Abstract
While genome-wide association studies are valuable in identifying CRC survival predictors, the benefit of adding blood DNA methylation (blood-DNAm) to clinical features, including the TNM system, remains unclear. In a multi-site population-based patient cohort study of 2116 CRC patients with baseline blood-DNAm, we analyzed survival predictions using eXtreme Gradient Boosting with a 5-fold nested leave-sites-out cross-validation across four groups: traditional and comprehensive clinical features, blood-DNAm, and their combination. Model performance was assessed using time-dependent ROC curves and calibrations. During a median follow-up of 10.3 years, 1166 patients died. Although blood-DNAm-based predictive signatures achieved moderate performances, predictive signatures based on clinical features outperformed blood-DNAm signatures. The inclusion of blood-DNAm did not improve survival prediction over clinical features. M1 stage, age at blood collection, and N2 stage were the top contributors. Despite some prognostic value, incorporating blood DNA methylation did not enhance survival prediction of CRC patients beyond clinical features.
Similar content being viewed by others
Introduction
Colorectal cancer (CRC) is one of the most common cancers and one of the most common causes of cancer-related deaths globally, accounting for more than 9% of all cancer-related deaths1. The prognosis and therapy management of CRC rely on the TNM stage system, with a relative 5-year survival over 90% for localized-stage CRC but dropping below 15% for distant-stage CRC2. Nevertheless, the current TNM stage system is insufficient for accurately predicting survival and guiding clinical management, especially among stage II–III patients, resulting in potential over- or undertreatment3,4. Consequently, there is a growing need to establish more accurate novel prognostic signatures in predicting survival of CRC patients.
DNA methylation (DNAm) is a crucial epigenetic modification whose genome-wide analysis allows exploration of potentially valuable biomarkers for predicting prognosis in CRC5,6,7. Predictive signatures based on high-dimensional tumor DNAm, such as DNAm from resected tumor tissue and circulating tumor DNA (ctDNA)8, using machine-learning approaches have been increasingly proposed. However, the added value in discriminatory ability provided by tumor DNAm-derived signature to traditional clinical variables was unsatisfactory9. Additionally, it is not possible to examine the postoperative DNAm profile following the removal of the tumor. DNAm profiles from peripheral whole blood present alternative opportunities to develop predictive signatures and use them to monitor survival over an extended period. DNAm-based scores derived from peripheral whole blood, such as a DNAm mortality risk score and the age acceleration of PhenoAge and GrimAge, have been identified as strongly associated with all-cause mortality10. Given that these DNAm scores have been designed for the general population and not specific to CRC patient populations, their associations with CRC-specific mortality were weaker than their associations with all-cause mortality11,12,13. It is furthermore unclear whether and to what extent blood DNAm signatures that are specifically derived for predicting survival of CRC patients may add prognostic value to predictive models based on established prognostic clinical factors.
This study aimed to develop and evaluate blood-DNAm-based prognostic signatures in a large cohort of colorectal cancer patients recruited from a multi-site, population-based prospective study. Comprehensive clinical variables were available, and most blood samples were collected from 1-month before surgery to 1-year post-surgery. This design enabled us to assess the added value of DNAm profiles alongside clinical variables, particularly for monitoring survival after surgery. To minimize potential biases from rapid inflammatory changes shortly after surgery, and to address the clinical need for improved therapeutic decision-making in CRC patients with intermediate TNM stages (II–III), we created two specific subsets for further investigation. Subset 1 included patients whose blood was collected at least 1 month after surgery, and subset 2 focused on CRC patients with intermediate TNM stages. This approach allowed us to thoroughly examine the potential predictive value of blood DNAm in these critical patient groups. To ensure the robustness of the predictive signatures, we employed rigorous nested leave-sites-out cross-validation (nLSOCV) and eXtreme Gradient Boosting (XGBoost). These methods were applied to the total CRC cohort and the two subsets, assuring high reliability of the developed prognostic signatures.
Results
Characteristics of study population
The baseline characteristics of study population are summarized Table 1. Median age at CRC diagnosis and blood sample collection were 69 and 70 years, and a slight majority of patients were male (58.9%). The blood samples were collected ≥1-month after surgery in approximately half of all patients (49.2%, Supplementary Fig. 1) and from more than half of patients with intermediate TNM stages (TNM stage II–III, 67.8%). The tumor was located in the distal colon and rectum in 66.2% of patients. The majority of patients were never smokers (40.8%) or former smoker (43.2%). During a median follow-up of 10.3 years, 1166 patients died, of whom 595 died from CRC. We designed two subsets to investigate the potential predictive value of blood DNAm in patients whose blood was collected ≥1-month after surgery (subset 1, N = 1042) and CRC patients with intermediate TNM stages (subset 2, N = 1434). The distribution of characteristics was similar among two subsets and the total CRC cohort.
Model performance
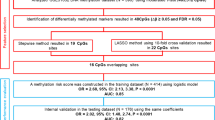
Predictive models for survival of CRC patients were developed using XGBoost with a 5-fold nLSOCV (Fig. 1) scheme across four feature groups: Model 1: traditional clinical features including TNM stage, Model 2: comprehensive clinical features including major tumor markers, Model 3: blood-DNAm, and Model 4: the inclusion of blood-DNAm with comprehensive clinical features. These four models were developed based on three datasets: the total CRC patient cohort, subset 1, and subset 2.
Prognostic predictive signatures for survival of CRC patients were developed using three datasets: the total CRC cohort and two subsets stratified by blood collection time and TNM stage. Predictive models were developed in each dataset based on four feature groups using 5-fold nested leave-sites-out cross-validation. In each outer-loop, the training set for Model 1 used traditional clinical features including age, sex, and TNM stages. The training sets for Models 2 and 3 underwent feature selection with their corresponding feature groups. The training set for Model 4 incorporated features selected from Models 2 and 3. Selected features were used to construct predictive signatures using XGBoost, with hyperparameter tuning conducted in inner-loops. Performance evaluation involved discrimination and calibration indicators, aggregating results from each test set in the outer loops. Abbreviation: CRC colorectal cancer, CV cross-validation, EN elastic net, XGBoost eXtreme Gradient Boosting.
Table 2 shows the performance of predictive models for overall survival of CRC patients trained on the total CRC cohort (N = 2116). At 1-, 3-, 5-, and 10-year follow-ups, 187 (8.8%), 499 (23.6%), 716 (33.8%), and 1087 (51.4%) of the patients had died in the total CRC cohort. Although Model 3, which was based on blood-DNAm, had some predictive value with a 10-year concordance index (C-index) of 0.64 and time-dependent areas under the receiver operating characteristics curve (AUROCs) ranging from 0.67 to 0.71, it performed worse compared to Models 1 and 2. Models 1 and 2, which were both based on clinical features, achieved higher 10-year C-indexes (0.74–0.75) and AUROCs at all time points (0.78–0.83). Model 4, which combined blood-DNAm with comprehensive clinical features, showed similar performance to Models 1 and 2, with a 10-year C-index of 0.75 and AUROCs ranging from 0.80 to 0.84. The statistical tests in each outer-loop supported these comparisons (Supplementary Table 1). These findings indicate that adding blood DNAm did not improve the prognostic performance compared to models based solely on clinical features. The Kaplan-Meier (KM) curves for the dichotomized signature of each model in the total CRC cohort are displayed in Supplementary Fig. 2. The prognostic performance, time-dependent ROC curve, and time-dependent calibration curve of the models in each outer-loop of the total CRC cohort are displayed in Supplementary Table 2 and Supplementary Figs. 3, 4.
Table 3 shows the performance of predictive models for overall survival of CRC patients trained on subset 1 (N = 1042) and subset 2 (N = 1434). At 1-, 3-, 5-, 10-year follow-ups, 80 (7.7%), 223 (21.4%), 321 (30.8%), and 501 (48.1%) patients, respectively, had died in subset 1, while in subset 2, the corresponding numbers were 80 (5.6%), 254 (17.7%), 404 (28.2%), and 672 (46.9). Among patients with intermediate stage, the 10-year AUROCs for Model 1, 2 and 4 were lower, ranging from 0.69 to 0.75, compared to the total CRC cohort, ranging from 0.79 to 0.82. Similar performance patterns were observed in both subsets as in the total CRC cohort, where adding blood DNAm to comprehensive clinical features did not significantly improve the predictive performance over traditional clinical features. The KM curves for dichotomized signature of each model in subset 1 are displayed in Supplementary Fig. 5, the comparison of prognostic performances, performance and calibrations, time-dependent ROC curve, and time-dependent calibration curve of the models in each outer-loop of subset 1 are displayed in Supplementary Tables 3, 4, and Supplementary Figs. 6, 7. Correspondingly, those in each outer-loop of subset 2 are displayed in Supplementary Tables 5, 6, and Supplementary Figs. 8–10.
Supplementary Table 7 shows the performances and calibrations of four predictive models for CRC-specific survival in the total CRC cohort and two subsets. Similar comparison patterns to overall survival were observed for CRC-specific survival in the total CRC cohort and the two subsets, and no improvements in performance were observed by adding blood DNAm to clinical features for the prediction of CRC-specific survival. The Kaplan-Meier (KM) curves for the dichotomized signature of each model in the total CRC cohort and the two subsets are displayed in Supplementary Figs. 11–13. The comparison of prognostic performances, performance, and calibrations of the models in each outer-loop of the total CRC cohort and two subsets are displayed in Supplementary Tables 8–13.
Model interpretation and feature importance
Figure 2 displays the top 20 features contributing to the prognostic signature based on Model 4, which combined both comprehensive clinical features and blood DNAm, in the total CRC cohort. The SHAP (SHapley Additive exPlanation) analysis suggests that M1 stage, age at blood collection, and N2 stage were the top contributors to overall survival prediction in CRC patients. The Charlson comorbidity index (CCI) and cg20352849 exhibited much higher contributions compared to T4 and T2 stages. Additionally, cg03067296 and cg07573085, along with retirement/early retirement status, showed similar contributions to the T4 and T2 stages.
The SHAP analysis is based on a model that incorporates blood DNA methylation along with comprehensive clinical features in the total CRC cohort. On the Y-axis, features are listed in descending order of importance (from top to bottom). On the X-axis, the left represents the SHAP value, and the right shows the number of times features were selected in the outer loops. Colors represent feature values, with red indicating high values and blue indicating low values. Each point corresponds to an individual patient. For features such as the M1 stage, patients with the M1 stage (red) contribute to a high SHAP value, indicating an elevated risk of death. Abbreviations: CCI Charlson Comorbidity Index, SHAP SHapley Additive exPlanations.
Discussion
In this multi-site large population-based prospective cohort study, incorporating comprehensive clinical features and DNAm data from peripheral whole blood, we found that the inclusion of these features did not lead to a significant enhancement of predictive signatures for survival in CRC patients when compared to those developed with traditional clinical variables only, including age, sex, and TNM stage. M1 stage, age at blood collection, and N2 stage emerged as the top contributors to overall survival prediction in CRC patients. Similar performance of predictive signatures was observed in patients whose blood samples were taken ≥1-month post-surgery and in those with intermediate TNM stages.
Interest in identifying prognostic DNAm biomarkers for survival in CRC patients has seen a steep rise, with the hope that biomarkers derived from novel omics-technologies may hold potential as valuable supplements to established prognostic criteria; however, there is still insufficient evidence to establish their utility in clinical practice5. A recent systematic review and external validation have highlighted the insufficient performance and limited generalizability of published prognostic DNAm biomarkers derived from tumor tissue9. These limitations could be attributed to relatively small sample sizes, improper handling of missing data, and a lack of evaluation of calibration. In addition, for most of the machine-learning-based epigenome-wide research, a robust method to validate models, such as LSOCV and external validation14, has not been applied15, an observation that was also made in studies concerning ctDNA and tumor methylation8,9,15,16. To our knowledge, no prior study has investigated the prognostic value of DNAm signatures derived from peripheral whole blood in CRC patients. Additionally, there are no available public DNAm datasets that provide DNA methylation array data from whole blood samples for external validation17. Most prior studies have focused on ctDNA8,18,19,20,21,22, with only one study examining FOXO3 blood DNA methylation, which found no association between FOXO3 CpGs and survival in CRC patients23.
In the current study, we developed prognostic signatures using DNAm profiles from peripheral whole blood, integrating comprehensive clinical features to assess the added value of blood DNAm compared to traditional clinical features on model performance. To ensure unbiased generalizability evaluation, we investigated a cohort with a large sample size and employed a nested LSOCV strategy, iteratively training and validating the model on different sites, maintaining independent training and testing datasets. This nested LSOCV approach simulated real-world scenarios, offering an optimal bias-variance trade-off and leveraging the full richness of data during training, including out-of-distribution samples for testing sites. The prognostic signature based on blood DNAm alone showed insufficient performance, with a time-dependent AUROC ranging from 0.67-0.71 for either short-term or long-term follow-up, and provided no added value beyond traditional clinical features, including age, sex, and TNM stage. Similarly, tumor DNAm showed poor performance in the aforementioned systemic review and external validation study9, while the effectiveness of ctDNA methylation is still unclear. An improvement in performance from combining ctDNA methylation with clinical features over TNM staging was noted; however, as this combination was not directly compared to clinical features alone, the source of the improvement—whether clinical features or ctDNA methylation—remains unclear8. Additionally, we explored prognostic signatures in CRC patients with intermediate TNM stage, requiring further risk stratification due to the survival paradox dilemma3,4,24. The predictive value of blood-DNAm profiles remained poor, offering no added value compared to traditional clinical features, and furthermore, it even decreased the discriminatory capability when combined with comprehensive clinical features.
Examining DNAm profiles in peripheral whole blood, spanning from pre-surgery to post-surgery periods, presents a unique opportunity to explore their potential as a tool for monitoring postoperative CRC prognosis. The comparable (albeit rather limited) discriminative performance of predictive signatures based on blood DNAm was evident across both the total CRC cohort and postoperative CRC patients, indicating that DNAm may maintain its predictive capability even after surgical tumor removal. While blood-based DNAm profiles do change post-surgery and during adjuvant therapy 25,26, suggesting their potential inclusion in predicting CRC response to therapy7,27, the number of patients with DNAm profiles in the current study determined during specific treatment windows, such as the period between surgery and chemotherapy, was limited. This hindered more differentiated assessment of when the prognostic value of blood DNAm might be most notable during treatment. Further research in large cohorts of CRC patients undergoing repeat longitudinal blood sampling is needed to address this important point. Additionally, considering that tumor recurrence may alter blood DNAm profiles, developing a signature for timely monitoring could offer early detection of CRC recurrence19.
Despite machine learning’s “black box” reputation, interpreted through the SHAP method, the M1 stage demonstrated the highest contribution, significantly surpassing other features, followed by age and N2 stage. This underscores the ongoing importance of traditional clinical features in survival prediction28. CCI ranking as the 4th contributor emphasizes the significant role of comorbidity in predicting survival29,30. Additionally, cg20352849, located in the south shelf of the PLCD3 gene, showed a much higher contribution than T4 and T2 stage, indicating its potential value in survival prediction. PLCD3, a phospholipase C family member, which hydrolyze phosphatidylinositol 4,5-diphosphate (PIP2) into diacylglycerol (DAG) and inositol-1,4,5-triphosphate (IP3), initiating Ca2+ release and activating protein kinase C (PKC)31. Its role in survival potentially involves the Wnt signaling pathway32.
A major strength of our study is its development of predictive signatures for survival in CRC patients, drawing from a large-scale, multi-site, prospective cohort with long-term follow-up and comprehensive clinical variables, including DNAm in blood samples taken over an extended time window and characterization of major tumor subtypes. Additionally, we adapted nested LSOCV and SHAP methods for high generalizability and interoperability. In particular, in contrast to many previous studies, we assessed the predictive value of blood DNAm in models including the best established clinical predictors of prognosis, including TNM stage, enabling assessment of incremental prognostic value beyond those predictors.
However, several limitations should be addressed. Firstly, approximately half of the blood samples were collected within 1 month after surgery, during which DNAm profiles are likely to have been influenced by surgery-related immune and inflammatory factors. This may have compromised the precision of the predictive signature for the entire CRC cohort and prompted us to provide separate analyses for a subset of patients whose blood samples were taken ≥1-month post-surgery. In addition, collection of blood samples at a single point of time prevented longitudinal assessment of changes in DNA methylation as a predictor of colorectal cancer patient survival. Secondly, the limited sample size of CRC patients with blood DNAm before surgery prevented a detailed assessment of the predictive value of presurgery blood DNAm. Thirdly, despite the overall large sample size and multi-site nature of our study, all CRC patients were recruited exclusively from the Rhine-Neckar region in southwest Germany which may limit generalizability. Therefore, external validation in different populations from other countries or with other ethnic composition is necessary. Lastly, some essential prognostic factors, such as lymphovascular invasion (LVI) and presurgery carcinoembryonic antigen (CEA) levels, were not included due to missing values exceeding 50% in this study.
In conclusion, in our multi-site large-scale population-based prospective cohort study, signatures incorporating comprehensive clinical features and blood DNA methylation did not enhance prediction performance compared to algorithms based only on traditional clinical features, including age, sex and TNM stage. This also applied to subsets of patients whose blood samples were taken ≥1-month post-surgery and patients with intermediate TNM stage. M1 stage, age at blood collection, and N2 stage emerged as the top contributors to survival prediction. This rigorously validated finding suggests a limited role for blood DNA methylation in predicting survival in CRC patients. Further research should evaluate the potential use of blood DNA methylation signatures for predicting and monitoring treatment response and CRC recurrence.
Methods
Study design and population
Our analysis is based on the DACHS (German name: Darmkrebs: Chancen der Verhütung durch Screening) study, an ongoing population-based case-control study with comprehensive follow-up of CRC cases conducted in the Rhine-Neckar region in southwestern Germany since 200333,34,35. Briefly, the DACHS study began in 2003 and covered CRC cases from a population of approximately two million people. Eligible participants aged 30 years or older who received a first diagnosis of CRC (ICD-10 codes C18-C20) were recruited from 22 hospitals providing CRC surgery in the study region. Following recruitment by the clinics, personal interviews by trained interviewers were conducted with patients and controls to collect information on lifetime and current exposure to CRC risk and prognostic factors, and blood and tumor samples were collected. Comprehensive follow-up with respect to treatments and overall and disease-specific survival over 10 years after diagnosis was conducted by collecting information from the patients’ treating physicians, record linkage with population registries, and collection of causes of death from public health authorities.
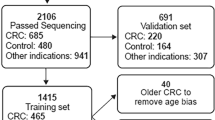
For this analysis, we included 2116 CRC patients who were diagnosed between 2003 and 2010 and from whom DNAm from peripheral whole blood, comprehensive clinical and follow-up data regarding survival outcomes were available (Supplementary Fig. 14). Additionally, we designed two subsets to investigate the potential predictive value of blood DNAm: patients whose blood was collected at least 1 month after surgery (subset 1, N = 1042) and CRC patients with intermediate TNM stages (subset 2, N = 1434).
The DACHS study was approved by the ethics committees of the Medical Faculty of the University of Heidelberg (#310/2001, 06 December 2001), and the Medical Chambers of Baden-Württemberg and Rhineland-Palatinate. Written informed consent was obtained from all participants.
DNA methylation preprocessing
Peripheral blood samples were collected after the interview and stored at −80 °C. DNA extraction and DNAm assessment based on Infinium MethylationEPIC BeadChip Kit (Illumina, Inc, San Diego, CA, USA), which covers over 850 thousand CpG probes, was conducted according to standard procedures. Details of quality control for samples and CpG probes are displayed in Supplementary Fig. 1436,37,38. Samples that did not meet the quality control criteria, including mismatched sex, low intensity, call-rate <95% on autosome, mean detected p-value > 0.01, duplicates, and lacking records of time of blood collection were excluded. Individual CpG probes that did not meet the quality control criteria, including (1) a detection p > 0.01 in any sample; (2) a bead count <3 in at least 5% of samples; (3) not CpG sites; (4) single nucleotide polymorphisms (SNPs)39; (5) align to multiple locations40,41; and (6) targeting sex chromosomes, were filtered out. Noob correction and beta-mixture quantile (BMIQ)42 were applied to normalize beta values (ranging from 0 to 1, i.e., from completely unmethylated to completely methylated) and batch correction were applied before machine learning analysis.
Machine learning procedure
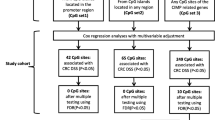
We constructed the prognostic signatures for overall survival, as well as CRC-specific survival. Survival time was defined as the period from the date of blood collection to the date of death or cancer-related death, or the last follow-up. Living participants were censored at the end of each follow-up period. A 5-fold nLSOCV scheme (Fig. 1), in which XGBoost was applied to develop predictive models for survival of CRC patients, was used separately for the total CRC cohort and the two subsets. CRC patients were split into five groups of approximately equal size according to their hospitals and institutions (Supplementary Table 14). Each group, in turn, served as the test set, with the remaining four subsets being the training set. In the outer-loop, a two-step filtering process was employed for selecting features. Firstly, we conducted Cox regression analysis on CpG sites that have passed quality control measures, aiming to select CpG sites associated with overall survival. This Cox regression adjusted for age at blood collection, sex, TNM stage, smoking status and alcohol consumption. Subsequently, we identified and selected the 5000 CpG sites with the lowest Benjamini-Hochberg adjusted p values (BH-adjusted p-values) for the corresponding Wald test. The second step aimed to selected predictive features with the elastic net (EN) approach43. The predictive features were selected based on comprehensive clinical features and all 5,000 survival-related CpGs selected in the previous step. XGBoost, which consistently achieves state-of-the-art performance in model prediction44, was then applied with selected predictive features. Both EN and XGBoost underwent another 5-fold cross-validation for hyperparameter tuning with grid search, called inner-loop. Performance evaluation involved discrimination and calibration indicators, aggregating results from each test set in the outer loops.
We developed 4 predictive models with the nLSOCV scheme using specific feature groups: (1) Model 1: traditional clinical features including age at blood collection, sex and separately stages of tumor size and invasion (T), lymph nodes involvement (N), and distant metastasis (M) ; (2) Model 2: comprehensive clinical features, incorporating additional features including age at diagnosis, tumor location, family history of CRC, resection edge, tumor differential grading, histological type, CCI29, body mass index (BMI), smoking status, pack-years of cigarette consumption, alcohol consumption, average lifetime physical activity, diet quality score45, occupational position, employment situation, MSI, KRAS, BRAF, CIMP46, neo-chemotherapy, neo-radiotherapy, adjuvant-chemotherapy, adjuvant-radiotherapy, relapse, metastasis, and surgery, adjuvant-chemotherapy and adjuvant-radiotherapy after relapse and metastasis (Supplementary Table 15); (3) Model 3: processed blood DNAm; and (4) Model 4: features after two-step filtering in each outer-loop, incorporating blood CpGs from Model 3 with clinical features from Model 2. Missing data of clinical features were imputed with the missforest method47. MissForest is a non-parametric, iterative imputation technique that utilizes the Random Forest algorithm. This method inherently implements a multiple imputation approach by averaging across numerous unpruned classifications or regression trees, enabling the handling of multivariate data, including both continuous and categorical variables, simultaneously.
Model evaluation and visualization
We evaluated the model’s performance at several specific follow-up times post-surgery, including at 1, 3, 5, and 10 years, to identify potential drift over time48. The model’s discriminatory ability was assessed using the Kaplan-Meier (KM) curve for signatures dichotomized by the median value and time-dependent ROC curves, along with calculating the AUROC. The ROC curve illustrates the trade-off between the true positive rate (TPR) and false positive rate (FPR) across various decision thresholds at specific times. A higher AUROC value indicates better predictive accuracy of the model. To align our findings with studies that employ the C-index, which captures the model’s capability to differentiate among predictions regarding risk, event occurrence, and time in a single metric but may reflect overconfidence in model discrimination49, we also measured the C-index within a 10-year time window. The time-dependent AUROCs and C-indexes in each outer-loop of different models were compared. The calibration performance was examined using the time-dependent Brier score and the integrated Brier score (IBS). The Brier score, which ranges from 0 to 1, measures the average squared difference between observed survival status at specific times and the predicted probabilities of survival, with a lower score indicating more accurate predictions. The IBS evaluates the overall accuracy of survival predictions over a specified time window of 10 years, reflecting the squared differences between the observed and predicted survival curves. Calibration curves were plotted; ideally, a perfectly calibrated model would exhibit a curve that closely aligns with the 45-degree diagonal line. The strict setting of nLSOCV ensured that the test set remained unseen during training, limiting predictive models’ performance but enhancing overall.
We used SHAP analysis to identify features with critical contribution in predicting risk of death in XGBoost models50. Higher SHAP values correspond to a higher risk of death. A summary plot was used to visualize the features’ contribution to predictive model.
Statistical analysis
Descriptive statistics were used to characterize the distribution of baseline variables for the total CRC cohort, subset 1 and subset 2. Categorical covariates were summarized using absolute and relative frequencies, while median and interquartile range were presented for continuous variables.
All statistical analyses were performed using R language program (version 4.1.2) with R Studio (version 1.4.1717; Boston, USA). We utilized several R packages for different aspects of the analysis: ChAMP38, limma36, and minfi37 for DNAm preprocessing; missForest47 for data imputation; survival and survminer51 for Cox regression; grpreg43 for developing the EN model; xgboost and survXgboost for developing the XGBoost model44; mlr3, mlr3proba, and mlr3extralearners for hyperparameter tuning52; pec53, Riskregression48, and compareC54 for model evaluations; and SHAPforxgboost50 for SHAP evaluation. Statistically significant p-values were defined as those with two-tailed p < 0.05, after BH correction for multiple comparisons where necessary.
Data availability
The datasets generated and analyzed during the current study are not publicly available due ethical and legal restrictions but are available from the corresponding author on reasonable request.
Code availability
The code is available from the corresponding author upon reasonable request.
References
Sung, H. et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71, 209–249 (2021).
Siegel, R. L., Wagle, N. S., Cercek, A., Smith, R. A. & Jemal, A. Colorectal cancer statistics, 2023. CA Cancer J. Clin. 73, 233–254 (2023).
Kim, M. J. et al. Survival paradox between stage IIB/C (T4N0) and stage IIIA (T1-2N1) colon cancer. Ann. Surg. Oncol. 22, 505–512 (2015).
Kannarkatt, J., Joseph, J., Kurniali, P. C., Al-Janadi, A. & Hrinczenko, B. Adjuvant chemotherapy for stage II colon cancer: a clinical dilemma. J. Oncol. Pract. 13, 233–241 (2017).
Draht, M. X. G. et al. Prognostic DNA methylation markers for sporadic colorectal cancer: a systematic review. Clin. Epigenetics 10, 35 (2018).
Jung, G., Hernandez-Illan, E., Moreira, L., Balaguer, F. & Goel, A. Epigenetics of colorectal cancer: biomarker and therapeutic potential. Nat. Rev. Gastroenterol. Hepatol. 17, 111–130 (2020).
Müller, D. & Győrffy, B. DNA methylation-based diagnostic, prognostic, and predictive biomarkers in colorectal cancer. Biochim. Biophys. Acta Rev. Cancer 1877, 188722 (2022).
Luo, H. et al. Circulating tumor DNA methylation profiles enable early diagnosis, prognosis prediction, and screening for colorectal cancer. Sci. Transl. Med. 12, eaax7533 (2020).
Yuan, T. et al. CpG-biomarkers in tumor tissue and prediction models for the survival of colorectal cancer: a systematic review and external validation study. Crit. Rev. Oncol. Hematol. 193, 104199 (2024).
Gao, X. et al. Whole blood DNA methylation aging markers predict colorectal cancer survival: a prospective cohort study. Clin. Epigenetics 12, 184 (2020).
Zhang, Y. et al. DNA methylation signatures in peripheral blood strongly predict all-cause mortality. Nat. Commun. 8, 14617 (2017).
Levine, M. E. et al. An epigenetic biomarker of aging for lifespan and healthspan. Aging 10, 573–591 (2018).
Lu, A. T. et al. DNA methylation GrimAge strongly predicts lifespan and healthspan. Aging 11, 303–327 (2019).
Bradshaw, T. J., Huemann, Z., Hu, J. & Rahmim, A. A guide to cross-validation for artificial intelligence in medical imaging. Radio. Artif. Intell. 5, e220232 (2023).
Yuan, T. et al. Machine learning in the identification of prognostic DNA methylation biomarkers among patients with cancer: a systematic review of epigenome-wide studies. Artif. Intell. Med. 143, 102589 (2023).
Yang, X. et al. Predicting disease-free survival in colorectal cancer by circulating tumor DNA methylation markers. Clin. Epigenetics 14, 160 (2022).
Xiong, Z. et al. EWAS Data Hub: a resource of DNA methylation array data and metadata. Nucleic Acids Res. 48, D890–d895 (2020).
Symonds, E. L. et al. Circulating epigenetic biomarkers for detection of recurrent colorectal cancer. Cancer 126, 1460–1469 (2020).
Jin, S. et al. Efficient detection and post-surgical monitoring of colon cancer with a multi-marker DNA methylation liquid biopsy. Proc. Natl Acad. Sci. USA 118, e2017421118 (2021).
Hallermayr, A. et al. Somatic copy number alteration and fragmentation analysis in circulating tumor DNA for cancer screening and treatment monitoring in colorectal cancer patients. J. Hematol. Oncol. 15, 125 (2022).
Mo, S. et al. Early detection of molecular residual disease and risk stratification for stage I to III colorectal cancer via circulating tumor DNA methylation. JAMA Oncol. 9, 770–778 (2023).
Bachet, J. B. et al. Circulating tumour DNA at baseline for individualised prognostication in patients with chemotherapy-naïve metastatic colorectal cancer. An AGEO prospective study. Eur. J. Cancer 189, 112934 (2023).
Yu, C. et al. Association of FOXO3 blood DNA methylation with cancer risk, cancer survival, and mortality. Cells 10, 3384 (2021).
Auclin, E. et al. Subgroups and prognostication in stage III colon cancer: future perspectives for adjuvant therapy. Ann. Oncol. 28, 958–968 (2017).
Sadahiro, R. et al. Major surgery induces acute changes in measured DNA methylation associated with immune response pathways. Sci. Rep. 10, 5743 (2020).
Robinson, N. et al. Anti-cancer therapy is associated with long-term epigenomic changes in childhood cancer survivors. Br. J. Cancer 127, 288–300 (2022).
Fatemi, N. et al. DNA methylation biomarkers in colorectal cancer: Clinical applications for precision medicine. Int. J. Cancer 151, 2068–2081 (2022).
Bibault, J. E., Chang, D. T. & Xing, L. Development and validation of a model to predict survival in colorectal cancer using a gradient-boosted machine. Gut 70, 884–889 (2021).
Charlson, M. E., Pompei, P., Ales, K. L. & MacKenzie, C. R. A new method of classifying prognostic comorbidity in longitudinal studies: development and validation. J. Chronic Dis. 40, 373–383 (1987).
Hahn, E. E. et al. Understanding comorbidity profiles and their effect on treatment and survival in patients with colorectal cancer. J. Natl Compr. Cancer Netw. 16, 23–34 (2018).
Danielsen, S. A. et al. Phospholipase C isozymes are deregulated in colorectal cancer-insights gained from gene set enrichment analysis of the transcriptome. PLoS ONE 6, e24419 (2011).
Hajebi Khaniki, S., Shokoohi, F., Esmaily, H. & Kerachian, M. A. Analyzing aberrant DNA methylation in colorectal cancer uncovered intangible heterogeneity of gene effects in the survival time of patients. Sci. Rep. 13, 22104 (2023).
Brenner, H., Chang-Claude, J., Seiler, C. M., Rickert, A. & Hoffmeister, M. Protection from colorectal cancer after colonoscopy: a population-based, case-control study. Ann. Intern Med. 154, 22–30 (2011).
Hoffmeister, M. et al. Statin use and survival after colorectal cancer: the importance of comprehensive confounder adjustment. J. Natl Cancer Inst. 107, djv045 (2015).
Carr, P. R. et al. Estimation of absolute risk of colorectal cancer based on healthy lifestyle, genetic risk, and colonoscopy status in a population-based study. Gastroenterology 159, 129–138.e129 (2020).
Ritchie, M. E. et al. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Fortin, J. P., Triche, T. J. Jr. & Hansen, K. D. Preprocessing, normalization and integration of the Illumina HumanMethylationEPIC array with minfi. Bioinformatics 33, 558–560 (2017).
Tian, Y. et al. ChAMP: updated methylation analysis pipeline for Illumina BeadChips. Bioinformatics 33, 3982–3984 (2017).
Zhou, W., Laird, P. W. & Shen, H. Comprehensive characterization, annotation and innovative use of Infinium DNA methylation BeadChip probes. Nucleic Acids Res. 45, e22 (2017).
Pidsley, R. et al. Critical evaluation of the Illumina MethylationEPIC BeadChip microarray for whole-genome DNA methylation profiling. Genome Biol. 17, 208 (2016).
Alcala, N. et al. Integrative and comparative genomic analyses identify clinically relevant pulmonary carcinoid groups and unveil the supra-carcinoids. Nat. Commun. 10, 3407 (2019).
Teschendorff, A. E. et al. A beta-mixture quantile normalization method for correcting probe design bias in Illumina Infinium 450 k DNA methylation data. Bioinformatics 29, 189–196 (2013).
Breheny, P. & Huang, J. Group descent algorithms for nonconvex penalized linear and logistic regression models with grouped predictors. Stat. Comput. 25, 173–187 (2015).
Chen, T. & Guestrin, C. In Proc. 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining 785–794 (Association for Computing Machinery, 2016). https://doi.org/10.1145/2939672.2939785
Carr, P. R. et al. Healthy lifestyle factors associated with lower risk of colorectal cancer irrespective of genetic risk. Gastroenterology 155, 1805–1815.e1805 (2018).
Hoffmeister, M. et al. Colonoscopy and reduction of colorectal cancer risk by molecular tumor subtypes: a population-based case-control study. Am. J. Gastroenterol. 115, 2007–2016 (2020).
Stekhoven, D. J. & Buhlmann, P. MissForest-non-parametric missing value imputation for mixed-type data. Bioinformatics 28, 112–118 (2012).
Lazic, S. E. Medical risk prediction models: with ties to machine learning. J. R. Stat. Soc. Ser. A 185, 425–425 (2021).
Blanche, P., Kattan, M. W. & Gerds, T. A. The c-index is not proper for the evaluation of t-year predicted risks. Biostatistics 20, 347–357 (2019).
Just, A. C. et al. Gradient boosting machine learning to improve satellite-derived column water vapor measurement error. Atmos. Meas. Tech. 13, 4669–4681 (2020).
Therneau, T. M. & Grambsch, P. M. Modeling Survival Data: Extending The Cox Model. (Springer New York, 2000).
Lang, M. et al. mlr3: a modern object-oriented machine learning framework in R. J. Open Source Softw. 4, 1903 (2019).
Mogensen, U. B., Ishwaran, H. & Gerds, T. A. Evaluating random forests for survival analysis using prediction error curves. J. Stat. Softw. 50, 1–23 (2012).
Kang, L., Chen, W., Petrick, N. A. & Gallas, B. D. Comparing two correlated C indices with right-censored survival outcome: a one-shot nonparametric approach. Stat. Med. 34, 685–703 (2015).
Acknowledgements
DKFZ clinician scientist program (Z.F.). German Research Council BR 1704/6-1, BR 1704/6-3, BR 1704/6-4, CH 117/1-1, HO 5117/2-1, HO 5117/2-2, HE 5998/2-1, HE 5998/2-2, KL 2354/3-1, KL 2354 3-2, RO 2270/8-1, RO 2270/8-2, BR 1704/17-1, and BR 1704/17-2 (H.B., M.H.). Interdisciplinary Research Program of the National Center for Tumor Diseases (NCT), Germany (H.B., M.H.). German Federal Ministry of Education and Research 01KH0404, 01ER0814, 01ER0815, 01ER1505A, 01ER1505B and 01KD2104A (H.B., M.H.). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript. We thank the study participants and the interviewers who collected the data. our study team at the Division of Clinical Epidemiology, German Cancer Research Center, Heidelberg, Germany for providing technical assistance, with no compensation outside of salary. We also thank the following hospitals and cooperating institutions that recruited patients for this study: Chirurgische Universitätsklinik Heidelberg, Klinik am Gesundbrunnen Heilbronn, St Vincentiuskrankenhaus Speyer, St Josefskrankenhaus Heidelberg, Chirurgische Universitätsklinik Mannheim, Diakonissenkrankenhaus Speyer, Krankenhaus Salem Heidelberg, Kreiskrankenhaus Schwetzingen, St Marienkrankenhaus Ludwigshafen, Klinikum Ludwigshafen, Stadtklinik Frankenthal, Diakoniekrankenhaus Mannheim, Kreiskrankenhaus Sinsheim, Klinikum am Plattenwald Bad Friedrichshall, Kreiskrankenhaus Weinheim, Kreiskrankenhaus Eberbach, Kreiskrankenhaus Buchen, Kreiskrankenhaus Mosbach, Enddarmzentrum Mannheim, Kreiskrankenhaus Brackenheim and Cancer Registry of Rhineland-Palatinate, Mainz.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
Z.F. contributed to concept and design, development of methodology, acquisition of data (acquired and managed patients, provided facilities, etc.), analysis and interpretation of data (e.g., statistical analysis, biostatistics, computational analysis), writing of the manuscript, critical review and revision of the manuscript. D.E. contributed to development of methodology, acquisition of data (acquired and managed patients, provided facilities, etc.), analysis and interpretation of data (e.g., statistical analysis, biostatistics, computational analysis), critical review, and revision of the manuscript. T.Y. contributed to analysis and interpretation of data (e.g., statistical analysis, biostatistics, computational analysis), critical review, and revision of the manuscript. B.C.K. contributed to analysis and interpretation of data (e.g., statistical analysis, biostatistics, computational analysis), critical review, and revision of the manuscript. M.H. contributed to analysis and interpretation of data (e.g., statistical analysis, biostatistics, computational analysis), critical review and revision of the manuscript, administrative, technical, or material support (i.e., reporting or organizing data, constructing databases). H.B. contributed to concept and design, development of methodology, acquisition of data (acquired and managed patients, provided facilities, etc.), analysis and interpretation of data (e.g., statistical analysis, biostatistics, computational analysis), writing of the manuscript, critical review and revision of the manuscript, administrative, technical, or material support (i.e., reporting or organizing data, constructing databases), study supervision.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fan, Z., Edelmann, D., Yuan, T. et al. Developing survival prediction models in colorectal cancer using epigenome-wide DNA methylation data from whole blood. npj Precis. Onc. 8, 191 (2024). https://doi.org/10.1038/s41698-024-00689-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41698-024-00689-5
- Springer Nature Limited