Abstract
Due to the high affinity and specificity of antibodies toward antigens, various antibody-based applications have been developed. Recently, variable antigen-binding domains of heavy-chain antibodies (VHH) have become an attractive alternative to conventional fragment antibodies due to their unique molecular characteristics. As an antibody-generating strategy, synthetic VHH libraries (including humanized VHH libraries) have been developed using distinct strategies to constrain the diversity of amino acid sequences. In this study, we designed and constructed several novel synthetic humanized VHH libraries based on biophysical analyses conducted using the complementarity determining region-grafting method and comprehensive sequence analyses of VHHs deposited in the protein data bank. We obtained VHHs from the libraries, and hit clones exhibited considerable thermal stability. We also found that VHHs from distinct libraries tended to have different epitopes. Based on our results, we propose a strategy for generating humanized VHHs with distinct epitopes toward various antigens by utilizing our library combinations.
Similar content being viewed by others
Introduction
An antibody recognizes a unique target molecule called an antigen. Due to the high affinity and specificity of antibodies toward antigens, various antibody-based applications have been developed in the biotechnological and medical fields, such as detection reagents for the target molecule, diagnostic tools, and antibody-based drugs for human therapy. Antibody-based proteins have become an important class of biologic therapeutics1. Currently, nearly 1200 antibody therapeutics are in clinical studies, and ~ 175 therapeutics are in regulatory review or have been approved2.
In 1993, functional antibodies devoid of light chains and composed of only the heavy chain were discovered in the serum of Camelus dromedarius3. The antibodies, referred to as heavy-chain antibodies (HCAbs), are unique because they lack the entire light chain and the first heavy chain constant region. The variable antigen-binding domain of a HCAb, referred to as the variable domain of the heavy-chain antibody (VHH), is generally functional as a single domain despite its small size of only 15 kDa4,5. While conventional antibodies tend to form a flat surface or a groove as a paratope, the convex paratopes of VHHs have a smaller antigen-binding interface. Thus, VHHs tend to bind to concave-shaped epitopes, such as enzyme catalytic sites6,7,8. VHHs are easily produced by bacterial expression systems due to their small size and single domain, and their production cost is much lower than that of immunoglobulin G. In addition, the small size of VHHs leads to rapid extravasation followed by deep-tissue penetration and rapid blood clearance. Based on these characteristics, VHHs are an attractive alternative to antigen-binding fragments from conventional antibodies (e.g., Fabs and scFvs) in biotechnological, diagnostic, and therapeutic applications8,9.
Synthetic VHH libraries coupled with display technologies such as phage display and yeast display have been developed as an antibody generating strategy10,11. Synthetic libraries do not require an immunization step, which decreases the period of time required for selection. Isolation of VHHs from synthetic libraries is also an attractive option when immunization is not feasible due to high toxicity or non-immunogenicity of the target antigen.
Several VHH-based antibody drugs, caplacizumab and ozoralizumab have been developed for human therapy, to treat acquired thrombotic thrombocytopenic purpura and rheumatoid arthritis, respectively12. Both VHH based drugs shows the superior therapeutic efficacy relying on multivalent format to accomplish long serum half life13. As both the VHH drug were humanized, antibodies derived from non-human origins must be subjected to the humanization process including CDR-grafting onto a humanized framework and resurfacing focusing on hallmark VHH-specific amino acids in the framework-2 region to avoid potential immunogenicity and this is a time-consuming and laborious step14. Humanized synthetic VHH libraries, which consist of humanized VHHs, facilitate the isolation of antigen-specific humanized VHHs because they do not require the humanization process. To date, several humanized VHH libraries have been developed15,16. They were designed using distinct strategies to constrain the diversity of amino acid sequences in the complementarity determining region (CDR), and successful isolation of functional VHHs for various targets was reported.
In this study, we designed several novel synthetic humanized VHH libraries based on biophysical analyses and comprehensive sequence analyses of VHHs deposited into the protein data bank (PDB). We obtained VHHs from the constructed libraries and analyzed their molecular characteristics. Based on the analyses, we discuss future strategies for effectively producing useful humanized VHHs with the desired epitopes.
Results
Sequence analysis of CDR loops
As the basis of humanized single domain antibody libraries, we hypothesized that VHHs in PDBs would be highly stable and biochemically well-behaved variants. Therefore, we obtained 600 VHH structures and sequences from the SAbDab database17,18. We then analyzed the CDR length of VHH sequences in the dataset to design the length of CDRs of synthetic libraries imitating the natural VHH (Fig. 1). The results clearly showed that almost all VHHs in the dataset had CDR1 containing 10 amino acid residues (Fig. 1A). About two-thirds of the VHHs had CDR2 of 17 amino acid residues, while the other one-third contained 16 amino acid residues (Fig. 1B). The length of CDR1 and CDR2 was highly conserved among the VHHs, but the length of CDR3 was longer and highly diverse, which is in agreement with results of a previous report19 (Fig. 1C). Although high diversity in the lengths of CDR3 was observed in the VHHs, lengths of 12, 15, and 18 residues were the most common.
Based on these results, we designed three independent libraries of VHHs with CDR3 length of 12, 15, and 18 each. To design the CDR sequences for each library, we calculated the position-specific frequency of amino acid residues in each CDR of VHHs with the same CDR3 length and designed a diversity of sequences in each CDR for each library. We used Weblogo-style views to show the sequence probability of each position (Fig. 2, Supplementary Figs. S1, S2).
CDR grafting to evaluate the humanized framework
To select a humanized framework for the libraries, we performed CDR grafting experiments. In CDR grafting, CDRs of the donor VHHs were grafted onto frameworks of different VHHs, and antigen binding affinity and thermal stability were evaluated for each generated synthetic VHH. We selected three anti-Epidermal growth factor (EGFR) VHHs (7D12, 9G8, EgA1)20,21 as donor VHHs and two clinically evaluated humanized VHHs, anti-IL6R VHH from vobarilizumab and anti-vWF VHH from caplacizumab22,23, as candidates for the humanized framework (Fig. 3).
We prepared the synthetic VHHs (wild type, WT) as well as parental VHHs (Vob, Capla) as recombinant proteins and assessed thermal stability using differential scanning calorimetry (DSC) (Fig. 4, Table 1). Comparison between 7D12 and its variants revealed a dramatic decrease in the melting temperature (Tm) of 7D12-Vob (52.2 °C) compared with 7D12 WT (63.6 °C) or 7D12-Capla (63.3 °C). Similarly, EgA1-Vob had a significantly lower Tm (68.5 °C) than EgA1 WT (77.8 °C) or EgA1-Capla (78.3 °C). In contrast, 9G8-Vob and 9G8-Capla had higher Tms (79.7 °C and 83.1 °C, respectively) compared with 9G8 WT (72.4 °C). Although these different tendencies were observed, in all cases the synthetic VHHs bearing the caplacizumab framework had a higher Tm than the synthetic VHHs bearing the vobarilizumab framework.
Summary of the biophysical analysis results for CDR grafting of VHHs. The values of (A) melting temperature (Tm) and (B) dissociation constant (KD) were plotted for parental and synthetic VHHs. Triangle symbols represent synthetic VHHs in which the CDR sequences were grafted on the vobarilizumab framework, and square symbols represent VHHs in which the CDR sequences were grafted on the caplacizumab framework.
Subsequently, we analyzed the affinity of each synthetic VHH to EGFR by measuring surface plasmon resonance (SPR) (Fig. 4, Table 2). Compared to 7D12 WT, 7D12-Vob showed an eightfold lower association rate constant and a 50-fold higher dissociation rate constant, resulting in a 350-fold worse dissociation constant. 7D12-Capla had a twofold lower association rate constant and 20-fold higher dissociation rate constant, resulting in a 30-fold worse dissociation constant compared with 7D12 WT. 9G8-Vob exhibited a 300-fold worse dissociation constant compared to 9G8-Capla and 9G8 WT, while 9G8-Capla and 9G8 WT had similar affinities. The affinity decrease for EgA1-Vob was only about 30–60-fold compared with EgA1 WT or EgA1-Capla.
Library design and construction of synthetic libraries
The results of CDR grafting analyses indicated that the framework from caplacizumab had high compatibility with long CDR3. Given that the affinity of all the CDR grafted synthetic anti-EGFR VHH whose CDR3 length were 15, 18, and 21 considerably diminished compared to original VHHs, we considered that CDR3 longer than 15 amino acids should be grafted on caplacizumab framework. Importantly, based on the crystal structures20,24, CDR3 in all three anti-EGFR VHHs and caplacizumab appear to form intramolecular interactions with the framework region. However, results of a previous study indicated that CDR3 with a length of 12 amino acids formed an extended conformation in which CDR3 and the framework did not interact with each other25. Given our previous study that revealed a vobarilizumab structure with an extended CDR3 loop, we hypothesized that the vobarilizumab framework would be compatible with VHHs containing a CDR3 of 12 amino acids, which does not require intramolecular interaction to maintain stable CDR conformation26. Taken together with the sequence analyses of the VHHs derived from SAbDab, we designed the following three synthetic libraries with the same length CDR1 (10 residues) and CDR2 (17 residues) but with variable length CDR3: (i) hNb18, which had an 18 amino acid long CDR3 loop with the caplacizumab framework; (ii) hNb15, which possessed a 15 amino acid long CDR3 loop with the caplacizumab framework and an L52F mutation according to the sequence analysis from the database; and (iii) hNb12, which had a 12 amino acid long CDR3 loop with the vobarilizumab framework. The design of each library is depicted in Figs. 5 and 6 and Supplementary Figs. S3, S4, S5, and S6. We severely restricted the diversity of the amino acids to make the actual antibody repertoire in the library close to the design as much as possible considering the maximum efficiency of bacterial transformation. The DNA encoding each designed library sequence was synthesized and inserted into a modified pLUCK phagemid vector27 and electroporated into TG-1 cells. The electroporated cells were incubated on TYE plates overnight, collected using 2 × TY medium, and stored as glycerol stocks. The resultant library sizes for hNb18, hNb15, and hNb12 were 2.4 × 109, 6.8 × 109, and 1.0 × 109, respectively.
Library screening and characterization of the obtained VHHs
To evaluate library quality, we isolated VHHs from the constructed libraries using a model antigen, EGFR. After five rounds of selection, we chose eight clones for further analyses based on phage enzyme-linked immunosorbent assay (ELISA) and convergence of the amino acid sequences (Table 3).
The DNA encoding selected VHHs was cloned into an expression vector, and the VHHs were prepared as recombinant proteins using an Escherichia coli expression system. We then measured the thermal stability of the VHHs using DSC (Fig. 7, Table 4). All of the VHHs exhibited high thermal stability, with Tms that reached about 60 °C or higher. Among them, 15EG-A7 had the highest thermal stability among the hit VHHs (Tm: 83 °C). These results illustrate the efficacy of our design strategy to construct the synthetic library based on sequences from PDBs to produce thermally stable VHHs. Intriguingly, one of the VHHs, 12EG-B5, had two melting peaks despite its single domain nature, suggesting that it might undergo multiple denaturation steps in which the partially denatured structure remains.
Subsequently, we analyzed the interactions with the designated antigen EGFR, using SPR. The binding (as measured by the KD) of hit clones to EGFR ranged from 80 to 600 nM (Fig. 8, Table 5). 15EG-A9 exhibited a clear binding response, but the kinetic parameters could not be determined due to too fast dissociation representing box-shaped response curve.
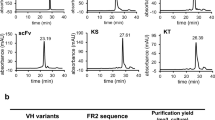
Results of surface plasmon resonance analysis for Hit clones from each library. Representative sensorgrams of the interaction analysis for recombinant hit VHHs with EGFR are summarized. Raw data and fitting curves are represented with blue and black colored lines, respectively. The sensorgram for EG-A9 could not be fitted.
Epitope binning of hit VHHs
To investigate the epitopes of each VHH, we performed dual injection assays28,29 using VHHs obtained from the libraries and EGFR, which enabled us to determine whether the epitopes overlapped with each other. In this assay, we injected one VHH into the immobilized EGFR on a CM5 sensor chip in a dose-dependent manner (0, 0.5, 2 µM) and measured the binding response (Binding 1) (Fig. 9A). Right after the first injection, we injected a mixture of the same VHH and EGF or another VHH (0.5 µM), and measured the binding response of the mixture (Binding 2) to investigate the overlap of epitopes among VHHs and also binding competitiveness with EGF. When the two proteins can bind to the EGFR simultaneously, Binding 2 should exhibit the same response regardless of the concentration of the first injected VHH, which we refer to as uncompetitive binding (Fig. 9B). When the two proteins competitively bind to the EGFR (i.e., competitive binding), Binding 2 should decrease as the concentration of VHH an increases (Fig. 9C).
Schematic images of dual injection analysis for dual injection. (A) An expected binding response curve in the dual injection assay. (B) Expected binding response curves when the two proteins can bind to the immobilized EGFR simultaneously. (C) Expected binding response curves when the two proteins cannot bind to the immobilized EGFR simultaneously. The binding response curves are aligned to zero at the report point just before the second injection.
Figure 10 summarize the results of combinations of VHHs and EGF that showed competitive binding as well as those of EG02 and EG-B5, which are representative of uncompetitive binding. These results indicate that the VHHs obtained from the same library tended to recognize the same epitope whereas VHHs from distinct libraries are likely to have different epitopes.
Results of dual injection assays for clones possessing competitive epitopes. The results of dual injection for the clone combinations showing competitive binding are summarized. The combination of EG02 and EG-B5 is shown as a model of uncompetitive binding. The combinations of VHHs or EGF that have overlapping binding sites are surrounded by colored circles.
Discussion
In this study, we constructed three novel synthetic humanized VHH libraries based on the comprehensive analysis of VHH sequences from PDB. The VHHs obtained from the libraries showed considerable thermal stability. In addition, our results suggested that VHHs from distinct libraries tends to have different epitopes. These results illustrate the value of our libraries for isolating humanized VHHs against various epitopes. Although we only screened the libraries against EGFR in this study, we expect that these libraries would also facilitate the generation of VHHs recognizing various epitopes on other antigens.
The obtained VHHs exhibited relatively higher thermal stability compared to previously reported synthetic libraries designed in a similar manner based on PDB sequences11, which indicates that the CDRs in our library design were well-tuned with scaffold sequences in the context of thermal stability. In contrast, the affinity of VHHs toward antigen was not comparable to that of typical VHHs obtained based on immune libraries. This may be because the CDR sequences were not optimized to acquire high affinity due to the constraint of bias toward molecular stability. It is challenging to exhaustively include all possible sequence variety due to the limitation of library size. Additional strategies such as a set of additional affinity-biased libraries or affinity maturation strategies to customize VHHs isolated from our library, further optimized selection conditions, and changing theoretical library size would yield VHHs with higher affinity toward antigens.
Researchers previously proposed CDR3 modeling based on VHH sequences30, structural classification of CDR3, and structure-sequence correlations25. Further library design considering CDR3 conformations such as extended or bending loops and based on structure-sequence correlations would generate libraries containing VHHs with different molecular characteristics. Indeed, Murakami et al. previously described the construction of a humanized VHH library that considered structure features of CDR loops15, and several other synthetic humanized single domain antibody libraries have been constructed16,31. Given that each library, including our libraries, has distinct constraints of CDR sequences, VHHs in each library may have different preferences for recognition epitopes. Therefore, detailed studies of molecular recognition for the VHHs from the libraries are needed to provide information about preferable epitopes of VHHs in each library. Such investigations will lead to strategies for choosing varied humanized libraries depending on the target epitope or its structural characteristics.
We designed three independent libraries that contained CDR3 of different lengths. However, some of the VHHs selected from the libraries had CDR3 lengths that differed from those that we designed. These clones would be derived from the error of DNA synthesis. Nevertheless, even if the length of the CDR3 differs from that of the library design, the selection should work to isolate hit clones.
In summary, we constructed three novel three synthetic humanized VHH libraries, which allowed the rapid and parallel selection of humanized VHHs against a variety of target antigens. Indeed, we have employed these libraries against 9 different proteins and have successfully isolated more than 2 unique binders for each target, which will be described in subsequent manuscripts. We believe that these libraries have the potential to produce useful humanized VHHs for both basic research and medical applications.
Methods
Data curation of VHH structures and sequences from the PDB
We extracted the structures that were annotated as VHHs in SabDab17,18. To remove duplicates and build a non-redundant dataset, we performed clustering of the VHH structures using Cluster Database at High Identity with Tolerance (CD-HIT)32,33 with a threshold of 100% sequence identity. Because some PDB files contained non-VHH structures, such as Fabs or engineered VHHs, we removed such antibodies and built the non-redundant datasets of single-domain antibodies (sdAbs). The antibody sequences in the datasets were annotated with the antigen receptor numbering and receptor classification (ANARCI) tool34 using the ImMunoGeneTics information system (IMGT) numbering scheme.
Library construction
The DNA fragments encoding humanized VHHs containing biased amino acid sequences that we designed were synthesized using the Geneart Combinatorial Library service (Thermo Fisher Scientific, Waltham, MA, USA). The synthetic DNA libraries were ligated using linearized pLUCK vector27 with an N-terminus pelB leader sequence and a C-terminal Myc-tag or Myc-His-tag sequence. The ligated products were purified and desalted using AMpure XP (Beckman Coulter, Brea, CA, USA). The purified phagemids were electroporated into E. coli TG1 electrocompetent cells (Lucigen, Middleton, WI, USA) at 1800 V, 200 Ω, and 25 µF for approximately 4–5 ms. Immediately thereafter, we added 980 µL of pre-warmed (37 °C) recovery medium supplied with the competent cells to each cuvette. After the cuvettes were incubated at 37 °C for 40 min at 200 rpm, the culture was plated on 16 large agar TYE dishes (100 µg mL–1 ampicillin, 1% glucose) and incubated overnight at 37 °C. We collected the colonies that emerged using a 2×TY medium and then added glycerol to make glycerol stocks containing 30% glycerol. The aliquots of the glycerol stock containing the libraries were stored at − 80 °C.
VHH selection from the constructed libraries
Antibody selection was conducted as previously described35. Briefly, E.coli cells were grown from an aliquot of the glycerol stock, and phage production was induced by infection with VCSM13 helper phage. After overnight incubation, phages were precipitated from the supernatant with polyethylene glycol/NaCl and resuspended in phosphate buffered saline (PBS)36. The VHH-phages were screened against recombinant EGFR immobilized on an immunotube (Thermo Fisher Scientific), and five rounds of selection were conducted. The phage ELISA was conducted using a 96-well microtiter plate. After selection, the single cloned VHH-phages were added to wells containing immobilized EGFR, followed by blocking. After the incubation, the wells were washed with PBS-Tween, and then anti-Myc tag antibody conjugated with horseradish peroxidase was added. Subsequently, we added tetra-methyl-benzidine (TMB), and we stopped the reaction with TMB stop buffer (Cosmo Bio, Tokyo, Japan). The signal for each well was measured by using Pheraster microplate reader (BMG Labtech, Ortenberg, Germany).
Expression and purification of recombinant proteins
Recombinant proteins were expressed and purified as previously described26. Briefly, the DNA sequences encoding each VHH were cloned into the expression vector pRA2, and E. coli BL21(DE3) cells (Merck, Rahway, NJ, USA) were transformed with the plasmids. Protein expression was induced by the addition of isopropyl β-d-1-thiogalactopyranoside, and the protein was extracted using sonication. The soluble fraction was separated by centrifugation and purified with immobilized metal chelate affinity chromatography (IMAC) followed by size exclusion chromatography (SEC). The DNA sequence encoding the extracellular domain of EGFR was cloned into pFastBac1 vector (Thermo Fisher Scientific) and expressed using the Bac-to-Bac baculovirus expression system (Thermo Fisher Scientific) according to the manufacture’s protocol. The recombinant EGFR was purified from the culture supernatant by IMAC followed by SEC. The protein concentration was determined by measuring absorbance at 280 nm using Nanodrop (Thermo Fisher Scientific) and calculating the value based on the computed extinction coefficient of each protein from its amino acid sequence.
DSC
The thermal stability of the VHHs was measured by DSC using a MicroCal PEAQ-DSC Automated system (Malvern Panalytical, Malvern, UK). The protein samples (1 mg mL–1 in PBS buffer) were heated from 20 to 110 °C at a scanning rate of 1.0 °C min–1. The data were analyzed using MicroCal PEAQ-DSC software.
SPR
The interactions of the VHHs with EGFR were analyzed by SPR using a Biacore 8K instrument (Cytiva, Marlborough, MA, USA). EGFR was immobilized on a Cytiva series S CM5 sensor chip according to the manufacturer’s standard amine coupling protocol at around 1000 RU. The VHHs were injected onto the immobilized EGFR in two-fold serial dilution with 120 s association time and 300 s dissociation time. The surface was regenerated with two 30 s injections of 10 mM Glycine–HCl pH 2.5. The measurements were carried out using PBS buffer supplemented with 0.005% Tween20. Data were analyzed using Biacore Insight Evaluation Software.
Dual injection assay
The recombinant EGFR was immobilized on a Cytiva CM5 sensor chip using the standard amine-coupling protocol at around 1200 RU. First, we injected one VHH onto the sensor chip for 120 s, and then we injected the mixture for 150 s. The chip surface was regenerated with two 30 s injections of 10 mM Glycine–HCl pH 2.5. The measurements were performed with PBS-Tween buffer at 25 °C. For the assay, a carrier-free recombinant human EGF (Catalog #: 236-EG, R&D Systems, Minneapolis, MN, USA) was reconstituted at 100 µg mL–1 in sterile PBS and then diluted to 0.5 µM with the SPR running buffer.
Data availability
All the structural data used in this study are available in Protein Data Bank (https://www.rcsb.org/) and all the data generated and/or analyzed during the current study are available from the corresponding author upon reasonable request.
References
Goulet, D. R. & Atkins, W. M. Considerations for the design of antibody-based therapeutics. J. Pharm. Sci. 109, 74–103 (2020).
Kaplon, H., Crescioli, S., Chenoweth, A., Visweswaraiah, J. & Reichert, J. M. Antibodies to watch in 2023. MAbs 15, 410 (2023).
Hamers-Casterman, C. et al. Naturally occurring antibodies devoid of light chains. Nature 363, 446–448 (1993).
Muyldermans, S. Nanobodies: Natural single-domain antibodies. Annu. Rev. Biochem. 82, 775–797 (2013).
Muyldermans, S. A guide to: Generation and design of nanobodies. FEBS J. 288, 2084–2102 (2021).
De Genst, E. et al. Molecular basis for the preferential cleft recognition by dromedary heavy-chain antibodies. Proc. Natl. Acad. Sci. U.S.A. 103, 4586–4591 (2006).
Zavrtanik, U., Lukan, J., Loris, R., Lah, J. & Hadži, S. Structural basis of epitope recognition by heavy-chain camelid antibodies. J. Mol. Biol. 430, 4369–4386 (2018).
Muyldermans, S. Applications of nanobodies. Annu. Rev. Anim. Biosci. 9, 401–421 (2021).
Jovčevska, I. & Muyldermans, S. The therapeutic potential of nanobodies. BioDrugs 34, 11–26 (2019).
Zimmermann, I. et al. Synthetic single domain antibodies for the conformational trapping of membrane proteins. Elife 7, 34317 (2018).
Sevy, A. M. et al. Structure- and sequence-based design of synthetic single-domain antibody libraries. Protein Eng. Des. Sel. 33, 1–13 (2020).
Jin, B. K., Odongo, S., Radwanska, M. & Magez, S. Nanobodies: A review of generation, diagnostics and therapeutics. Int. J. Mol. Sci. 24, 5994 (2023).
Tsumoto, K. & Takeuchi, T. Next-generation anti-TNFα agents: The example of ozoralizumab. BioDrugs 38, 341–351 (2024).
Vincke, C. et al. General strategy to humanize a camelid single-domain antibody and identification of a universal humanized nanobody scaffold. J. Biol. Chem. 284, 3273–3284 (2009).
Murakami, T. et al. Construction of a humanized artificial VHH library reproducing structural features of camelid VHHs for therapeutics. Antibodies 11, 10 (2022).
Moutel, S. et al. NaLi-H1: A universal synthetic library of humanized nanobodies providing highly functional antibodies and intrabodies. Elife 5, 16228 (2016).
Schneider, C., Raybould, M. I. J. & Deane, C. M. SAbDab in the age of biotherapeutics: Updates including SAbDab-nano, the nanobody structure tracker. Nucleic Acids Res. 50, D1368–D1372 (2022).
Dunbar, J. et al. SAbDab: The structural antibody database. Nucleic Acids Res. 42, D1140–D1146 (2014).
Mitchell, L. S. & Colwell, L. J. Comparative analysis of nanobody sequence and structure data. Proteins Struct. Funct. Bioinform. 86, 697–706 (2018).
Schmitz, K. R., Bagchi, A., Roovers, R. C., Van Bergen En Henegouwen, P. M. P. & Ferguson, K. M. Structural evaluation of EGFR inhibition mechanisms for nanobodies/VHH domains. Structure 21, 1214–1224 (2013).
Roovers, R. C. et al. Efficient inhibition of EGFR signalling and of tumour growth by antagonistic anti-EGFR Nanobodies. Cancer Immunol. Immunother. 56, 303–317 (2007).
Duggan, S. Caplacizumab: First global approval. Drugs 78, 1639–1642 (2018).
Kerschbaumer, A. et al. Efficacy of pharmacological treatment in rheumatoid arthritis: A systematic literature research informing the 2019 update of the EULAR recommendations for management of rheumatoid arthritis. Ann. Rheum. Dis. 79, 744–759 (2020).
Lee, H. T. et al. High-resolution structure of the vWF A1 domain in complex with caplacizumab, the first nanobody-based medicine for treating acquired TTP. Biochem. Biophys. Res. Commun. 567, 49–55 (2021).
Kuroda, D. & Tsumoto, K. Structural classification of CDR-H3 in single-domain VHH antibodies. Methods Mol. Biol. 2552, 61–79 (2023).
Kinoshita, S. et al. Molecular basis for thermal stability and affinity in a VHH: Contribution of the framework region and its influence in the conformation of the CDR3. Protein Sci. 31, e4450 (2022).
Maenaka, K. et al. A stable phage-display system using a phagemid vector: Phage display of hen egg-white lysozyme (HEL), Escherichia coli alkaline, phosphatase, and anti-HEL monoclonal antibody, HyHEL10. Biochem. Biophys. Res. Commun. 218, 682–687 (1996).
He, M., Bianchi, M. E., Coleman, T. R., Tracey, K. J. & Al-Abed, Y. Exploring the biological functional mechanism of the HMGB1/TLR4/MD-2 complex by surface plasmon resonance. Mol. Med. 24, 1–9 (2018).
Fleetwood, F. et al. Simultaneous targeting of two ligand-binding sites on VEGFR2 using biparatopic Affibody molecules results in dramatically improved affinity. Sci. Rep. 4, 1–14 (2014).
Sircar, A., Sanni, K. A., Shi, J. & Gray, J. J. Analysis and modeling of the variable region of camelid single-domain antibodies. J. Immunol. 186, 6357–6367 (2011).
Hairul Bahara, N. H., Chin, S. T., Choong, Y. S. & Lim, T. S. Construction of a semisynthetic human VH single-domain antibody library and selection of domain antibodies against α-crystalline of mycobacterium tuberculosis. J. Biomol. Screen. 21, 35–43 (2016).
Fu, L., Niu, B., Zhu, Z., Wu, S. & Li, W. CD-HIT: Accelerated for clustering the next-generation sequencing data. Bioinformatics 28, 3150–3152 (2012).
Li, W. & Godzik, A. Cd-hit: A fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22, 1658–1659 (2006).
Dunbar, J. & Deane, C. M. ANARCI: Antigen receptor numbering and receptor classification. Bioinformatics 32, 298–300 (2016).
Valenciano-Bellido, S. et al. Targeting hemoglobin receptors IsdH and IsdB of Staphylococcus aureus with a single VHH antibody inhibits bacterial growth. J. Biol. Chem. 299, 104927 (2023).
Lee, C. M. Y., Iorno, N., Sierro, F. & Christ, D. Selection of human antibody fragments by phage display. Nat. Protoc. 2, 3001–3008 (2007).
Acknowledgements
This work was funded in part by the Japan Society for the Promotion of Science (Grant Nos. 21H05090 to M.N.; 19H05766 and 20H02531 to K.T.), the JST-Mirai Program (Grant No. JP MJMI21G6 to M.N.), the JST-CREST Program (Grant No. JP MJCR20H8 to K.T.), and the Japan Agency for Medical Research and Development (Grant Nos. JP 18ak0101100 and 19am0401010to M.N., 20mk0101170 to K.T.).
Author information
Authors and Affiliations
Contributions
M.N. and K.T. conceptualized and administrated the research. S.K. conducted experiments and analyzed data. M.N. wrote the main manuscript and S.K. prepared figures. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
We declare that we filed a patent regarding the synthetic libraries described in this paper. The applicant is our institute, The University of Tokyo, and inventors are; Makoto Nakakido, Seisho Kinoshita, and Kouhei Tsumoto. Application number is JP2024-095995 and status of application is pending.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Nakakido, M., Kinoshita, S. & Tsumoto, K. Development of novel humanized VHH synthetic libraries based on physicochemical analyses. Sci Rep 14, 19533 (2024). https://doi.org/10.1038/s41598-024-70513-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-70513-4
- Springer Nature Limited