Abstract
The interaction between genotype and environment (GEI) significantly influences plant performance, crucial for breeding programs and ultimately boosting crop productivity. Alongside GEI, breeders encounter another hurdle in their quest for yield improvement, notably adverse and negative correlations among pivotal traits. This study delved into the stability of white sugar yield (WSY), root yield (RY), sugar content (SC), extraction coefficient of sugar (ECS), and the interplay among essential traits including RY, SC, alpha amino nitrogen (N), sodium (Na+), and potassium (K+) across 15 sugar beet hybrids and three control varieties. The investigation spanned two locations over two consecutive years (2022–2023), employing a randomized complete block design with four replications to comprehensively analyze these factors. The analysis of variance highlighted the significant effects of environment, genotype, and GEI at the 1% probability level. Notably, the AMMI analysis of GEI revealed the significance of the first component for WSY, RY, and SC, with the first two components proving significant for ECS. Within the linear mixed model (LMM), WSY, RY, SC, and ECS demonstrated significant effects from both genotype and GEI. In the WAASB biplot, genotypes 10, 8, 17, 6, 13, 14, 15, 7, 12, and 16 exhibited stability in WSY, while genotypes 9, 10, 6, 14, 7, 8, 13, 12, 18, and 15 displayed stability in RY. Additionally, genotypes 10, 15, 12, 13, 16, 17, 6, and 14 were stable for SC, and genotypes 8, 10, 7, 6, 13, 12, 16, 17, 15, 14, and 18 showcased stability in ECS, boasting above-average yield values. In the genotype by yield × trait (GYT) biplot, genotypes 15, 18, and 16 emerged as top performers when combining RY with SC, Na+, N, and K+, suggesting their potential for inclusion in breeding programs.
Similar content being viewed by others
Introduction
Ensuring global food security has emerged as the primary scientific obstacle1. At present, the world's population is roughly seven billion individuals, with projections indicating a rise to around 10 billion by 2050 and possibly 11 billion by the century's end2. Naturally, as the population expands, the demand for food will also escalate significantly3. Although the global pressure on food production has intensified, advancements in crop cultivation and increased efficiency have thus far met the demand4. Nevertheless, projections suggest that food production must increase by approximately 70% by 2050 to sustain over 9 billion people5,6. However, serious concerns arise regarding the capacity of current agricultural varieties and methods to meet the growing demand for plant crops4. Conversely, the degradation of agricultural areas due to erosion, groundwater pollution from excessive fertilizer and pesticide use, yield reductions in numerous regions, and the depletion of natural resources pose significant threats to food security7,8,9. Furthermore, observed and anticipated climate changes exacerbate these challenges and lead to decreased crop yields in many global regions10,11. Consequently, various agricultural and environmental research disciplines face unprecedented challenges and necessitate approaches with minimal environmental impact to achieve global food security, ultimately ensuring stability and performance enhancement.
Developing new variations to further enhance yield potential in high-yield environments12,13,14 is a viable approach to addressing the food security challenge15,16. This strategy offers the advantage of continued yield improvement even under less favorable growth conditions17,18. Within the field of Agricultural Sciences, the study of sugar beet breeding holds great significance for the global food industry19,20. Although sugar beet does not directly contribute to human nutrition, it ranks as the second most cultivated crop worldwide and plays a crucial role in the production of white sugar for human consumption19,20. With an annual root production of approximately 278 million tons21, sugar beet exclusively serves the sugar industry and accounts for nearly 20% of the world's annual sugar output22,23,24,25,26. Over the past few decades, researchers and breeders have achieved substantial progress in enhancing the quantitative and qualitative performance of sugar beet through systematic breeding programs and biotechnological methods20. Notably, the sugar content has increased from eight percent to 18 percent in contemporary varieties. Genetic modification has been actively employed to confer desirable traits such as viral and fungal disease resistance, improved RY, seed monogermity, and bolting resistance14,16,20,27,28,29,30. Given the profound impact of these traits on growth and yield, their improvement is of utmost importance to meet future sugar demand. However, the multifaceted nature of sugar beet yield, influenced by genotype, environment, and their interaction, poses challenges to stability in product crop production28. GEI plays a crucial role in determining genotype yield stability across different environments13,14,31. This phenomenon describes how genetic traits can manifest differently depending on specific environmental conditions28, resulting in phenotypic variations among genotypes with identical genetic makeup in different environments27. The challenge of increasing yields in breeding poses a significant obstacle. To address this challenge, breeders can identify genotypes that consistently excel across diverse environmental conditions through extensive testing in varied locations14. This method aids in selecting genotypes with broad adaptability and stability31,32, ultimately ensuring stability in yield enhancement.
Advanced statistical methods have been developed to thoroughly examine and unveil patterns of GEI. The additive main effects and multiplicative interaction (AMMI) model, a statistical approach combining variance analysis and principal component analysis (PCA)33, plays a vital role in this realm. By dissecting GEI effects and visualizing patterns, the AMMI model aids in pinpointing genotypes with consistent yield across diverse environments34. Despite its benefits, the AMMI model also presents limitations35,36. To address the non-additive nature and decomposition challenges of the linear mixed model (LMM) structure, the best linear unbiased predictions (BLUP) model emerges as a complementary tool, offering estimates of average yields for high-yielding genotypes. Consequently, the AMMI and BLUP models' features are amalgamated, giving rise to the stability index of weighted average absolute scores of BLUPs (WAASB), facilitating the selection of superior genotypes35. The fusion of the AMMI model's power and the BLUP model's predictive accuracy enhances the efficiency of both models in exploring GEI35. However, as the introduction of a new variety necessitates considering both yield stability and productivity simultaneously, the WAASB index evolves into the WAASBY index. This index acts as a metric for phenotypic stability and quantitative yield, aiding in the selection of genotypes with high yield potential while mitigating GEI, thus optimizing their inclusion in cultivar introduction programs27.
The challenge of GEI is not the sole obstacle in the breeding process aimed at enhancing performance37. Another significant challenge faced by breeders is the unfavorable and negative correlation among key traits38,39,40. In the context of sugar beet, WSY is influenced by a combination of quantitative and qualitative traits such as RY, SC, N, Na+, and K+ levels41,42,43, exhibiting a spectrum of correlations from negative to positive44,45. These trait associations pose challenges for sugar beet breeding programs46, requiring breeders to navigate improvements across multiple traits while managing the implications of negative trait correlations. To address this complexity, the graphical method of GYT biplot operates on the premise that yield reigns as the paramount trait, with other key traits gaining significance only when linked to high yield. Judging the superiority of a genotype is contingent upon its ability to combine yield with other traits37. Yield emerges as the crucial trait determining genotype efficiency, with other traits proving valuable solely when correlated with optimal yield levels. Consequently, in selecting superior cultivars, the amalgamation of yield with traits holds more weight than evaluating cultivars based on individual traits alone. An ideal genotype strikes a balance by exhibiting optimal levels across traits to counteract negative trait correlations and perform well across various environments47. The primary objective of this study was to pinpoint superior genotypes among a group of promising sugar beet genotypes. The investigation delved into the impact of GEIs on the quantitative yield stability of sugar beet genotypes and assessed the interplay of key traits influencing yield. Ultimately, superior genotypes were selected based on these key traits.
Materials and methods
Plant materials
In this study, a total of 18 sugar beet genotypes were employed, comprising 15 hybrids and three controls (Table 1). The hybrids were developed at the Sugar Beet Seed Institute (SBSI), Karaj, Alborz, Iran, to incorporate resistance genes against diseases such as rhizomania, rhizoctonia, and cyst nematodes into genotypes with desirable quantitative and qualitative traits. Prior assessments through field and molecular evaluations confirmed the genetic resistance of these hybrids against diseases.
Research sites and experimental design
Phenotypic assessments of experimental genotypes were conducted over two consecutive crop years (2022 and 2023) at two agricultural research stations in Karaj and Kermanshah. These sites were selected based on varying ecological features, including differences in altitude, latitude and longitude (Table 2), atmospheric temperature and precipitation (Table 2), and physical and chemical characteristics of soil (Table 3). Each research station employed a randomized complete block design with four replications. Individual genotypes were planted in separate plots measuring three square meters, arranged in six rows, each 10 m long, with a row spacing of 50 cm. Planting took place between April 10th and April 20th in both years. Throughout each cropping season, standard agricultural practices such as weed control, irrigation, fertilization, and other management activities were carried out according to expert recommendations. Regular monitoring and preventive measures against sugar beet pests and diseases were diligently implemented at each research station. Harvesting occurred from October 20th to 30th every 2 years, focusing on the roots of the four central rows. Roots were harvested, counted, and weighed, with one meter of roots removed from both ends of the rows for evaluation. The experiments adhered to international, national, and institutional guidelines at all stages to ensure proper research conduct.
Data collection
Following field evaluations, the experimental genotypes were transported to the quality control laboratory of the SBSI for assessing their quality traits. The roots underwent a washing process, and a pulp sample was prepared utilizing an automated machine. This sample was subsequently preserved in a freezer at −18 °C. At the appropriate time, 26 g of the frozen samples were extracted and combined with 177 ml of lead (II) hydroxide acetate for three minutes using a mixer. The resulting solution was then sieved to obtain a clear liquid, which was utilized in the Betalyser device an automated system for sugar beet quality analysis to measure the levels of SC, N, Na+, and K+. To compute the WSY for each genotype, molasses sugar (MS) and white sugar content (WSC) were calculated using Eqs. (1) and (2). These values, in conjunction with the RY of each experimental genotype, were utilized in a formula to determine the WSY. Based on the obtained values, ECS was estimated by Eq. 4 Throughout the experiments, strict adherence to international, national, and institutional guidelines was maintained at every stage.
where MS is molasses sugar (%), K+ is potassium (meq 100 g−1), Na+ is sodium (meq 100 g−1), alpha-amino-N is nitrogen (meq 100 g−1), WSC is white sugar content (%), SC is sugar content (%), WSY is white sugar yield (t ha−1), RY is root yield (t ha−1) and ECS is extraction coefficient of sugar (%).
Statistical analysis
Prior to conducting any analysis, the Grubbs test48 was initially applied to the data, assuming normality. Additionally, the Bartlett test49 was utilized to assess the homogeneity of variance of experimental error across different years and locations. Once the uniformity of variance of experimental error was confirmed, a combined analysis of variance was conducted. This analysis involved a random effect for year, with fixed effects attributed to location and genotype. The analysis focused on WSY, RY, SC and ECS, as calculated using Eq. (5).
where \({Y}_{ijkl}\) is the response measured on the \({ijkl}\)th experimental unit (plot), \(\mu\) is the overall mean, \({Y}_{i}\) is the effect of the ith year, \({L}_{j}\) is the effect of the jth location, \({YL}_{ij}\) is the interaction effect of the ith level of Y with the jth level of L, \({B}_{\left(ij\right)k}\) is the effect of the jth block within the ith location, \({T}_{l}\) is the effect of the lth treatment, \({YT}_{il}\) is the interaction effect of the ith level of Y with the lth level of T, \({LT}_{jl}\) is the interaction effect of the jth level of L with the lth level of T, \({YLT}_{ijl}\) is the interaction effect of the ith level of Y with the jth level of L and the lth level of T, and \({BT}_{\left(ij\right)kl}\) is experimental error.
The first step in combining the features of AMMI and BLUP models was to calculate the scores of these models for each genotype under examination. Subsequently, these scores were merged using the WAASB, as outlined in Eq. (6). This process yielded a consolidated score that incorporated components from both the AMMI and BLUP models.
where (Eq. 6) \({\text{WAASB}}_{\text{i}}\) is the weighted average of absolute scores of the ith genotype or environment; \({\text{IPCA}}_{\text{ik}}\) is the absolute score of the ith genotype or environment in the kth interaction principal component (IPC); and \({\text{EP}}_{\text{k}}\) is the magnitude of the variance explained by the kth IPC. Given that the attainment of high-performance, stable genotypes is a primary objective, the individual contributions of GEI were examined using a BLUP matrix. Following this analysis, the WAASBY, which measures both yield average and stability, was calculated for each genotype as per Eq. (7)50.
where \({\text{WAAS}BY}_{\text{i}}\) is the superiority index with different weights between yield and stability for the \({g}\text{th}\) genotype; \({\theta }_{Y}\) and \({\theta }_{S}\) are the weights for yield and stability, respectively; \({rG}_{g}\) and \({rW}_{g}\) are the rescaled values of the \({g}\text{th}\) genotype for yield and WAASB, respectively.
Pearson's correlation coefficient51 was utilized to assess the level of correlation among the different traits being evaluated. The values of this coefficient range from 1 to −1. A correlation coefficient of 1 signifies a complete positive association between two traits, indicating that when one trait increases or decreases, the other will also increase or decrease correspondingly. Conversely, as the coefficient approaches zero, the strength of the positive association diminishes, eventually leading to no linear association when the correlation coefficient is zero. Conversely, as the coefficient moves towards −1, the strength of the inverse association between the two traits intensifies, culminating in a correlation coefficient of −1 denoting a complete inverse association. A perfect inverse association implies that as one trait increases, the other decreases and vice versa. To explore the interaction between genotype, yield, and trait, the original data were initially standardized using Eq. (8)37.
where \({P}_{ij}\) is the standardized value of genotype i for trait or yield-trait combination j in the standardized table, \({T}_{ij}\) is the original value of genotype i for trait or yield-trait combination j, \({\overline{T} }_{j}\) is the mean across genotypes for trait or yield-trait combination j, and \({S}_{j}\) is the standard deviation for yield-trait combination j. The biplot representing the GYT was constructed using the first and second principal components (PCs) derived from the singular value decomposition of the standardized data. In this model, the singular value decomposition breaks down the GYT data into genotype eigenvalues, yield-trait combination eigenvalues, and singular values. This process is based on Eq. (9) as proposed by Yan and Frégeau-Reid37.
where \({\zeta }_{\text{i}1}\) and \({\zeta }_{\text{i}2}\) are the eigenvalues for PC1 and PC2, respectively, for genotype i; \({\tau }_{1\text{j}}\) and \({\tau }_{2\text{j}}\) are the eigenvalues for PC1 and PC2, respectively for trait j, and \({\varepsilon }_{\text{ij}}\) is the residual from fitting the PC1 and PC2 for genotype i on trait j; \({\uplambda }_{1}\) and \({\uplambda }_{1}\) are the singular values for PC1 and PC2, respectively. α is the singular value partitioning factor. The GYT biplot is created by plotting \(d{\lambda }_{1}^{\alpha }{\zeta }_{\text{i}1}\) against \(d{\lambda }_{2}^{\alpha }{\zeta }_{\text{i}2}\) for genotypes, and \({\lambda }_{1}^{1-\alpha }{\tau }_{1\text{j}}/\text{d}\) against \({\lambda }_{2}^{1-\alpha }{\tau }_{2\text{j}}/\text{d}\) for yield-trait combinations. This analysis centers on four primary patterns: (1) investigating the relationships among various yield-trait combinations, (2) determining the optimal genotype for each yield-trait combination, (3) assessing and comparing genotypes by ranking them using a superiority index to identify strengths and weaknesses, and (4) ranking genotypes based on an ideal hypothetical genotype.
Results and discussion
Genotype–environment interaction analysis
The insignificance of the G statistic in the Grubbs test was determined based on the normality of the experimental data regarding WSY, RY, SC, and ECS. Additionally, the non-significant chi-square values obtained from Bartlett's test for these traits further affirmed the consistency of experimental error variance across four environments (two location over 2 years). Once the hypotheses were validated, a combined analysis of variance was conducted to delineate main effects and quantify interactions among different sources of variation. The mean square of the main effect of genotype exhibited a significant disparity at the 1% probability level for all four traits—WSY, RY, SC and ECS. Interaction effects of year-location, year-location- genotype at the 1% probability level for all traits, and year- genotype at the 5% probability level for WSY and at the 1% probability level for SC showed notable differences (Table 4). These significant variations observed in experimental years, locations, and genotypes can be attributed to changes in environmental conditions and genetic makeup, leading to distinct performances of WSY, RY, SC, and ECS across different environments. This underscores the importance of assessing genotypes in diverse environments due to the GEI, wherein genotypes respond differently to various locations. The study by Sadeghzadeh Hemayati, et al.28 highlighted the substantial impact of the environment and its interaction with the genetic composition of different genotypes on the phenotypic expression of WSY in sugar beet genotypes. The research by Saremirad and Taleghani14 emphasized that the GEI significantly influences the quantitative and qualitative aspects of sugar yield in sugar beet hybrids, underscoring the importance of considering this interaction in breeding new hybrids. Estimating GEI provides valuable insights for decision-making regarding adaptability in different environmental conditions, contributing to the development of stable cultivars suited to specific environments. Employing multivariate statistical methods can enhance the understanding of GEI in agricultural research27.
To enhance the reliability of multi-environment experimental analyses and gain a more precise understanding of GEIs, the data analysis of WSY, RY, SC, and ECS was conducted using a combination of AMMI and BLUP models. The significant impact of the environment, observed at a 1% probability level for all four traits, highlighted variations in the environmental conditions across the study areas. Genotype effects were also significant for all studied traits at a 1% probability level, indicating diverse performances in WSY, RY, SC, and ECS among the examined genotypes. GEI exhibited significance for all four traits at the 1% probability level (Table 5), demonstrating that experimental genotypes respond differently to changing environmental conditions. This interaction determines the strengths and weaknesses of each genotype based on environmental factors, leading to fluctuations in genotype rankings and resulting in variations in WSY, RY, SC, and ECS across different environments. These findings underscore the necessity of considering diverse interactions in agricultural research to achieve favorable outcomes in agricultural research development39,52.
Considering the significant impact of GEI on all assessed traits—WSY, RY, SC, and ECS—a multiplicative effects analysis was conducted to identify stable genotypes based on the AMMI's model. The analysis of the multiplicative effect of GEI on WSY revealed that the first IPC, significantly different at the 1% probability level, contributed to 87.30% of the total square of GEI, while the two non-significant IPC accounted for 10.10% and 2.60%, respectively. Similarly, for RY, the first IPC was significant at the 1% probability level, with the second and third IPCs showing no significant differences. Their respective contributions to explaining the sum of squares of GEI were 87.70%, 9.20%, and 3.20%. Analysis of SC highlighted one significant IPC at the 1% probability level and two non-significant IPCs, explaining 85.20%, 11.30%, and 3.50% of the variation. In terms of ECS, two significant IPCs elucidated 98.20% of the GEI variations, with shares of 79.40% and 18.80% each (Table 6). Previous studies by Omrani, et al.53, Sadabadi, et al.54, Mostafavi and Saremirad55, Rajabi, et al.13, and Sadeghzadeh Hemayati, et al.28 have demonstrated varying contributions of PCs in explaining GEI. Combining the power of the AMMI model with the predictive accuracy of the BLUP model is recommended to mitigate biases and provide a comprehensive understanding of GEIs, as indicated in this study and previous research.
The likelihood ratio test affirmed the results of the AMMI analysis, indicating significant effects of genotype, environment, and GEI at a 1% probability level across all four traits: WSY, RY, SC, and ECS (Table 5). The noteworthy impact of GEI underscores how the genetic composition of a genotype, coupled with environmental conditions specific to the field, jointly shape the phenotypic expression of these traits. Given the continuous influence of multiple genes associated with quantitative traits, genomic regions, and environmental factors on these traits56, genes governing yields and their components are particularly sensitive to environmental variations and demonstrate a complex interaction between control regions of quantitative traits and the environment. This interplay can either facilitate or impede responses to selection, emphasizing the importance of breeders carefully managing these effects57. Utilizing BLUP analysis to estimate genetic variance components can lead to more robust and accurate results, improving breeding program outcomes50. The REML/BLUP method was employed to estimate genetic variance components for the studied traits, attributing the highest phenotypic variance in WSY and RY to genetic variance, while SC and ECS were predominantly influenced by environmental variance (Fig. 1). A higher genetic diversity-to-environment ratio in WSY and RY enhances selection efficiency, enabling the identification and selection of favorable genotypes more accurately58. GEI variance had the smallest impact on phenotypic variance across all traits, suggesting limited influence on trait expression variability among sugar beet genotypes across different environments. When GEI variance is low compared to environmental variance, genetic variation has a reduced effect on trait expression, implying that phenotypic trait differences among environments are less genetically influenced59. This insight is crucial for breeders when selecting traits. The outcomes of studies on winter sugar beet cultivation and related research underline the primary influence of the environment on WSY, followed by genotype contributions, while GEI plays a lesser role in trait expression variation32. These findings shed light on the differential impacts of environmental conditions and genetic factors on trait expression in sugar beet genotypes, guiding breeders in their selection processes.
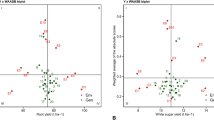
While the first two IPCs from the AMMI model typically explain the diversity of most genotypes, certain genotypes may require additional IPCs for a comprehensive analysis27. To address this, the WAASB index stands as a quantitative metric of sustainability, accommodating these variations. The biplot in Fig. 2 illustrates performance traits (WSY, RY, SC, and ECS) on the horizontal axis and WAASB index values on the vertical axis. Within this biplot, the vertical line at the center represents the average overall performance across test environments. Genotypes and environments to the right of this line indicate higher performance values than the average, whereas those on the left side exhibit lower performance values. The horizontal axis at the middle signifies the average WAASB index. The biplot is divided into four quadrants at the intersection of this axis with the vertical axis (average performance), enabling classification of genotypes based on their adaptability to distinct environments. In the first quadrant of the WSY biplot (Fig. 2A), genotypes 1, 2, and 3 and environments E3 and E4, for RY (Fig. 2B), genotypes 1, 2, and 3 and environments E3 and E4, for SC (Fig. 2C), genotypes 1, 2, 3, 4, and 11 and environments E1 and E2, and for ECS (Fig. 2D), genotypes 2, 3, and 1 and environments E2 and E3 demonstrated high WAASB values and performances below the overall average, indicating instability and fluctuation in performance, as well as below-average values. These genotypes are deemed unsuitable for cultivation due to their unstable nature. In contrast, genotypes like 18 and environment E2 in the second quadrant of the WSY biplot (Fig. 2A), genotype 16 and environment E2 in the second quadrant of the RY biplot (Fig. 2B), genotypes 7 and 18 and environments E3 and E4 in the second quadrant of the SC biplot (Fig. 2C), and environments E1 and E2 in the second quadrant of the ECS biplot (Fig. 2D) exhibited high WAASB and performance values surpassing the average. These genotypes, responsive to environmental conditions, possess good adaptability and exceptional performance under favorable conditions, making them suitable for cultivation in environments conducive to sugar beet growth. Environments in the second quadrant warrant special attention due to their above-average productivity and ability to distinguish genotypes effectively. In the WSY biplot (Fig. 2A) genotypes 9, 4, 5, and 11, in the RY biplot (Fig. 2B) genotypes 4, 5, 17, and 11, in the SC biplot (Fig. 2C) genotypes 8, 9, and 5, and in the ECS biplot (Fig. 2D) genotypes 9, 5, 11, and 4 were positioned in the third quadrant. These genotypes displayed lower WAASB index values, indicating stability or reduced sensitivity to environmental influence, alongside lower performance values. Essentially, these genotypes exhibited lower performance but higher stability, signifying a trade-off between performance and stability. On the other hand, genotypes 10, 8, 17, 6, 13, 14, 15, 7, 12, and 16, along with environment E1, were situated in the fourth quadrant of the WSY biplot (Fig. 2A), genotypes 9, 10, 6, 14, 7, 8, 13, 12, 18, and 15, along with environment E1, in the fourth quadrant of the RY biplot (Fig. 2B), genotypes 10, 15, 12, 13, 16, 17, 6, and 14 in the fourth quadrant of the SC biplot (Fig. 2C), and genotypes 8, 10, 7, 6, 13, 12, 16, 17, 15, 14, and 18 in the fourth quadrant of the ECS biplot (Fig. 2D) showed low WAASB index values and yields exceeding the average. These genotypes, recognized for their stable and optimal performance, exhibited minimal sensitivity to environmental conditions and consistently strong performance. Environments in the fourth quadrant demonstrated high productivity and low WAASB values compared to the other experimental environments. The WAASB index, derived from AMMI decomposition on the BLUP matrix, offers an advantage over the AMMI model by encompassing all IPCs scores to determine stability based on GEIs comprehensively. This index considers the total GEI variance, aiding in the identification of stable genotypes. In cases where the first IPC alone cannot effectively identify stable genotypes due to multiple IPCs influencing GEI, the WAASB index is recommended for its ability to capture the variance of GEI comprehensively. Studies have shown that a BLUP-based mixed model, like the one incorporating the WAASB index, often outperforms fixed-effects AMMI models in accuracy and stability assessments60,61,62,63. The WAASB stability index has proven effective in identifying genotypes with favorable and consistent performance across various plant species such as wheat64, soybean65, lentils66, rice67, corn63, and sugar beet27,28,31,52, yielding valuable and reliable results in plant breeding studies.
To enhance the accuracy of performance and stability evaluations, experimental genotypes were ranked based on WAASBY index scores (Fig. 3). The WAASBY index scores, depicted in a heat map format, represent various ratios of WAASB index to yield for WSY (Fig. 3A), RY (Fig. 3B), SC (Fig. 3C), and ECS (Fig. 3D). Genotype rankings using the WAASBY can vary depending on the ratio of WAASB to performance. The first component, situated to the left of the diagonal line, corresponds to the ratio of WAASB to yield, emphasizing environmental sustainability, whereas the second component, on the right side of the diagonal line, relates to performance. This ranking system assigns a ratio of 0.100 to stability and another ratio of 0.100 to performance. Moving across one unit from left to right diminishes the environmental stability component by 5% and increases the performance component, ultimately resulting in the genotypes being ranked solely based on performance (0.100). Based on the WAASBY scores, experimental genotypes were categorized into four groups. The first group (green) for WSY comprised genotypes 1 and 11, for RY included genotypes 1, 2, 3, 4, 5, 11, and 17, for SC involved genotypes 1, 3, and 11, and for ECS comprised genotypes 1, 2, 3, 4, 9, and 11, representing stable genotypes with optimal performance. The second group (red) encompassed genotypes with unfavorable performance and instability, such as genotypes 2, 3, 4, and 5 for WSY, 6, 9, 13, and 16 for RY, 2, 4, 5, 8, and 9 for SC, and 5, 7, and 8 for ECS. The third group (blue) included genotypes with good performance value but instability, comprising genotypes 6, 7, 9, 13, 14, 15, 16, 17, and 18 for WSY, 7, 8, 10, and 12 for RY, 6, 10, 12, and 16 for SC, and 6, 10, and 12 for ECS. The fourth group (black) consisted of genotypes with stable performance despite unfavorable values, like genotypes 8, 10, and 12 for WSY, 14, 15, and 18 for RY, 7, 13, 14, 15, 17, and 18 for SC, and 13, 14, 15, 16, 17, and 18 ECS. The WAASBY, serving as a selection index balancing stability and performance, enables breeders to choose genotypes based on varying stability and performance ratios aligned with their breeding objectives. Given the global significance of sugar beet production, accounting for approximately 30% of the world's sugar demand7, and the growing concerns regarding climate change and the shift towards renewable fuels68, developing genotypes with optimal quantitative and qualitative performance resilient to diverse environmental conditions becomes pivotal28,31,69. The WAASBY, with its emphasis on both yield and stability, facilitates the selection of high-yielding and stable genotypes, aligning with strategic breeding goals31,70,71.
Genotype by yield × trait interaction analysis
The results of Pearson's correlation analysis for various traits in experimental sugar beet genotypes are depicted in Fig. 4. The most notable finding was a strong positive correlation, significant at the 1% probability level, between N and SC, reaching a coefficient of 0.54. This correlation implies that as the SC increases, so does the N content. Additionally, K+ exhibited a positive but weak correlation with N (r = 0.34) and with SC (r = 0.24), both significant at the 1% probability level. Conversely, RY displayed negative correlations with all traits, including SC, Na+, K+, and N. Particularly, a substantial negative correlation, significant at the 1% probability level, was noted between RY and K+ (r = −0.27), indicating a decrease in K+ content with higher RY. On the other hand, weak and non-significant negative correlations were observed between RY and SC (r = −0.03), RY and N (r = −0.02), and RY and Na+ (r = −0.11). Similar associations between RY and SC have been documented in previous studies19,72. In the study by Hassani, et al.27, positive correlations were found between N and K+, Na+ and N, SC and K+, Na+ and K+, while negative correlations were identified between RY and K+, RY and SC, RY and N, and RY and Na+. These interrelations among key sugar beet traits introduce complexities into breeding programs, necessitating breeders to carefully manage the trade-offs induced by negative correlations among these crucial traits. Utilizing graphical method such as GYT analyses can provide a comprehensive and effective approach to navigating these trait interactions in sugar beet breeding programs.
To dissect the genetic influences impacting crop productivity, a GYT graphical analysis method was implemented. The outcomes revealed that the first and second PCs elucidated 60.61% and 33.73%, respectively, cumulatively explaining 94.34% of the variations in RY data. The substantial coverage by the first two PCs, exceeding 70% of the data variance, underscores the relative validity of the biplot diagrams derived from this study in delineating GYT73. In cases where the sum of the first and second PCs cannot account for most of the data variance, it signifies the intricate nature of the dataset, although this does not render the biplot invalid74. Shojaei, et al.75 demonstrated that approximately 50% of the GYT variation can be attributed to the first two PCs. In a study by Faheem, et al.47, the combined contribution of the first and second PCs reached nearly 85%, with the PC1 explaining around 74% and the second PC2 elucidating approximately 11% of the total data variations. In a separate experiment, Hassani, et al.27 found that the first and second PCs accounted for 50.53% and 34.96%, respectively, collectively representing 85.49% of the alterations in RY data.
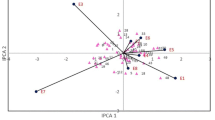
Undesirable N, Na+, and K+ levels are recognized as impurities within sugar beet roots, impacting both the quality and yield of the resultant sugar. N in sugar beet roots, embodying soluble amino acids and amide groups, significantly influences sugar extraction efficiency76. Elevated N levels can impede sugar extraction while also playing a crucial role in the preservation and quality of sugar beet during storage. Hence, managing N levels becomes critical, influenced by factors like temperature, storage duration, and the diverse genetic makeup of sugar beet varieties77. Na+ and K+ in sugar beet roots act as molasses substances, enhancing sucrose solubility while dampening crystallization, ultimately compromising sugar beet quality and extraction efficiency78,79,80,81. Excessive levels of these elements can lead to their accumulation in molasses, impacting quality and limiting specific applications. Therefore, corrective strategies should aim to simultaneously enhance RY while reducing impurity concentrations. Exploring the correlations between yield and trait combinations can unveil intricate associations, guiding future experiments towards developing novel genotypes. In the graphical representation, acute angles among the vectors indicate positive correlations between yield-trait combinations, reflecting performance as a primary component within these combinations37. Strong correlations among yield-trait pairings suggest a high consistency in genotype rankings across these combinations. Notably, there was a weak positive correlation between RY/N and RY*SC, alongside a negative correlation with RY/Na+. Conversely, there existed a significant positive correlation between RY*SC, RY/K+, and RY/Na+ (Fig. 5A). These positive associations between RY and SC with other compounds imply the potential to enhance SC by minimizing N, Na+, and K+ levels in a given genotype. However, the negative correlation between RY/N and RY/Na+ complicates the breeding process, necessitating meticulous planning to address all three impurities effectively.
(a) The tester vector view of the genotype by root yield × trait biplot to show associations among the yield-trait combinations, (b) The which-won-where view of the genotype by root yield × trait biplot to highlight genotypes with outstanding profiles, (c) The average tester coordination view of the genotype by root yield × trait biplot to rank the genotypes based on their overall superiority and their strengths and weaknesses, (d) The genotype ranking view of the genotype by root yield × trait biplot to rank the genotypes based on ideal genotype.
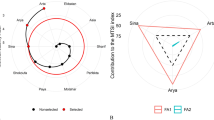
In biplot of Fig. 5B, a polygon is evident, connecting genotypes 15, 18, 16, 3, 2, and 1, positioned farthest from the coordinate origin. Perpendicular lines were extended from the origin to the sides of this polygon, aiding in the classification of GYT combinations. Genotypes situated above specific yield-trait combinations signify their strong performance value concerning those specific traits, indicating superior performance in those trait combinations. Notably, genotypes 15 and 12 excelled in the RY/Na+ combination, genotype 18 shone in the RY*SC and RY/K+ combinations, while genotypes 16 and 7 stood out in the RY/N combination. Conversely, genotypes located in sections devoid of combinations indicate their lack of desirability across all trait combinations, portraying them as weaker performers. The polygonal biplot facilitated the categorization of yield-trait combinations into three distinct groups: RY/Na+ in the first group, RY*SC and root RY/K+ in the second group, and RY/N in the third group (Fig. 5B).
The ranking of genotypes based on the yield-trait combination was assessed using the biplot diagram of average tester coordinates (Fig. 5C) and the superiority index (Table 7). Within the tester's coordinate biplot, the average axis, marked by an arrow passing through the coordinate origin, represents the mean of all yield-trait combinations. The axis perpendicular to the average axis gauges the genotypes' equilibrium concerning various traits. Notably, genotype 18, followed by genotype 16, exhibited the highest average yield and were identified as the top-performing genotypes, while genotypes 1 and 2 displayed the lowest average yield. The outcomes derived from the average tester coordinates biplot were further corroborated by the superiority index. Multiple studies, including those by Faheem, et al.47, Hassani, et al.27, Shojaei, et al.75, and Yan and Frégeau-Reid37, alongside the current study's findings, attest to the utility of the average tester coordinates biplot in the GYT analysis, providing valuable insights into genotype performance. The identification of an ideal hypothetical genotype is predicated on achieving a balance between traits and maximizing performance. An ideal genotype ideally combines the highest yield with optimal balance, with superior genotypes being those closest to this hypothetical ideal, and less desirable genotype being further away. Genotypes 18, 10, 12, 16, and 7 emerged as top performers due to their proximity to the ideal genotype, while genotypes 1 and 2 were deemed unfavorable for being farthest from this ideal (Fig. 5D). Hassani, et al.27 utilized the GYT graphical method to explore trait associations, identify superior genotypes, yielding insightful and practical results.
Conclusion
In this study, the effectively utilized the strengths of the AMMI method alongside the BLUP method within an LMM framework to assess genotypic stability and GEIs in sugar beet, as facilitated by the WAASB. This integrated approach not only addressed limitations inherent in the AMMI model but also provided a comprehensive view of the complexities of GEIs, which are crucial for understanding the adaptability and performance of sugar beet genotypes across varying environments. The findings reveal that environmental factors significantly influence the expression of key traits such as WSY, RY, SC, and ECS, leading to different responses by genotypes under diverse conditions. Notably, Genotypes 16, 10, 9, and 6 demonstrated both stability and high yield across all evaluated traits, highlighting their potential as superior genotypes for sugar beet breeding programs. These genotypes consistently showed high RY and SC, with low impurities, which are desirable traits for enhancing sugar beet production. Moreover, Genotypes 15, 18, and 16 were particularly outstanding in their performance across the combinations of RY*SC, RY/Na+, RY/K+, and RY/N, indicating their suitability for specific breeding goals aimed at improving performance and reducing impurities. Given these results, future breeding efforts should focus on these genotypes to develop sugar beet varieties that can deliver high performance and stability in diverse environmental conditions. Continued exploration of these genotypes will aid in fine-tuning breeding strategies and potentially introduce more resilient and productive sugar beet varieties.
Data availability
The datasets used and/or analyzed during the current study available from the corresponding author on reasonable request.
References
Cassman, K. G. What do we need to know about global food security?. Glob. Food Sec. 2, 81–82 (2012).
United Nations. World Population Prospects 2019: Highlights. (Department of Economic and Social Affairs, Population Division, 2019).
Saremirad, A. & Mostafavi, K. Genetic diversity study of sunflower (Helianthus annus L.) genotypes for agro-morphological traits under normal and drought stress conditions. Plant Prod. 43, 227–240. https://doi.org/10.22055/ppd.2020.27588.1671 (2020).
Voss-Fels, K. P., Stahl, A. & Hickey, L. T. Q&A: Modern crop breeding for future food security. BMC Biol. 17, 1–7 (2019).
FAO. How to feed the world 2050: High-level expert forum. (Food and Agriculture Organization of the United Nations, 2024).
United Nations. World Population Prospects: the 2017 Revision. (United Nations. Department of International Economic, 2017).
FAO. (Food and Agriculture Organization of the United Nation, 2018).
Ferber, D. (American Association for the Advancement of Science, 2001).
Smith, M. D. et al. Seafood prices reveal impacts of a major ecological disturbance. Proc. Natl. Acad. Sci. 114, 1512–1517 (2017).
Asseng, S. et al. Wheat yield potential in controlled-environment vertical farms. Proc. Natl. Acad. Sci. 117, 19131–19135 (2020).
Field, C. B. & Barros, V. R. Climate Change 2014–Impacts, Adaptation and Vulnerability: Regional Aspects. (Cambridge University Press, 2014).
Evans, L. & Fischer, R. Yield potential: Its definition, measurement, and significance. Crop Sci. 39, 1544–1551 (1999).
Rajabi, A., Ahmadi, M., Bazrafshan, M., Hassani, M. & Saremirad, A. Evaluation of resistance and determination of stability of different sugar beet (Beta vulgaris L.) genotypes in rhizomania-infected conditions. Food Sci. Nutr. 11, 1403–1414. https://doi.org/10.1002/fsn3.3180 (2023).
Saremirad, A. & Taleghani, D. Utilization of univariate parametric and non-parametric methods in the stability analysis of sugar yield in sugar beet (Beta vulgaris L.) hybrids. J. Crop Breed. 14, 49–63 (2022).
Reynolds, M. et al. Addressing research bottlenecks to crop productivity. Trends Plant Sci. 26, 607–630 (2021).
Taleghani, D., Rajabi, A., Saremirad, A. & Darabi, S. Estimation of gene action and genetic parameters of some quantitative and qualitative characteristics of sugar beet (Beta vulgaris L.) by line × tester analysis. Crop Breed. 15, 201–212 (2024).
Bustos, D. V., Hasan, A. K., Reynolds, M. P. & Calderini, D. F. Combining high grain number and weight through a DH-population to improve grain yield potential of wheat in high-yielding environments. Field Crops Res. 145, 106–115 (2013).
Reynolds, M. et al. Achieving yield gains in wheat. Plant Cell Environ. 35, 1799–1823 (2012).
Saremirad, A., Hamdi, F. & Taleghani, D. Evaluation of genetic diversity in sugar beet (Beta vulgaris L.) hybrids in terms of yield, qualitative and germination traits. Appl. Field Crops Res. 35, 87–67. https://doi.org/10.22092/aj.2023.357194.1580 (2023).
Taleghani, D., Rajabi, A., Hemayati, S. S. & Saremirad, A. Improvement and selection for drought-tolerant sugar beet (Beta vulgaris L.) pollinator lines. Results Eng. 13, 100367 (2022).
FAO. (Food and Agriculture Organization, 2021).
Akyüz, A. & Ersus, S. Optimization of enzyme assisted extraction of protein from the sugar beet (Beta vulgaris L.) leaves for alternative plant protein concentrate production. Food Chem. 335, 127673 (2021).
Lammens, T., Franssen, M., Scott, E. & Sanders, J. Availability of protein-derived amino acids as feedstock for the production of bio-based chemicals. Biomass Bioenergy 44, 168–181 (2012).
Tenorio, A. T., Schreuders, F., Zisopoulos, F., Boom, R. & Van der Goot, A. Processing concepts for the use of green leaves as raw materials for the food industry. J. Clean. Prod. 164, 736–748 (2017).
Tomaszewska, J. et al. Products of sugar beet processing as raw materials for chemicals and biodegradable polymers. RSC Adv. 8, 3161–3177 (2018).
Monteiro, F. et al. Genetic and genomic tools to asssist sugar beet improvement: The value of the crop wild relatives. Front. Plant Sci. 9, 74–89 (2018).
Hassani, M., Mahmoudi, S. B., Saremirad, A. & Taleghani, D. Genotype by environment and genotype by yield × trait interactions in sugar beet: analyzing yield stability and determining key traits association. Sci. Rep. 13, 23111. https://doi.org/10.1038/s41598-023-51061-9 (2024).
Sadeghzadeh Hemayati, S. et al. Evaluation of white sugar yield stability of some commercially released sugar beet cultivars in Iran from 2011–2020. Seed Plant J. 38, 339–364. https://doi.org/10.22092/spj.2023.362024.1305 (2022).
Taleghani, D., Hosseinpour, M., Nemati, R. & Saremirad, A. Study of the possibility of winter sowing of sugar beet (Beta vulgaris L.) early cultivars in Moghan region, Iran. Iran. Soc. Crops Plant Breed. Sci. 24, 319–334 (2023).
Taleghani, D. & Saremirad, A. Evaluation of the sugar beet (Beta vulgaris L.) half-sib lines response to drought stress. Crop Sci. Res. Arid Regions 5, 81–104 (2023).
Taleghani, D., Rajabi, A., Saremirad, A. & Fasahat, P. Stability analysis and selection of sugar beet (Beta vulgaris L.) genotypes using AMMI, BLUP, GGE biplot and MTSI. Sci. Rep. 13, 10019. https://doi.org/10.1038/s41598-023-37217-7 (2023).
Taleghani, D. et al. Genotype × environment interaction effect on white sugar yield of winter-sown short-season sugar beet (Beta vulgaris L.) cultivars. Seed Plant J. 38, 53–69. https://doi.org/10.22092/spj.2022.360021.1275 (2022).
Gauch, H. Statistical Analysis of Regional Yield Trials: AMMI Analysis of Factorial Designs. (Elsevier Science Publishers, 1992).
Senguttuvel, P. et al. Evaluation of genotype by environment interaction and adaptability in lowland irrigated rice hybrids for grain yield under high temperature. Sci. Rep. 11, 15825. https://doi.org/10.1038/s41598-021-95264-4 (2021).
Olivoto, T. et al. Mean performance and stability in multi-environment trials I: combining features of AMMI and BLUP techniques. Agron. J. 111, 2949–2960 (2019).
Rodrigues, P. C., Monteiro, A. & Lourenço, V. M. A robust AMMI model for the analysis of genotype-by-environment data. Bioinformatics 32, 58–66 (2016).
Yan, W. & Frégeau-Reid, J. Genotype by yield∗ trait (GYT) biplot: A novel approach for genotype selection based on multiple traits. Sci. Rep. 8, 1–10 (2018).
Yan, W. Crop Variety Trials: Data Management and Analysis. (Wiley, 2014).
Yan, W. & Kang, M. S. GGE Biplot Analysis: A Graphical Tool for Breeders, Geneticists, and Agronomists. (CRC Press, 2002).
Yan, W. et al. Development and evaluation of a core subset of the USDA rice germplasm collection. Crop Sci. 47, 869–876 (2007).
Cook, D. & Scott, R. The Sugar Beet Crop: Science into Practice. (Champan and Hall Press, 1993).
Kunz, M., Martin, D. & Puke, H. Precision of beet analyses in Germany explained for polarization. Zuckerindustrie 127, 13–21 (2002).
Reinfeld, E., Emmerich, G., Baumgarten, C., Winner & Beiss, U. Zur Voraussage des Melassez zuckersaus Ruben Analysen Zucker. (Chapman & Hall, World Crop Series, 1974).
Rašovský, M., Pačuta, V., Ducsay, L. & Lenická, D. Quantity and quality changes in sugar beet (Beta vulgaris Provar. Altissima Doel) induced by different sources of biostimulants. Plants (Basel) https://doi.org/10.3390/plants11172222 (2022).
Tsialtas, J. T. & Maslaris, N. Sugar beet root shape and its relation with yield and quality. Sugar Tech. 12, 47–52. https://doi.org/10.1007/s12355-010-0009-5 (2010).
Taleghani, D., Rajabi, A., Sadeghzadeh Hemayati, S. & Saremirad, A. Improvement and selection for drought-tolerant sugar beet (Beta vulgaris L.) pollinator lines. Results Eng. 13, 100367. https://doi.org/10.1016/j.rineng.2022.100367 (2022).
Faheem, M., Arain, S. M., Sial, M. A., Laghari, K. A. & Qayyum, A. Genotype by yield × trait (GYT) biplot analysis: A novel approach for evaluating advance lines of durum wheat. Cereal Res. Commun. 51, 447–456. https://doi.org/10.1007/s42976-022-00298-7 (2023).
Grubbs, F. E. Procedures for detecting outlying observations in samples. Technometrics 11, 1–21 (1969).
Bartlett, M. S. Properties of sufficiency and statistical tests. Proc. R. Soc. Lond. Ser. A-Math. Phys. Sci. 160, 268–282 (1937).
Olivoto, T., Lúcio, A. D., da Silva, J. A., Sari, B. G. & Diel, M. I. Mean performance and stability in multi-environment trials II: Selection based on multiple traits. Agron. J. 111, 2961–2969 (2019).
Sedgwick, P. Pearson’s correlation coefficient. Bmj 345, 54 (2012).
Sadeghzadeh Hemayati, S. et al. Study of genotype-environment interaction effect on sugar yield of sugar beet (Beta vulgaris L.) hybrids. Crop Sci. Res. Arid Regions 5, 345–364. https://doi.org/10.22034/csrar.2023.346833.1248 (2023).
Omrani, S., Omrani, A., Afshari, M., Bardehji, S. & Foroozesh, P. Application of additive main effects and multiplicative interaction and biplot graphical analysis multivariate methods to study of genotype-environment interaction on safflower genotypes grain yield. J. Crop Breed. 11, 153–163 (2019).
Sadabadi, M. F., Ranjbar, G., Zangi, M., Tabar, S. & Zarini, H. N. Analysis of stability and adaptation of cotton genotypes using GGE Biplot method. Trakia J. Sci. 16, 51–61 (2018).
Mostafavi, K. & Saremirad, A. Genotype-environment interaction study in corn genotypes using additive main effects and multiplicative interaction method and GGE-biplot method. J Crop Prod. 14, 1–12. https://doi.org/10.22069/ejcp.2022.17527.2293 (2021).
Said, A. A. et al. Genome-wide association mapping of genotype-environment interactions affecting yield-related traits of spring wheat grown in three watering regimes. Environ. Exp. Bot. 194, 104740 (2022).
Falconer, D. S. The problem of environment and selection. Am. Nat. 86, 293–298 (1952).
Saremirad, A., Bihamta, M. R., Malihipour, A., Mostafavi, K. & Alipour, H. Genome-wide association study in diverse Iranian wheat germplasms detected several putative genomic regions associated with stem rust resistance. Food Sci. Nutr. 9, 1357–1374. https://doi.org/10.1002/fsn3.2082 (2021).
Saltz, J. B. et al. Why does the magnitude of genotype-by-environment interaction vary?. Ecol. Evol. 8, 6342–6353. https://doi.org/10.1002/ece3.4128 (2018).
Hilmarsson, H. S., Rio, S. & Sánchez, J. I. Y. Genotype by environment interaction analysis of agronomic spring barley traits in Iceland using AMMI, factorial regression model and linear mixed model. Agronomy 11, 499 (2021).
Piepho, H., Möhring, J., Melchinger, A. & Büchse, A. BLUP for phenotypic selection in plant breeding and variety testing. Euphytica 161, 209–228 (2008).
Piepho, H.-P. Best linear unbiased prediction (BLUP) for regional yield trials: A comparison to additive main effects and multiplicative interaction (AMMI) analysis. Theor. Appl. Genet. 89, 647–654 (1994).
Yue, H. et al. Genotype by environment interaction analysis for grain yield and yield components of summer maize hybrids across the Huanghuaihai region in China. Agriculture 12, 602 (2022).
Verma, A. & Singh, G. Stability index based on weighted average of absolute scores of AMMI and yield of wheat genotypes evaluated under restricted irrigated conditions for peninsular zone. Int. J. Agric. Environ. Biotechnol. 13, 371–381 (2020).
Abdelghany, A. M. et al. Exploring the phenotypic stability of soybean seed compositions using multi-trait stability index approach. Agronomy 11, 2200 (2021).
Sellami, M. H., Pulvento, C. & Lavini, A. Selection of suitable genotypes of lentil (Lens culinaris Medik.) under rainfed conditions in south Italy using multi-trait stability index (MTSI). Agronomy 11, 1807 (2021).
Sharifi, P., Erfani, A., Abbasian, A. & Mohaddesi, A. Stability of some of rice genotypes based on WAASB and MTSI indices. Iran. J. Genet. Plant Breed. (IJGPB) 9, 113 (2020).
Salazar-Ordóñez, M., Pérez-Hernández, P. P. & Martín-Lozano, J. M. Sugar beet for bioethanol production: An approach based on environmental agricultural outputs. Energy Policy 55, 662–668 (2013).
Rajabi, A., Ahmadi, M., Bazrafshan, M., Hassani, M. & Saremirad, A. Evaluation of resistance and determination of stability of different sugar beet (Beta vulgaris L.) genotypes in rhizomania-infected conditions. Food Sci. Nutr. 11, 1403–1414. https://doi.org/10.1002/fsn3.3180 (2022).
Lee, S. Y. et al. Multi-environment trials and stability analysis for yield-related traits of commercial rice cultivars. Agriculture 13, 256 (2023).
Nataraj, V. et al. WAASB-based stability analysis and simultaneous selection for grain yield and early maturity in soybean. Agron. J. 113, 3089–3099 (2021).
Nasri, R., Kashani, A., Paknejad, F., Sadeghi, S. M. & Ghorbani, S. Correlation and path analysis of qualitative and quantitative yield in sugar beet in transplant and direct cultivation method in saline lands. Agron. Plant Breed. 8, 213–226 (2012).
Cruz, C., Regazzi, A. & Carneiro, P. Modelos Biométricos Aplicados ao Melhoramento (UFV, 2012).
Yan, W. & Tinker, N. A. An integrated biplot analysis system for displaying, interpreting, and exploring genotype× environment interaction. Crop Sci. 45, 1004–1016 (2005).
Shojaei, S. H. et al. Comparison of genotype× trait and genotype× yield-trait biplots in sunflower cultivars. Int. J. Agric. Environ. Food Sci. 7, 136–147 (2023).
Martínez-Arias, R., Müller, B. U. & Schechert, A. Near-infrared determination of total soluble nitrogen and betaine in sugar beet. Sugar Tech. 19, 526–531. https://doi.org/10.1007/s12355-016-0496-0 (2017).
Gippert, A.-L. et al. Unraveling metabolic patterns and molecular mechanisms underlying storability in sugar beet. BMC Plant Biol. 22, 430. https://doi.org/10.1186/s12870-022-03784-6 (2022).
Aljabri, M. et al. Recycling of beet sugar byproducts and wastes enhances sugar beet productivity and salt redistribution in saline soils. Environ. Sci. Pollut. Res. 28, 45745–45755. https://doi.org/10.1007/s11356-021-13860-3 (2021).
Makhlouf, B. S. I., Khalil, S. R. A. E. & Saudy, H. S. Efficacy of humic acids and chitosan for enhancing yield and sugar quality of sugar beet under moderate and severe drought. J. Soil Sci. Plant Nutr. 22, 1676–1691. https://doi.org/10.1007/s42729-022-00762-7 (2022).
Muir, B. M. Sugar Beet Cultivation, Management and Processing. 837–862 (Springer, 2022).
Xie, X. et al. Potassium determines sugar beets’ yield and sugar content under drip irrigation condition. Sustainability 14, 12520 (2022).
Acknowledgements
The authors express their gratitude to the technicians at the Sugar Beet Seed Institute (SBSI), Karaj, Iran, for their assistance in conducting the experiments.
Author information
Authors and Affiliations
Contributions
"S.S. and H.O. wrote the main manuscript text and M.R. prepared Figs. 1, 2 and 3. A. N read and check manuscript. All authors reviewed the manuscript."
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Ramazi, M., Omidi, H., Sadeghzadeh Hemayati, S. et al. Unraveling genotypic interactions in sugar beet for enhanced yield stability and trait associations. Sci Rep 14, 20815 (2024). https://doi.org/10.1038/s41598-024-71139-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-71139-2
- Springer Nature Limited