Abstract
The consensus molecular subtypes (CMS) of colorectal cancer (CRC) is the most widely-used gene expression-based classification and has contributed to a better understanding of disease heterogeneity and prognosis. Nevertheless, CMS intratumoral heterogeneity restricts its clinical application, stressing the necessity of further characterizing the composition and architecture of CRC. Here, we used Spatial Transcriptomics (ST) in combination with single-cell RNA sequencing (scRNA-seq) to decipher the spatially resolved cellular and molecular composition of CRC. In addition to mapping the intratumoral heterogeneity of CMS and their microenvironment, we identified cell communication events in the tumor-stroma interface of CMS2 carcinomas. This includes tumor growth-inhibiting as well as -activating signals, such as the potential regulation of the ETV4 transcriptional activity by DCN or the PLAU-PLAUR ligand-receptor interaction. Our study illustrates the potential of ST to resolve CRC molecular heterogeneity and thereby help advance personalized therapy.
Similar content being viewed by others
Introduction
CRC is a leading cause of cancer-related death worldwide with over 1.85 million diagnosed cases and 850000 deaths annually1. Despite a decline in mortality rates due to personalized treatments in recent years2, the extensive inter-patient and intra-tumor heterogeneity of CRC still pose substantial treatment challenges3. This heterogeneity manifests at genomic, epigenomic and transcriptomic levels, and in the composition of the tumor microenvironment (TME)4.
In 2015, the CRC subtyping consortium proposed a classification of CRC into four CMS, derived from large-scale gene expression datasets5. Despite its widespread use, its clinical impact is still limited due to its reliance on bulk-sequencing, which cannot accurately categorize mixed or transitional CMS phenotypes, nor precisely define the cellular composition and microenvironment of tumors. Recently, scRNA-seq was applied to CRC samples, revealing CMS features at the cellular level and the coexistence of multiple CMS in individual patients6,7,8,9,10. However, the spatial distribution of the different CMS and their interactions with their respective TMEs remain poorly understood.
ST technologies can address these limitations by measuring gene expression levels throughout tissue space, integrating morphology, spatial localization and transcriptomic profile. In oncology, ST has been employed to study breast cancer11, prostate cancer12 and melanoma13, among others. To date, its application to CRC has been mostly to support results obtained from other technologies, without specifically addressing the CMS of CRC14,15,16,17.
Here, we applied ST to analyze 14 samples from seven CRC patients, aiming to deepen our understanding of the spatial properties and heterogeneity of CMS. By mapping cell type composition spatially, linking distinct molecular and morphological features to different CMS, and investigating predicted intercellular interactions in CMS2 carcinomas, we highlighted the capacity of ST to support the future development of personalized treatment strategies for CRC.
Results
ST and deconvolution reliably reveal the spatial cell type distribution in CRC
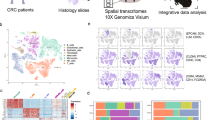
We used 10x Genomics VISIUM to process fresh-frozen resection samples from CRC tumors of seven individuals, obtained from different anatomical locations, and exhibiting varying metastatic status, growth patterns, and immune cell (IC) infiltration levels (Fig. 1a, Table 1). We considered two serial sections per patient to generate technical replicates, resulting in a total of 20,733 Visium spots, each of which contained an average of 3,738 unique genes (Supplementary Fig. S1a). Technical reproducibility among the replicates, along with inter-patient heterogeneity, were revealed via the UMAP projection of the transcriptomic profiles of the aforementioned spots (Fig. 1b). A pathologist independently examined the samples and assigned each spot to its corresponding anatomical compartment based on tissue type and cellular morphology (Fig. 1c, Methods).
a Study outline displaying the anatomical localization of our set of CRC samples, their spatial transcriptomics processing and the deconvolution-based approach to characterize spatial features of CMS. Figures created with BioRender.com. b UMAP embedding of the gene expression measurements per spot split by technical replicates. Colors represent the different patients. c UMAP embedding of the gene expression measurements per spot colored by pathologist’s annotations. In addition, a bar plot displays the proportions of these annotations per sample. IC: immune cells. d Proportions of major cell classes per sample as estimated by the results of the deconvolution approach. The right hand side of the plot displays the number of analyzed spots per sample. e Enrichment/depletion plot describing the association between cell type abundance as predicted by the deconvolution (x-axis) and the different anatomical regions as annotated by the pathologist (y-axis). The dot size represents the enrichment score (Methods), while the color represents enrichment (red) or depletion (blue). IC immune cells. f–h Spatial mapping of the predicted number of mature enterocytes type I, stem-like transient amplifiers (TA) and CD4+ T cells per spot matching the pathologists’ tissue annotation and expected cell type localization as illustrated for sample S6_Rec_Rep2.
To determine the cellular composition per spot, we used Cell2Location18 and a recently published CRC scRNA-seq dataset6 as reference (Supplementary Table S1, Methods). We found highly comparable proportions between replicates when considering major cell types (Fig. 1d). In contrast, proportions varied greatly across individuals: for instance, unlike all other patients, S7_Rec/Sig samples mainly contained non-neoplastic tissue (Table 1), and tumor cells only comprised around 5%. Upon assessing the deconvolution results by computing the spatial correlation of cell subtype abundance among technical replicates, we found high stability with Pearson’s correlation coefficients over 0.9, except for a low-quality sample (Supplementary Fig. S1b, Methods).
We next evaluated whether the deconvolution-predicted cell type abundances were located in their respective anatomical compartments using the pathologist’s annotations as reference (Supplementary Fig. S1c). As expected, non-neoplastic intestinal cells were the most abundant in non-neoplastic epithelium (89%), while T and B cells were the prevalent types in the immune cell aggregates (83% and 68%). In tumor-annotated spots, tumor cells (36%), T cells (26%), and B cells (25%) were the predominant types. At the cell subtype level, we observed significant enrichment of non-neoplastic mucosal cells, such as mature enterocytes type 1 and 2, goblet cells and stem-like transiently amplifying cells in spots labeled as non-neoplastic epithelium, lamina propria or mixed (Fig. 1e, Methods). Tumor cells, CD19+CD20+ B cells and CD8+ T cells were mainly enriched in spots classified as tumor or tumor-stroma mixed. CD4+ T-cells and other immune cells were mostly found in IC aggregates and stromal regions with high IC content. The agreement between the pathologist’s annotations and deconvolution results was also evident when visualizing the individual samples in more detail (Fig. 1f–h, Supplementary Figs. S2 and S3).
In summary, the estimated cell type abundances were consistent across technical replicates, and their spatial distribution aligned with the pathologist’s assessment for all samples.
Spatially resolved consensus molecular subtyping of CRC
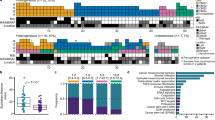
We further utilized the deconvolution results and pathologist’s annotations to spatially characterize the TME and CMS (Supplementary Figs. S4 and S5). CMS2 tumor cell proportions were predominant in patient samples S2_Col_R (94%), S4_Col_Sig (98%), S5_Rec (81%), and S6_Rec (90%) (Fig. 2a). A mixed abundance of CMS1 and CMS2 tumor cells was identified in patients S1_Cec (49% and 41%) and S3_Col_R (65% and 29%). Additionally, CMS3 tumor cells were detected in the S1_Cec (10%) and S5_Rec (16%) patients. In the non-neoplastic S7_Rec/Sig sample, the few spots exhibiting a tumor cell signature were mainly classified as CMS3 (60%). The prevalence of CMS4 was low and showed a multifocal distribution that overlapped with anatomical regions presenting an invasive phenotype. To characterize the TME composition, we next computed immune and stromal cell proportions (Fig. 2b–e). Mixed CMS1-CMS2 tumors exhibited higher T and B cell proportions, particularly CD8+ T and CD19+CD20+ B cells, consistent with the immune-rich phenotype associated with CMS15. Myofibroblasts were the dominant stromal cell type in mixed CMS1-CMS2 tumors, while the stromal cell types in CMS2 neoplasms were more heterogeneous. This is consistent with previous scRNA-seq studies reporting myofibroblast prevalence in CMS1 tumors6,7.
a–e Cell type proportions per sample as estimated by the results of the deconvolution. The number of spots containing an abundance of at least 20% of the specified cell types is also displayed. NK natural killers, Mac Macrophages, cDCs conventional dendritic cells. f Enrichment/depletion assessment of selected cell types (x-axis) in CMS2 and mixed CMS1-CMS2 tumors in the different tissue compartments defined by the pathologist’s spot classification (y-axis). IC immune cells. g–j Spatial mapping of the predicted abundance of CMS2 and CMS3 tumor cells and the module scores of the iCMS2-upregulated and the gastric metaplasia signatures overlaid with the pathologist’s tissue annotation in the S5_Rec_Rep1 sample. k Per spot Pearson’s cross-correlation across all the samples between TF activities and CMS cell abundances. For visualization purposes, the 10 most highly correlated TFs in absolute value per CMS are shown. l Per spot Pearson’s cross-correlation across all the samples between pathway activities and CMS cell abundances. m–o Spatial mapping of the predicted abundance of the CMS1 cells, the JAK-STAT pathway activity and the MAPK pathway activity overlaid with the pathologist’s tissue annotation in the S3_Col_R sample. p–r Spatial mapping of the predicted abundance of the CMS2 cells, the WNT pathway activity and the VEGF pathway activity overlaid with the pathologist’s tissue annotation in the S2_Col_R_Rep1 sample. s, t Spatial mapping of the predicted transcriptional activity of the MYC and E2F4 TFs overlaid with the pathologist’s tissue annotation in the S5_Rec_Rep1 sample. Note the colocalization with CMS2 tumor cell abundance (Fig. 2g).
We next associated these results with histological and morphological features by computing cell subtype enrichment in the pathologist-defined tissue compartments (Fig. 2f, Methods). CMS1 and CMS2 signatures were associated with tumor-annotated spots, while CMS3 signatures were confined to non-neoplastic mucosa. In CMS2-dominant tumors, immune cells were mostly found in the stroma, whereas in mixed CMS1-CMS2 tumors, CD19+CD20+ B and CD8+ T cells were also present in the neoplastic tissue. Irrespective of the CMS phenotype, SPP1+ macrophages and myofibroblasts were enriched in stromal fibrotic regions, echoing recent findings showing that proportions of these populations influence prognosis beyond CMS classification7.
We also connected our deconvolution-based CMS classification with the recently introduced IMF classification, which integrates intrinsic epithelial subtypes, microsatellite instability status, and fibrosis8. The predicted CMS2 abundance correlated significantly with the intrinsic epithelial subtype CMS2 (iCMS2) signature score (Fig. 2g, h, Supplementary Fig. S6). CMS2 - iCMS2 correspondence was additionally supported by mutational profiles (Table 1), anatomical location (Table 1), microsatellite instability status (Supplementary Fig. S7), and tubular adenoma and crypt bottom marker associations (Supplementary Fig. S8). Further, CMS3 signals were associated with key molecular features of iCMS3, including gastric metaplasia (Fig. 2i, j, Supplementary Fig. S9), upper crypt signals, and sessile serrated lesion markers (Supplementary Fig. S10).
Interestingly, we demonstrated that ST can spatially resolve known CMS-associated molecular features (Fig. 2k, l, Methods), such as the correlation between CMS1 tumor cell abundance and activity of the immune-related pathways JAK-STAT19 (Fig. 2m, n), TNFα20 and NFkB. Additionally, activation of the MAPK pathway (Fig. 2o), which is characteristic of the hypermutated CMS121, was observed. For CMS2 tumor cells, we identified their known association with the activation of the WNT and VEGF pathways22 (Fig. 2p–r) and higher expression of MYC- and E2F4-regulated genes5 (Fig. 2s, t).
Hence, our deconvolution-based approach spatially mapped the different CMS and TME cell types to their expected tissue compartments and associated them with key molecular and histological features.
ST reveals inter-patient and intra-patient heterogeneity of CRC tumors
To assess and further characterize the inter-patient heterogeneity among CMS2 tumors7,23, we extracted tumor-annotated spots from the CMS2-dominant carcinomas: S2_Col_R, S4_Col_Sig, S5_Rec, and S6_Rec (Supplementary Fig. S11). Although CMS2 cells dominated these spots with abundances ranging from 65% to 84% (Supplementary Fig. S12a, b), differential gene expression, pathway, and TF activity analyses (Fig. 3a–d, Supplementary Table S2) unveiled significant inter-patient differences. For instance, we found overrepresented mTORC1 signaling genes in tumors from the S4_Col_Sig and S5_Rec patients, but differentially expressed genes within this pathway suggested alternative signaling cascades (Supplementary Table S3). Notably, NUPR1, a promoter of metastasis through activation of the PTEN/AKT/mTOR pathway24, was highly expressed only in CMS2 tumor cells from the S4_Col_Sig patient (Fig. 3b). Tumor spots from the S2_Col_R and S4_Col_Sig patients showed lower EGFR signaling (Fig. 3c, Supplementary Fig. S12c), while FOXM1 displayed higher transcriptional activity in patient S6_Rec (Fig. 3d, Supplementary Fig. S12d).
a–d UMAP embeddings of the gene expression measurements in tumor annotated spots which were colored by different criteria: a per patient, b per the expression of the NUPR1 gene, c per activity of the EGFR pathway and d per activity of the FOXM1 TF. e Cell type proportions in the tumor-surrounding spots per sample as estimated by the results of the deconvolution approach. The number of tumor-surrounding spots for the different samples is also displayed. TA transient amplifiers. f Differential pathway activity computed on pseudo-bulk RNA-seq generated from the tumor-surrounding spots for the different samples. g–j Spatial mapping of the predicted abundance of CMS1, CMS2, CD19+CD20+ B cells and CD8+ T cells overlaid with the pathologist’s tissue annotation in the S3_Col_R sample. k Overlay of the spatial mapping of the clustering at subspot enhanced resolution of the tumor-annotated spots with the pathologist’s tissue annotations in the S5_Rec_Rep1 sample. l Spatial mapping and violin plots per group of the TGFb pathway activity at the enhanced subspot resolution in the S5_Rec_Rep1 sample. A Kruskal–Wallis statistical test was performed to assess whether the pathway activities in the different subclusters originated from the same distribution (p-value).
Inter-patient transcriptomic differences in CMS2 tumors can arise from inherent heterogeneity, anatomical origin and the composition and architecture of the TME. The latter can be uniquely assessed using ST. By selecting the spots surrounding CMS2 tumors, we assessed differential pathway activity among patients (Fig. 3e, f, Methods). The S5_REC patient exhibited a depletion of myofibroblasts (Supplementary Fig. S12e), potentially explaining its lower TGFβ pathway activity25. In S4_Col_Sig, the higher proportion of SPP1+ macrophages (Supplementary Fig. S12g), may contribute to an immunosuppressive TME26, in line with its lower activities in immune response-associated pathways such as NFκB and TNFα. The proportions and spatial distributions of these specific cell types are crucial as they drive clinical outcomes, with higher proportions linked to poorer prognosis7.
The assessment of the CMS1/CMS2 mixed sample S3_Col_R highlights the power of ST to characterize the CMS heterogeneity within a patient’s tumor and its associated morphologic features. CMS1-dominated regions displayed a solid growth pattern and immune-rich profile, whereas CMS2-dominated regions were associated with a tubular growth pattern and were immune-deprived (Fig. 3g–j, Supplementary Fig. S4d), in accordance with previous studies on these molecular subtypes27.
We subsequently addressed the intra-tumor heterogeneity in tumors displaying a pronounced CMS2 phenotype. To illustrate this, we categorized tumor-annotated spots from the S2_Col_R_Rep1 sample into peripheral, intermediate, and central tumor areas (Methods). As expected, genes involved in epithelial-mesenchymal transition (EMT) and angiogenesis, such as SPARC28, were significantly upregulated in the tumor boundary (Supplementary Fig. S13a, c, Supplementary Table S3, Methods). In contrast, the central tumor area showed an increased activity in hypoxic response and cholesterol homeostasis pathways, putatively driven by the upregulation of genes like SCD (Supplementary Fig. S13b, d, Supplementary Table S3). SCD upregulation was previously associated with the metabolic reprogramming necessary to promote metastasis of CRC cancer cells29. We finally sub-clustered the tumor-annotated spots extracted from S5_Rec_Rep1 (Fig. 3k, Methods) and identified regions with differentially expressed genes, biological processes, and pathway activities (Supplementary Figs. S14 and S15, Supplementary Table S4). Notably, CMS2-associated WNT and VEGF pathways displayed a more consistent distribution of their activities across tumor regions as compared to the activity of EGFR and MAPK pathways. Similarly, subcluster 1 demonstrated increased TGFβ pathway activity, suggesting tumor regions with higher proliferation and metastatic potential30 (Fig. 3l).
Together, our results demonstrate how ST unveils inter- and intra-tumor heterogeneity, TME architecture and spatial patterns of key molecular processes in CRC.
ST charts cell-to-cell communication processes involved in CMS2 tumor progression
The power of ST is that it reveals the cellular organization of the tissue at the molecular level, and thereby allows the study of cell communication events. We therefore explored these processes at the tumor-stroma interface and investigated their potential involvement in the tumor progression of the CMS2 subtype.
To study conserved biological processes across our CMS2 tumor samples (S2_Col_R; S4_Col_Sig; S5_Rec; S6_Rec), we merged and clustered their spots based on TF activity profiles (Fig. 4a–c, Supplementary Fig. S16, Methods). This approach revealed higher similarity as compared to gene expression-based clustering, and was hence used for our downstream analysis. Cluster 0, hereafter referred to as the tumor cluster, contained mainly spots annotated as tumor (49%) and tumor&stroma_IC med to high (26%) across replicates and patients (Fig. 4d, Supplementary FIg. S17a). Cluster 1, hereafter referred to as the TME cluster, predominantly included stromal annotated spots (63% as stroma_fibroblastic_IC med to high and 20% as tumor&stroma_IC med to high), neighboring the tumor in every sample (Fig. 4d, Supplementary Fig. S17a). As expected, MYC and E2F4 were highly activated TFs in the tumor cluster, while TFs such as JUN and ETS1, were identified in the TME cluster (Fig. 4b, c, Supplementary Fig. S17b).
a–c UMAP embedding of the TF activity profiles for our set of CMS2 samples. The spots were colored following different criteria: a per cluster group, b per activity of the MYC TF, and c per activity of the ETS1 TF. d Number of spots belonging to the different categories of pathologist’s annotations and clusters as inferred from the TF activity profiles. IC immune cells. e Misty results showing the potential importance of ligands (rows) expression on TF (columns) activity. The ligand-TFs relationships with an importance score over 1 are represented as black slots and were further investigated. Bold red characters were used to highlight the name of the ligands and TFs referred to in the main text. f Top predicted ligand-receptor interactions at the tumor stroma interface. The left panel shows the source of the interaction (ligands) and the right the target (receptors). g Signaling cascades potentially linking ligands (V shape) to their downstream TF targets (triangles) according to Misty predictions. The downstream signaling cascades go first through the top predicted receptors (diamonds) and then to intermediary signaling proteins (ellipses). The color of the nodes indicates the average expression of these genes in the TME cluster. Network edges can represent stimulatory (arrows) or inhibitory (squares) interactions. The edges representing interactions referred to in the main text are highlighted in black.
We then estimated the potential influence of ligands highly expressed in the tumor and TME compartments on the transcriptional activity of stroma-enriched TFs using Misty31 (Fig. 4e, Methods). We connected the most consistent ligand-TF associations to putative upstream signaling by predicting inter-cellular ligand-receptor interactions at the tumor-stroma interface and their known signaling pathways (Fig. 4f, g). To validate the ST-derived signaling events and to identify the involved cell types, we additionally estimated TF activity and ligand-receptor interactions in CMS2 patients from the Lee et al.’s scRNA-seq dataset6 (Fig. 5a–d, Supplementary Fig. S18a, Methods).
a Average TF activity per cell type. The percentage of cells of a given type where the TF is active is represented by the size of the circle. NK: natural killers, Mac: macrophages, cDCs: conventional dendritic cells, ECs: endothelial cells. b–d Ligand-receptor interactions between the different cell types overlapping with the interactions predicted in our ST data. The left panel shows the source of the interaction (ligands) and the right the target (receptors): b target cell types are myeloid cells, c target cell types are the major stromal cell populations, and d target cell types are the different CMS tumor cell types. Mac macrophages, cDCs conventional dendritic cells. e–g Overlay of the DCN gene expression, the predicted ETV4 TF activity and the metastasis score with the pathologist’s tissue annotations in the S2_Col_R_Rep1 sample. A red square highlights the tumor region exhibiting invasive morphological traits. h–j Overlay of the RNF43 gene expression, the predicted JUN TF activity and the metastasis score with the pathologist’s tissue annotations in the S6_Rec_Rep2 sample.
Our results suggested that decorin (DCN), a proteoglycan secreted by stromal cells, triggers a protective pathway inhibiting tumor progression in the CMS2 subtype. DCN interacts with receptors like EGFR, IGF1R and MET, promoting their degradation and impairing downstream signaling, as described in previous studies32. The DCN-EGFR-SRC-STK11, DCN-EGFR-PRKDC-HMGB1-HOXD9 and DCN-MET-STAT3 signaling axis may modulate the transcriptional activity of ETV4, MEIS1 and SPI1 respectively, as supported by our findings in the ST and scRNA-seq data (Figs. 4e–g, 5a–f, Supplementary Fig. S18b–d). Increased activity of these TFs is associated with greater tumor invasiveness33,34,35. The spatial mapping of ETV4 transcriptional activity revealed overall low levels within the tumor, excepting for a region exhibiting invasive morphological traits and higher macrophage infiltration (Fig. 5e, f, Supplementary Figs. S2b and S18f). Our findings capture DCN’s effects on these macrophages through its interaction with the TLR2 and TLR4 receptors (Figs. 4f, 5b, Supplementary Fig. S18e). In summary, our results highlight DCN’s pivotal role in tumor suppression, particularly in CMS2 regions with elevated invasiveness potential (Fig. 5g).
Moreover, our data indicated that the CMS2-associated RNF43, a transmembrane protein, might influence several TFs within the TME, including JUN and TEAD4 (Figs. 4e, 5h, i, Supplementary Fig. S19a, b). Notably, these TFs are involved in tumor progression and associated with WNT signaling36,37. We predicted an RNF43-FZD2 interaction targeting stromal cell populations (Figs. 4f, 5c, Supplementary S19c, d), and signaling cascades connecting these elements, such as the FZD2-DVL3 and the YAP-TEAD4 interactions (Fig. 4g). In summary, elevated RNF43 expression increases WNT receptor degradation, affecting downstream transcriptional activity, and potentially indicating anatomical regions with lower metastatic activity (Fig. 5j).
In addition, we identified other ligand-TF pairs potentially modulating CMS2 tumor progression. For instance, the THBS2-CD36 interaction, known to inhibit angiogenic processes38, may modulate STAT1 activity (Fig. 4e, f, Supplementary Fig. S19e–h). The expression of MMP1, a matrix metalloproteinase involved in cancer progression through degradation of the extracellular matrix39, was predicted to have an effect on the activity of the FOS TF (Fig. 4e, Supplementary Fig. S18g, h). The PLAU-PLAUR interaction was identified between myofibroblasts and macrophages or conventional dendritic cells (Figs. 4f, 5b, Supplementary Fig. S18i–k), consistent with prior studies in prostate cancer, associating this interaction with macrophage infiltration and tumor progression40. Moreover, we found that chemokine CXCL14 could influence MAF transcriptional activity (Fig. 4e), which was shown to regulate the immunosuppressive function of tumor-associated macrophages41. Interestingly, a CXCL14-based peptide has previously been suggested as a potential cancer treatment42.
In conclusion, our results generate mechanistic hypotheses on how highly expressed ligands in CMS2 tumors and their TME may trigger signaling cascades modulating TFs involved in cancer progression.
Deconvolution-based subtyping, heterogeneity and cell communication events confirmed in independent CRC cohort
To corroborate our findings, we analyzed an independent ST dataset14, comprising four primary CRC tumors exhibiting morphological features indicative of CMS2, along with their corresponding liver metastases. The samples were obtained from two untreated (Unt) and two neoadjuvant chemotherapy-treated patients (Tre).
We first applied our deconvolution-based approach to profile this dataset (Fig. 6a). Major cell type proportions revealed a reduced tumor content of approximately 4% in ST-colon2_Unt, ST-colon3_Tre, and ST-liver3_Tre samples, in accordance with their histology. All samples, including the liver metastases, predominantly exhibited a CMS2 phenotype, with over 80% of tumor cells mapped to this subtype (Fig. 6b, c, Supplementary Fig. S20). In agreement with our previous results, CMS3 signatures were restricted to the non-neoplastic mucosa and CMS4 signals were minor and multifocally distributed. The CMS1 presence was almost negligible in these samples. Notably, substantial CMS2 and iCMS2 signals overlapped with the liver tumor histology, suggesting a conservation of the CMS phenotype in metastasis (Fig. 6d, Supplementary Figs. S21–S22). We further characterized these samples by analyzing the relative abundance of the different types of T cells, B cells, myeloid cells and the main stromal cells (Supplementary Fig. S23).
a Proportions of major cell classes per sample as estimated by the results of the deconvolution. The right hand side of the plot displays the number of analyzed spots per sample. b CMS tumor cell type proportions per sample as estimated by the results of the deconvolution approach. The number of spots containing an abundance of at least 20% of tumor cells subtypes is also displayed. c, d Overlay of the spatial mapping of the predicted CMS2 tumor cell abundance with the pathologist’s tissue annotations in the ST-colon1_Unt and ST-liver1_Unt samples. e Per spot Pearson’s cross-correlation across all the samples between pathway activities and CMS cell abundances. f Per spot Pearson’s cross-correlation across all the samples between TF activities and CMS cell abundances. For visualization purposes, the 10 most highly correlated TFs in absolute value per CMS are shown. g Overlay of the spatial mapping of the predicted WNT pathway activity with the pathologist’s tissue annotations in the ST-colon1_Unt sample. h Overlay of the spatial mapping of the predicted MYC TF activity with the pathologist’s tissue annotations in the ST-colon2_Unt sample. i Overlay of the spatial mapping of the predicted MAPK pathway activity with the pathologist’s tissue annotations in the ST-liver1_Unt sample. j Overlay of the spatial mapping of the predicted NR2C2 TF activity with the pathologist’s tissue annotations in the ST-liver2_Unt sample. k Misty results showing the potential importance of ligands (rows) expression on TF (columns) activity when considering the samples from primary CRC tumors. The ligand-TFs relationships with an importance score over 1 are represented as black slots and were considered as relevant. The ligands and TFs discussed in the results sections are highlighted in red. l, m Overlay of the spatial mapping of the RNF43 gene expression and the predicted TEAD1 TF activity with the pathologist’s tissue annotations in the ST-colon4_Tre sample. n, o Overlay of the spatial mapping of the DCN gene expression and the predicted ETV4 TF activity with the pathologist’s tissue annotations in the ST-liver4_Tre.
Next, we spatially mapped CRC-associated molecular features and assessed their correlation with the CMS cell abundance jointly in primary and hepatic metastatic tumors, focusing on the prevalent CMS2 subtype (Fig. 6e, f, Methods). As a result, we verified the activation of WNT and VEGF pathways in CMS2-rich regions and confirmed the activity of MYC and E2F4 transcription factors in CMS2 tumors (Fig. 6g, h, Supplementary Fig. S24a, b). Moreover, we noticed a link between the estimated CMS2 cells and the activity of the MAPK pathway and NR2C2 TF (Fig. 6i, j). This finding is consistent with our primary sample set (Fig. 2k, l) and of particular interest as their role in CMS2 tumors is not clearly defined.
We also used the external dataset to validate selected cell-to-cell communication processes previously identified, specifically the ligand-TF regulations. Using primary CRC tumors, we confirmed the modulation of JUN and TEAD family transcriptional activity by RNF43 expression, and the potential influence of DCN on ETV4 activity (Fig. 6k–m, Supplementary Fig. S24c, d). We also confirmed the potential downstream impact of the CXCL14 chemokine on MAF’s transcriptional activity (Fig. 6k, Supplementary Fig. S24e, f). Notably, we found that ETV4 and JUN’s transcriptional activity regulation by DCN and RNF43, respectively, was preserved in the liver metastatic samples (Fig. 6n, o, Supplementary Fig. S24g–i). These findings align with a recent study describing the protective role of DCN in hepatic metastasis of CRC43 and may provide new insights into the underlying molecular mechanisms.
Overall, the main findings of our study were indeed validated in an independent ST CRC dataset.
Discussion
The clinical need for accurate CRC patient stratification led to the development of several gene expression-based classification systems, such as the CMS5 or the IMF8. The CMS classification system is broadly used and has helped to understand the different molecular mechanisms underlying CRC and disease prognosis44. Nevertheless, CMS intra-tumor heterogeneity hampers its clinical application, underlining the necessity of further characterizing the cellular composition and architecture of CRC and its microenvironment.
To complement our understanding of CRC CMS, we combined ST and scRNA-seq via cell type deconvolution, elucidating subtype-inherent transcriptomic and morphological features. This allowed us to map CMS1 and CMS2 tumor cells to neoplastic areas exhibiting distinct morphological features. In contrast, CMS3 signatures were confined to the non-neoplastic mucosa, which might be related to their normal-like expression patterns5. The EMT-associated CMS4 signals were minimal and overlapped with invasive tumor regions, in line with previous studies referring to CMS4 as a transcriptional state of stromal cells rather than tumor-like epithelial cells10,45. This reduced signal made it challenging to observe typical CMS4 molecular features such as TGFb pathway activation in our integrated analysis (Figs. 2l and 6e), though such features are evident in individual samples (Supplementary Fig. S25a). Across various samples, we observed a co-existence of the different subtypes in line with recent findings suggesting that CRC is more accurately represented by a transcriptomic continuum than by discrete subtypes7. Indeed, the bulk RNA-based classification of our analyzed samples emphasizes the significant influence of the surrounding tissue on tumor classification (Supplementary Fig. S25b, c). The S6_Rec patient samples illustrate this, with small tumor islands enveloped by large stroma bundles, leading to a CMS2 classification via deconvolution but a CMS4 assignment by CMScaller46. This morphology hampers the separation of the tumor components in bulk RNA-seq data, whereas ST can provide their detailed assessment. The CMS4 classification of stroma-rich tumors is in accordance with previous studies linking CMS4 signatures with marker genes of cancer-associated fibroblast and other stromal cells47. Similarly, the external ST-colon4_Tre sample, classified as CMS2 by deconvolution but CMS3 by CMScaller, raises concerns about the impact of non-neoplastic mucosa, which contains CMS3 signals, on bulk-based CMS classification systems.
Overall, our results underline the potential of ST in CRC characterization beyond bulk- or scRNA-seq, enabling the spatial correlation of morphological tumor, stroma and non-neoplastic tissue patterns with corresponding transcriptomic features. Nevertheless, limitations inherent to our deconvolution-based approach should be acknowledged. Firstly, the choice of the scRNA-seq reference can significantly impact the deconvolution results. We compared the results yielded by two similarly annotated reference datasets6 in Supplementary Note 1. The overall results were highly comparable, but some discrepancies were observed for particular cell types, e.g. CMS1 tumor cells. Factors such as the differences in the genetic background between both cohorts could contribute to these discrepancies. Secondly, and regardless of the used reference, the deconvolution partially failed to map stromal cells on their expected anatomical location, especially in the S3_Col_R sample. This can be attributed to the absence of specific stromal cell types in the reference or due to a decrease in deconvolution sensitivity in regions with lower transcripts per spot, as a result from tissue properties or technical variabilities (Supplementary Fig. S26). Finally, the current size of 10x VISIUM spots makes region-specific assignment challenging, as seen in samples from the S6_Rec patient, where its unique morphology complicates pure tumor spot annotation. This may cause interpatient tumor expression differences due to residual stromal cells. It possibly explains the elevated FOXM1 transcriptional activity and the mixed CMS2 and stromal-related signatures in cluster 6, unique to S6_Rec in our TF activity-based clustering (Figs. 3d and 4a–d).
We also explored the ability of ST to scrutinize ligand-receptor interactions at the tumor-stroma interface, which might trigger signaling pathways critical for tumor progression. Our results encompass a range of novel and well-known tumor growth-inhibiting as well as -activating signatures, such as the potential regulation of the ETV4 transcriptional activity by DCN or the PLAU-PLAUR ligand-receptor interaction. While these predictions may guide the identification of potential therapeutic targets, they require further investigation as our methodology of spatially modeling TF activity based on ligand gene expression may not necessarily reflect direct causal regulations. Along the same line, the ligand-receptor analysis could also capture indirect gene expression associations. For instance, we consistently predicted the RNF43-FZD2 interaction targeting stromal cell populations in both ST and scRNA-seq data. However, this interaction is mostly reported to occur in the intracellular domain of RNF43 in tumor cells48, with few studies reporting a potential extracellular interaction49.
To support our key findings, we used an independent ST CRC dataset. Interestingly, our deconvolution approach delineated the primary, but also the metastatic carcinomas, as CMS2. In these liver tumors, we captured the CMS2 main molecular features and preserved cell communication events as the modulation of the transcriptional activity of ETV4 by DCN. This suggested that the CMS2 phenotype was largely retained after migration of the primary CRC cells to sites of metastasis.
In conclusion, our study illustrates the value of integrating ST and scRNA-seq in analyzing CRC and its CMS, providing insights into spatial cellular organization within tumors and their TME. Although the small patient cohort limits the scope of our study, we envision that our proof-of-concept work demonstrates ST’s potential to inform patient-specific treatment strategies. More refined patient stratification could be achieved by jointly considering cell composition, spatial distribution and morphological features. In addition, understanding intra-tumor spatial heterogeneity can unveil anatomically restricted or region-specific progression-related processes, fueling the development of novel therapies, such as targeted or combination treatments. As ST technologies evolve in resolution, affordability, and clinical validation, we anticipate its application to larger CRC cohorts, paving the way towards personalized oncology.
Methods
Collection of CRC samples
Human CRC tissues (<8 months storage) and annotated data were obtained and experimental procedures were performed within the framework of the non-profit foundation HTCR (Human Tissue and Cell Research) Foundation50. This includes written informed consent from all donors and the approval by the ethics commission of the Faculty of Medicine in the Ludwig Maximilian University of Munich (Number 025-12) and the Bavarian State Medical Association (Number 11142). Sampling and handling of any patient material was performed in accordance with the ethical principles of the Declaration of Helsinki. Tissues were cut on a Cryostat (CryoStar NX70, Thermo Scientific) at 10 um. A pathologist performed quality and comparability assessment of fresh-frozen material using a hematoxylin-eosin (H&E) stained slide.
Sample preparation
RNA from all samples was extracted using the Arcturus® PicoPure® RNA Isolation Kit (Applied Biosystems™, KIT0204). For cell lysis, a 10 um section of the sample was resuspended in a 200 ul extraction buffer. Total RNA was extracted following the instructions of the manual. RNA integrity number (RIN) was assessed using the 2100 Bioanalyzer system (Agilent Technologies, Inc.) with an Agilent RNA 6000 Pico Kit (Agilent Technologies, Inc., 5067-1513). Samples with RIN above 7.0 were used.
Tissue optimization was carried out according to the manufacturer’s instructions (VISIUM Spatial Tissue Optimization User Guide_RevC). Image acquisition was performed on the Hamamatsu NanoZoomer S 360 C13220 series at 40x magnification and the coverslip was removed afterwards by immersing the slide in a 3x Saline-Sodium Citrate buffer. The stained tissue sections were permeabilized using a time course to test for the optimal permeabilization time. After performing a fluorescent cDNA synthesis, the tissue was removed. Finally, the fluorescent cDNA was imaged using a Zeiss Axio Scan.Z1 with a Plan Apochromat 20×/0.8 M objective, an ET-Gold FISH filter (ex 538–551 nm/em 556–560 nm) and 100 ms exposure time.
For the gene expression analysis, 10 um thick sections of the samples were placed with a random distribution over four chilled 10x Genomics VISIUM Gene Expression slides containing four capture areas each. The sections were similarly stained with H&E and subsequently imaged as described above. To release the mRNA, the sections were permeabilized for 30 min as defined by tissue optimization. For further processing, the cDNA was amplified according to the manufacturer’s protocol (CG000239_VisiumSpatialGeneExpression_UserGuide_RevC). Double indexed libraries were prepared. The libraries were quality controlled using a 2100 Bioanalyzer system with Agilent High Sensitivity DNA Kit (Agilent Technologies, Inc., 5067-4626) and quantified with Qubit™ 1X dsDNA HS Assay Kit (Invitrogen, Q33230) on a Qubit 4 Fluorometer (Invitrogen, Q33238). The libraries were loaded onto the NovaSeq 6000 (Illumina) at a concentration of 250 pM. A NovaSeq S1 v 1.5 or SP v 1.5 Reagent Kit (100 cycles) (Illumina, 20028319 and 20028401) was used. For paired end-dual indexed sequencing, the following read protocol was used: read 1: 28 cycles; i7 index read: 10 cycles; i5 index read: 10 cycles; and read 2: 90 cycles. All libraries were sequenced at a minimum of 50000 reads per covered spot.
Raw sequencing data were demultiplexed using the mkfastq function from Space Ranger (v. 1.2.0). Demultiplexed data were mapped to the human reference GRCh38 with spaceranger count. Spots under tissue folds, artifacts and at the tissue boundary were manually removed using the 10X Loupe browser (v. 5.1.0).
Histopathological annotations and spot categorization
H&E stained tissue sections were annotated by the pathologist using QuPath software (v. 0.2.3)51. Spot categorization was performed by the pathologist using the 10X Loupe browser (v. 5.1.0). Categories and corresponding criteria are listed in Supplementary Table S5.
Grading of CMS signatures
Grading of CMS signatures in the tumor tissue was performed semi-quantitatively according to the number of spots with positive signature and the percentage of positive cells per spot. This grading was done in an individual replicate per patient (S1_Cec_Rep1, S2_Col_R_Rep1, S3_Col_R_Rep1, S4_Col_Sig_Rep1, S5_Rec_Rep1, S6_Rec_Rep2 and S7_Rec/Sig_Rep1) according to the scheme detailed in Supplementary Table S6.
ST data pre-processing
We used the Seurat52, Scanpy53 and SingleCellExperiment54 packages to load the output of the Space Ranger pipeline and process the ST data. We evaluated the quality of the ST data by determining the average number of reads, UMIs and genes per spot covered by tissue and compared it with those from spots non covered by tissue. We found substandard quality for the S1_Cec_Rep2 sample as revealed by its low numbers of unique molecular identifier (UMI) counts and genes in spots covered by tissue (Supplementary Fig. S1). Consequently, this sample was either treated carefully or excluded from integrative analysis. For each individual sample, we filtered out spots for which the number of UMI counts detected were below 500 or above 45000. In addition, spots containing a fraction of more than 0.5 mitochondrial genes were not considered in the analysis. We normalized the UMI counts from the remaining spots using SCTransform55.
Sample integration, batch correction and dimensionality reduction
To jointly represent the CRC samples in the same low dimensional space (UMAP embedding), correct from batch effects and integrate samples and technical replicates for downstream analysis, we used Harmony56. We ran Harmony with default parameters allowing a maximum number of 20 interactions (max.iter.harmony = 20) and correcting per individual samples. Of note, Harmony was either applied to batch-correct for all the spots derived from all the samples or to batch-correct only the tumor annotated spots from a subset of samples (CMS2 tumor samples).
Deconvolution of the ST datasets
ST datasets derived from 10x Genomics VISIUM technology currently lack single cell resolution. Therefore, the gene expression values detected per spot originate from a variable number of different cells, i.e. every spot can be considered as a mini-bulk RNAseq dataset. Consequently, a deconvolution approach is required to estimate the different cell types and their proportions across spots.
To this end, we used the recently proposed Cell2Location (v 0.0.5)18 method. Cell2location first creates gene expression signatures of cell types from a scRNA-seq reference. We adopted as scRNA-Seq reference a comprehensive dataset from a recent publication exploring the cellular landscape of the different CRC subtypes and their microenvironment6. The annotations from the original publication at the cell subtype level (Supplementary Table S1) were used to generate the signature using the run_regression function with the following parameters: n_epochs = 100, minibatch_size = 1024, learning_rate = 0.01 and train_proportion = 0.9. These signatures are subsequently used to assess cell type abundances in the ST data using the run_cell2location with selection_specificity = 0.20. This parameter determines the number of genes used to establish the signature per cell type (Supplementary Table S1). Additional parameters were set as follows: n_iter = 40000, cells_per_spot = 8, factors_per_spot = 9, combs_per_spot: 5, mean = 1/2 and sd = 1/4.
Consistency of deconvolution results between technical replicates
To evaluate the consistency of the deconvolution between technical replicates, we batch-corrected their transcriptomic profiles using Harmony56 as described above. Then, we clustered the Harmony embeddings using the Louvain algorithm as encoded in the FindClusters function from the Seurat package. We chose a series of large resolution parameters (ranging from 1 to 2 increasing by 0.1 steps) to obtain fine-grain clusters that can match with anatomical regions displaying similar cell type distribution patterns across replicates. Finally, we computed the mean number of UMIs estimated by Cell2Location per cell type and cluster, and applied Pearson’s correlation to evaluate their similarity between technical replicates.
Enrichment/depletion of cell types in different anatomical regions
The enrichment (depletion) in the abundance of the deconvolution-estimated cell types in different pathologist-assigned tissue categories was assessed following a similar procedure to be one described in Andersson et al.11. Briefly, the estimated cell type proportions per spot were 10 000 times randomly shuffled with respect to their spatial location. Then, we computed the average cell type proportions per permutation and tissue type. The mean value of differences between the real and the permuted average proportions divided by the standard deviation of these differences was used as the enrichment score for the different tissue categories.
Pathway activity
We estimated pathway activity per spot and at subspot resolution (see section Clustering and enhanced gene expression at the sub spot level) using PROGENy57. PROGENy computes pathway activity by accounting for the expression of genes which are more responsive to perturbations on those pathways. The PROGENy model comprises 14 pathways, namely: Wnt, VEGF, Trail, TNFα, TGFβ, PI3K, p53, NFkB, MAPK, JAK/STAT, Hypoxia, Estrogen, Androgen and EGFR. In our setup, we ran PROGENy using the top 500 most responsive genes per pathway.
In addition, we also computed pathway activities in pseudo-bulk generated from our ST samples (see section Pseudo-bulk generation). We again used the top 500 most responsive genes per pathway. In this case, we set the scale parameter to TRUE to allow direct comparison of pathway activities between samples.
Transcription factor activity
We computed TF activity per spot using the Viper58 algorithm coupled with regulons extracted from DoRothEA59. In DoRothEA, every TF–target interaction is assigned a confidence score based on the reliability of its source, which ranges from A (most reliable) to E (least reliable). In this study, we selected interactions with confidence scores A, B and C and computed the activity for TFs with at least four different targets expressed per spot.
The activity profiles of the different TFs were additionally used to cluster the spots from our four CMS2 tumor samples. To do so, the TF activity scores from these samples were first merged and subsequently scaled and centered. Then, the standard procedure to compute clustering using the Seurat package was followed. Briefly, we computed a Principal Component Analysis (PCA) dimensionality reduction on the scaled TF activities per spot followed by the computation of the 20 nearest neighbors. Finally, we applied the Louvain algorithm with a resolution parameter of 0.5 to group the spots into different clusters according to their TF activity profile. We identified TF with a differential activity profile among the different clusters using Receiver Operating Characteristic (ROC) analysis as implemented in the Seurat’s FindAllMarkers function. We only considered TF whose activity was computed in at least 25% of the spots per cluster and with a log2 fold-change greater than 1.
Of note, we used the same procedure to compute TF activity per cell on the scRNA-seq dataset from Lee et al.6.
Canonical correlation analysis
We used the cc function from the CCA package60 to compute canonical correlation between the cell type proportions per spot and pathway or TF activity per spot. This canonical correlation analysis was first performed for every individual CRC sample. To capture global correlations across samples, we performed an integrative analysis by merging spots coming from all the different samples (excluding S1_Cec_Rep2) into matrices and computing the canonical correlation on them.
Selection of tumor surrounding spots
We applied the GetTissueCoordinates function from the Seurat package to get the spatial coordinates of the spots in the different CRC samples. We subsequently computed the Euclidean distance between every pair of spots. Finally, we selected as tumor-surrounding-spots those lying within a distance smaller or equal to 2 from a tumor annotated spot. Spots fulfilling these criteria but annotated as tumors were discarded.
Pseudo-bulk generation
We generated pseudo-bulk from the ST samples using the sumCountsAcrossCells function from the Scater package61. Here, counts were normalized by the total number of reads (counts per million normalization). We used the filterByExpr function from the edgeR package62 to filter out genes with less than 50 counts per sample.
Definition of different anatomical regions in tumor annotated spots
The distance between every tumor annotated spot and non-tumor annotated spots was calculated as described in section Selection of tumor surrounding spots. We then defined the different tumor anatomical regions for the S2_Col_R_Rep1 sample based on the following criteria:
-
Peripheral Tumor: tumor spots in direct contact with at least a non-tumor annotated spot. Their Euclidean distance to a non-tumor annotated spot is smaller than 2.
-
Central Tumor: tumor spots in the most solid and internal region of the tumor. Their Euclidean distance to a non-tumor annotated spot is greater than 2.5.
-
Intermediary Tumor: tumor spots that we consider as a transition region between the inner and outer tumor. Their Euclidean distance to a non-tumor annotated spot is greater or equal to 2 and smaller than 2.5.
Clustering and enhanced gene expression at the sub spot level
We applied BayesSpace63 to cluster at the subspot level and increase the gene expression resolution of our CMS2 tumor annotated spots in the S5_Rec_Rep1 sample. To do so, BayesSpace uses the neighborhood structure in spatial transcriptomic data. Of note, the preprocessing of the ST raw data was conducted following the recommendations of BayesSpace authors. This procedure is slightly different from the one described in previous sections. Briefly, the ST data was processed using the SingleCellExperiment package and raw counts were log normalized using the logNormCounts function from the Scuttle package61. Then, the Scran64 package was used to model the variance of the log-expression profiles for each gene and select the 2000 most variable genes. We performed a PCA using the Scater61 package.
Using BayesSpace, we subsequently computed the spatial clustering and the enhanced clustering with default parameters, excepting the jitter_scale parameter which was set to 3. Finally, we enhanced the gene expression of all the genes expressed in the considered spots using the enhanceFeatures function with default parameters.
Differential gene expression analysis
The CMS2 tumor regions extracted from the different samples were integrated into the same Seurat52 object. We used the Wilcoxon Rank Sum test to identify differentially expressed genes between the groups of spots coming from different patients as implemented in the Seurat’s FindAllMarkers function. We set a log2 fold-change threshold of 0.25 and only positive markers were retrieved. Some specific criteria were followed for the analyses conducted in section 2.3:
-
To describe inter-patient heterogeneity, the differential gene expression analysis was performed between the different patients (two replicates per patient considered). We filtered results by only considering genes that are overexpressed in tumor annotated spots versus non-tumor annotated spots. To do so, we took advantage of the pathologist’s annotations and used the Seurat’s FindMarkers with the same parameters described above for the FindAllMarkers function. Ribosomal and mitochondrial genes were removed due to the fact that they can be overrepresented in tumor necrotic regions.
-
To describe intra-tumor heterogeneity, the differential expression analysis was carried out between the different anatomical regions of the tumor in the S2_Col_R_Rep1 sample (see section Definition of different anatomical regions in tumor annotated spots) with no further considerations.
-
Another differential gene expression analysis was conducted on the enhanced gene expression between the different enhanced clusters generated by BayesSpace (see section Clustering and enhanced gene expression at the sub spot level) on the S5_Rec_Rep1 sample. We selected for further analysis genes with an adjusted p-value smaller than 0.01 in the Wilcoxon Rank Sum test. Ribosomal and mitochondrial genes were excluded from the analysis.
Gene set overrepresentation analysis
Differentially expressed genes were subsequently used for gene set overrepresentation analysis using the Hallmark annotations from MSigDB65. The Hallmark gene sets contain 50 well-defined biological states or processes. We used the enricher function from the clusterProfiler66 package to carry out the analysis. We set a minimal size of the genes annotated for testing to five, excepting for the analysis between different patients where it was set to three. Background genes were adjusted accordingly to the global set of genes expressed in the different contexts.
Ligand modulation of TF activity
As a first step and taking as reference the TF activity-based clustering, we selected ligands which are overexpressed in the tumor and TME with respect to the other anatomical regions across all our CRC samples. To do so, we applied the Seurat’s FindMarkers function with a log2 fold-change threshold of 0.5 and only positive markers were retrieved. We matched our set of overexpressed genes against the set of proteins annotated as ligands in the Omnipath67 database. Additionally, we filtered out ligands that are not detected in at least 10% of the tumor and TME spots in every individual sample.
In the second place, we chose TFs with a higher differential activity profile in the TME regions across all the samples according to the clustering approach described in section Transcription factor activity. In particular, we selected those TFs that are considered as markers of the TME cluster when using the Seurat’s FindAllMarkers function (AUC ≥ 0.75).
We then applied Misty31 to investigate the potential effect of the expression of the selected ligands in modulating the transcriptional activity of the chosen TFs. Specifically, we created an intrinsic view (intraview) describing ligand gene expression and a local niche view (juxtaview) using TF activity with a neighbor.thr = 2 aiming at capturing effects in the direct neighborhood of each spot. This criteria is based on the fact that many cancer relevant ligands are membrane bound and that the majority of secreted ligands cannot travel long distances. Following this approach, Misty was first individually applied to every sample. Then, the individual results were collected and aggregated using Misty’s collect_results function in order to obtain the most robust common signals across samples. Ligand-TF associations with an aggregated importance greater than 1 were considered for further analysis. Of note, when running Misty on the external dataset, the ST-colon3-Tre and ST-liver3-Tre samples were excluded from the analysis due to their reduced tumor content.
Prediction of ligand-receptor interactions
We used LIANA68 to estimate the most likely ligand-receptor interactions between the different spatial clusters defined by their TF activity profiles. It is to note that the interactions were computed for every pair of clusters, but for subsequent analysis and visualization we focused on the interactions between the clusters labeled as 0 (Tumor) and 1 (TME). LIANA computes an aggregated score for every potential ligand-receptor interaction based on the results of different methods. In our particular case, we ran LIANA with default settings and used OmniPath67 as a source of prior knowledge in human ligand-receptor interactions. For further analysis, we considered interactions involving Misty’s predicted ligands with an aggregated rank smaller than 0.01, as this value can be seen as analogous to a p-value69. We also ran LIANA on the scRNA-seq dataset from Lee et al.6 using the same procedure.
Inference of signaling networks
We used a network-based approach to infer the most likely signaling cascades linking LIANA’s predicted ligand-receptor interactions to their targeted TFs according to Misty’s predictions. To do so, we first built an intra-cellular signaling network by retrieving protein-protein interactions from Omnipath67. Then, for every ligand, we selected their predicted receptors and targeted TFs. We subsequently connected every receptor to every corresponding TF by selecting the shortest path between them in the signaling network. All the resultant shortest paths were merged into a network together with the previously predicted ligand-receptor interactions. Finally, for every gene in the predicted network, we computed its average expression in the TME cluster, as defined by TF activity profiles (see section Transcription factor activity), across all the CMS2 samples. Cytoscape70 was utilized for the visualization of the network.
Metagenes/module scores
We computed module scores for different sets of genes using the Seurat’s AddModuleScore function. We detail below the particular gene sets used:
-
The list of up-regulated genes in iCMS2 and iCMS3, as well as, the markers involved in gastric metaplasia were extracted from the study where the IMF classification system was introduced8.
-
The list of genes associated with tubular adenomas or with sessile serrated lesions were extracted from Chen et al.71.
-
We fetched the crypt bottom and upper markers from Kosinski et al.72.
-
We retrieved a list of genes linked to metastatic processes from CancerSEA73.
Prediction of microsatellite status
We inferred microsatellite instability status by running Microsatellite instability Absolute single sample Predictor (MAP)74 on pseudo-bulk generated from our ST samples (see section Pseudo-bulk generation). They were classified as microsatellite instable (MSI) or microsatellite stable (MSS).
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The output of Space Ranger, including processed count data matrices and histological images, for the ST data generated in this study is available at https://doi.org/10.5281/zenodo.7551712. In addition, this repository also contains the spot categorization made by the pathologist. The processed scRNA-seq and metadata used for the deconvolution and for further characterization of the cell communication processes are available via the GEO database under the accession codes GSE132465 and GSE1447356. The processed data from the external ST CRC dataset used to support our findings was downloaded from http://www.cancerdiversity.asia/scCRLM14.
Code availability
The scripts containing all the code used to generate the results presented in this study are available at https://github.com/alberto-valdeolivas/ST_CRC_CMS. Their associated notebooks containing additional results and information about the versions of the different packages used are available at https://doi.org/10.5281/zenodo.7440182. Finally, Intermediary object files to reproduce the analysis are available at https://doi.org/10.5281/zenodo.7551712.
References
Biller, L. H. & Schrag, D. Diagnosis and treatment of metastatic colorectal cancer: a review. JAMA 325, 669–685 (2021).
Wang, W. et al. Molecular subtyping of colorectal cancer: recent progress, new challenges and emerging opportunities. Semin. Cancer Biol. 55, 37–52 (2019).
Okita, A. et al. Consensus molecular subtypes classification of colorectal cancer as a predictive factor for chemotherapeutic efficacy against metastatic colorectal cancer. Oncotarget 9, 18698–18711 (2018).
Chan, D. K. H. & Buczacki, S. J. A. Tumour heterogeneity and evolutionary dynamics in colorectal cancer. Oncogenesis 10, 1–9 (2021).
Guinney, J. et al. The consensus molecular subtypes of colorectal cancer. Nat. Med. 21, 1350–1356 (2015).
Lee, H.-O. et al. Lineage-dependent gene expression programs influence the immune landscape of colorectal cancer. Nat. Genet. 52, 594–603 (2020).
Khaliq, A. M. et al. Refining colorectal cancer classification and clinical stratification through a single-cell atlas. Genome Biol. 23, 1–30 (2022).
Joanito, I. et al. Single-cell and bulk transcriptome sequencing identifies two epithelial tumor cell states and refines the consensus molecular classification of colorectal cancer. Nat. Genet. 54, 963–975 (2022).
Cañellas-Socias, A. et al. Metastatic recurrence in colorectal cancer arises from residual EMP1+ cells. Nature 611, 603–613 (2022).
Chowdhury, S. et al. Implications of intratumor heterogeneity on consensus molecular subtype (CMS) in colorectal cancer. Cancers 13, 4923 (2021).
Andersson, A. et al. Spatial deconvolution of HER2-positive breast cancer delineates tumor-associated cell type interactions. Nat. Commun. 12, 1–14 (2021).
Berglund, E. et al. Spatial maps of prostate cancer transcriptomes reveal an unexplored landscape of heterogeneity. Nat. Commun. 9, 1–13 (2018).
Hunter, M. V., Moncada, R., Weiss, J. M., Yanai, I. & White, R. M. Spatially resolved transcriptomics reveals the architecture of the tumor-microenvironment interface. Nat. Commun. 12, 1–16 (2021).
Wu, Y. et al. Spatiotemporal immune landscape of colorectal cancer liver metastasis at single-cell level. Cancer Discov. 12, 134–153 (2022).
Peng, Z., Ye, M., Ding, H., Feng, Z. & Hu, K. Spatial transcriptomics atlas reveals the crosstalk between cancer-associated fibroblasts and tumor microenvironment components in colorectal cancer. J. Transl. Med. 20, 302 (2022).
Qi, J. et al. Single-cell and spatial analysis reveal interaction of FAP fibroblasts and SPP1 macrophages in colorectal cancer. Nat. Commun. 13, 1742 (2022).
Zhang, R. et al. Spatial transcriptome unveils a discontinuous inflammatory pattern in proficient mismatch repair colorectal adenocarcinoma. Fundam. Res. https://doi.org/10.1016/j.fmre.2022.01.036 (2022).
Kleshchevnikov, V. et al. Cell2location maps fine-grained cell types in spatial transcriptomics. Nat. Biotechnol. https://doi.org/10.1038/s41587-021-01139-4 (2022).
Mevizou, R., Sirvent, A. & Roche, S. Control of tyrosine kinase signalling by small adaptors in colorectal cancer. Cancers 11, 669 (2019).
Nunez, S. K. et al. Identification of gene co-expression networks associated with consensus molecular subtype-1 of colorectal cancer. Cancers 13, 5824 (2021).
García-Aranda, M. & Redondo, M. Targeting receptor kinases in colorectal cancer. Cancers 11, 433 (2019).
Rebersek, M. Consensus molecular subtypes (CMS) in metastatic colorectal cancer - personalized medicine decision. Radiol. Oncol. 54, 272–277 (2020).
Orouji, E. et al. Chromatin state dynamics confers specific therapeutic strategies in enhancer subtypes of colorectal cancer. Gut 71, 938–949 (2022).
Martin, T. A. et al. NUPR1 and its potential role in cancer and pathological conditions (Review). Int. J. Oncol. 58, 21 (2021).
Shi, X., Young, C. D., Zhou, H. & Wang, X. Transforming growth factor-β signaling in fibrotic diseases and cancer-associated fibroblasts. Biomolecules 10, 1666 (2020).
Lin, Y., Xu, J. & Lan, H. Tumor-associated macrophages in tumor metastasis: biological roles and clinical therapeutic applications. J. Hematol. Oncol. 12, 76 (2019).
Thanki, K. et al. Consensus molecular subtypes of colorectal cancer and their clinical implications. Int Biol. Biomed. J. 3, 105–111 (2017).
Naito, T. et al. Mesenchymal stem cells induce tumor stroma formation and epithelial‑mesenchymal transition through SPARC expression in colorectal cancer. Oncol. Rep. 45, 104 (2021).
Ran, H. et al. Stearoyl-CoA desaturase-1 promotes colorectal cancer metastasis in response to glucose by suppressing PTEN. J. Exp. Clin. Cancer Res. 37, 54 (2018).
Syed, V. TGF-β Signaling in Cancer. J. Cell. Biochem. 117, 1279–1287 (2016).
Tanevski, J., Flores, R. O. R., Gabor, A., Schapiro, D. & Saez-Rodriguez, J. Explainable multiview framework for dissecting spatial relationships from highly multiplexed data. Genome Biol. 23, 97 (2022).
Neill, T., Schaefer, L. & Iozzo, R. V. Decorin: a guardian from the matrix. Am. J. Pathol. 181, 380–387 (2012).
Deves, C. et al. Analysis of select members of the E26 (ETS) transcription factors family in colorectal cancer. Virchows Arch. 458, 421–430 (2011).
Gİrgİn, B., KaradaĞ-Alpaslan, M. & KocabaŞ, F. Oncogenic and tumor suppressor function of MEIS and associated factors. Turk. J. Biol. 44, 328–355 (2020).
Du, B., Gao, W., Qin, Y., Zhong, J. & Zhang, Z. Study on the role of transcription factor SPI1 in the development of glioma. Chin. Neurosurg. J. 8, 7 (2022).
Nie, X., Liu, H., Liu, L., Wang, Y.-D. & Chen, W.-D. Emerging Roles of Wnt Ligands in Human Colorectal Cancer. Front. Oncol. 10, 1341 (2020).
Guillermin, O. et al. Wnt and Src signals converge on YAP-TEAD to drive intestinal regeneration. EMBO J. 40, e105770 (2021).
Koch, M. et al. CD36-mediated activation of endothelial cell apoptosis by an N-terminal recombinant fragment of thrombospondin-2 inhibits breast cancer growth and metastasis in vivo. Breast Cancer Res. Treat. 128, 337–346 (2011).
Page-McCaw, A., Ewald, A. J. & Werb, Z. Matrix metalloproteinases and the regulation of tissue remodelling. Nat. Rev. Mol. Cell Biol. 8, 221–233 (2007).
Zhang, J., Sud, S., Mizutani, K., Gyetko, M. R. & Pienta, K. J. Activation of urokinase plasminogen activator and its receptor axis is essential for macrophage infiltration in a prostate cancer mouse model. Neoplasia 13, 23–30 (2011).
Liu, M. et al. Transcription factor c-Maf is a checkpoint that programs macrophages in lung cancer. J. Clin. Invest. 130, 2081–2096 (2020).
Hara, T. & Tanegashima, K. CXCL14 antagonizes the CXCL12-CXCR4 signaling axis. Biomol. Concepts 5, 167–173 (2014).
Reszegi, A. et al. The protective role of decorin in hepatic metastasis of colorectal carcinoma. Biomolecules 10, 1199 (2020).
Fontana, E., Eason, K., Cervantes, A., Salazar, R. & Sadanandam, A. Context matters-consensus molecular subtypes of colorectal cancer as biomarkers for clinical trials. Ann. Oncol. 30, 520–527 (2019).
Dunne, P. D. et al. Challenging the cancer molecular stratification dogma: intratumoral heterogeneity undermines consensus molecular subtypes and potential diagnostic value in colorectal cancer. Clin. Cancer Res. 22, 4095–4104 (2016).
Eide, P. W., Bruun, J., Lothe, R. A. & Sveen, A. CMScaller: an R package for consensus molecular subtyping of colorectal cancer pre-clinical models. Sci. Rep. 7, 1–8 (2017).
Herrera, M. et al. Cancer-associated fibroblast-derived gene signatures determine prognosis in colon cancer patients. Mol. Cancer 20, 73 (2021).
Zhong, Z. A., Michalski, M. N., Stevens, P. D., Sall, E. A. & Williams, B. O. Regulation of Wnt receptor activity: Implications for therapeutic development in colon cancer. J. Biol. Chem. 296, 100782 (2021).
Tsukiyama, T. et al. Molecular role of RNF43 in canonical and noncanonical Wnt signaling. Mol. Cell. Biol. 35, 2007–2023 (2015).
Thasler, W. E. et al. Charitable state-controlled foundation human tissue and cell research: ethic and legal aspects in the supply of surgically removed human tissue for research in the academic and commercial sector in Germany. Cell Tissue Bank. 4, 49–56 (2003).
Bankhead, P. et al. QuPath: open source software for digital pathology image analysis. Sci. Rep. 7, 1–7 (2017).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–3587.e29 (2021).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Amezquita, R. A. et al. Orchestrating single-cell analysis with Bioconductor. Nat. Methods 17, 137–145 (2020).
Hafemeister, C. & Satija, R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol. 20, 296 (2019).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Schubert, M. et al. Perturbation-response genes reveal signaling footprints in cancer gene expression. Nat. Commun. 9, 20 (2018).
Alvarez, M. J. et al. Functional characterization of somatic mutations in cancer using network-based inference of protein activity. Nat. Genet. 48, 838–847 (2016).
Garcia-Alonso, L., Holland, C. H., Ibrahim, M. M., Turei, D. & Saez-Rodriguez, J. Benchmark and integration of resources for the estimation of human transcription factor activities. Genome Res. 29, 1363–1375 (2019).
Gonzalez, I., Déjean, S., Martin, P. & Baccini, A. CCA: AnRPackage to extend canonical correlation analysis. J. Stat. Softw. 23, 1–14 (2008).
McCarthy, D. J., Campbell, K. R., Lun, A. T. L. & Wills, Q. F. Scater: pre-processing, quality control, normalization and visualization of single-cell RNA-seq data in R. Bioinformatics 33, 1179–1186 (2017).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Zhao, E. et al. Spatial transcriptomics at subspot resolution with BayesSpace. Nat. Biotechnol. https://doi.org/10.1038/s41587-021-00935-2 (2021)
Lun, A. T. L., McCarthy, D. J. & Marioni, J. C. A step-by-step workflow for low-level analysis of single-cell RNA-seq data with Bioconductor. F1000Res. 5, 2122 (2016).
Liberzon, A. et al. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Wu, T. et al. clusterProfiler 4.0: a universal enrichment tool for interpreting omics data. Innovation 2, 100141 (2021).
Türei, D. et al. Integrated intra- and intercellular signaling knowledge for multicellular omics analysis. Mol. Syst. Biol. 17, e9923 (2021).
Dimitrov, D. et al. Comparison of methods and resources for cell-cell communication inference from single-cell RNA-Seq data. Nat. Commun. 13, 1–13 (2022).
Kolde, R., Laur, S., Adler, P. & Vilo, J. Robust rank aggregation for gene list integration and meta-analysis. Bioinformatics 28, 573–580 (2012).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Chen, B. et al. Differential pre-malignant programs and microenvironment chart distinct paths to malignancy in human colorectal polyps. Cell 184, 6262–6280.e26 (2021).
Kosinski, C. et al. Gene expression patterns of human colon tops and basal crypts and BMP antagonists as intestinal stem cell niche factors. Proc. Natl Acad. Sci. USA. 104, 15418–15423 (2007).
Yuan, H. et al. CancerSEA: a cancer single-cell state atlas. Nucleic Acids Res. 47, D900–D908 (2019).
Seo, M.-K., Kang, H. & Kim, S. Tumor microenvironment-aware, single-transcriptome prediction of microsatellite instability in colorectal cancer using meta-analysis. Sci. Rep. 12, 6283 (2022).
Acknowledgements
This work was supported by The Roche Postdoctoral Fellowship (RPF) programme. We acknowledge the support of the non-profit foundation HTCR, which holds human tissue on trust, making it broadly available for research on an ethical and legal basis. We thank Daniel Dimitrov, Ricardo Omar Ramirez Flores and Dario Zimmerli for productive scientific discussions around the topics covered in this manuscript.
Author information
Authors and Affiliations
Contributions
A.V., K.H., T.B., B.J. and P.S. planned and designed the study. A.V., B.A. and K.H. wrote the manuscript with the input and feedback from the remaining authors. B.A., N.G., M.R. and N.K. conducted the sample preparation and laboratory experiments. A.V., A.J.L. and E.G. carried out the data analysis. K.H., A.L. and M.D.T. performed pathology assessments and assisted bioinformatics data interpretation. D.T., L.V., S.B., I.W., B.P., E.Y., M.D.T., M.B., S.R., J.S.R. and M.S. provided guidance on the data analysis direction and the biological findings. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
A.V., B.A., E.G., N.G., M.R., S.B., I.W., B.P., L.V., E.Y., M.B., M.S., N.K., B.J., P.S., T.B. and K.H. are currently employed by F. Hoffmann-La Roche Ltd. A.J.L. and D.T. were previously employed by F. Hoffmann-La Roche Ltd. A.J.L. is currently employed by Idorsia Pharmaceuticals Ltd. D.T. is currently employed by University of Bern. A.L. is currently employed by Genentech, Inc. M.D.T. was previously employed by Genentech, Inc and is currently employed by Gilead Sciences, Inc. J.S.R. has received funding from GSK and Sanofi and fees from Travere Therapeutics and Astex Pharmaceuticals. The authors declare that they have no other competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Valdeolivas, A., Amberg, B., Giroud, N. et al. Profiling the heterogeneity of colorectal cancer consensus molecular subtypes using spatial transcriptomics. npj Precis. Onc. 8, 10 (2024). https://doi.org/10.1038/s41698-023-00488-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41698-023-00488-4
- Springer Nature Limited