NPC) markers and additional division markers as seen in cluster 13, specifically HMGB2 and TOP2A70,71, therefore depicting a non-quiescent (i.e., dividing) neuronal progenitor stem cell population. We also find that the cultures contain ~14% fibroblasts (clusters # 3, 9, 11; n = 1415 cells) by the variable expression of COL1A174, arising from remaining meninges and perivascular system75. Interestingly, one fibroblast population (#11) showed a very weak expression of neuronal markers (e.g., SOX11 and ENC1) (subcluster shown in Supplementary Fig. 4d, e). This is a unique scenario for which we could not find any mention in the literature. In fact, the inclusion of SOX11 (a transcription factor) in fibroblasts (postnatal skin fibroblasts) is used to differentiate the cells into cholinergic neurons76. Identification of clusters #12 and 4 proved more challenging. Cluster #4 (residing adjacent to clusters 0–2) expresses RGs markers (SLC1A3, VIM, NES, PROM1; similar to cluster #1), along with expression of MFGE8, GJA1, and ID3; astrocytic markers in the mouse60,77. However, this cluster (or any other) presented no other astrocytic markers, notably GFAP (Supplementary Fig. 5a)78. This deficiency was further confirmed by the lack of immunostaining for the protein, whereas control rat cultures showed extensive labeling (Supplementary Fig. 5b, left and middle panels, c). Furthermore, cluster #4 shows weak, albeit significant, expression of RELN (neuronal marker). This unexpected transcriptome signature led us to hypothesize that this population represents an earlier differentiation state than RGs, namely RG-Progenitors (RGPs)79,80. Indeed, ~65% of cells within cluster #4 showed enriched levels of SFRP1; a prominent marker for RGP (Fig. 3d). Lastly, cluster #12 (n = 135 cells) shows strikingly overlapping transcriptional signatures with cluster #0, but with higher expression of inhibitory neuronal markers (SOX11 and ENC1), though with no other GABAergic markers. Its location between clusters # 0, 1, and 6 suggests it to represent a slightly more NPC-RG differentiated state toward inhibitory neurons81. Together, we find that the quail embryonic cultures provide a very rich repertoire of cell types82, ideal for studying stem cells of the brain. Notably, despite the embryonic origin of our cultures, we find very low amounts (and in very few cells scattered across all clusters) of SOX2 and PAX6—markers of very early developmental stages as reported for the chicken embryo (Supplementary Fig. 6)45 (and see discussion).
Viral screening in vitro and rationale evolution of viral capsid to infect quail brain cells
We proceeded to screen for transduction of primary quail cultures by various viruses. We focused on common adeno-associated viruses currently in use: AAV1, AAV2 (and variants), AAV9, AAV-PhP-B83, Baculovirus (baculo.)84, lentivirus (lenti.)26,27, and an avian AAV (A3V)33. We produced YFP-expressing viruses (AAVs, A3V and lenti.) and purchased a commercial baculovirus expressing GCaMP6 (genetically-encoded calcium indicator85) (see Methods). We infected the cultures in parallel to cultured mammalian cells (HEK293t cells), followed by an assessment of fluorescence after three DIV (Fig. 4a). Whereas all viruses robustly infected and yielded expression in mammalian cells (except A3V, which showed no infection of HEK293t, as described earlier33), we observed very sparse infection and weak expression in quail cells. Of note, the lack of infectivity of our cultures by A3V is surprising because of its reported infectivity of chicken brain cells33. We do not know the reason behind these observations; however, following trials with three different viral batches (DNAs were kindly provided by its developers, see Acknowledgements), we opted to stop pursuing this variant. Nevertheless, and importantly, a few positively-infected neurons (assessed by morphology) were observed following infection by AAVs, specifically AAV1, AAV2-SL1, and AAV-PhP-B and by lenti., though to a slightly lesser degree (Fig. 4). Baculovirus infection was highly toxic to cultured cells (Fig. 4, bottom row).
Various AAV serotypes, lentivirus, and baculovirus show very limited transduction efficacy of quail cultures. Infection efficacy was dually assessed on cultured cells (HEK293T cells, top lines) and quail cultures (bottom lines). AAVs (including A3V) express CAG-eYFP, Baculovirus expresses CAG-jGCaMP7f, and the lentivirus expresses CMV-eGFP. The strong expression of fluorescent markers within HEK293T cells (except for A3V, which showed no infection of HEK293T cells83) demonstrates the viability of the viruses used. Despite the latter, very few infected cells (mostly neurons) were detected in quail cultures. Note the cytotoxicity of the baculovirus.
These low infection efficiencies motivated us to try to tailor these viral vectors for quail cells. To do so, we first examined the entry routes of AAVs into mammalian cells and noted that most AAVs do so by binding to non-specific proteoglycans at the membranes of cells86. However, AAV1 and AAV2 (and AAV5) also require the AAV-receptor (AAVR, also known as Dyslexia-associated KIAA0319-like protein) as a co-receptor for entry into cells87,88,89. Fortunately, our sc.mRNA seq. data revealed that our cultures express this transcript (at least to the method’s detection limit), especially in microglia and neurons (Fig. 5a). The AAVR contains five polycystic kidney disease (PKD) repeat domains, from which PKD2 is recognized by AAV1 and AAV2 (Fig. 5b)87,88. Sequences of the PKD2 domain from various species show very high homology (Fig. 5c), though the very few differences between the sequences of quail and mammalian AAVR (qAAVR and mAAVR, respectively) occur specifically in residues that are essential for the interactions with AAV1 and AAV2, namely Q432 and K464 (mAAVR numbering) (Fig. 5b, inset, c and Supplementary Table 3). Of note, the divergent residues in qAAVR do not appear to be completely random, but rather mirror the residues found in the viral capsid. For instance, whereas mAAVR contains a lysine (K464) that interacts with a threonine (T593) found in capsids of AAV1 protomers (Fig. 5b, green and magenta, respectively), the qAAVR has a threonine (T204) instead. Similarly, while the mAAVR contains a glutamine (Q432) that interacts with an arginine of the AAV2 capsid, the qAAVR mirrors AAV2’s capsid with an arginine of its own (R172). We thereby hypothesized that the mirroring residues (T-T for AAV1 and R-R for AAV2) may disturb the interaction between qAAVR and AAV1 and -2 viral capsids. To restore these interactions, we rationally mutated the residues in the capsids of AAV1 or AAV2 to match the residues found in the mammalian AAVR (instead of modifying the receptor, which would require transgenesis). Thus, this limited our mutagenesis towards residues that solely contain one interacting partner (Supplementary Table 3). We thereby produced AAV1 with a single mutation (T593K, denoted AAV1*) and two AAV2-variants (AAV2 and AAV2-SL190, an optimized AAV2 variant) bearing two mutations (R471Q and T592K), denoted AAV2* and AAV2-SL1*, respectively (see Supplementary Table 3 and Methods).
a Feature plot of KIAA0319L, the quail AAV-receptor (qAAVR). Mean expression (log2 fold change) is noted at the left of the plot and the percent of cells per each cluster are depicted below the plot. b (left) Cartoon illustration of the binding of the AAVR (gray transmembrane domain and PKD domains in lavender) by an AAV (magenta icosahedron). The PKD2 domain is highlighted (dark lavender) because of its essential role in the binding the AAV’s (AAV1 and AAV2) capsid86,89. (right) Atomic structure (by cryo-EM) of AAV1’s capsid (magenta) bound to PKD2 (lavender) (pdb: 6JCQ87). Key residues in PKD2 that interact with the capsid monomer are highlighted in yellow. The K464 residue (green) in PKD2 interacts with residue T593 (pink) found in the capsid of a second AAV1 protomer. Inset: binding of the receptor (three yellow highlights) by multiple AAV capsid monomers (magenta) shows the very close proximity between K464 (green) and T593 (pink). c PKD2’s protein sequence alignment between different species shows high conservation (lavender). Residues in the AAVR that directly interact with the AAV1 capsid are highlighted (bold yellow). Two essential residues differ between the quail and mammalian receptor (qAAVR and mAAVR, respectively)—R432 and K464 (bold red). Chicken AAVR (chAAVR).
We initially infected primary rat neuronal cultures with AAV1* and found that infection efficacy was diminished approximately twofold (Supplementary Fig. 7a, b), supporting the importance of this residue for AAV’s ability to infect mammalian cells87. We then infected the quail cultures with AAV1* and by AAV2* and AAV2-SL1* and found that, whereas the AAV2-variants yielded no infection of cells in multiple experiments (Supplementary Fig. 7c–e), AAV1* exhibited an order of magnitude (~14 fold) improvement in transfection efficiency of cultured cells (Fig. 6a). Moreover, AAV1*-infected cells showed approximately twofold brighter YFP-fluorescence (Fig. 6a, b). We patched YFP-positive cells with neuronal morphology (Fig. 6c, inset) and found that these exhibited very prominent action potential firing, robust synaptic activity, and expected resting membrane potential, as shown above (Fig. 6c, traces and see Fig. 2), whereas cells with non-neuronal morphology (Fig. 6d, inset) did not fire action potentials, but maintained a healthy and hyperpolarized resting membrane potential, and displayed some electrical activities, reminiscent of glia cells (Fig. 6d, black traces) (e.g., refs. 91,92,93). These demonstrate that AAV1* infected both neurons and non-neuronal cells (whether glia or neuronal precursors) and—importantly—infection did not induce any detectable cytotoxic effects (attested by intact morphology and electrical properties), unlike infection by baculovirus (see Fig. 4).
a Micrographs of quail cultures infected by AAV1* (left) or by AAV1 (right) (both of identical titer—3.3 × 1012). The amount of YFP-positive cells (infected) and fluorescence intensity are summarized in (b). b AAV1* infected significantly more cells compared to AAV1 by the same titer –AAV1* (Mean ± SEM: 396.5 ± 9.5, AAV1 – 27.25 ± 5.7, T-test, p < 0.0001). Fluorescence intensity is significantly higher in cells infected with AAV1* compared to AAV1 (AAV1* - 48.7 ± 1.66, AAV1 – 24.06 ± 1.02, T-test, p < 0.0001). c, d Electrophysiological recordings of AAV1*-infected cells (i.e., expressing YFP). c- Recording from a mature neuron (inset) reveals excitatory postsynaptic currents (top trace) and action potential firing and plateaus (bottom trace, n = 5), whereas amorphic cells (d, inset) do not fire (non-firing cells) and exhibit very health and hyperpolarized resting membrane potentials (n = 4). Small membrane activity could be easily seen, reminiscent of glial activities. e Micrograph showing cultures infected by AAV1*-CAG-GCaMP6m and two representative cells showing propagating calcium activities (right). f Calcium-dynamics divided into three subgroups: oscillatory (top trace), spiking (middle), and long-lasting calcium plateaus (bottom), summarized in the heatmap in g (N = 5, n = 64). Data were presented as mean ± SEM, p values are indicated, and noted by *p < 0.05; **p < 0.01; ***p < 0.001; ****p < 0.0001.
We then packaged GCaMP6m94 within AAV1* and infected 9 DIO cultures (grown for a week in vitro) to assess calcium activity in cultured cells. AAV1* transduction of GCaMP in these cells was not cytotoxic, and calcium activity could be easily monitored in many cells (Fig. 6e). We could distinguish a variety of different calcium-dynamics, such as slow and long-lasting calcium waves (reminiscent of glial activity95,96) and very sharp and transient ca2+-spikes, likely action potential firing (Fig. 6f, g). Together, our results demonstrate the positive, though not exclusive86, role of the pairing residues between AAVR and AAV1 for the infection of cells in vitro87. Importantly, AAV1* shows improved infection efficiency of quail cells in vitro and is highly suitable for delivering optical probes, such as GCaMP6m, for monitoring activity from various cell types in culture.
AAV1* infects chicken neurons
Quails are close relatives of chickens (Gallus gallus), which are a more widely used experimental model. We, therefore, sought to examine the infection efficiency of AAV1* of chicken neurons. As noted above, chicken AAVR (chAAVR) only shows one divergent residue from mammalian AAVR (T464 instead of K464) (Fig. 5c). We first applied our culturing method to chicken embryos and found our method to be similarly suitable for this species, extending the usability of our protocol towards primary chicken cultures (Supplementary Fig. 8a). We infected chicken cultures with AAV1, AAV1*, AAV2*, and AAV2-SL1*. AAV1 showed higher transfection efficiency of cells in chicken cultures, and as seen in quail cultures, AAV1* significantly increased infection efficiency (Supplementary Fig. 8a–c, AAV1*: 79 ± 16.7, AAV1: 42.7±4.4 cells/frame; p = 0.055, T-test). AAV2-variants were non-infectious (Supplementary Fig. 8d–f).
In vivo AAV1* transduction of neurons in young and adult quails
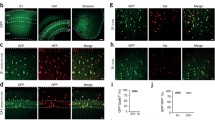
To assess AAV1* transduction in vivo, we injected AAV1* encoding CAG-eYFP into the Wulst of five young (4–5 weeks old) quails and, in parallel, injected AAV1 encoding CAG-eYFP into the Wulst of five aged-matched quails. All injections were at similar depths, volumes, and titers (see Supplementary Table 5). After 7 to 8 weeks (waiting periods were based on previous reports14,18,32,36,37 and see Fig. 1), brains were removed, fixed, and sectioned (Methods). Consistent with our in vitro results, AAV1* showed significantly higher infection efficiency (~5-fold) compared to AAV1 (Fig. 7a–c; AAV*—509.2 ± 107.04, AAV1—101.9 ± 18.4 cells/mm2, p = 0.005, T-test). This side-by-side comparison in young quails supports our initial observations following injections of AAV1 into adult quails’ brains, in which instances we only found one animal to express YFP (Supplementary Fig. 1). Note that the highest density observed in animals injected with AAV1 was equivalent to the lowest density obtained by AAV1* (Fig. 7c, compare with all examples provided in Supplementary Fig. 9). Infected cells also tended to exhibit higher YFP-fluorescence compared to AAV1, though this result did not reach significance (Fig. 7c, AAV1*—207.8 ± 5.9, AAV1—173.4 ± 12.04 a.u.). AAV1* also showed better dispersion than AAV1 (Supplementary Fig. 8 and Supplementary Table 4). Most infected cells appeared to be neurons (assessed by morphology and features such as spines), despite the ubiquitous promoter employed (CAG97) and the pan-tropism observed for AAV1* in culture (see Fig. 6). Very few cells appeared glial (Fig. 7 and Supplementary Figs. 9,10, arrowheads). Lastly, we also observed that injections of AAV1* yielded expression in three adults (2 months old) quails, whereas AAV1 showed expression in only one animal (Supplementary Table 4 and Supplementary Fig. 9). Together, these observations demonstrate that AAV1* significantly outperforms AAV1 for delivery of genetically-encoded tools to neurons of the Japanese quails in vivo, thereby making it—to the best of our knowledge—the first tailored-AAV for this bird species.
a Coronal brain slices of young quails (4 weeks old), produced after 7 weeks from animals injected, side-by-side, with 1 μl of AAV1*.or AAV1 (b); expressing CAG-eYFP. c AAV1* provides a higher density of infected cells (mostly neurons, top)- AAV1* (Mean ± SEM: 509.2 ± 107.04, AAV1—101.9 ± 18.4, T-test, p < 0.001). AAV1*-infected neurons also show a tendency for higher expression within each cells (bottom)- AAV1* - 207.8 ± 5.9, AAV1—173.4 ± 12.04. Each data point represents one animal (n = 5). The complete data set is provided in Supplementary Fig. 9. Note that the highest density observed in animals injected with AAV1 is equivalent to the lowest density obtained by AAV1* (c, top). Data were presented as mean ± SEM, p values are indicated, and noted by *p < 0.05; **p < 0.01; ***p < 0.001, n.s. non-significant.
Discussion
Avian species provide unique opportunities for studying the brain (e.g., refs. 2,7,13,19). The Japanese quail, in particular, presents several advantages over other avian species currently in use, for instance, faster sexual maturity, ground dwelling, and a high reproductive rate15. We have recently used this model to explore spatial navigation by use of electrodes, only to find that quails contain head-direction cells, as in other mammalian species, but we could not detect place or border cells in their hippocampal formation20. This is striking, as spatially-modulated cells are at the core of mammalian navigation and have also been recently described in another species of a food-caching bird98. We thereby suggest that this model provides distinctive opportunities to elucidate how the brain achieves spatial specialization across different species99. Despite the latter, genetic and molecular tools15 (or transgenic lines6,23,26,27,28) for interrogating the brains of quails are much less abundant or completely absent18,31,100. These are behind the motivation to concentrate our efforts on the design of viral vectors for the transduction of quail brain cells.
Scrutiny of the literature reveals that the use of viruses for brain research in avian models is relatively uncommon7,14,32,33,34,35,37, especially in comparison to mammalian models (e.g., refs. 83,101). In the case of quails, there are even fewer descriptions, and in most, if not all, instances, retroviruses were employed for transgenesis28,31,102,103,104. Thus whether—and which—viruses can directly infect and transduce brain cells of the quail have yet to be demonstrated. We were particularly interested in testing whether AAVs can be used for the transduction of neurons in quails, for the reasons that AAVs are small, replication-deficient DNA viruses (safe to handle) and, despite their limited payload (<4.7 kb), are minimally toxic to cells, provide long-lasting expression of the gene(s) and can be produced at very high titers (~1013)101. Importantly, many AAV serotypes show better spread in the brain and improved tropism towards neurons compared to other viruses (e.g., lenti.). To quickly sift through a long list of viruses, we developed a primary culturing protocol from the brains of quail embryos (Fig. 2). We chose the E9 developmental stage (equivalent to E10 or HH36 in chicken105) for producing the cultures based on previous reports that have suggested that neurogenesis should be completed by this developmental stage44,45; in which case there should be many neurons with smaller amounts of mature glial cells46. We deemed this ideal, as we were mainly interested in examining the suitability of various AAVs to infect neurons. Nevertheless, and although our results do show very few mature glia cells (e.g., mature astrocytes and oligodendrocytes are completely absent; Fig. 3c and Supplementary Fig. 5a), we found a very large population of neuronal precursor cells instead, with a small proportion of mature neurons (GABAergic and Glutamatergic- 969/9561 cells; ~10%) (Fig. 3). These numbers are not reflected by the use of NeuroTrace (a Nissl stain–commonly employed for staining neurons49,50), but are supported by the use of MAP2-staining (~18%) (Fig. 1c, d and Supplementary Fig. 4f). Thus, and although beyond the scope of this work, we suggest that NeuroTrace may also stain neuronal precursor cells (our survey of the literature suggests that this has not been systematically explored). Together, our results indicate that neurogenesis is not complete by E9 in the quail and this is supported by reports showing the abundance of neuronal progenitors at similar embryonic stages in cultures from chicks106, as well as by in vivo reports of chicken embryos in which neurogenesis peaks at ~E7, but continues past this stage as late as E12, in various brain regions107. Furthermore, Radial glia (i.e., progenitors of neurons108) can be observed in later developmental stages, as late as E11. However, most previous studies employed various staining techniques, unlike our sequencing results, and these are hard to compare side-by-side (especially since some staining techniques may be promiscuous- e.g., NeuroTrace; above and Fig. 1c). The only available sc.mRNA seq. dataset is of the closely-related chicken species, but of much earlier embryonic stages (HH7 or E1)45. Thereby, it is less suitable for comparison with the developmental stage examined in this report. For instance, although we find some very early progenitor cell markers, such as PAX6 and SOX11, in our datasets (Supplementary Fig. 6), these are found in very low amounts in our cultured cells compared to chicken embryos45.
Lastly, a comparison of our sc.mRNA seq. data with reported transcriptomes from other animal models (e.g., refs. 58,61,65,69,109) reveals differences between species, such as lack of astrocytes in our cultures, the abundance of precursors of RG77, as well as a unique fibroblast population with the expression of neuronal markers (e.g., SOX11) (Supplementary Fig. 4d, e). Thus, embryonic quail brains appear to present a unique treasure trove of cell types, which may be of interest to developmental and comparative biologists.
AAV1-infection of cells
Quail cells in culture are somewhat resistant to infectivity by most viruses tested (Fig. 4). This observation was somewhat unexpected as (1) several AAVs have been shown to efficiently infect avian neurons in vivo14,32, (2) A3V has been tailored for infection of chicken neurons57, and (3) the specific lenti. we have examined has been previously employed for transgenesis of quail cells (albeit germ cells)26. The reason(s) behind this resistance is unknown but, in the case of AAV1 and AAV2, we suspected divergence in the receptor of the viruses, i.e., the AAVR89.
It is well established that the infection route of AAV is highly complex and requires a variety of membrane proteins that serve as co-receptors for the virus86,110,111. However, some variants (notably AAV1, 2 and 5) also require the AAVR89. We mined our sc.mRNA seq. data and indeed find our cultures to express qAAVR in a variety of cell types, but at low levels (Fig. 5a). These observations are highly consistent with the receptor’s reported “Low cell type specificity” expression patterns (see full details in ref. 112). Interestingly, transcripts of qAAVR are found at slightly higher levels in microglia (Fig. 5a). Nevertheless, even in this cluster, mRNA of this transcript is only detected in ~30% of the cells (Fig. 5a). Whether these low levels are representative of low protein levels is unknown, and we could not address this by immunostaining as this receptor lacks a suitable antibody. In fact, the only available antibody we could find (HPA072692) shows the expression of the receptor intracellularly and is unable to detect receptors on cell membranes112. These prevented us from addressing whether there is a correlation between receptor levels and AAV1*-infectivity.
Scrutiny of the receptor’s protein sequence demonstrates that the mammalian and the quail AAVRs diverge at key residues responsible for the binding of the AAV1 and AAV2’s capsids (Fig. 5c)87,88,89,101. We also note that different bird species have different AAVRs, and that these slight differences could be part of the reason why some AAVs may be suitable for one species, but not others14,32,39,40. In support, the AAV-receptor in Zebra finches is more similar to that of rodents (Fig. 5c), which coincides with reports showing the efficiency of AAVs in transducing Zebra finches’ neurons36,37,38.
Our mirroring mutagenesis scheme significantly improved the infection efficiency of AAV1 in vitro and in vivo, however, it did not enable AAV2-variants to infect the cultures (Figs. 6, 7 and Supplementary Figs. 6, 8). In vitro, AAV1* infected a variety of cell types (Supplementary Fig. 11a, b), and this is consistent with the notion that AAV1 is not exclusively neurotropic113,114,115,116,117,118 (unlike other AAVs, for instance, AAV2118). Correspondingly, in vivo, AAV1* infects both neurons and glia, although we did observe a slight preference towards neurons (Fig. 7 and Supplementary Figs. 9, 10). Based on these observations, we expected AAV1* to better infect chicken cultures owing to the higher homology between the chAAVR and mAAVR and that the single divergent residue, precisely residue K464 (mammalian numbering, Fig. 5c) which is involved in interaction with the viral capsid (Supplementary Table 3). Indeed, AAV1* outperformed AAV1 in transducing cultured chicken cells (Supplementary Fig. 8c). Together, these strongly support the important role of the AAVR in the infectivity of quail brain cells despite the necessity for added co-receptors89. Together, these imply that our unique strategy (in contrast to viral evolution methods that require highly specialized labs83) should be compatible with tailoring AAVs for other bird species.
In conclusion, our study provides a detailed description of the development of a unique quail-tailored AAV1—starting from the development of a culturing protocol, through molecular characterization of the embryonic cellular landscape of the quail’s brain and, finally, to rationale engineering of AAV1’s capsid exhibiting improved transduction capabilities of quail’s brain cells in vitro and in vivo. Our efforts thereby expand the available toolbox for interrogating the brains of a new animal model, which should likely increase interest in this unique avian model.
Methods
Animal ethics
Animal experimentations were approved by the Technion Institutional Animal Care and Use Committee (permit no. IL-157-11-17 and IL-19-10-143) and all experiments strictly followed the approved guidelines.
Primary quail neuronal cultures
Neuronal cultures were prepared from forebrains of seven, 8- or 9-day quail embryos. In a laminar hood, shells from post-fertilized eggs were gently removed (by breaking the shell at its upper tip). Embryos were isolated from the yolk and placed in Dulbecco’s Modified Eagle Media/Nutrient Mixture F-12 (denoted dissociation medium). Tissues were then enzymatically and mechanically dissociated as previously described in ref. 47,119, with several key modifications (see Supplementary Table 1). Briefly, skin, skull, and meninges were removed by scissors and tweezers, and forebrains were isolated and placed in a 15 ml conical tube containing dissociation media enriched with Papain (30 U/ml) and DNAse 1 (57 U/ml) and placed at 37 °C for 30 min. Following incubation, the solution was gently removed without disturbing the forebrains. Digested forebrains were washed three times with PBS and after the last wash, PBS was completely removed and replaced by 2 ml of plating medium consisting of neurobasal medium (Gibco, Cat. 21103049) supplemented with 2% B-27 (Gibco, Cat. 17504044), 1% Pen/Strep and 0.25% Glutamax (Gibco, Cat. 35050061). Then, tissue was manually dissociated by gentle trituration of solution with a fire-polished glass pipette (x15), followed by a single trituration with a 1000 μl plastic tip (to ensure complete dissociation). This solution was then applied onto a 40 μm cell strainer (placed on a 50 ml conical tube and pre-washed by 1 ml plating medium). The additional plating medium was added to the strainer after passing the entire solution to elute the remaining cells from the strainer. The filtered solution, containing the dissociated cells, was transferred to sterile tissue culture grade plastic plates (Corning, 60 mm, Cat. 430166) precoated with PDL, and these were placed for one hour in an incubator (37 °C, 5% CO2). Following incubation, the medium was removed and replaced by a fresh plating medium (prewarmed in the incubator). Plates were placed back in the incubator and half the media was replaced every 2 days by a fresh and prewarmed plating medium.
Mammalian cell culture and transfection
HEK293T (Human Embryonic Kidney cells, ATCC #CRL-1573) were maintained in DMEM (containing 10% FBS and 1% l-glutamine) in 100 mm Corning cell culture dishes. Cells were purchased from the American Tissue Culture Collection (ATCC) and are regularly tested for mycoplasma. These cells were used to examine the infectivity of viruses (see Fig. 4) and produce viruses (see below—viral production section).
DNA constructs and mutagenesis
Helper plasmid pAdDeltaF6 (expressing adenovirus E4, E2A, and VA; Cat. 112867); Rep/Cap plasmids—pAAV2/1, pAAV2/9, pAAV2/SL1, PHP-eB, 7m8 (Cat. 112862, 112865, 81070, 103005, and 64839, respectively), and Transfer plasmid—pAAV-CAG-eYFP (Cat. 104055) were purchased from Addgene. For the production of Avian adeno-associated virus (A3V), all plasmids (Helper, A3V rep/cap, Transfer-RSV-eGFP) were generously provided by Prof. Watanabe (Kyoto University Japan)33. For lentivirus production, we used pFUGW, the HIV-1 packaging vector Δ8.9 (pΔNR/8120), the VSVG envelope glycoprotein vector (pVSVG) and FR(GCaMP6S-p2A-nls-tdTom) as described earlier26, and these were a kind gift from Prof. Lois C. (Caltech, USA). Baculovirus (BacMam) containing CAG-GCaMP7s was purchased from Montana Molecular. Ltd (Montana, USA). Point mutations in AAV rep/cap plasmids were introduced by PCR; using PFU polymerase (Promega, United States). The PCR program consisted of 18 cycles of 55 °C annealing temperature and 68 °C (for 20 min) extension. (For a list of primers—see Supplementary Table 6).
Electrophysiology
Patch clamp recordings were obtained by MultiClamp 700B and Digidata 1440 A (Molecular Devices), and performed as previously described in ref. 121. Briefly, cells were voltage-clamped at −70 mV. Borosilicate glass capillaries (i.e., pipettes) were pulled to resistances of 3–7 MΩ and were filled with an internal solution containing (in mM): 135 K-gluconate, 10 NaCl, 10 HEPES, 2 MgCl2, 2 Mg2+-ATP, 1 EGTA, pH = 7.3. Recordings were done in extracellular recording solution containing (in mM): 138 NaCl, 1.5 KCl, 2.5 CaCl2, 10 d-glucose, 5 HEPES, 0.05 glycine, pH = 7.4. Gap-free recording protocol was used to assess spontaneous activity in the culture. For assessing intrinsic excitability, cells were current clamped by injected currents to −60 mV. The current steps were at 50 pA increments and the number of action potentials was calculated. We analyzed only complete action potentials (not differential potentials). Data were analyzed using the Clampfit software (Molecular Devices, USA).
Histochemistry
Quail cultures (grown directly on plastic plates) were fixed by 4% paraformaldehyde (PFA) for 15 min, permeabilized with PBS containing 5% FBS (fetal bovine serum) and 0.1% triton X-100. NeuN or MAP2-immunostaining were performed by overnight incubation of fixed cells with anti-NeuN antibody (mouse, clone A60, Millipore CAT# MAB377) (1:1000 in PBS) or with anti-MAP2 (mouse, cat # MA5-12826, Thermo Fisher) (1:500 in PBS) at 4 °C, with 3% FBS and 0.1% triton X-100. The next day, plates were washed three times with PBS and stained by secondary antibody (Rhodamine Red-X goat Anti-mouse IgG, Jackson Laboratories, CAT# 115-295-003) (1:200), in 3% FBS and 0.1% triton X-100 for 1 h at room temperature. Plates were then washed by PBS and stained with DAPI (1:1000). NeuroTrace 640/660 staining (Thermo Fisher) was performed as previously described in ref. 122. Briefly, cultures were fixed as described above, permeabilized with PBS containing 0.1% triton X-100 for 10 min, washed, and followed by incubation for 30 min with PBS containing NeuroTrace stain (1:100). Then, cells were washed by PBS and stained with DAPI (1:1000). GFAP-staining (1:500) was performed by using anti-GFAP (mouse, clone G-A-5, Calbiochem, CAT# IF03L), followed by staining with a secondary antibody (1:500) (Alexa Fluor-488 goat anti-mouse IgG, Jackson Laboratories, CAT# 115-545-003).
Virus production
We have produced viruses by the iodixanol method, as we previously described in ref. 123. Briefly, HEK293T (Human Embryonic Kidney cells, ATCC #CRL-1573) were grown in Dulbecco’s Modified Eagle Media/Nutrient Mixture F-12 (supplemented with 10% Fetal bovine serum and 1% l-glutamate) on 10 ml tissue culture plates (Corning, Cat. 430167), at 37 °C and 5% CO2. Cells were grown to 50 to 70% confluency and transfected with three viral plasmids (Helper, rep/cap, and transfer) using Polyethyleneimine (PEI) at a ratio of 8.1, 5.4, and 13.5 ug of DNA, respectively. After 6–8 h, growth media was replaced by serum-free media with 1% Glutamax. Media was collected after 48 and 72 h following transfection (collected media was preserved in −80 °C). The collected media was then filtered and concentrated via the Iodixanol step gradient method124 to a final volume of 100–500 μl. Viral titer was determined by qPCR. Titers below 1010 were discarded. Viral titers of the viruses used: AAV1 WT—3.13 × \({10}^{12}\), AAV1*TK—5.25 × \({10}^{12}\), AAV2-SL1—3.92 × \({10}^{11}\), AAVphpB—6.3 × \({10}^{11}\), AAV2-7m8—4.1 × \({10}^{10}\), AAV9—9.3 × \({10}^{12}\), Baculovirus—1.3 × \({10}^{13}\), A3V—3 × \({10}^{10}.\)
Cryosectioning
After 4 to 8 weeks following viral injections, anesthetized animals underwent whole animal perfusion fixation, as previously described in ref. 125. Briefly, 4% PFA was perfused to the entire body of the animal via the vascular system through the heart of the quail. Whole brains were then dissected and placed in 4% PFA overnight. The following day preserved tissue was treated with 30% sucrose solution and embedded in OCT for cryosectioning. Brains were sliced (40 μm sections) via a cryostat.
Fluorescence imaging
Imaging was performed on a Zeiss Laser Scanning Confocal Microscope (LSM-880 or 900-Airy2; Zeiss, Germany), as previously described in refs. 126,127. Experiments were conducted using a water immersion objective lens 20x [a water Plan-Apochromat objective lens; 20x/1.0 DIC D = 0.17 (UV) VIS-IR M27 75 mm] with a focal spot diameter of 0.5 μm (D = 1.2 × λ/NA). Brain slices were imaged serially and automatically tiled for reconstructing brain hemispheres. Live cells (neurons and glia) were imaged in a standard extracellular imaging solution containing (in mM): 138 NaCl, 1.5 KCl, 1.2 MgCl2, 2.5 CaCl2, 10 d-glucose, 10 HEPES, pH 7.4. GFP/GCaMP were excited using a 488 nm laser; Neu- 561 nm; NeuroTrace- 640 nm, and DAPI- 405 nm. Cultures were imaged after 3 days from viral infection. Change in fluorescence (ΔF/F) was calculated by (Ft–F0)/F0, where Ft is measured fluorescence (in arbitrary units, a.u.) at a given time t and F0 is initial baseline fluorescence, typically calculated from averaging the 10 first images, representing the basal fluorescence. In cells showing activity, baseline fluorescence was taken during times of inactivity (or at the trough of fluorescence signals). ΔF/F = 1 describes an increase of 100%, equivalent to a twofold increase in fluorescence. We did not need to encounter near division-by-zero artifacts and bleaching was minimal (most times not observed). Therefore, no corrections were made to fluorescent signals. For analysis of the percent of cells transfected by various AAVs (e.g., Fig. 6b), we imaged multiple (>5) large fields of view, in which we calculated the percent of infected cells by dividing the number of YFP-positive cells by the total number of cells. We averaged the percentages from multiple experiments, namely from different and independent cultures. Classification of cell types was obtained by staining (DAPI and NeuroTrace, see above Histochemistry). NeuroTrace-positive were counted as neurons, whereas NeuroTrace-negative denoted non-neuronal cells. The total number of cells was calculated by counting DAPI-positive nuclei (bright blue staining) in each field of view. This was obtained automatically by the ImageJ software. Assessment of cell density (per mm2) in vivo was determined as previously described in ref. 32, namely by imaging identical fields of view (i.e., identical sizes of images see Fig. 7 and Supplementary Fig. 8, typically 0.5 mm × 0.5 mm) in which we counted the number of visible fluorescent somata (we did not count processes as these could project to several planes of view). This was repeated in multiple animals, as noted in the legends.
Stereotaxic viral injections in vivo
Adult quails were anesthetized by a mixture of 2% isoflurane and 1% nitrous oxide. A small craniotomy was made above the Wulst (on modified stereotaxic equipment), followed by the removal of dura by tweezers and injections of 0.5 or 1 μL of each virus (at 50 nl/10 s) at multiple depths. Craniotomies were sealed by silicon and C&B MetaBond (Parkell, USA), as previously described in ref. 128. Briefly, silicon was first applied, and when completely dried, a C&B MetaBond was applied.
Single-cell mRNA sequencing
Cultures produced from 9-day-old embryos, grown for a week DIV, had their medium replaced with dissection-dissociation medium supplemented with 20 U/ml papain. The plate was incubated for 10 min at 37 °C, 5% CO2. Cells were then pipetted thoroughly and moved to a 15 ml canonical tube for gentle centrifugation (5 min at 300 rcf). The supernatant was removed and replaced with 1 ml DMEM with 10% FBS. Cells were then filtered through a cell strainer (40 µm nylon filter, LifeGene). Cells were then centrifuged again, and the medium was replaced as before. Cells were counted using a hemocytometer. Survival assay was performed with trypan blue. Cells were diluted to obtain 1200 cells/µl and 40 µl were processed with 10x scRNAseq with NextGEM v3 (10x Genomics), analyzed further by cell ranger (10x Genomics) and the Seurat pipeline (Version 4)62,129,130,131,132,133,134.
Library preparation and data generation for single-cell mRNA sequencing
One RNA single-cell library was prepared according to the 10X manufacture protocol (Chromium Next GEM Single Cell 3’ Library & Gel Bead Kit v3.1, PN-1000268) using 20,000 input cells. Single-cell separation was performed using the Chromium Next GEM Chip G Single Cell Kit (PN-1000120). The RNAseq data were generated on Illumina NextSeq2000, P2 100 cycles (R1-28bp, R2-90bp, I1-10bp, I2-10bp) (Illumina, cat no. 20046811).
Data processing bioinformatic analysis
Data was analyzed the data using R (Version 4.0.1) and the Seurat R package. Following QC results, we excluded from the data set cells with <200 and >2500 genes (potential cell duplets) and a mitochondrial gene percentage of >30%. In addition, genes detected in less than three cells were filtered out. Following these steps, 9561 cells were left for downstream analysis. Subsequently, PCA was used with all genes as the input and identified significant principal components (PCs) based on the ElbowPlot function. Ten PCs were selected as the input for uniform manifold approximation and projection (UMAP). Cells were clustered by the FindClusters function selecting a resolution of 0.5, yielding 15 different clusters (#0–14). Expression levels were determined based on the log2 fold change of various differentially expressed genes.
Statistics and reproducibility
All data are presented as mean ± SEM. The number of cells are indicated by n, and the number of experiments by N. Statistical significance (Sigmaplot 11 or Prism 8) was obtained by Student T-test (for two-group comparison) or one-way ANOVA for multiple group comparison with post hoc Tukey test. *p < 0.05; **p < 0.01, and ***p < 0.001; n.s., non-significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All datasets were deposited in a public data repository (https://doi.org/10.5061/dryad.vq83bk3xx; single-cell mRNA seq. data at NCBI GEO, accession GSE227334). All other data and/or materials are available from the corresponding author on reasonable request.
References
Clayton, N. S. & Emery, N. J. Avian models for human cognitive neuroscience: a proposal. Neuron 86, 1330–1342 (2015).
Nieder, A., Wagener, L. & Rinnert, P. A neural correlate of sensory consciousness in a corvid bird. Science 369, 1626–1629 (2020).
Yartsev, M. M. The emperor’s new wardrobe: rebalancing diversity of animal models in neuroscience research. Science 358, 466–469 (2017).
Olkowicz, S. et al. Birds have primate-like numbers of neurons in the forebrain. Proc. Natl Acad. Sci. USA 113, 7255–7260 (2016).
Lormant, F. et al. Research note: role of the hippocampus in spatial memory in Japanese quail. Poult. Sci. 99, 61–66 (2020).
Seidl, A. H. et al. Transgenic quail as a model for research in the avian nervous system – a comparative study of the auditory brainstem. J. Comp. Neurol. 521, 5–23 (2013).
Cohen, Y. et al. Hidden neural states underlie canary song syntax. Nature 582, 539–544 (2020).
Mouritsen, H. Long-distance navigation and magnetoreception in migratory animals. Nature 558, 50–59 (2018).
McCabe, B. J. Visual imprinting in birds: behavior, models, and neural mechanisms. Front. Physiol. 10, 658 (2019).
Brodin, A. The history of scatter hoarding studies. Philos. Trans. R. Soc. B Biol. Sci. 365, 869–881 (2010).
Stacho, M. et al. A cortex-like canonical circuit in the avian forebrain. Science 369, eabc5534 (2020).
Lipkind, D. et al. Songbirds work around computational complexity by learning song vocabulary independently of sequence. Nat. Commun. 8, 1247 (2017).
Daou, A. & Margoliash, D. Intrinsic neuronal properties represent song and error in zebra finch vocal learning. Nat. Commun. 11, 952 (2020).
Thiele, N., Hildebrandt, K. J. & Köppl, C. Gene delivery to neurons in the auditory brainstem of barn owls using standard recombinant adeno-associated virus vectors. Curr. Res. Neurobiol. 1, 100001 (2020).
Serralbo, O. et al. Transgenesis and web resources in quail. eLife 9, e56312 (2020).
Ben-Tov, M., Duarte, F. & Mooney, R. A neural hub for holistic courtship displays. Curr. Biol. S0960-9822(23)00245-2 https://doi.org/10.1016/j.cub.2023.02.072 (2023).
Martinho, A. & Kacelnik, A. Ducklings imprint on the relational concept of “same or different”. Science 353, 286–288 (2016).
Sid, H. & Schusser, B. Applications of gene editing in chickens: a new era is on the horizon. Front. Genet. 9, 456 (2018).
Amador, A. & Mindlin, G. B. Synthetic birdsongs as a tool to induce, and Iisten to, replay activity in sleeping birds. Front. Neurosci. 15, 835 (2021).
Ben-Yisahay, E. et al. Directional tuning in the hippocampal formation of birds. Curr. Biol. 31, 2592–2602.e4 (2021).
Finkelstein, A. et al. Three-dimensional head-direction coding in the bat brain. Nature 517, 159–164 (2015).
Le Douarin, N. A biological cell labeling technique and its use in experimental embryology. Dev. Biol. 30, 217–222 (1973).
Huss, D. et al. A transgenic quail model that enables dynamic imaging of amniote embryogenesis. Development 142, 2850–2859 (2015).
Tian, L. et al. Imaging neural activity in worms, flies and mice with improved GCaMP calcium indicators. Nat. Methods 6, 875–881 (2009).
Boyden, E. S., Zhang, F., Bamberg, E., Nagel, G. & Deisseroth, K. Millisecond-timescale, genetically targeted optical control of neural activity. Nat. Neurosci. 8, 1263–1268 (2005).
Lois, C., Hong, E. J., Pease, S., Brown, E. J. & Baltimore, D. Germline transmission and tissue-specific expression of transgenes delivered by lentiviral vectors. Science 295, 868–872 (2002).
Scott, B. B. & Lois, C. Generation of tissue-specific transgenic birds with lentiviral vectors. Proc. Natl Acad. Sci. USA 102, 16443–16447 (2005).
Poynter, G. & Lansford, R. Generating transgenic quail using lentiviruses. In Methods in Cell Biology. (ed. Bronner-Fraser, M.) Ch. 15 87, 281–293 (Elsevier, 2008).
Kumar, M., Keller, B., Makalou, N. & Sutton, R. E. Systematic determination of the packaging limit of lentiviral vectors. Hum. Gene Ther. 12, 1893–1905 (2001).
Lee, J., Ma, J. & Lee, K. Direct delivery of adenoviral CRISPR/Cas9 vector into the blastoderm for generation of targeted gene knockout in quail. Proc. Natl Acad. Sci. USA 116, 13288–13292 (2019).
Mizuarai, S. et al. Production of transgenic quails with high frequency of germ-line transmission using VSV-G pseudotyped retroviral vector. Biochem. Biophys. Res. Commun. 286, 456–463 (2001).
Rook, N. et al. AAV1 is the optimal viral vector for optogenetic experiments in pigeons (Columba livia). Commun. Biol. 4, 1–16 (2021).
Matsui, R., Tanabe, Y. & Watanabe, D. Avian adeno-associated virus vector efficiently transduces neurons in the embryonic and post-embryonic chicken brain. PLoS ONE 7, e48730 (2012).
Woodcock, M. E., Idoko-Akoh, A. & McGrew, M. J. Gene editing in birds takes flight. Mamm. Genome 28, 315–323 (2017).
Ahmadiantehrani, S. & London, S. E. A reliable and flexible gene manipulation strategy in posthatch zebra finch brain. Sci. Rep. 7, 43244 (2017).
Xiao, L. et al. A basal ganglia circuit sufficient to guide birdsong learning. Neuron 98, 208–221.e5 (2018).
Roberts, T. F., Gobes, S. M. H., Murugan, M., Ölveczky, B. P. & Mooney, R. Motor circuits are required to encode a sensory model for imitative learning. Nat. Neurosci. 15, 1454–1459 (2012).
Düring, D. N. et al. Fast retrograde access to projection neuron circuits underlying vocal learning in songbirds. Cell Rep. 33, 108364 (2020).
Markowitz, J. E. et al. Mesoscopic patterns of neural activity support songbird cortical sequences. PLoS Biol. 13, e1002158 (2015).
Liberti, W. A. et al. Unstable neurons underlie a stable learned behavior. Nat. Neurosci. 19, 1665–1671 (2016).
Shimizu, T. & Karten, H. J. Immunohistochemical analysis of the visual wulst of the pigeon (Columba livia). J. Comp. Neurol. 300, 346–369 (1990).
Berlin, S. & Isacoff, E. Y. In Biochemical Approaches for Glutamatergic Neurotransmission (eds. Parrot, S. & Denoroy, L.) 293–325 (Springer, 2018).
Kumar, A. & Mallick, B. N. Long-term primary culture of neurons taken from chick embryo brain: a model to study neural cell biology, synaptogenesis and its dynamic properties. J. Neurosci. Methods 263, 123–133 (2016).
Tsai, H. M., Garber, B. B. & Larramendi, L. M. 3H-thymidine autoradiographic analysis of telencephalic histogenesis in the chick embryo: II. Dynamics of neuronal migration, displacement, and aggregation. J. Comp. Neurol. 198, 293–306 (1981).
Williams, R. M., Lukoseviciute, M., Sauka-Spengler, T. & Bronner, M. E. Single-cell atlas of early chick development reveals gradual segregation of neural crest lineage from the neural plate border during neurulation. eLife 11, e74464 (2022).
Kumar, A. & Mallick, B. N. Long-term primary culture of neurons taken from chick embryo brain: a model to study neural cell biology, synaptogenesis and its dynamic properties. J Neurosci Methods 263, 123–133 (2016).
Berlin, S. et al. A family of photoswitchable NMDA receptors. eLife 5, e12040 (2016).
Mullen, R. J., Buck, C. R. & Smith, A. M. NeuN, a neuronal specific nuclear protein in vertebrates. Dev. Camb. Engl. 116, 201–211 (1992).
Susaki, E. A. et al. Versatile whole-organ/body staining and imaging based on electrolyte-gel properties of biological tissues. Nat. Commun. 11, 1982 (2020).
Mai, H. et al. Scalable tissue labeling and clearing of intact human organs. Nat. Protoc. 17, 2188–2215 (2022).
Tanay, A. & Regev, A. Single cell genomics: from phenomenology to mechanism. Nature 541, 331–338 (2017).
Tekin, H. et al. Effects of 3D culturing conditions on the transcriptomic profile of stem-cell-derived neurons. Nat. Biomed. Eng. 2, 540–554 (2018).
Perez, J. D. et al. Subcellular sequencing of single neurons reveals the dendritic transcriptome of GABAergic interneurons. eLife 10, e63092 (2021).
Jerber, J., Haldane, J., Steer, J., Pearce, D. & Patel, M. Dissociation of neuronal culture to single cells for scRNA-seq (10x Genomics) v1. https://doi.org/10.17504/protocols.io.bh32j8qe (2020).
Ilicic, T. et al. Classification of low quality cells from single-cell RNA-seq data. Genome Biol. 17, 29 (2016).
Luecken, M. D. & Theis, F. J. Current best practices in single-cell RNA-seq analysis: a tutorial. Mol. Syst. Biol. 15, e8746 (2019).
Morris, K. M. et al. The quail genome: insights into social behaviour, seasonal biology and infectious disease response. BMC Biol. 18, 14 (2020).
Fan, X. et al. Spatial transcriptomic survey of human embryonic cerebral cortex by single-cell RNA-seq analysis. Cell Res. 28, 730–745 (2018).
Loo, L. et al. Single-cell transcriptomic analysis of mouse neocortical development. Nat. Commun. 10, 134 (2019).
Ximerakis, M. et al. Single-cell transcriptomic profiling of the aging mouse brain. Nat. Neurosci. 22, 1696–1708 (2019).
Eze, U. C., Bhaduri, A., Haeussler, M., Nowakowski, T. J. & Kriegstein, A. R. Single-cell atlas of early human brain development highlights heterogeneity of human neuroepithelial cells and early radial glia. Nat. Neurosci. 24, 584–594 (2021).
Kaczmarczyk, L. et al. Slc1a3-2A-CreERT2 mice reveal unique features of Bergmann glia and augment a growing collection of Cre drivers and effectors in the 129S4 genetic background. Sci. Rep. 11, 5412 (2021).
Martínez-Cerdeño, V. & Noctor, S. C. Neural progenitor cell terminology. Front. Neuroanat. 12, 104 (2018).
Vogel, K. R. et al. In vitro modeling of experimental succinic semialdehyde dehydrogenase deficiency (SSADHD) using brain-derived neural stem cells. PLoS ONE 12, e0186919 (2017).
Johnson, M. B. et al. Single-cell analysis reveals transcriptional heterogeneity of neural progenitors in human cortex. Nat. Neurosci. 18, 637–646 (2015).
Tutukova, S., Tarabykin, V. & Hernandez-Miranda, L. R. The role of neurod genes in brain development, function, and disease. Front. Mol. Neurosci. 14, 662774 (2021).
Wu, S.-X. et al. Pyramidal neurons of upper cortical layers generated by NEX-positive progenitor cells in the subventricular zone. Proc. Natl Acad. Sci. USA 102, 17172–17177 (2005).
Dilly, G. A., Kittleman, C. W., Kerr, T. M., Messing, R. O. & Mayfield, R. D. Cell-type specific changes in PKC-delta neurons of the central amygdala during alcohol withdrawal. Transl. Psychiatry 12, 1–10 (2022).
Zhong, S. et al. A single-cell RNA-seq survey of the developmental landscape of the human prefrontal cortex. Nature 555, 524–528 (2018).
Harkin, L. F. et al. Distinct expression patterns for type II topoisomerases IIA and IIB in the early foetal human telencephalon. J. Anat. 228, 452–463 (2016).
Kimura, A., Matsuda, T., Sakai, A., Murao, N. & Nakashima, K. HMGB2 expression is associated with transition from a quiescent to an activated state of adult neural stem cells: HMGB2 expression in the adult hippocampus. Dev. Dyn. 247, 229–238 (2018).
Masuda, T. et al. Spatial and temporal heterogeneity of mouse and human microglia at single-cell resolution. Nature 566, 388–392 (2019).
Zheng, J. et al. Single-cell RNA-seq analysis reveals compartment-specific heterogeneity and plasticity of microglia. iScience 24, 102186 (2021).
Skelly, D. A. et al. Single-cell transcriptional profiling reveals cellular diversity and intercommunication in the mouse heart. Cell Rep. 22, 600–610 (2018).
Derk, J., Jones, H. E., Como, C., Pawlikowski, B. & Siegenthaler, J. A. Living on the edge of the CNS: meninges cell diversity in health and disease. Front. Cell. Neurosci. 15, 245 (2021).
Liu, M.-L. et al. Small molecules enable neurogenin 2 to efficiently convert human fibroblasts to cholinergic neurons. Nat. Commun. 4, 2183 (2013).
Batiuk, M. Y. et al. Identification of region-specific astrocyte subtypes at single cell resolution. Nat. Commun. 11, 1220 (2020).
Bushong, E. A., Martone, M. E., Jones, Y. Z. & Ellisman, M. H. Protoplasmic astrocytes in CA1 stratum radiatum occupy separate anatomical domains. J. Neurosci. 22, 183–192 (2002).
Esteve, P., Crespo, I., Kaimakis, P., Sandonís, A. & Bovolenta, P. Sfrp1 modulates cell-signaling events underlying telencephalic patterning, growth and differentiation. Cereb. Cortex 29, 1059–1074 (2019).
Beattie, R. & Hippenmeyer, S. Mechanisms of radial glia progenitor cell lineage progression. FEBS Lett. 591, 3993–4008 (2017).
Haslinger, A., Schwarz, T. J., Covic, M. & Chichung Lie, D. Expression of Sox11 in adult neurogenic niches suggests a stage-specific role in adult neurogenesis. Eur. J. Neurosci. 29, 2103–2114 (2009).
Hong, Y. J. & Do, J. T. Neural lineage differentiation from pluripotent stem cells to mimic human brain tissues. Front. Bioeng. Biotechnol. 7, 400 (2019).
Chan, K. Y. et al. Engineered AAVs for efficient noninvasive gene delivery to the central and peripheral nervous systems. Nat. Neurosci. 20, 1172–1179 (2017).
Wang, C. Y. & Wang, S. Astrocytic expression of transgene in the rat brain mediated by baculovirus vectors containing an astrocyte-specific promoter. Gene Ther. 13, 1447–1456 (2006).
Dana, H. et al. High-performance calcium sensors for imaging activity in neuronal populations and microcompartments. Nat. Methods 16, 649–657 (2019).
Dhungel, B. P., Bailey, C. G. & Rasko, J. E. J. Journey to the center of the cell: tracing the path of AAV transduction. Trends Mol. Med. 27, 172–184 (2021).
Zhang, R. et al. Divergent engagements between adeno-associated viruses with their cellular receptor AAVR. Nat. Commun. 10, 3760 (2019).
Zhang, R. et al. Adeno-associated virus 2 bound to its cellular receptor AAVR. Nat. Microbiol. 4, 675–682 (2019).
Pillay, S. et al. An essential receptor for adeno-associated virus infection. Nature 530, 108–112 (2016).
Tervo, D. G. R. et al. A designer AAV variant permits efficient retrograde access to projection neurons. Neuron 92, 372–382 (2016).
Woo, D. H. et al. Activation of astrocytic μ-opioid receptor elicits fast glutamate release through TREK-1-containing K2P channel in hippocampal astrocytes. Front. Cell. Neurosci. 12, 319 (2018).
McNeill, J., Rudyk, C., Hildebrand, M. E. & Salmaso, N. Ion channels and electrophysiological properties of astrocytes: implications for emergent stimulation technologies. Front. Cell. Neurosci. 15, 644126 (2021).
Zhong, S. et al. Electrophysiological behavior of neonatal astrocytes in hippocampal stratum radiatum. Mol. Brain 9, 34 (2016).
Chen, T.-W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013).
Shigetomi, E. et al. Imaging calcium microdomains within entire astrocyte territories and endfeet with GCaMPs expressed using adeno-associated viruses. J. Gen. Physiol. 141, 633–647 (2013).
Umpierre, A. D. et al. Microglial calcium signaling is attuned to neuronal activity in awake mice. eLife 9, e56502 (2020).
Jun-ichi, M. et al. Expression vector system based on the chicken β-actin promoter directs efficient production of interleukin-5. Gene 79, 269–277 (1989).
Payne, H. L., Lynch, G. F. & Aronov, D. Neural representations of space in the hippocampus of a food-caching bird. Science 373, 343–348 (2021).
Apostel, A. & Rose, J. Avian navigation: head direction cells in the quail hippocampus. Curr. Biol. 31, R781–R783 (2021).
Singer, E. Expanding neuroscience’s menagerie of model animals. Simons Foundation https://www.simonsfoundation.org/2019/06/21/expanding-neurosciences-menagerie-of-model-animals/ (2019).
Nectow, A. R. & Nestler, E. J. Viral tools for neuroscience. Nat. Rev. Neurosci. 21, 669–681 (2020).
Scott, B. B. & Lois, C. Production of transgenic birds using lentiviral vectors. In Mammalian and Avian Transgenesis — New Approaches (eds. Pease, S. & Lois, C.) 221–230 (Springer, 2006).
Zhang, Z. et al. Transgenic quail production by microinjection of lentiviral vector into the early embryo blood vessels. PLoS ONE 7, e50817 (2012).
Poynter, G., Huss, D. & Lansford, R. Japanese quail: an efficient animal model for the production of transgenic avians. Cold Spring Harb. Protoc. 2009, pdb.emo112 (2009).
Ainsworth, S. J., Stanley, R. L. & Evans, D. J. R. Developmental stages of the Japanese quail. J. Anat. 216, 3–15 (2010).
Pettmann, B., Louis, J. C. & Sensenbrenner, M. Morphological and biochemical maturation of neurones cultured in the absence of glial cells. Nature 281, 378–380 (1979).
Lever, M., Brand-Saberi, B. & Theiss, C. Neurogenesis, gliogenesis and the developing chicken optic tectum: an immunohistochemical and ultrastructural analysis. Brain Struct. Funct. 219, 1009–1024 (2014).
Striedter, G. F. & Beydler, S. Distribution of radial glia in the developing telencephalon of chicks. J. Comp. Neurol. 387, 399–420 (1997).
Nowakowski, T. J. et al. Spatiotemporal gene expression trajectories reveal developmental hierarchies of the human cortex. Science 358, 1318–1323 (2017).
Lisowski, L., Tay, S. S. & Alexander, I. E. Adeno-associated virus serotypes for gene therapeutics. Curr. Opin. Pharmacol. 24, 59–67 (2015).
Nance, M. E. & Duan, D. Perspective on adeno-associated virus capsid modification for Duchenne muscular dystrophy gene therapy. Hum. Gene Ther. 26, 786–800 (2015).
AlbaNova site, Stockholm. Protein atlas for: ENSG00000142687-KIAA0319L. https://www.proteinatlas.org/ENSG00000142687-KIAA0319L/single+cell+type (2023).
Nieuwenhuis, B. et al. Optimization of adeno-associated viral vector-mediated transduction of the corticospinal tract: comparison of four promoters. Gene Ther. 28, 56–74 (2021).
O’Carroll, S. J., Cook, W. H. & Young, D. AAV targeting of glial cell types in the central and peripheral nervous system and relevance to human gene therapy. Front. Mol. Neurosci. 13, 618020 (2021).
Koerber, J. T. et al. Molecular evolution of adeno-associated virus for enhanced glial gene delivery. Mol. Ther. 17, 2088–2095 (2009).
Merienne, N., Douce, J. L., Faivre, E., Déglon, N. & Bonvento, G. Efficient gene delivery and selective transduction of astrocytes in the mammalian brain using viral vectors. Front. Cell. Neurosci. 7, 106 (2013).
Wang, C., Wang, C.-M., Clark, K. R. & Sferra, T. J. Recombinant AAV serotype 1 transduction efficiency and tropism in the murine brain. Gene Ther. 10, 1528–1534 (2003).
Hadaczek, P. et al. Widespread AAV1- and AAV2-mediated transgene expression in the nonhuman primate brain: implications for Huntington’s disease. Mol. Ther. Methods Clin. Dev. 3, 16037 (2016).
Berlin, S. & Isacoff, E. Optical control of glutamate receptors of the NMDA-kind in mammalian neurons, with the use of photoswitchable ligands. Neuromethods https://doi.org/10.1007/978-1-4939-7228-9_10 (2018).
Naldini, L. et al. In vivo gene delivery and stable transduction of nondividing cells by a lentiviral vector. Science 272, 263–267 (1996).
Kellner, S. et al. Two de novo GluN2B mutations affect multiple NMDAR-functions and instigate severe pediatric encephalopathy. eLife 10, e67555 (2021).
Verlengia, G. et al. Engineered HSV vector achieves safe long-term transgene expression in the central nervous system. Sci. Rep. 7, 1507 (2017).
Rozales, K. et al. Differential roles for DNAJ isoforms in HTT-polyQ and FUS aggregation modulation revealed by chaperone screens. Nat. Commun. 13, 516 (2022).
Sena-Esteves, M. & Gao, G. Purification of recombinant adeno-associated viruses (rAAVs) by iodixanol gradient centrifugation. Cold Spring Harb. Protoc. 2020, pdb.prot095612 (2020).
Gage, G. J., Kipke, D. R. & Shain, W. Whole animal perfusion fixation for rodents. J. Vis. Exp. https://doi.org/10.3791/3564 (2012).
Hussein, W. & Berlin, S. Red photoactivatable genetic optical-indicators. Front. Cell. Neurosci. 14, 113 (2020).
Berlin, S. et al. Photoactivatable genetically encoded calcium indicators for targeted neuronal imaging. Nat. Methods 12, 852–858 (2015).
Cetin, A., Komai, S., Eliava, M., Seeburg, P. H. & Osten, P. Stereotaxic gene delivery in the rodent brain. Nat. Protoc. 1, 3166–3173 (2006).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e29 (2021).
Skelly, D. A. et al. Single-cell transcriptional profiling reveals cellular diversity and intercommunication in the mouse heart. Cell Rep. 22, 600–610 (2018).
Su, C. et al. Single-cell RNA sequencing in multiple pathologic types of renal cell carcinoma revealed novel potential tumor-specific markers. Front. Oncol. 11, 719564 (2021).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
Zeisel, A. et al. Cell types in the mouse cortex and hippocampus revealed by single-cell RNA-seq. Science 347, 1138–1142 (2015).
Chen, R., Wu, X., Jiang, L. & Zhang, Y. Single-cell RNA-seq reveals hypothalamic cell diversity. Cell Rep. 18, 3227–3241 (2017).
Acknowledgements
We would like to thank Prof. Watanabe D (Department of Biological Sciences, Graduate School of Medicine, Kyoto University) for providing us with the vectors to produce the A3V virus. The research submitted is in partial fulfillment of a doctoral degree for S.Z. Funding— Support was provided by the Rappaport Family Thematic grant (S.B. and Y.G.).
Author information
Authors and Affiliations
Contributions
S.Z.: Data curation, software, formal analysis, visualization, writing—original draft, writing—review and editing; M.A.: Data curation and formal analysis; I.C.: Data curation, formal analysis; R.H.: Data curation, project administration; S.R.: Data curation; Y.G.: Conceptualization, resources, data curation, supervision, funding acquisition, validation, investigation, and writing—review and editing; S.B.: Conceptualization, resources, data curation, software, formal analysis, supervision, funding acquisition, validation, investigation, visualization, methodology, writing—original draft, project administration, and writing—review and editing.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Communications Biology thanks Fernando García-Moreno, Rusty Lansford and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editors: Christian Wozny and George Inglis. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zoabi, S., Andreyanov, M., Heinrich, R. et al. A custom-made AAV1 variant (AAV1-T593K) enables efficient transduction of Japanese quail neurons in vitro and in vivo. Commun Biol 6, 337 (2023). https://doi.org/10.1038/s42003-023-04712-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s42003-023-04712-6
- Springer Nature Limited