Abstract
Recurrent fusions of receptor tyrosine kinases (RTKs) are often driving events in tumorigenesis that carry important diagnostic value and are potentially targetable by the increasing number of tyrosine kinase inhibitors (TKIs). Here, we characterized the spectrum of 1324 RTK fusions with intact kinase domains in solid tumors by DNA-based high-throughput sequencing. Overall, the prevalence of RTK fusions were 4.7%, with variable frequencies and diverse genomic structures and fusion partners across cancer types. Cancer types, such as thyroid cancers, urological cancers and neuroendocrine tumors are selective in the RTK fusions they carry, while others exhibit highly complex spectra of fusion events. Notably, most RTKs were promiscuous in terms of the partner genes they recombine with. A large proportion of RTK fusions had one of the breakpoints localized to intergenic regions. Comprehensive genomic profiling revealed differences in co-mutational patterns pre- and post-TKI treatments across various RTK fusions. At baseline, multiple cases were detected with co-occurring RTK fusions or concomitant oncogenic mutations in driver genes, such as KRAS and EGFR. Following TKI resistance, we observed differences in potential on- and off-target resistance mutations among fusion variants. For example, the EML4-ALK v3 variant displayed more complex on-target resistance mechanisms, which might explain the reduced survival outcome compared with the v1 variant. Finally, we identified two lung cancer patients with MET+ and NTRK1+ tumors, respectively, who responded well to crizotinib treatment. Taken together, our findings demonstrate the diagnostic and prognostic values of screening for RTK fusions using DNA-based sequencing in solid tumors.
Similar content being viewed by others
Introduction
Many fusion genes are drivers of tumorigenesis, and have important diagnostic and prognostic values in informing clinical action1. In particular, fusions of receptor tyrosine kinases (RTKs) represent an important class of oncogenic events that are selected for during cancer initiation and progression. Although found at a lower rate in solid tumors compared with hematologic malignancies, large-scale genomic studies have identified important RTK fusions across a wide range of cancer types2,3,4,5. Given the potential druggability of RTKs, extensive characterization of the landscape of RTK fusions would likely facilitate new drug development and expand the therapeutic options for cancer patients.
Traditional methods, such as fluorescence in situ hybridization (FISH) and quantitative real-time polymerase chain reaction (RT-PCR) or PCR, are highly sensitive in detecting fusion genes. However, such low-throughput methods are time- and cost-ineffective and also suffer from its limitations in detecting rare fusion events. Recent advances in massively parallel sequencing and bioinformatics methods have revealed the complexity of genetic fusions in cancer. A number of genomic approaches have been commonly applied for the detection of fusion events, including whole genome sequencing (WGS), RNA sequencing (RNA-seq), and targeted DNA sequencing. However, in the clinical setting, targeted DNA sequencing has its unique advantages in increased sensitivity and also overcoming the challenges of fusion detection using formalin-fixed paraffin-embedded (FFPE)-derived tumors.
In this study, we sought to characterize the prevalence and the spectrum of RTK fusions in patients with diverse solid tumors who underwent hybridization capture-based DNA-targeted sequencing. We also examined co-mutations and potential resistance mechanisms at baseline and following TKI treatment, respectively.
Results
Prevalence of RTK fusions across diverse cancers
We examined the frequencies of RTK fusions in a large cohort of Chinese patients across a diverse range of solid tumor types (Supplementary Fig. 1) whose tumor and/or ctDNA samples underwent targeted profiling. Only those fusion events that retained the intact kinase domain were included in the analysis.
The overall prevalence of RTK fusions detected in this cohort was 4.7% (n = 1324), with varying frequencies across different cancer types (Fig. 1a and Table S1). RTK fusions were detected at high frequencies in thyroid cancers (7.8%), lung cancers (7.1%), neuroendocrine tumors (3.7%), urological cancers (3.6%), and gastric cancers (2.2%). In line with previous reports, our analysis recapitulated the key targetable oncogenic fusion events in lung cancers, with a 4.2% frequency of ALK fusions, 1.3% of RET fusions, and 1.2% of ROS1 fusions (Fig. 1a and Supplementary Table 1). Multiple additional but relatively rare oncogenic fusions have been described in lung cancers, including fusions of the FGFR, NTRK, MET, and ErbB family genes6,7,8. In our lung cancer cohort, we also observed a 0.2% of FGFR family (mostly FGFR3) fusions and 0.02% of NTRK1/3 gene fusions. Rearrangements of MET and the ErbB family RTKs (EGFR, ERBB2, ERBB3, and ERBB4) in lung cancer are less well documented and mostly as case reports6,9. A total of three (0.02%) lung cancer patients harbored MET fusions (Fig. 1a and Supplementary Table 1), against which crizotinib has reportedly demonstrated clinical activity10,11. On the other hand, the prevalence of ErbB family gene fusions in our lung cancer cohort was non-negligible reaching 0.23%, with the majority of these patients carrying EGFR (0.11%) and ERBB2 (0.07%) fusions.
a Heatmap showing the prevalence of rearrangements of specific RTKs in different cancers, LUC lung cancer, CRC colorectal cancer, GAC gastric cancer, BRC breast cancer, HEPC hepatobiliary cancer, PAC pancreatic cancer, OVC ovarian cancer, STS soft tissue sarcoma, CEC cervical cancer, ESC esophageal cancer, URC urinary cancer, HNC head and neck cancer, SKCM skin cutaneous melanoma, NET neuroendocrine tumor, PRC prostate cancer, THC thyroid cancer. b ROS1, ERBB2, ALK, and MET fusions showed increased associations with the female sex. c ALK, ROS1, and RET fusions showed increased associations with younger age.
Besides lung cancer, several other cancer types also displayed a broad array of RTK fusions, particularly gastric cancers, colorectal cancers and breast cancers (Fig. 1a and Supplementary Table 1). The top oncogenic RTK fusions in gastric cancer were of the FGFR family genes (total, 1.4%; FGFR2, 1.2%) and ErbB family genes (total, 0.67%; ERBB2, 0.41%, EGFR, 0.26%), with a smaller percentage of patients also harboring EPHA2 (0.15%), ALK (0.15%), MET (0.10%), RET (0.10%), and ROS1 (0.07%) fusions. The overall prevalence of RTK fusions in colorectal cancers was 1.0%, with the top frequent fusion genes being RET (0.34%), ALK (0.19%), and NTRK1 (0.10%). In breast cancers, the most prevalent fusion genes were FGFR family genes (total, 0.48%; FGFR2, 0.34%; FGFR1, 0.14%), ErbB family genes (total, 0.47%; EGFR, 0.20%; ERBB2, 0.27%), ALK (0.27%) and ROS1 (0.07%).
By contrast, a restricted spectrum of RTK fusions was observed in thyroid cancers, urological cancers, and neuroendocrine tumors. Of the 77 patients with thyroid cancers, six (7.8%) RTK fusion events were identified, which were exclusively RET fusions. Urological cancers also had a high proportion (15 out of 417; 3.6%) of RTK fusions, with the majority (12; 80%) of patients carrying FGFR2/3 fusions. Similarly, ALK fusions (2.2%) accounted for three of the five fusion-positive cases with neuroendocrine tumors.
RTK fusions were detected in both the tumor specimens (n = 963) and circulating cell-free DNA (cfDNA) samples from a variety of body fluids, including plasma (n = 268), pleural effusion (n = 79), cerebrospinal fluid (n = 6), and additional liquid biopsy samples from ascites and pericardial effusions (n = 8). No apparent differences were observed between the overall frequencies detected in the tumor and cfDNA samples (Supplementary Fig. 2A and Supplementary Tables 2–3). In addition, comparing the RTK frequencies in lung cancer between tumor specimens and cfDNA samples, only a slight enrichment of RET fusions in the cfDNA samples were detected (false discovery rate (FDR) adjusted q = 0.04; Supplementary Fig. 2B). No other significant differences were identified.
Significant associations of sex and age with the occurrence of specific RTK fusions were also observed. Overall, ROS1 (63% vs. 46%, Fisher’s exact test P < 0.0001, FDR-adjusted q = 0.001), ERBB2 (69% vs. 46% Fisher’s exact test P = 0.007, FDR-adjusted q = 0.04), ALK (50% vs. 46%, Fisher’s exact test P = 0.01, FDR-adjusted q = 0.05), and MET (89% vs. 46%; Fisher’s exact test P = 0.02, FDR-adjusted q = 0.06) fusions were more frequently detected in females (Fig. 1b). Taking into account potential cancer type differences, we found that ROS1 (63% vs. 44%; Fisher’s exact test P < 0.0001) and ALK fusions (50% vs. 44%, Fisher’s exact test P = 0.006) remain more prevalent in female lung cancer patients (Supplementary Fig. 3A). No differences were found comparing the sex distributions among different variants of ALK (Supplementary Fig. 3B), RET (Supplementary Fig. 3C) or ROS1 (Supplementary Fig. 3D). In addition, fusions in ALK (overall, Fisher’s exact test P < 0.0001; lung cancer, P < 0.0001), ROS1 (overall, Fisher’s exact test P < 0.0001; lung cancer, P < 0.0001) and RET (overall, Fisher’s exact test P = 0.006; lung cancer, P < 0.0001) were associated with an earlier disease onset (Fig. 1c and Supplementary Fig. 4A). Further analysis stratified age differences in ALK fusion variants and showed that the EML4-ALK v1 variant was associated with an earlier disease onset (Fisher’s exact test P = 0.008), while the v3 variant was more likely to occur at an older age (Fisher’s exact test P = 0.02, Supplementary Fig. 4B). No age differences among ROS1 or RET fusion variants were found (Supplementary Fig. 4C, D).
Genomic structures and partner genes of RTK fusions
The genomic structures and partner genes of the most commonly altered RTK fusion genes (ALK, RET, ROS1, FGFR3/2, EGFR, MET, and NTRK1) were illustrated in Fig. 2. While EML4 accounted for 66.5% (610/917) of the ALK fusion events in this cohort, a remarkably diverse array of ALK fusion partners were detected (Fig. 2a). A total of 227 distinct fusion partners of ALK were detected. Among these, some recurrent ALK partners included STRN (1.0%), TOGARAM2 (0.5%), KIF5B (0.4%), and KLC1 (0.4%), and 46 fusion partners occurred only once. In addition, there were a considerable number (15.4%, 141/917) of intergenic rearrangements (i.e., having one breakpoint localized to intergenic regions (IGR)). The EML4-ALK fusion gene is generated by an inversion on chromosome 212. For the majority (65.5%, 201/307) of non-EML4 fusions, including 74% of intergenic rearrangements, they were also clustered on chromosome 2. Notably, recombination of ALK could occur with a vast range of chromosomal regions with fusion partners scattered across the genome (Fig. 2a). ALK fusions in non-lung cancer patients were more commonly fused with non-EML4 partners (Bonferroni’s post-test, P < 0.0001; Fig. 2a). Of the 31 ALK fusion events detected in non-lung cancers, including gastrointestinal and gynecological cancers, 11 patients (35.5%) were detected with EML4-ALK fusions. STRN represented the second most common ALK fusion partner, and were detected in two cases of colorectal cancer and one case of hepatobiliary cancer. All ALK fusions took place at the 5’ end of the protein and breakpoints in intron 19 were highly recurrent, accounting for 95% overall, 96% in lung cancers and 81% in non-lung cancers, producing fusion products fused to exon 20 of the ALK gene (Fig. 2a). No clear consensus sequences were found around the breakpoints of the ALK gene across different cancer types (Supplementary Fig. 5A, B) or fusion variants (Supplementary Fig. 5C–F). Consistent with v1 and v3 being the most common ALK fusion variants, breakpoints in EML4 mostly clustered around exons 6 and 13 (Supplementary Fig. 5G). Other recurrent breakpoints included exon 3 of STRN and 5’UTR of TOGARAM2 (Supplementary Fig. 5H). Interestingly, the breakpoints of ALK fusion partners, EML4 or non-EML4, were surrounded by AT-rich regions (Supplementary Fig. 5I, J).
a–h From left to right: Distributions of common fusion partners; Circos plot showing the genomic rearrangement events in different cancer types; Comparisons of fusion partner frequencies in different cancer types, P values using chi-square tests are as indicated; Comparisons of RTK breakpoints in different cancer types in a ALK, b RET, c ROS1, d FGFR3, e FGFR2, f EGFR, g MET, and h NTRK1 positive samples. IGR intergenic regions.
Fifty distinct RET 5’ fusion partners were identified, with KIF5B being the most common fusion partner, accounting for 44% (131/298) of RET fusions, which was followed by CCDC6 (52/298, 18%) and NCOA4 (16/298, 5%; Fig. 2b). Other recurrent fusion partners included RASGEF1A (1.3%), CCDC186 (0.7%), JCAD (0.7%), and KIF13A (0.7%; Fig. 2b). All recurrent fusion partners of RET, except for JCAD, has been previously reported. Of the two JCAD-RET cases, one had lung adenocarcinoma and the other were diagnosed with soft tissue sarcoma of the cervix, and both had the 5’ UTR of JCAD fused to the RET gene. RET fusions were mostly caused by rearrangements with nearby genes or intergenic regions on chromosome 10 (Fig. 2b). In addition to lung cancers, RET fusions were also frequently detected in colorectal and thyroid cancers. The frequencies of the recurrent fusion partners differed across the cancer types (Fig. 2b). For example, KIF5B-RET was highly frequent but almost exclusively found in lung cancers (127/262, 48.5%; Bonferroni’s post-test, P < 0.001). It was found in only 18.8% (3/16) of colorectal cases and one case of neuroendocrine tumors, but none of the eight thyroid cases. On the other hand, while the frequency of NCOA4-RET was low (2.3%, 6/262) in lung cancer, it was the most frequent RET fusion (44%, 7/16) in colorectal cancer (Bonferroni’s post-test, P < 0.0001). In addition, CCDC6-RET was only detected in non-colorectal cancers. The most frequent breakpoints across all cancers took place in RET intron 11, resulting in a fusion product involving RET exon 12 (overall rate, 78.9%, 235/298). Across all RET fusions, regardless of cancer types (Supplementary Fig. 6A, B) or fusion variants (Supplementary Fig. 6C–E), GC-rich sequences were found around the breakpoints of RET. For the common RET fusion partners, breakpoints in KIF5B, CCDC6 and NCOA4 most commonly occurred in exons 15, 1, and 7, respectively (Supplementary Fig. 6F–H), and were enriched with AT-rich sequences (Supplementary Fig. 6I–K).
Unlike ALK and RET, whose fusion partners were largely nearby genes residing on the same chromosome, ROS1 fusions were more scattered across the entire genome (Fig. 2c). Only 20.9% (48/230) of ROS1 rearrangements were generated by translocations in chromosome 6. Overall, thirty-nine distinct ROS1 fusion partners were detected. In lung cancers, fusions with CD74 (83/216, 38.4%), EZR (28/216, 13.0%), SDC4 (26/216, 12.0%), and SLC34A2 (26/216, 12.0%) were highly frequent (Fig. 2c). Of these, the generation of CD74-RET, SDC4-RET and SLC34A2-RET fusions involved translocation with chromosome 5, 20, and 4, respectively. TPM3-ROS1, which was another relatively common form of ROS1 fusions in lung cancers (8/216, 3.7%), was generated by translocation with chromosome 1. Other recurrent ROS1 fusions included MYH9-ROS1 (2/216, 0.9%) and MYO5C-ROS1 (2/216, 0.9%), involving translocation with chromosome 22 and 15, respectively. In the non-lung cancer cases (n = 14), ROS1 fusions mainly involved chromosome 5 and 6 (Fig. 2c). Non-lung cancers were more likely to harbor rare ROS1 fusions (Bonferroni’s post-test, P = 0.002). CD74-ROS1 was detected in one case each of cervical and urinary cancers, and SDC4-ROS1 were detected in one case of soft tissue sarcoma. Similar to the wide genomic distributions of its fusion partners, breakpoints of ROS1 also spanned across multiple introns, mostly from intron 31 to intron 34 (Fig. 2c). Unlike ALK or RET, the sequences around the breakpoints of ROS1 were highly AT-rich (Supplementary Fig. 7A–F). For common ROS1 partner genes, breakpoints in CD74, EZR, SDC4 and SLC34A2 were predominantly located in exons 6, 10, 2, 13, respectively (Supplementary Fig. 8A, B). TPM3 breakpoints were mostly located in exons 7 and 8 (Supplementary Fig. 8B). Sequences around the breakpoints of CD74 were GC-rich (Supplementary Fig. 8C), whereas sequences around all other ROS1 partners tended to be AT-rich (Supplementary Fig. 8D–F).
FGFR and ErbB family fusions were also rather common in a multitude of cancers. Of the FGFR family, FGFR2 and FGFR3 fusions accounted for 39% and 51% of the total FGFR fusion events, respectively. FGFR3 fusions were common in lung and urinary cancers, but were also found in a wide variety of cancer types, particularly cancers of the gastrointestinal tract (Fig. 2d). Remarkably, regardless of cancer types, FGFR3 fusions were almost exclusively in the form of FGFR3-TACC3 (94%, 47/50), generated by translocations in chromosome 4. Breakpoints in FGFR3 were either in intron 17 or exon 18. The most frequent (19/47, 40.4%) breakpoints involved intron 17 of FGFR3 and intron 11 of TACC3, resulting in the F17:T11 fusion variant. Regions around the breakpoints of FGFR3 and TACC3 were enriched with GC-rich sequences (Supplementary Fig. 9A–E). In contrast to FGFR3, fusion partners of FGFR2 were highly diverse although mostly located on chromosome 10 (Fig. 2e). The two recurrent FGFR2 fusions were FGFR2-BICC1 in two cases of hepatobiliary cancers (Bonferroni’s post-test, P = 0.006) and FGFR2-TACC2 in one case each of breast and gastric cancers. Interestingly, one recurrent intergenic FGFR2 fusion was identified solely in patients with gastric cancer (6/22, 27.3%), being fused to intergenic regions upstream of WDR11. Similar to FGFR3, the majority of FGFR2 breakpoints were located in intron 17. No specific sequence patterns were found around the breakpoints of FGFR2 and its partner genes (Supplementary Fig. 9F–H).
Of the ErbB family, EGFR and ERBB2 were the most frequently rearranged genes. The majority of ErbB fusions were non-recurrent and occurred downstream of the kinase domain. Recurrent EGFR fusions included CCT6A-EGFR fusions in two cases of lung cancer (2/13, 15.4%) and intergenic EGFR-SEC61G fusions, occurring at intergenic regions downstream of SEC61G, in three cases of gastric cancers and one case of lung cancer (Fig. 2f). EGFR-VSTM2A fusion was detected in one case of colorectal cancer and another case occurring upstream of the VSTM2A gene in lung cancer. Fusion partners were mostly localized to chromosome 7 (12/30, 40%), whereas breakpoints in EGFR were rather widely distributed as shown in Fig. 2f. The breakpoints for 3’ end EGFR fusions were predominantly in intron 24 (10/30, 33.3%) and for 5’end EGFR fusions, intron 17 (3/30, 10%; Fig. 2f). No clear consensus was found for the sequences around the breakpoints of EGFR, although those of its partner genes tended to be AT-rich (Supplementary Fig. 10A–C). Recurrent ERBB2 fusions included ERBB2-PGAP3 in a total of three cases of gastric, colorectal, and cervical cancers and GRB7-ERBB2 in one case each of lung and cervical cancers. The majority of ERBB2 rearrangements occurred in chromosome 17 (31/36, 86.1%). Similar to EGFR, fusions in ERBB2 occurred at many breakpoints across the gene, with exon 27 being the most frequent spot (7/36, 19.4%). Regions around the ERBB2 breakpoints were enriched with GC-rich sequences and no clear sequence consensus was observed near the breakpoints of its partner genes (Supplementary Fig. 10D, E).
MET and NTRK fusions occurred at relatively low frequencies, but were detected in a number of cancer types surveyed. MET partners were all fused to the 5’ end of the protein and mostly localized to chromosome 7 (Fig. 2g). No recurrent MET fusions were identified, some of the fusion partners included HLA-DRB1-MET (lung cancer), CD47 (lung cancer), CAPZA2 (gastric cancer), AKAP9 (hepatobiliary cancer), CLIP2 (hepatobiliary cancer), and SLC25A19 (ovarian cancer). Of these, the HLA-DRB1-MET fusion and CLIP2-MET fusion were previously reported in cases of lung cancer11 and glioneuronal cancer13, respectively. For NTRK fusions, all of the four NTRK1 + colorectal cases harbored a recurrent TPM3-NTRK1 fusion (overall, 4/7, 57.1%; Bonferroni’s post-test, P = 0.04). Three non-recurrent NTRK1 fusions were each detected in two cases of lung cancer (SQSTM1-NTRK1 and IRF2BP2-NTRK1) and one case of breast cancer (EFNA1-NTRK1). Of these non-recurrent fusions, SQSTM1-NTRK1 and IRF2BP2-NTRK1 have been previously reported14. NTRK3 fusions were detected in only two cases in the entire cohort, one ABHD17C-NTRK3 fusion in a case with urinary cancer and one TTC23-NTRK3 fusion in a lung cancer case. Although no clear consensus sequences were found in regions near the breakpoints of MET and NTRK, the NTRK gene showed marked enrichment of the C base at the location of the breakpoint (Supplementary Fig. 10F–I).
Analysis of co-mutations in RTK fusion-carriers prior to TKI treatment
For patients with sufficient clinical records, including treatment regimens and timelines, we next investigated the mutations co-occurring with the respective RTK fusions prior to and following targeted TKI treatments. The top genes frequently co-mutated with RTK fusions prior to TKI treatment were illustrated in Fig. 3a and Supplementary Fig. 11. At the level of each RTK genes, we found that fusions in EGFR, ERBB2 and MET were highly likely to be accompanied by an amplification of the respective RTK gene. Specifically, copy number gain occurred in all cases of EGFR and ERBB2 fusion-positive cases, except for one case of EGFR fusion+ hepatobiliary cancer. The most frequently altered gene was TP53, with varying frequencies across different RTK fusion genes (Fig. 3a, b). ALK fusions were associated with a relatively lower frequency of TP53 co-mutations (35%, Bonferroni’s post-test, P = 0.05), whereas over 90% of the ErbB family fusions carried an additional TP53 mutation (Bonferroni’s post-test, P = 0.01).
a Oncoplot showing the most frequently co-mutated genes across different RTK fusion-positive samples. b Different RTK fusions are associated with varying spectra of concomitant mutations, P values using Bonferroni’s post-test are as indicated. c Estimated clonality comparing different fusion genes or fusion variants.
In addition, different RTK fusions showed substantial differences in the pattern of their concomitant mutations (Fig. 3b). The overall co-mutation rates in ALK + cases were low, with significantly lower frequencies of PIK3CA (1%, Bonferroni’s post-test, P = 0.003) and APC (1%, Bonferroni’s post-test, P = 0.08) alterations compared with other RTK fusion+ cases. RET fusions were associated with a higher rate of PTEN alterations (12%, Bonferroni’s post-test, P = 0.02), whereas ROS1 fusions were associated with a higher rate of RB1 (12%, Bonferroni’s post-test, P = 0.006) in comparison with other RTK fusion+ cases. FGFR fusions were characterized by high frequencies of PIK3CA (22%, Bonferroni’s post-test, P = 0.01) and KRAS (39%, Bonferroni’s post-test, P < 0.001) alterations. Finally, ErbB family fusions had higher incidences of alterations in CTNNB1 (27%, Bonferroni’s post-test, P = 0.008) and NF1 (18%, Bonferroni’s post-test, P = 0.09).
Given that multiple oncogenic driver genes were among the top altered genes in our TKI-naive RTK fusion+ cohort, we further assessed the exclusivity and relevance of concomitant driver mutations (Supplementary Table 4). Remarkably, we identified a non-negligible number of concomitant driver mutations. First of all, RTK fusions themselves were not mutually exclusive. We identified three cases that were ALK +/RET+, two cases with ALK +/ROS1+, two cases with RET +/ROS1+, and one case with ALK +/NTRK3+. In addition, while the functionality of some non-canonical RTK fusions remained to be determined, concomitant driver mutations were detected in many cases with highly recurrent RTK fusions (Supplementary Table 4). Specifically, we identified three lung cancer patients with EML4-ALK, who each also harbored an activating mutation in KRAS, and loss of function mutations in BRCA2 and PTEN, respectively. Six additional lung cancer patients with KIF5B-RET (n = 3), CCDC6-RET (n = 2) or SDC4-ROS1 (n = 1) were also detected with activating mutations in NRAS, EGFR, and PIK3CA, as well as loss of function mutations in BRCA1 and PTEN. Moreover, two FGFR3-TACC3-positive cases also harbored KRAS-activating mutations. No concomitant driver mutations in BRAF were identified. Of these 18 patients with concomitant drivers, seven (38.9%) harbored these driver mutations in separate subclones, and six had RTK fusion and the other driver co-occurring as clonal events. In the remaining five patients, all RTK fusions were clonal events, in which cases the respective concomitant drivers were subclonal (Supplementary Table 4). In addition, we evaluated the clonality of additional RTK fusions with no detectable co-drivers and found no clear differences in clonality between common recurrent fusions and rare fusion events, other than an increase in the frequency of clonal RET fusions in cases with rare fusion partners (Fig. 3c).
Potential on-target and off-target resistance mechanisms following TKI treatment
Next, based on evaluable clinical data, we investigated the co-mutational spectrum following targeted TKI treatment aiming to explore the potential resistance mechanisms. In a total of 78 ALK + patients who were treated with ALK TKIs, we detected on-target resistance mechanisms in 30.5% (14/46) and 43.8% (14/32) of patients following crizotinib and multi-TKI treatment, respectively (Fig. 4a and Supplementary Fig. 12A). Moreover, patients treated with multiple TKIs were more likely to acquire more than one on target ALK resistance mutations than those treated with crizotinib alone (21.9% vs. 2.2%; Fig. 4a). An increasing trend of RTK fusions following targeted therapies was also observed (Supplementary Fig. 12B), which likely reflects clonal evolution imposed by drug selection. Different ALK variants also seemed to respond differently to ALK inhibition. Patients carrying the EML4-ALK v3 variant had worse progression-free survival (PFS) outcomes compared with those with the v1 variant, both following crizotinib treatment (Hazards ratio (HR) = 2.46, 95%CI = 1.21–4.98, P < 0.01; Fig. 4b) and multi-TKI treatment (HR = 2.76, 95%CI = 0.71–10.72, P = 0.13; Supplementary Fig. 13A). Differences in PFS outcome might be attributed to a more complex spectrum of on-target resistance (Fig. 4c), as well as a higher incidence of acquiring multiple resistance mutations following TKI treatment (Fig. 4d) in the V3 variant. On the other hand, ALK + patients with other EML4-ALK variants or non-EML4 partners, including previously unreported MEMO1-ALK and WRD43-ALK fusions, also responded well to first-line crizotinib treatment, with a median PFS of 11 months (Supplementary Fig. 13B). In addition, there were significant increases in MAPK pathway gene alterations (crizotinib, FDR-adjusted P = 0.07; multi-TKI, FDR-adjusted P = 0.001), as well as higher frequencies of PI3K pathway, STAT3 and MET alterations following ALK inhibition (Fig. 4e), which might be associated with off-target TKI resistance.
a Proportions of patients carrying on-target ALK resistance mutations following ALK TKI treatment. b Kaplan–Meier estimates of PFS comparing patients with the two major EML4-ALK variants following first-line crizotinib. c Lollipop plots mapping the on-target resistance mutations in different ALK fusion variants following crizotinib or multi-TKI treatments. d Resistance mutations in patients who acquired multiple on-target ALK mutations following TKI treatment. e Potential off-target resistance mechanisms of ALK fusions (MAPK pathway included EGFR, NF1/2, BRAF, RAF1, KRAS, and NRAS mutations; PI3K pathway included PIK3CA, PTEN, AKT2, and RICTOR mutations). f Proportions of patients carrying on-target ROS1 resistance mutations following TKI treatment. g Lollipop plots mapping the on-target resistance mutations in different ROS1 fusion variants following crizotinib or multi-TKI treatments. h Potential off-target resistance mechanisms of ROS1 fusions.
Similarly, in ROS1 + patients, on-target ROS1 resistance mutations were identified in 27.3% (3/11) and 80% (4/5) of cases treated with crizotinib and multi-TKI, respectively (Fig. 4f). The most common on-target resistance mutations in this study cohort was p.G2032R, which could be induced by both first-line crizotinib and second- or subsequent-line TKIs (Fig. 4g). Finally, comparisons of TKI-naive and -treated patients revealed increases in NF1/2 (FDR-adjusted P = 0.06), PDGFR and PBRM1 alterations following TKI resistance (Fig. 4h).
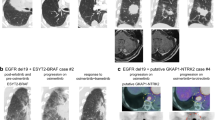
In addition to the top altered fusion genes, we also identified a number of cases with relatively uncommon fusions that were treated with TKI therapies. In particular, we identified two lung cancer cases, one (PT218) with an HLA-DRB1-MET fusion gene and the other (PT230) with an IRF2BP2-NTRK1 fusion gene. Both fusion genes were clonal mutations and no additional known oncogenic mutations were identified, suggesting that the fusion genes were the sole driving events in these tumors. Targeted RNA sequencing confirmed the expression of the HLA-DRB1-MET fusion (Supplementary Fig. 14). Following crizotinib treatment, the MET + patient remained disease-free for 6 months, and the NTRK1 + patient reached a remarkable PFS of 18.5 months.
Discussion
Understanding of recurrent genetic fusions, particularly those of RTKs, may facilitate drug development. An increasing number of TKIs have been developed to target RTK gene aberrations and provide the most promising therapeutic effects for cancer patients. In this study, we described the RTK fusion events in 1324 patients across diverse solid tumors using DNA-based next-generation sequencing profiling. We were able to characterize the prevalence and genomic structures of RTK fusions. While a number of cancer types showed a wide spectrum of RTK fusions, certain cancer types, such as thyroid cancer, urological cancers, and neuroendocrine tumors, are very selective in the RTK fusion genes they carry. The association between these RTK fusion genes and the cancer type makes them highly valuable for diagnostic purposes. In addition, we found that except for the highly reported fusion genes, such as EML4-ALK, KIF5B-RET, CD74-ROS1, and FGFR3-TACC3, the majority of fusion events were largely non-recurrent. Nearly all RTK genes, with the exception of FGFR3, were highly promiscuous in their fusion partner, which might be scattered across the genome. Our study also suggested that different RTK fusions and their respective variants might display varying spectra of concomitant and acquired resistance mutations prior to and following TKI treatments, respectively, and consequently impact the patients’ response to TKIs. The above findings highlight the importance of screening using high-throughput sequencing technologies.
Although RTK fusions had largely been considered to occur mutually exclusively to other oncogenic driver mutations, several studies have provided evidence supporting the co-existence of dual drivers15,16. In addition, patients with dual drivers may exhibit variable response to single-agent targeted therapies. The best-studied example is concomitant EGFR-activating mutations and ALK fusions. However, in such patients, it remains unclear whether they responded equally well to single-agent TKIs or combination TKI therapies are needed for prolonged survival benefit15,16,17,18,19,20. The choice of therapy may depend on the relative abundance or activation levels of the two drivers16,18. Taking advantage of high-throughput sequencing technologies, we identified several cases (~7% of RTK fusion+ baseline samples) with dual driver alterations. Concomitant driver mutations included activating mutations in classic oncogenes (e.g., KRAS, NRAS, EGFR, and PIK3CA), as well as loss-of-function mutations in tumor suppressors (e.g., BRCA1/2 and PTEN). Notably, such driver events co-existed with the RTK fusion either in the same clone or as distinct subclones, which might further influence disease progression and treatment outcome. In addition, we showed that different RTKs varied in their repertoire of concomitant driver and other somatic mutations, and may serve as potential intrinsic resistance mechanisms to targeted therapies. However, due to the retrospective nature of our study, which has inadequate clinical follow-up data, further investigations are necessary to dissect the effect of concurrent alterations on clinical response to RTK fusion-targeted inhibitors.
As mentioned, our study reported a considerable number of rare RTK fusions; some of which have previously been reported in sporadic cases, for other uncommon fusions, their driver roles might require additional confirmation in the absence of treatment outcomes. Despite the mostly non-recurrent nature of RTK fusions, numerous studies, including those of our own, have shown that patients with rare fusion events can be successfully targeted by TKI treatment. For example, favorable responses to TKI therapies have been demonstrated against the less common ALK fusion genes, such as STRN-ALK, CUX1-ALK, and GCC2-ALK21,22,23. In line with previous reports, we showed that patients with non-v1/v3 variants of ALK, including previously unreported ALK fusions, might also derive long-term clinical benefit from TKI treatment. Similarly, we also reported two relatively uncommon MET and NTRK1 fusions, against which crizotinib was shown to be effective. These results might support the use of DNA-based sequencing strategies for fusion screening to inform clinical actions.
It is worth noting that while both DNA- and RNA-based sequencing approaches are commonly used in the research and diagnostic settings, each has their own unique advantages and disadvantages. DNA-based approach is more clinically applicable and allows for the detection of ctDNA using liquid biopsies. Although “hotspot” breakpoints and common partner genes exist for most RTK fusions, whole genome-based sequencing approach does not rely on the design and performance of targeted panels and would enable an even coverage of all potential structural alterations, particularly those that occur in the intronic regions. By contrast, targeted approach is more cost-effective and offers a greater sequencing depth, and consequently higher sensitivity at regions of strong clinical relevance. On the other hand, RNA-based approach would depend on the quality of the sample but has a unique advantage in detecting functional fusion events as compared to DNA-based methods. For example, it has been shown that a considerable portion of rare or IGR fusion events as detected by DNA-based sequencing approach are common fusion genes at the RNA level24,25. In our study, we confirmed our DNA-based finding in a case with a rare HLA-DRB1-MET fusion who had responded to crizotinib by using targeted RNA sequencing. The limitations and challenges of DNA- and RNA-based sequencing approaches can likely be overcome by a complementary approach combining the two methods.
In addition to rare fusion genes, different fusion variants may also influence clinical response or the development of resistance to TKI therapies. In line with studies on ALK-positive lung cancers, which have showed the variable clinical outcome of patients with different EML4-ALK fusion variants26,27, we also observed prolonged PFS outcome in patients harboring the v1 variant compared with those carrying the v3 variant. Interestingly, we found that the v3 variant were more likely to acquire two or more on-target resistance mutations than the v1 variant. In addition, the spectrum of resistance mutations in the v3 variant was more complex. The difference in their ability to acquire on-target resistance mutations might explain the preferential clinical outcome associated with the v1 variant.
Numerous preclinical and clinical studies have led to the rapid expansion of first- and next-generation TKIs for aberrations in RTKs. Given the functional conservation of RTKs, many of which can be targeted by multi-kinase inhibitors with activity against various targets. For example, crizotinib has demonstrated activity across a wide range of targets, including ALK, RET, ROS1, NTRK, and MET, in our study cohort. However, the activities of multi-kinase inhibitors can vary widely based on their selectivity for the targets of interest. Recent advances in the highly selective TKIs have largely increased the proportion of patients that can benefit from TKI therapies. NTRK inhibitors, including larotrectinib and entrectinib, represent the best examples that are associated with remarkable response rates (>75%) on NTRK-positive tumors regardless of cancer types28,29,30. Next-generation NTRK inhibitors have shown promising results in overcoming acquired resistance to first-generation inhibitors, and are currently undergoing clinical development31,32. Other examples of selective RTK inhibitors include those against RET protein, BLU-667 (pralsetinib) and LOXO-292 (selpercatinib), which also demonstrated potent activity against RET fusions and activating mutations across a multitude of cancer types33,34,35.
In addition, FGFR inhibitors are being rapidly developed in the clinic. Erdafitinib, an oral pan-FGFR inhibitor, was the first FGFR-selective compound approved by FDA for the second-line treatment of metastatic urothelial carcinoma with an FGFR2 or FGFR3 alteration36. More recently, pemigatinib was approved for previously treated cholangiocarcinoma with FGFR2 fusions or rearrangements37, with FoundationOne®CDx as the companion diagnostics. Our study also supported the use of NGS to screen for genetic fusion events. In the TCGA cohort of 285 cancer patients that underwent comprehensive RNA-seq5, only one patient (0.35%) was found to carry a FGFR2 fusion. Similarly in our multi-cancer cohort, we identified a total of 89 FGFR fusion-positive patients, accounting for 0.40% of the cancer patients overall.
Taken together, our study described the landscape of RTK fusions and their associated mutational spectrum in a large cohort of cancer patients with diverse cancers. The largely non-recurrent and complex nature of RTK fusions, the existence of concomitant somatic mutations, together with the differences in acquired resistance mutations among different fusion variants all emphasize the need for high-throughput sequencing technologies to fully capture fusion events and better inform clinical diagnosis and treatment strategies. Our findings support the diagnostic and prognostic values of RTK fusions and highlight the importance of RTK fusion screening in relevant cancer types and future clinical trials to facilitate the clinical development of therapeutic strategies to target these aberrations.
Methods
Study cohort and sample collection
The study retrospectively reviewed the clinico-genomics database of Geneseeq Technology Inc., China consisting of cancer patients who were routinely treated at multiple hospitals, including the Third Affiliated Hospital of Sun Yat-sen University and the National Cancer Center/National Clinical Research Center for Cancer/Cancer Hospital, between August 2015 and January 2020. Targeted genomic sequencing which encompasses all exons and flanking intronic regions of the reported RTKs, as well as selected exon and intronic regions of the respective previously reported fusion partners, was performed on the tumor specimen and/or circulating cell-free DNA (cfDNA) from body fluids, including plasma, pleural effusion and cerebrospinal fluid. Sample processing and sequencing were performed in a CLIA-certified and CAP-accredited laboratory (Geneseeq Technology Inc., Nanjing, China). Patient information was retrospectively reviewed. For patients with sufficient clinical follow-up data, progression-free survival (PFS) was defined as the time from the beginning of TKI treatment to the date of progressive disease. PFS2 was defined as the time from the beginning of the respective second-line TKI treatment to the date of disease progression. Informed written consent was obtained from each subject or the subject’s family member upon sample collection according to the protocols approved by the ethics committee of each hospital.
Next-generation sequencing
Next-generation sequencing (NGS) was performed as previously described38. In brief, genomic DNAs from tissue or circulating cell-free DNA from body fluids were extracted. Customized xGen lockdown probes (Integrated DNA Technologies) targeting 425 cancer-relevant genes were used for hybridization enrichment. Libraries were on-beads PCR-amplified, purified, sized and quantified, and sequenced on an Illumina HiSeq4000 platform. The mean coverage depth was 143X for controls, 1341X for tissues, and 4185X for cfDNA samples.
Mutation calling
Trimmomatic39 was used for FASTQ file quality control. Leading/trailing low quality (quality reading below 20) or N bases were removed. Paired-end reads were then aligned to the reference human genome (build hg19), using the Burrows-Wheeler Aligner40 with the default parameters. PCR deduplication was performed using Picard and local realignment around indels and base quality score recalibration were performed using GATK341. Further, samples with mean dedup depth <30X were removed. Single nucleotide variants (SNVs) and indels were identified using VarScan242, with a minimum variant allele frequency threshold set at 0.01 and p-value threshold for calling variants set at 0.05 to generate Variant Call Format files. All SNVs/indels were annotated with ANNOVAR, and each SNV/indel was manually checked on the Integrative Genomics Viewer. Variants were further filtered with the following parameters: (i) minimum read depth = 20; (ii) minimum base quality = 15; (iii) minimum variant supporting reads = 5; (iv) variant supporting reads mapped to both strands; (v) strand bias no greater than 10%; (vi) if present in >1% population frequency in the 1000 g or ExAC database and vii) through an internally collected list of recurrent sequencing errors using a normal pool of 100 samples. The sequencing assay has been validated in compliance with CAP and CLIA with a limit of detection of 1% VAF. Copy number variations (CNVs) were called by FACETS43 (Fraction and Allele-Specific Copy Number Estimates from Tumor Sequencing) to obtain tumor purity-, ploidy-, and clonal heterogeneity-adjusted copy number data. Fusion events were called using the Delly fusion callying tool44 to identify the number of chimeric reads (sequencing paired ends mapped to different genes) and split reads (spanning a fusion breakpoint) from the targeted DNA-seq data. RTK fusions were filtered if (i) split reads <3 or paired reads <5, or (ii) lack of an intact kinase domain. All fusions were manually confirmed using the Integrative Genomics Viewer (IGV).
Clonality analysis
To infer the clonality of concomitant driver mutations, we used Pyclone45 to estimate the cancer cell fraction (CCF) of each mutation, with CCF > 0.6 considered as clonal mutations and CCF ≤ 0.6 considered as subclonal. For fusion events, CCF values were converted from the estimation of variant allele frequency of each fusion gene.
Statistical analysis
Comparisons of proportion between groups were done using the Fisher’s exact test. A two-sided p value <0.05 was considered significant. P values were adjusted for multiple group comparisons by Bonferonni’s post hoc test or corrected for multiple hypotheses testing using the false discovery rate (FDR) adjustment method as appropriate. Adjusted p value <0.1 was considered significant. For survival analyses, Kaplan–Meier curves were compared using the log-rank test, and hazard ratios (HRs) were calculated by Cox proportional hazards model. All statistical analyses were done in R (v.3.5.2).
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The data that support the findings of this study are available from the corresponding authors upon reasonable request. The raw sequencing data are available from GSA for human using the accession code HRA003240.
References
Mitelman, F., Johansson, B. & Mertens, F. The impact of translocations and gene fusions on cancer causation. Nat. Rev. Cancer 7, 233–245 (2007).
Lee, M. et al. ChimerDB 3.0: an enhanced database for fusion genes from cancer transcriptome and literature data mining. Nucleic Acids Res. 45, D784–D789 (2017).
Giacomini, C. P. et al. Breakpoint analysis of transcriptional and genomic profiles uncovers novel gene fusions spanning multiple human cancer types. PLoS Genet. 9, e1003464 (2013).
Hu, X. et al. TumorFusions: an integrative resource for cancer-associated transcript fusions. Nucleic Acids Res. 46, D1144–D1149 (2018).
Stransky, N., Cerami, E., Schalm, S., Kim, J. L. & Lengauer, C. The landscape of kinase fusions in cancer. Nat. Commun. 5, 4846 (2014).
Pan, Y. et al. Detection of novel NRG1, EGFR, and MET fusions in lung adenocarcinomas in the Chinese population. J. Thorac. Oncol. 14, 2003–2008 (2019).
Wang, R. et al. FGFR1/3 tyrosine kinase fusions define a unique molecular subtype of non-small cell lung cancer. Clin. Cancer Res. 20, 4107–4114 (2014).
Farago, A. F. et al. Clinicopathologic features of non-small-cell lung cancer harboring an NTRK gene fusion. JCO Precis. Oncol. 2018, https://doi.org/10.1200/po.18.00037 (2018).
Konduri, K. et al. EGFR fusions as novel therapeutic targets in lung cancer. Cancer Disco. 6, 601–611 (2016).
Zhu, Y. C. et al. Identification of a novel crizotinib-sensitive MET–ATXN7L1 gene fusion variant in lung adenocarcinoma by next generation sequencing. Ann. Oncol. 29, 2392–2393 (2018).
Davies, K. D. et al. Dramatic response to crizotinib in a patient with lung cancer positive for an HLA-DRB1-MET gene fusion. JCO Precis Oncol. 2017, PO.17.00117 (2017).
Sasaki, T., Rodig, S. J., Chirieac, L. R. & Jänne, P. A. The biology and treatment of EML4-ALK non-small cell lung cancer. Eur. J. Cancer 46, 1773–1780 (2010).
Chowdhury, T. et al. A glioneuronal tumor with CLIP2-MET fusion. npj Genomic Medicine 5, https://doi.org/10.1038/s41525-020-0131-6 (2020).
Okamura, R. et al. Analysis of NTRK alterations in pan-cancer adult and pediatric malignancies: implications for NTRK-targeted therapeutics. JCO Precis. Oncol. 1–20, https://doi.org/10.1200/po.18.00183 (2018).
Won, J. K. et al. Concomitant ALK translocation and EGFR mutation in lung cancer: a comparison of direct sequencing and sensitive assays and the impact on responsiveness to tyrosine kinase inhibitor. Ann. Oncol. 26, 348–354 (2015).
Yang, J. J. et al. Lung cancers with concomitant EGFR mutations and ALK rearrangements: diverse responses to EGFR-TKI and crizotinib in relation to diverse receptors phosphorylation. Clin. Cancer Res. 20, 1383–1392 (2014).
Kuo, Y. W., Wu, S. G., Ho, C. C. & Shih, J. Y. Good response to gefitinib in lung adenocarcinoma harboring coexisting EML4-ALK fusion gene and EGFR mutation. J. Thorac. Oncol. 5, 2039–2040 (2010).
Lou, N.-N. et al. Clinical outcomes of advanced non-small-cell lung cancer patients with EGFR mutation, ALK rearrangement and EGFR/ALK co-alterations. Oncotarget 7, 65185–65195 (2016).
Sahnane, N. et al. EGFR and KRAS mutations in ALK-positive lung adenocarcinomas: biological and clinical effect. Clin. Lung Cancer 17, 56–61 (2016).
Popat, S. et al. Lung adenocarcinoma with concurrent exon 19 EGFR mutation and ALK rearrangement responding to erlotinib. J. Thorac. Oncol. 6, 1962–1963 (2011).
Yang, Y. et al. A rare STRN-ALK fusion in lung adenocarcinoma identified using next-generation sequencing-based circulating tumor DNA profiling exhibits excellent response to crizotinib. Mayo Clin. Proc. Innov., Qual. Outcomes 1, 111–116 (2017).
Zhang, M. et al. CUX1-ALK, a novel ALK rearrangement that responds to crizotinib in non-small cell lung cancer. J. Thorac. Oncol. 13, 1792–1797 (2018).
Jiang, J. et al. GCC2-ALK as a targetable fusion in lung adenocarcinoma and its enduring clinical responses to ALK inhibitors. Lung Cancer 115, 5–11 (2018).
Yao, Y. et al. Characterizing kinase intergenic-breakpoint rearrangements in a large-scale lung cancer population and real-world clinical outcomes. ESMO Open 7, 100405 (2022).
Kang, J. et al. Complex ALK fusions are associated with better prognosis in advanced non-small cell lung cancer. Front. Oncol. 10, https://doi.org/10.3389/fonc.2020.596937 (2020).
Woo, C. G. et al. Differential protein stability and clinical responses ofEML4-ALK fusion variants to various ALK inhibitors in advancedALK-rearranged non-small cell lung cancer. Ann. Oncol. 28, 791–797 (2017).
Lin, J. J. et al. Impact of EML4-ALK variant on resistance mechanisms and clinical outcomes in ALK-positive lung cancer. J. Clin. Oncol. 36, 1199–1206 (2018).
Cocco, E., Scaltriti, M. & Drilon, A. NTRK fusion-positive cancers and TRK inhibitor therapy. Nat. Rev. Clin. Oncol. 15, 731–747 (2018).
Drilon, A. et al. Efficacy of larotrectinib in TRK fusion-positive cancers in adults and children. N. Engl. J. Med. 378, 731–739 (2018).
Doebele, R. C. et al. Entrectinib in patients with advanced or metastatic NTRK fusion-positive solid tumours: integrated analysis of three phase 1-2 trials. Lancet Oncol. 21, 271–282 (2020).
Drilon, A. et al. A next-generation TRK kinase inhibitor overcomes acquired resistance to Prior TRK kinase inhibition in patients with TRK fusion-positive solid tumors. Cancer Disco. 7, 963–972 (2017).
Drilon, A. et al. Repotrectinib (TPX-0005) is a next-generation ROS1/TRK/ALK inhibitor that potently inhibits ROS1/TRK/ALK solvent-front mutations. Cancer Disco. 8, 1227–1236 (2018).
Subbiah, V. et al. Precision targeted therapy with BLU-667 for RET-driven cancers. Cancer Disco. 8, 836–849 (2018).
Drilon, A. et al. Efficacy of selpercatinib in RET fusion-positive non-small-cell lung cancer. N. Engl. J. Med. 383, 813–824 (2020).
Wirth, L. J. et al. Efficacy of selpercatinib in RET-altered thyroid cancers. N. Engl. J. Med. 383, 825–835 (2020).
Loriot, Y. et al. Erdafitinib in locally advanced or metastatic urothelial carcinoma. N. Engl. J. Med. 381, 338–348 (2019).
Abou-Alfa, G. K. et al. Pemigatinib for previously treated, locally advanced or metastatic cholangiocarcinoma: a multicentre, open-label, phase 2 study. Lancet Oncol. 21, 671–684 (2020).
Fang, W. et al. Comprehensive genomic profiling identifies novel genetic predictors of response to anti-PD-(L)1 therapies in non-small cell lung cancer. Clin. Cancer Res. https://doi.org/10.1158/1078-0432.CCR-19-0585 (2019).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Koboldt, D. C. et al. VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res. 22, 568–576 (2012).
Shen, R. & Seshan, V. E. FACETS: allele-specific copy number and clonal heterogeneity analysis tool for high-throughput DNA sequencing. Nucleic Acids Res. 44, e131–e131 (2016).
Rausch, T. et al. DELLY: structural variant discovery by integrated paired-end and split-read analysis. Bioinformatics 28, i333–i339 (2012).
Roth, A. et al. PyClone: statistical inference of clonal population structure in cancer. Nat. Methods 11, 396–398 (2014).
Acknowledgements
We would like to thank all the patients and family members who gave their consent on presenting the data in this study, as well as the investigators and research staff at all hospitals and research sites involved. We are also grateful for all the support from the bioinformatics team of Nanjing Geneseeq Technology Inc., with special thanks to Drs. Jinfeng Zhang, Baihan Zhu, Xunbiao Liu, and Tingting Wu.
Author information
Authors and Affiliations
Contributions
Y.S., Z.W., and Z.C. conceived the project and supervised the research. T.W., L.W., Q.L., Y.M.S., S.Y., J.C.Y., and S.W. acquired and analyzed the data. T.W., L.W., Q.L., Y.M.S., J.C.Y., and S.W. participated in data interpretation. J.C.Y. and S.W. drafted the manuscript. All authors provided critical revision of the manuscript for important intellectual content.
Corresponding authors
Ethics declarations
Competing interests
Jiani C. Yin, Sha Wang, and Yang Shao are employees of Nanjing Geneseeq Technology Inc. All remaining authors have declared no conflict of interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, T., Wei, L., Lu, Q. et al. Landscape of potentially targetable receptor tyrosine kinase fusions in diverse cancers by DNA-based profiling. npj Precis. Onc. 6, 84 (2022). https://doi.org/10.1038/s41698-022-00325-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41698-022-00325-0
- Springer Nature Limited