Abstract
Broad-spectrum biocatalysts enzymes, Laccases, have been implicated in the complete degradation of harmful pollutants into less-toxic compounds. In this study, two extracellularly produced Laccases were purified to homogeneity from two different Ascomycetes spp. Trichoderma lixii FLU1 (TlFLU1) and Talaromyces pinophilus FLU12 (TpFLU12). The purified enzymes are monomeric units, with a molecular mass of 44 kDa and 68.7 kDa for TlFLU1 and TpFLU12, respectively, on SDS-PAGE and zymogram. It reveals distinct properties beyond classic protein absorption at 270–280 nm, with TlFLU1's peak at 270 nm aligning with this typical range of type II Cu site (white Laccase), while TpFLU12's unique 600 nm peak signifies a type I Cu2+ site (blue Laccase), highlighting the diverse spectral fingerprints within the Laccase family. The Km and kcat values revealed that ABTS is the most suitable substrate as compared to 2,6-dimethoxyphenol, caffeic acid and guaiacol for both Laccases. The bioinformatics analysis revealed critical His, Ile, and Arg residues for copper binding at active sites, deviating from the traditional two His and a Cys motif in some Laccases. The predicted biological functions of the Laccases include oxidation–reduction, lignin metabolism, cellular metal ion homeostasis, phenylpropanoid catabolism, aromatic compound metabolism, cellulose metabolism, and biological adhesion. Additionally, investigation of degradation of polycyclic aromatic hydrocarbons (PAHs) by purified Laccases show significant reductions in residual concentrations of fluoranthene and anthracene after a 96-h incubation period. TlFLU1 Laccase achieved 39.0% and 44.9% transformation of fluoranthene and anthracene, respectively, while TpFLU12 Laccase achieved 47.2% and 50.0% transformation, respectively. The enzyme structure–function relationship study provided insights into the catalytic mechanism of these Laccases for possible biotechnological and industrial applications.
Similar content being viewed by others
Introduction
Fungi belonging to the phylum Ascomycota are well known for rapid growth, utilization of diverse substrate ranges, and degradation of the xenobiotics1. Among these fungi, the families Hypocreaceae and Trichocomacea are not only studied primarily for their role as biological control agents against numerous pathogens2,3, but also for their versatile enzymatic system including Laccases, Peroxidases and Oxidases which are crucial for degrading lignin and other recalcitrant aromatic compounds of environmental concerns4. Among these groups of versatile extracellular enzymes, Laccases have continued to receive significant attention in the past decade because of high redox potentials, its metal tolerance properties5,6 and industrial applications, such as polyaromatic hydrocarbons (PAHs) bioremediation, delignification or second-generation ethanol production, pulp bleaching, dye decolorization and detoxification of xenobiotics7,8,9,10.
Laccase (Benzenediol oxygen oxidoreductases, EC 1.10.3.2) belongs to the multi-copper class of enzymes with the potential of oxidizing aromatic and non-aromatic compounds to an innocuous state, unlike other ligninolytic enzymes such as Manganese peroxidase (MnP) and Lignin peroxidase (LiP), which require hydrogen peroxide for activity11. Laccases ubiquitously found in fungi, higher plants, some insects, and a few bacteria12. The production of these enzymes is always triggered in response to environmental stress and nutrient depletion7. The physiological function of Laccase varies depending on the organisms; in plants these enzymes are involved in the lignification processes13, in insects, it aids the formation of cuticles during the sclerotization process14, in fungi, it helps the morphogenesis process (formation of spores, pigments of fruiting bodies)15,16, pathogenesis17, virulence18, lignin degradation, monomer cross-linking, polymer degradation19, and also the breakdown of aromatic rings19. The precise role of Laccase in bacteria remains unclear but few studies have shown its functions in industrial applications requiring activity at high pH and temperatures as robust biocatalysts20. Fungal Laccases are glycoproteins which exist as monomer, homodimer, heterodimer, and multimer forms, with four redox-active copper atoms used in oxidizing substrates using oxygen as electron acceptor and its subsequent reduction to form water molecule21. The molecular mass and isoelectric point (pl) of Laccases ranges from 50 to 100 kDa and 3–7, respectively22. Typical Laccase consists of three Cu centres (Type I, II and III) with UV–vis spectrum absorption between 280 to 610 nm and a shouldering at 330 nm23. Type I Cu is responsible for the intense blue colour with UV–vis absorption spectrum at 605 nm and detectable in electro paramagnetic resonance (EPR) spectrum with narrow hyperfine coupling characteristics due to an S-Cu LMCT transition (ligand-to-metal charge transfer)24,25. Type II, unlike type I, is colourless and EPR detectable, while Type III Cu, on the other hand, has a pair of Cu atoms which have weak absorbance in the UV–vis spectrum and lacks an EPR spectrum24. Both. Type II and III Cu, forms the trinuclear cluster of Laccases where dioxygen binds and four-electron reduction to water takes place26. Different fungal species with blue Laccase production characteristics and PAHs degradation potential includes Trametes versicolor27, Trichoderma viride (EXF8977), Penicillium chrysogenum (EXF1818) and Irpex lacteus (MUM 04.98)28, while the use of a monomeric yellow Laccase in Leucoagaricus gongylophorus (Lac1Lg) has been documented in the oxidization of anthracene29.
Despite the wide distribution and importance of Laccases, detailed studies on different Laccases in ascomycetes and their potential application in transforming organo-pollutants like PAHs into an innocuous state are scanty. Additionally, systematic spectral studies and bioinformatic analyses of ascomycete Laccases remain limited. Hence, this study aims to fill these knowledge gaps and contribute to the development of more effective strategies for PAH bioremediation using ascomycete Laccases by purifying and characterizing from Trichoderma lixii FLU1 and Talaromyces pinophilus FLU12, two ascomycetes isolated from benzo(b)fluoranthene activated sludge and known for degrading anthracene and fluoranthene. Furthermore, the substrate binding mechanism, homology modelling and prediction of their three-dimensional structure were determined.
Materials and methods
Reagents
ABTS (2,2-azino-bis-(3-ethyl-benzothiazoline-6-sulphonic acid)), Basal Salt Medium (BSM), anthracene, benzo (b) fluoranthene (BbF), 2,6-dimethoxyphenol (DMP), (NH4)2SO4 DEAE-cellulose and Sephadex G-100 were purchased from Merck (Burlington, MA, USA). All other chemicals used were of the highest purity and analytical grade.
Fungal strains and molecular identification
The isolation and identification of the two fungal strains, Trichoderma lixii FLU1 (TlFLU1) and Talaromyces pinophilus FLU12 (TpFLU12), used in this study have been reported previously30. Briefly, the strains were isolated from benzo(b)fluoranthene-enriched activated sludge which was later serially diluted and plated on potato dextrose agar (PDA) medium containing 100 mg/L of benzo(b)fluoranthene. The plates were incubated at 30 °C for 7 days and observed for the appearance of fungal colonies. The colonies that showed clear zones were selected as potential benzo(b)fluoranthene-degrading fungi, purified by repeated subculturing on fresh PDA plates and maintained on PDA slants at 4 °C. Due to the hydrophobic nature of benzo(b)fluoranthene, a superficial concentration of 100 mg/L was achieved on a PDA agar plate by evenly spreading it across the surface using a sterile glass rod and then left in the laminar hood for 2 h to allow the solvent used to prepare benzo(b)fluoranthene to evaporate which ensures a strict adherence of benzo(b)fluoranthene to the agar surface.
Later, the purified fungal strains were identified by morphological and molecular methods. The morphological characteristics of the fungal strains were examined by light microscopy and compared with standard references. The molecular identification was performed by extracting the genomic DNA from the fungal strains and amplifying the internal transcribed spacer (ITS) region of the ribosomal RNA gene by PCR. The PCR products were sequenced and compared with the sequences available in the GenBank database. The phylogenetic analysis was conducted by using the MEGA software.
Culture conditions for Laccase production
Each fungal strain was separately cultured in cotton-plugged Erlenmeyer flasks (250 mL) containing BSM and anthracene (200 mg/L) as an inducer to a final volume of 150 mL prior to the set-up. The media were sterilized by autoclaving at 121 °C for 15 min. Inoculation was done directly into individual Erlenmeyer flasks using two 20 mm mycelial disks of each strain before incubating at 30℃ and shaking at 180 rpm for 10 days in complete darkness (MRC laboratory instrument, Essex, U.K). The pH of the flask containing strain TlFLU1 was adjusted to 4 using 1 M HCl while that of TpFLU12 was adjusted to 7 using 1 M NaOH. The details of the specific components and concentration for the BSM media and PDA are presented in Table S1. The variation in the culture media pH was observed due to optimized condition during anthracene degradation with 100% degradation efficiency after 12 days.
Purification of Laccase
The purification was carried out according to the previously described method (Othman et al., 2018) with some modifications. Briefly, crude extract from TlFLU1 and TpFLU12 was individually centrifuged at 5,000 × g for 20 min at 4℃ to obtain a clear supernatant. Protein from each supernatant was sequentially precipitated to saturation with 60% (NH4)2SO4 under a gentle continuous stirring at 4℃ overnight. The protein pellets were recovered by centrifugation at 10,000 × g for 20 min, dissolved in sodium acetate buffer (50 mM, pH 5) and dialyzed against a large volume of the same buffer using 30 kDa cut off size dialysis tubing cellulose membrane (Merck, Burlington, MA, USA). The protein was further purified using DEAE liquid chromatography column (1.5 × 9 cm) at room temperature before eluting with 100 mM NaCl in 50 mM sodium acetate buffer (pH 7) at a flow rate of 0.5 ml/min. Six fractions showing Laccase enzymes activity of substrate ABTS were pooled and applied on Sephadex G-100 column (2.0 × 9 cm) size exclusion column before elution with 50 mM sodium acetate buffer (pH 7). All fractions with Laccase activity were pooled, desalted, filter-sterilized, and stored at 4℃ until further usage.
Protein quantification and enzyme activity
The protein concentrations were determined as described previously31 using bovine serum albumin (BSA) as standard. Laccase activity was determined according to previously described method32 with slight modifications. Briefly, the reaction mixture contained 100 μL of 50 mM ABTS and 800 μL of 20 mM sodium acetate buffer (pH 5) and an appropriately diluted purified enzyme (370 μg). The mixture was incubated at 30℃ for 15 min and the reaction was terminated with 40 μL of 20% trichloroacetic acid. One unit of Laccase is the amount of enzyme that produces 1 µM of oxidized product per minute (ε420 nm = 36,000 M−1 cm−1 for oxidized product).
Zymogram analysis
Non-denaturing polyacrylamide gel electrophoreses (Zymogram) were used to confirm the protein purity and activity33. The sample loading buffer was prepared without the addition of β-mercaptoethanol (non-reducing condition), and the samples (100 µg for TlFLU1 and 75 µg for TpFLU12 of total protein) were not heated before loading into the wells. To detect the Laccase activity after the electrophoresis, the gel was washed once with a mixture of equal parts of isopropanol-sodium acetate buffer (50 mM, pH 5.0) for 30 min and once with sodium acetate buffer for 30 min, to remove the SDS, fix the proteins in the gel, and to reduce the pH to 5.0. The gel was then transferred onto a glass plate before layering with ABTS-agar (20 mg of ABTS, 400 mg of agar, 40 mL of water; heated to dissolve agar). These layers were incubated at 25 °C until the appearance of green bands.
UV–visible absorption spectrum and FTIR analysis
To study the copper active centre of purified enzymes, 1 mL of enzymes (370 µg of total protein) was added to a 10 mm optical path quartz cuvette and absorbance was measured in a scanning mode wavelength from 200–800 nm using Agilent Cary 60 UV–Vis spectrophotometer (Agilent, Santa Clara, CA, USA)34. To monitor functional groups of the purified proteins (370 µg of total protein), FTIR analysis was performed as described previously30.
Catalytic properties of the purified Laccases
Laccase activity was assayed by using several substrates. Assays were performed in 50 mM of the appropriate buffer and pH, and the oxidized products were read at the appropriate wavelength, as follows: 5 mM ABTS (ε420 = 36.00 cm-1 mM−1), 5 mM 2,6-DMP (ε469 = 49.60 cm−1 mM−1), 5 mM guaiacol (ε465 = 12.10 cm−1 mM−1) and 5 mM caffeic acid (ε312 = 11.20 cm−1 mM−1). The optimum pH for Laccase activity was investigated by using purified enzyme prepared in a reaction mixture containing the appropriate substrate in buffers as follows: 50 mM citrate buffer pH 1–6, 50 mM acetate buffer pH 3–5, 50 mM phosphate buffer pH 6–8 and 50 mM Tris buffer at pH 7 and 11. The optimum temperature of Laccase activity was investigated by using each purified enzyme (370 µg of total protein) for TlFLU1 and TpFLU12, respectively) prepared separately in a reaction mixture containing 5 mM ABTS in 50 mM acetate buffer pH 5, which was incubated at different temperatures (5, 10, 20, 30, 40, 50, 60, 70, 80, 90 and 100 °C) for 1 h. Also, the pH stability and thermal stability of the purified Laccase were studied by determining the residual activity for each enzyme (370 µg of total protein) after 24 h incubation under the same assay conditions for the optimum pH and temperature in darkness using ABTS as substrate. The reaction was monitored at 420 nm using Agilent Cary 60 UV–Vis spectrophotometer (Agilent, Santa Clara, CA, USA). The effect of appropriate substrate concentration (ABTS, 2,6-DMP, guaiacol and caffeic acid) on initial reaction velocity catalysed by purified Laccase was also investigated at constant enzyme concentration (370 µg of total protein). Therefore, as described above, the purified enzyme was assayed using varying concentrations (1–5 mM) of each substrate in 50 mM acetate buffer pH 5 and 7 at 30 and 50 °C for Laccase from TlFLU1 and TpFLU12, respectively. Michaelis–Menten reciprocal plots were used to determine the kinetic constants (vmax, Km, kcat and kcat/Km) for appropriate substrate using the Graph-Pad Prism 8.0 (San Diego, USA) software. The residual enzyme activity at varied concentrations of metal ions (10, 50 and 100 mM: Al3+, As5+, Ca2+, Cd2+, Co2+, Cu2+, Fe2+, K+, Li+, Mg2+, Mn2+, Mo+, Na+, Ni2+ and Zn2+), Organic solvent (10, 50 and 90%: 1,4 Dioxane, acetone, acetonitrile, butanol, DMSO, ethanol, ethyl-acetate, isopropanol, methanol and toluene) and inhibitors (DTT, 0.01 and 0.1 mM; EDTA, 1 and 10 mM; NaN3, 0.05 and 0.1 mM, SDS 5 and 10 mM; Urea, 20, 50 and 100 mM) was investigated by pre-incubating the purified enzymes (370 µg of total protein) for 24 h.
In-gel trypsin digestion and identification of the purified Laccase in ESMS
Purified Laccase (100 μg) from TlFLU1 and TpFLU12 was loaded onto 10% SDS-PAGE and stained with Coomassie blue R250. Protein bands were excised, digested with trypsin and subjected to electrospray mass spectrometry (ES-MS) for amino acid fragment identification at the Central Analytical Facility of Stellenbosch University, Stellenbosch, South Africa. The raw files generated by the mass spectrometer were imported into Proteome Discoverer v1.4 and processed using the Sequest algorithm against the UniProt (www.uniprot.org) Trichoderma genus and Talaromyces genus database, limited to Laccases. The peptide sequences were inferred from these matches with validation using the Target-Decoy PSM validator node. The search results were further validated using Scaffold Q + 35.
Template-based structure prediction, homology modelling and Laccase ligand binding site prediction for Laccase
The three-dimensional structure of the protein was predicted by submitting the amino acid sequence to the I-TASSER servers36. LOMETS approach was used to search for templates of similar folds from the PDB library37,38. Ten templates were used to build homology models while the top model structure was selected to optimize using the PyMOL Molecular Graphics System program (Schrödinger LLC., USA). The model quality validation through the 3D protein structure assessment, Errat and Ramachandran plot was performed using the Schrödinger Maestro software (Schrödinger LLC., NY, USA). Structural homology alignment was constructed with Jalview 2.11.2.439. Additionally, the Laccase ligand binding sites residues, their molecular function, biological process, and cellular components were predicted using I-TASSER servers in accordance with the ancestor chart of the EMBL’s European Bioinformatics institute gene ontology (GO) terms.
Sequence alignment
Detection of Laccase amino acid sequences from TlFLU1 and TpFLU12 similarities with amino acid sequences of Laccase from Trichoderma species and Talaromyces species deposited at UniProtKB were analysed using the BLAST tool (http://uniprot.org/blast). A phylogenetic approach was used to determine the taxonomic classification while amino acid sequence alignment was performed using muscle (multiple sequence comparison by UMPGA) as implanted in MEGA 11. Alignments were examined manually and a phylogenetic tree was constructed with a neighbour-joining likelihood approach and bootstrap resampling of 1000 replicates using MEGA 1140.
Degradation of PAHs by Laccase
The two PAHs (fluoranthene and anthracene) were selected based on their designated profile as known or reasonably anticipated human carcinogens on US EPA’s priority pollutants list41. The assays were carried out as described previously42 with some modifications. Briefly, 5 mL reaction mixture contained anthracene and fluoranthene (200 mg/L) individually in a 15 mL centrifuge tube, Laccase (2 U/ml) and sodium acetate buffer (50 mM, pH 5 and 7 for TlFLU1 and TpFLU12, respectively). Tween 80 (1%) was added to the mixture to enhance PAH bioavailability and shaken continuously at 80 rpm to avoid PAHs precipitation at 30 °C in darkness for 96 h (MRC Lab suspension mixer SM-3600, U.K). The set up was performed in triplicate while samples were drawn every 24 h for residual PAHs quantification. 50 μl of 20% (v/v) TCA solution was constantly added to each draw sample before quantifying residual PAHs to stop further enzymatic oxidation. A control set up was carried out under the same experimental condition except for the Laccase being heated denatured. However, extraction and quantification of the residual PAHs was carried out as previously30. The mixture was then vigorously shaken for 5 min and allowed to stand for 20 min to enhance the separation of aqueous and organic phases using the standard extraction method. The organic phase was collected, dried over 10 g anhydrous Na2SO4 and evaporated to dryness at 40 °C under reduced pressure. The dried fractions were then redissolved with the same extraction solvent and diluted tenfold with ethyl-acetate before quantifying the residual anthracene in a scanning spectrum mode using Agilent Cary 60 UV–Vis spectrophotometer from a wavelength of 200 to 400 nm. The absorption spectrum of anthracene and fluoranthene shows peaks at λmax 286 and 265 nm respectively, using a quartz cuvette with an optical path of 10 mm. However, the reaction medium's anthracene and fluoranthene residual concentrations were determined through extrapolation from a standard curve with an R2 value of 0.972 and 0.958 respectively. Also, a recovery study was carried out in triplicate with a sterile sodium acetate buffer with heated-denatured Laccase to an appropriate volume of known concentrations of each PAH compounds as standard, under static conditions and subjected to PAH determination with an average recovery of 98.7 ± 0.53% and 97.9 ± 7.17%, respectively, for anthracene and fluoranthene.
Statistics analysis
All data were collected in triplicates. Data were statistically analysed using the Graph-Pad Prism 8.0 (San Diego, USA) software and expressed as the means ± standard error. For determining Km and vmax, the Michaelis–Menten equation was fitted directly to the experimental data using the nonlinear least-squares fitting procedure of the Graph-Pad Prism 8.0 software.
Results
Purification of Laccases
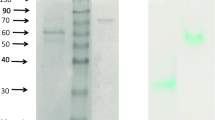
Table 1 shows the purification summary of Laccases from TlFLU1 and TpFLU12. Laccase from TlFLU1 was purified to 25.3-fold showing specific activity of 4003 U/mg of protein and a total yield of 7.0% while Laccase from TpFLU12 was purified to 5.6-fold showing specific activity of 790 U/mg of protein and a total yield of 5.5%. The purified Laccase from TlFLU1 showed a single band with molecular mass of 44 kDa (Fig. 1a, Fig. S1), while TpFLU-12 showed a molecular mass of 68.7 kDa (Fig. 1b, Fig. S2). The activity staining zymogram gel corresponding to the Laccase’s bands confirmed the nature of the enzymes (Fig. 1c, Fig. S3).
SDS-PAGE and native page showing single bands and in gel activity of Laccases. (A) Lane M: protein marker, Lane 1: purified Laccase from TlFLU1; (B) Lane M: protein marker, Lane 1–2 purified Laccase from TpFLU12; (C) Lane M: protein marker, Lane 1: In gel activity of purified Laccase from TlFLU1, Lane 2: In gel activity of purified Laccase from TpFLU12.
UV–visible and FTIR spectra of purified Laccases
The purified Laccases from TlFLU1 and TpFLU12 showed the absorption peak at wavelengths 270 nm and 600, respectively, measured by UV–visible spectrophotometer (Fig. 2). The FTIR spectra of Laccase from TlFLU1 showed the intense peaks at 3270 cm−1, 2148 cm−1, 1652 cm−1, 1066 cm−1 and 594 cm−1 attributing to –NH2 and –OH of amides A and B, OH/NH of Amide II, C=O stretching of α helix, C=C and C–N stretching of Amides II, respectively (Fig. 3, curve A, Table 2). The FTIR spectra of Laccase from TpFLU12 showed the peaks at 3260 cm−1, 2148 cm−1, 1646 cm−1 and 600 cm−1 attributing to –NH2 and –OH of amides A, B, OH/NH of Amide II, ß-sheets of peptide and C=C stretching and C-N stretching of Amides II, respectively (Fig. 3, curve B, Table 2).
Optimum pH, optimum temperature and stabillity
Purified Laccases from TpFLU1 and TpFLU12 showed optimum activity at pH 5 and pH 7, respectively (Fig. 4a). TpFLU1 Laccase showed 80–100% residual activity at pH 5–9 while TpFLU12 showed 60–100% residual activity at pH 1–5 after 24 h of incubation period prior to the assay (Fig. 4b). Furthermore, Laccases from TpFLU1 and TpFLU12 showed optimum activity at temperatures 30 °C and 50 °C, respectively (Fig. 4c). After 24 h of incubation at 10–30 °C, Laccases from TpFLU1 retained 100% activity and dropped to 90% at 50 °C, 70% at 60 °C, 50% at 70 °C and finally lost activity at 100 °C. TpFLU12 Laccase residual activity was found to be very much stable between a temperature 50–60 °C. The enzyme showed 80% residual activity at temperatures 30 °C and 80 °C while retained 85% residual activity at 70 °C (Fig. 4d).
Effect of metal ions, organic solvents and inhibitors on activity
Table 3 summarises the activity of Laccases in the presence of metal ions, organic solvents, and inhibitors. Laccase from TlFLU1 showed 100%, 121%, 100%, 100%, 102%, 98.0% and 107% relative activity in the presence of Ca2+, Cu2+, Fe2+, Mg2+, Mn2+, Na+ and Zn2+, respectively, at 10 mM concentration, however, the activity was decreased with increased metal ion concentrations. Laccase from TpFLU12 showed 104%, 100%, 101%, 119% and 100% relative activity in presence of Ca2+, Cd2+, Co2+, Cu2+and Na+ at concentration of 10 mM, while a relative activity of 100% was recorded in the presence of 50 mM Cu2+and Na+.
The relative activity of Laccase enzyme from TlFLU1 and TpFLU12 in the presence of organic solvents is shown in Table 4. Enzyme from TlFLU1 showed decreased relative activity with an increase in the organic solvents’ concentrations. 100% relative activity was recorded in the presence of acetone, acetonitrile, and ethyl-acetate while a relative activity of 127% was recorded in the presence of toluene at a concentration of 10%. The purified Laccase from TpFLU12 was more active in organic solvents up to 90% of concentration. Maximum relative activity of 100% was recorded in the presence of acetonitrile and ethyl-acetate at a concentration of 10%. Relative activity of 100% was recorded in the presence of acetone and ethyl-acetate at a 50 and 90% concentration, respectively.
The enzymatic acvity of Laccase from both the isolates, TlFLU1 and TpFLU12 were significantly inhibited by the tested inhibitors (Fig. 5). NaN3 significantly inhibited Laccases from both strains showing 81% and 97% inhibition at a concentration of 0.05 and 0.1 mM for enzyme from TlFLU1. In comparison, 97% and 100% inhibition were recorded at a concentration of 0.05 and 0.1 mM for enzyme from TpFLU12.
Kinetic parameters analysis
Table 5 shows the kinetics parameters values for Laccase from TlFLU1 and TpFLU12 when assayed with various substrates. Among all the phenolic compounds used as substrates, Laccase from TlFLU1 and TpFLU12 showed lowest Km values 10.8 µM and 8.20 µM, for substrate ABTs, followed by guaiacol with Km values of 12.7 µM and 11.6 µM while 13.03 µM and 14.91 µM for 2,6-DMP, respectively. Caffeic acid showed the lowest affinity as a substrate for both the enzymes from TlFLU1 and TpFLU12 with the Km values of 43 µM and 38.60 µM.
ES-MS analysis of tryptic digested proteins
ES-MS analysis of tryptic digested Laccase from TlFLU1 and TpFLU12 showed 12 and 18 protein clusters, respectively, when searched in UniProt Laccase database (Fig. S4a,b). The protein purified from TlFLU1 showed 100% similarity with 386 amino acids long Laccase from Trichoderma asperellum strain OX with Uniprot accession number A0A6V8QJQ4 while protein purified from TpFLU12 showed 100% similarity with 619 amino acids long Laccase from Talaromyces marneffei strain PM1 with Uniprot accession number A0A093UKS0 (Fig. S4c). Interestingly, no common proteins were detected between the Laccase from both strains at a 100% false discovery rate (FDR), and it quite raises concerns about relying soley on MS data sequence accuracy which might lead to unreliable models and inaccurate biophysical property predictions of the protein 3D structures. To address these limitations, a phylogenetic analysis using RSB PDB sequences of Laccases from both Trichoderma and Talaromyces families was conducted (Figs. S2 and S3). This approach provides a more robust understanding of the evolutionary relationships between the studied Laccases and their known counterparts, potentially offering a more reliable basis for modelling and prediction. Upon analysis, Laccase from TlFLU1 was observed to share the same cluster with that of Laccase from Trichoderma asperellum with Uniprot accession number A0A6V8QJQ4 (Fig. S5), while Laccase from TpFLU12 shared the same cluster with Laccase from Talaromyces marneffei PM1 with Uniprot accession number A0A093V1A3 (Fig. S6).
Homology sequence alignment
The homology sequence alignment of Laccases from TlFLU1 and TpFLU12 were constructed with PDB template IGW0 (Laccase from Melanocarpus albomyces) (Fig. S7) and 4X4KA (Laccase from Botrytis aclada) (Fig. S8), respectively, using LOMETS threading programs of the I-TASSER server. Laccases from TlFLU1 shared 13% sequence identity with the template IGW0 in the threading aligned region with the query sequence, 24% sequence identity of the whole template chains with the query sequence, 81% coverage of the threading alignment and 1.78 normalized Z-score of the threading alignments. Amino acids sequence WADNLINGYRLLNYWFNPDNPVNTLPNDPGVAVLNPRRWRHI (42 sequences) were observed as the consensus sequence between template IGW0. Laccase from TpFLU12 shared 16% sequence identity with the template 4X4KA in the threading aligned region with the query sequence, 18% sequence identity of the whole template chains with the query sequence, 73% coverage of the threading alignment and 5.85 normalized Z-score of the threading alignments. Amino acids sequence DTGTHWHGQCPQPTCINLGDGFLINGFIDHLTVIANDVVARYTPPGSDNPLDDSKNNDNLNPSEITG (67 sequences) were observed as the consensus sequence between template 4X4KA.
Template based structure prediction
Template-based structure modelling and ligand binding site residues of Laccases were predicted on iterative threading assembly refinement (I-TASSER) and shown in Fig. 6A–D and Table 6. The investigation revealed that the Laccase from TlFLU1 shares 13% sequence identity with the IGW0 template within the threading-aligned region and exhibits 24% sequence identity with the complete template chains. Furthermore, the threading alignment covers 81% of the sequence and the normalized Z-score amounts to 1.78 (Fig. 6A). Likewise, the Laccase derived from TpFLU12 shares 16% sequence identity with the 4X4KA template within the threading-aligned region and shows 18% sequence identity with the entire template chains. The threading alignment coverage encompasses 73% of the sequence, with a normalized Z-score of 5.85. Employing Jalview workbench alignment, 3-D models that conform to spatial restraints were generated, exhibiting high precision as evidenced by minimal restraint violations (Fig. 6B). Further, The I-TASSER server-generated predicted protein structures for TlFLU1 and TpFLU12, manifest a moderate degree of structural similarity based on the TM-score (0.48 ± 0.15 and 0.53 ± 0.15, respectively). However, these models exhibited moderate structural deviation, as indicated by the root mean square deviation (RMSD) values of 1.13 ± 0.5 Å and 1.15 ± 0.5 Å, respectively. Notably, the aligned sequences display a relatively low sequence identity (24% and 18%, respectively), implying a limited proportion of identical residues. Nevertheless, the alignment covered a substantial portion of the hit structure, as indicated by the coverage (Cov) values of 81% and 73%, respectively.
To evaluate the quality of the models, verified 3D visualization using polymol was employed. The assessment revealed that the protein model derived from TlFLU1 exhibited good 3D-1D scores for 43.95% of the total residues, while the model obtained for TpFLU12 demonstrated good scores for 90% of the total residues. Structural analysis of the TlFLU1 protein model unveiled the presence of four α-helices, with the absence of β-strands, β-α-β motifs, and β-hairpins. In contrast, the TpFLU12 model displayed 104 α-helices, 60 β-sheets, one β-α-β motif, and two β-hairpins. Also, the Ramachandran plots (not shown) of the final minimized model suggested that the protein residues from TlFLU1 was 9% most favourable, 12.11% additional allowed, 79% generously allowed and 0% disallowed, while that of TpFLU12 had 72.7% most favourable regions, 17.3% additional allowed regions, 0% generously allowed regions and 0% disallowed regions. When compared with the template and results obtained from COACH and COFACTOR, 21 catalytic residues such as Pro-177, Ala-178, Aap-179, Lys-180, Lys-182, Asp-183, Ser-258, Ala-259, Leu-260, His-261, Val-263, Arg-298, Thr-300, Leu-312, Asp-313, Ala-314, Pro-315, Ala-316, Asn-355, Trp-356 and Ile-358 were obtained for Laccase from TlFLU1 (Fig. 6C) while only Laccase from TpFLU12 showed three combinations of catalytic residues namely His-19, Ile-60 and Arg-62 with Cu2+ (T1) residues at the binding site (Fig. 6D). The 21 ligands binding sites residues of Laccase from TlFLU1 were shown to be associated with molecular functions related to copper ion binding, oxidoreductase activity and Laccase activity; biological process related to oxidation–reduction, lignin metabolism, phenylpropanoid catabolic, aromatic compound metabolism and cellulose metabolism; and cellular component related to extracellular structure; according to the gene ontology (GO). There were 3 binding residue sites (His-19, Ile-60 and Arg-62) of Laccase from TpFLU12 were predicted with molecular functions related to copper ion binding, oxidoreductase activity and oxygen oxidoreductase activity; biological process related to oxidation–reduction, biological adhesion, cellular metabolism, cellular metal ion homeostasis, aromatic compound metabolism; and cellular component related to extracellular structure (Table 6).
Biophysical properties
The amino acid sequences (386 and 619 amino acids for Laccases from TlFLU1 and TpFLU12, respectively) were submitted to ProtParam to predict their biophysical properties (Table 7). Molecular mass of 43.8 and 68.6 kDa, theoretical pI value of 9.33 and 5.41 were predicted for Laccases from TlFLU1 and TpFLU12, respectively. The extinction coefficient values of 68,995 M−1cm−1 and 85,050 M−1cm−1 assuming all pairs of Cys residues form cystines while 68,870 M−1cm−1 and 84,800 M−1cm−1 when assuming all Cys residues are reduced were exhibited by Laccase from TlFLU1 and TpFLU12, respectively. However, Laccase from TlFLU1 was predicted to have an estimated half-life of 4.4 h in mammalian reticulocytes (in vitro), > 20 h in yeast (in vivo) and > 10 h in E. coli (in vivo) while that of TpFLU12 Laccase was estimated to have a half-life of 1.1 h in mammalian reticulocytes (in vitro), > 3 min in yeast (in vivo) and > 10 h in E. coli (in vivo), confirming their stability in eukaryotic and prokaryotic cells. The predicted instability, aliphatic index, and grand average of hydropathicity (GRAVY) values of 39.54, 33.43; 76.58, 77.50 and -0.508, -0.578 for Laccases from TlFLU1 and TpFLU12, respectively, classify them as stable, thermophilic, and non-polar.
Degradation of PAHs by purified Laccase
A general reduction in the residual PAHs (Fluoranthene and anthracene) concentrations were observed after 96 h incubation period (Fig. 7A,B). For fluoranthene, a residual concentration of 61.0% of 200 mg/L initial concentration was recorded in TlFLU1 Laccase incubated tube while that of the TpFLU12 Laccase incubated tube accounted for 52.8% residual fluoranthene concentration in comparison with the control tube (no Laccase addition) where no reduction was recorded in its initial concentration (100%). Therefore, 39.0% and 47.2% transformation of fluoranthene was recorded by Laccases from TlFLU1 and TlFLU12, respectively (Fig. 7A). Also, during anthracene degradation, a residual concentration of 44.9% was recorded at 96 h incubation period in the TlFLU1 Laccase incubated tube while that of the TpFLU12 Laccase incubated tube showed a residual concentration of 50.0% at the same incubation period in comparison with the control tube (no Laccase addition) where no reduction was recorded in its initial concentration (100%). There was 55.1% and 50.0% transformation of fluoranthene was recorded by Laccases from TlFLU1 and TlFLU12, respectively (Fig. 7B).
Discussion
The capability of TlFLU1 and TpFLU12, investigated in this study, to produce active Laccases with a robust cold-adaptation and thermostability using anthracene as the sole carbon source corroborates previous study33, where the use of anthracene as a sole carbon source for growth and production of Laccase with high pH stability and temperature stability in an ascomycete fungus, Fusarium solani MAS2.
In this study, the obtained low purification fold with high protein-specific activities of purified Laccases from both strains following adoption of chromatographic techniques like ion-exchange (DEAE-cellulose) and gel filtration (Sephadex G-100) in chronological order is an indication that Laccases were the most abundant among the extracellular enzymes released into the media43. Similarly, the purification fold falls within the range of a previously reported purified Laccase from Ganoderma lucidum with a purification fold of 25.444. The absence of a peak near 600 nm in the UV–vis spectrum at 270 nm observed in Laccase from TlFLU1 suggests the presence of a type 2 copper centre with unique Laccase peptide interactions, which is characteristic of white Laccase enzymes45. Also, the absence of type 1 spectrum (blue Laccase) could be due to incomplete oxidation of copper ions which then results in the formation of one copper atom at the catalytic site in combination with metal ions like zinc, iron or manganese, which give it four active reaction sites46. Similarly, the presence of white/yellow Laccase in ascomycete fungi might may be due to the quenching of the type 1 Laccase (blue Laccase) through an exogenous bond of aromatic group harnessed from the carbon source used during production, which then acts as a redox mediator in its high redox potential25. In addition, the UV–vis spectrum absorption at 600 nm by Laccase from TpFLU12 is a characteristic of blue Laccase due to the type 1 spectrum peak24. Fourier-transform infrared spectroscopy (FTIR) analysis is an effective tool used in analysing proteins and polypeptides structures. In TpFLU-12 Laccase, the observed vibrational bands at wavenumber 1646 cm−1 attributing to β-sheets, absorption at 600 cm−1 (amides II) and amide A, B (bands > 3000 cm−1) is an indication of blue Laccase24 while the peaks at 594–1066 cm−1, 1652 cm−1 and > 3000 cm−1 in purified Laccase from TlFLU1 could be linked to α helix, amides II and amide A, B of white Laccase structures45. It is important to note that the peaks at 2142 cm−1 and 2148 cm−1 observed in both purified Laccases represent the presence of O–H stretching due to water interaction with the protein and its polypeptide structures.
Under denaturing and native conditions, the single band of both Laccases with a molecular mass of 44 and 68.7 kDa from TlFLU1 and TpFLU12, respectively, is consistent with other documented fungal Laccases (40–66 kDa) (Kumar et al. 2017). A Laccase with molecular mass of 45 kDa have been previously reported to be produced by Ceriporiopsis subvermispora (Jung et al. 2002). Another Laccases showing a molecular mass of ~ 67kD47 and with a molecular mass of 70 kDa from ascomycete fungus, Thielavia sp. HJ2248 are also reported previously, very close to Laccase purified from TpFLU12 (68.7 kDa) in this study.
In this study, the catalytic rate of Laccases and their stability were significantly influenced by pH and temperature which is of industrial advantage compared to previous reports. The optimum pH activity (5 and 7) and stability at a wide range of pH for both Laccases corroborate with the previous study49 where optimum pH activity of white Laccase was reported at pH 4 and stable from pH 2–7. Similarly, relative activities of two Laccase isoforms (Lacc1 and Lacc2) from Agaricus bisporus CU13 were optimum at pH of 5 and 7, respectively, while the stability was reported at pH 7 and pH 950. Here, optimum temperature of 30 °C and 50 °C were recorded for Laccase from TlFLU1 and TpFLU12, respectively, is corroborated with previous report51 where a temperature of 30 °C best-enhanced Laccase activity of a Laccase from white-rot fungus Trametes hirsute and maintained its 80% activity at a temperature between 20 to 40 °C. Likewise, an optimum temperature of 50 °C has been implicated in enhancing the maximum activity for purified Laccase from Lentinus tigrinus52. The temperature stability of purified Laccase from TlFLU1 (white Laccase), which is from 5 to 50 °C with over 90% residual activity after 24 h incubation period, is inconsonant with the previous report53, where thermo-stability reported between 50 and 70 °C for Laccase from Trichoderma harzianum strain HZN10. Notably, the observed difference in Laccase activity between TlFLU1 and TpFLU12 at 5 °C could be attributed to the inherent genetic and physiological differences between the two strains54,55,56. TlFLU1 might have a more robust cold-shock response mechanism that allows it to maintain higher Laccase activity at lower temperatures compared to TpFLU12. This aligns with a previous report, where Laccase from Thielavia terrastris Co3Bag1 retains 86% of its activity after 12 d storage at 4 ˚C due to cold-shock response57. Similarly, it has been reported that Laccases from fungi can retain activity at low temperature (4–7 °C)58. Conversely, the substantial loss of activity in TpFLU12 below 20 °C after 24 h of incubation could be since Laccase like many enzymes, has an optimal temperature range for activity59. At higher temperatures, fungi Laccases have been reported to possess characteristics of retaining more activity above 20 °C after 24 h57. Also, Psychrophilic Laccases, such as those produced by Kabatiella bupleuri G3 IBMiP which have been documented to have exhibit an optimal temperature range of 30–40 °C, with a significant decrease in activity below this ranges but still retains some activity even at 10 °C. This behaviour further attests to the substantial loss of activity in TpFLU12 below 20 °C after 24 h of incubation period.
The effect of different metal ion concentrations on Laccase stability is highly imperative in assessing the enzyme’s biotechnological applications. The significant low activity of Laccase from TlFLU1 in the presence of metal ions (Al3+ and K+) at a low concentration (10 mM) implies that these metal ions are activity inhibition stimulator which interferes with the electron transport system of Laccase activity. Similarly, a previous report linked Laccase inhibition with interaction of Fe2+ with the electron transport system of Laccase60. Laccase from Ganoderma lucidum showed complete activity inhibition in the presence of Fe2+ during the effect of metal ions on reactive dye decolourization61. 1 mM K+ reported to show 54% inhibition activity of Laccase from Lentinula edodes62. A detailed study on the inhibitory effect of metal ion concentration of 12.5 mM showed that Al3+ inhibit 55.9% and Mn2+ inhibit 63.5% Laccase activity on Laccase activity63. It is worth noting that the significant decrease in activity of Laccase from TlFLU1 at concentrations above 10 mM suggests that Laccase activity is a function of the metal ions concentration63. Conversely, the increase in activity of Laccase from TpFLU12 in the presence of high Ca2+, Cd2+, Co2+, Cu2+ and Na2+ concentrations suggest them as a potent Laccase activity activator due to the oxidation state (divalent cations) of the metal ions. It is opined that divalent cations of metal ions enhance Laccase activity and stability which aided in its binding to the carboxylic group of aspartic and glutamic acid residues for better complete substrate consumption and catalytic effect activation64. Also, the stability of Laccase activity to a high concentration of Cu2+ (100 mM) might be due to the type-2 copper-binding sites filled by the copper ions62.
The phenomenon of Laccase activity stability at a varying concentration of hydrophobic and hydrophilic organic solvents in this study could be ascribed to the stripping-off of the crucial bound-water monolayer from the enzyme molecule64. Also, the variability in the stability of purified Laccase from TpFLU12 observed at high organic solvents concentrations could be linked to each solvent's physical and chemical properties, the mixture pH, log P of the solvent and thermodynamic activity of water in mixed systems23,65. Unlike Laccase from TpFLU12, Laccase from TlFLU1 could not tolerate low water content systems containing organic solvents, which explains why high organic solvent concentrations inhibited its stability. A similar observation has been reported previously66, where Laccase stability was inhibited at high organic solvent concentrations after an hour incubation period which was credited to a low water content of the solvent.
The extent of the inhibitory effect of metalloenzymes and detergents varied greatly with their concentrations. The high inhibitory effect of NaN3 in both strains is due to internal electron transfer obstruction at the types 2 and 3 Laccase copper atom binding sites65. The high inhibition rate of the chelating agent (EDTA), denaturing agents DDT and SDS at varying concentrations could be justified by their interactions between the substrate and the Laccase active site leading to the unfolding of the compact protein structure during the catalytic enzyme activity48. Likewise, the high inhibition by NaN3 and EDTA is due to the chelating effect on the Cu atoms, which leads to inactivation of the Laccase12, while high inhibition by DTT suggests the strong inhibitory effect on the di-sulphide bonds present at the Laccase active domain33.
The obtained Km and Kcat values established the superior affinity of the purified Laccases towards substrate ABTS as compared to other phenolic substrates (2,6 DMP, caffeic acid and guaiacol). This observation agrees with the previous reports48, where strong affinity towards ABTS as substrate was documented during the characterization of Laccase from an ascomycetes fungi, Thielavia sp. In comparison with that of Laccase from TpFLU12 (blue Laccase), the observed high Km and vmax values for purified Laccase of TlFLU1 (white Laccase) in the presence ABTS, 2,6 DMP, caffeic acid and guaiacol confirm the characteristics of a yellow Laccase of fungi origin24,44.
The ES-MS generated unique Laccase peptides from Laccases from TlFLU1 and TpFLU12 and showed different numbers of protein clusters suggesting that these ascomycetes fungi produce two types of Laccases. The most likely explanation for this observation is that the number of Cys residues in the putative copper-binding site in the peptide sequence leads to the formation of two disulphide bridges which are known to be titratable in non-reducing conditions, a sole characteristic type-I copper (blue Laccase)67. Also, changes in the valence state of the Laccase Cu2+ residues and Fe2+ residues with a low spin electronic configuration account for the lack of blue colouration in white Laccase from TlFLU1 could be attributed to the differences in the numbers of protein clusters and unique Laccase peptides49. The variation in ascomycetes amino acids sequence number and structural differences in sensu stricto Laccases is linked to variation in the number of their encoding genes68.
Sequence homology analysis according to the LOMETS threading programs of the I-TASSER server revealed that variation in consensus sequence of Laccases from TlFLU1 and TpFLU12 which ranges between 42 and 67 suggests these sites as a major contributing factor to the protein stability due to non-covalent interactions69. Similarly, a previous report, demonstrated how 20 consensus mutation sequence cloned into Saccharomyces cerevisiae was linked to stabilizing Laccase with high-redox without compromising its catalytic activity70.
Prediction of binding site residues such as Pro-177, Ala-178, Aap-179, Lys-180, Lys-182, Asp-183, Ser-258, Ala-259, Leu-260, His-261, Val-263, Arg-298, Thr-300, Leu-312, Asp-313, Ala-314, Pro-315, Ala-316, Asn-355, Trp-356 and Ile-358, His-19, Ile-60 and Arg-62 in Laccases from both strains further strengthen the result of this study that the purified enzyme is Laccase with high catalytic activity since their high conservation across the amino acid sequences have been linked in the stabilization of the protein secondary structure leading to correct formation of the copper-binding site71. Similarly, it is opined that positively charged amino acids like Arg), His and Lys are responsible for the favourable conformation changes in Laccase leading to an increase in its catalytic activity69. It is previously demonstrated how binding residues such as Ala191, Pro192, Glu235, Leu363, Phe371, Trp373, Phe427, Leu429, Trp507 and His508 are crucial in the oxidation of phenolics during the structure–function studies of Melanocarpus albomyces Laccase72. Additionally, the Laccase gene ontology predicted molecular functions such as copper ion binding, oxidoreductase activity and Laccase activity, biological processes such as oxidation–reduction, lignin metabolism, phenylpropanoid catabolic, biological adhesion, cellular metabolism, cellular metal ion homeostasis, aromatic compound metabolism and cellulose metabolism and cellular component such as extracellular structure is not surprising because Laccases are known as oxidoreductases68, with Cu atoms in the active site, capable of lignin metabolism73,74, phenylpropanoid75, biological adhesion74, aromatic compound metabolism like fluorene76, cellular metal ion homeostasis77 and cellulose metabolism78.
The TM-score (0.48 ± 0.15 and 0.53 ± 0.15), and RMSD values (1.13 ± 0.5 Å and 1.15 ± 0.5 Å) observed in the model structure prediction for Laccases from TlFLU1 and TpFLU12, respectively, indicted a moderate degree of structural similarity with the PDB hit template used for model prediction since TM score close to 1 is classified as a perfect match with template structure, while a lower RMSD indicates a better structural alignment or match with template structure36,37. Despite covering a substantial portion of the hit structure, as indicated by the coverage (Cov) values of 81% and 73% for Laccases from TlFLU1 and TpFLU12, respectively, the aligned sequences displayed a relatively low sequence identity (24% and 18%). The most probable explanation for this observation can be linked to evolutionary divergences between the taxa of the template used and that of Laccases from TlFLU1 and TpFLU1279. Similarly, a previous study opined further that as species diverge evolutionarily, the sequence identity between their protein-coding regions may decrease due to changes in selective pressures and the fixation of different alleles80. Further, verify3D is a commonly used method for evaluating the quality of protein models. It compares the 3D environment of each residue in the model with high-resolution structures, assigning a 3D-1D score based on compatibility81. The predicted protein structure model obtained a high 3D-1D score in this study, indicating its excellent quality and similarity in overall shape and active site with the template structure82. The underlying assumption of verify3D is that a good 3D-1D score implies a similar structural arrangement to well-established protein structures83. Therefore, if the model's residues exhibit comparable structural characteristics and environments to those in known high-quality structures, it suggests that the model itself is likely of good quality.
Bioinformatics analysis by ProtParam further lends credence to this study finding that the two strains possess two different Laccase types. The predicted molecular weight of 44 and 68.6 kDa and theoretical pI values 5.22 and 6.32 agree with previous studies, where Laccase amino acids sequence of 566–600 aa, a molecular weight of 61.83–66.84 KDa and pl of pH 4–6 have been documented in Trichoderma Laccases68. While an amino acids sequence of 594 and with a molecular weight of 66.6 kDa close to this study has been reported in Talaromyces marneffei (strain ATCC 18,224/CBS 334.59/QM 7333). The predicted low instability indices (39.54 for TlFLU1 and 33.43 for TpFLU12) further gave an insight that the enzyme could be stable under harsh conditions. This prediction agrees with the previous report84 where low instability indices of 31.92 was linked to the Laccase from Kurthia huakuii stability under harsh conditions. Similarly, a report85 documented how low Laccase instability indices of 34.70 close to this study's prediction gave 40% of the Laccase relative activity at 70 °C after 60 min incubation period. In addition, the predicted aliphatic index values of the Laccase proteins ranged from 76.58 to 77.50 with a GRAVY index values mostly negative suggesting this protein to be thermophilic and hydrophilic nature. This prediction is in corroborated with previous reports86 where aliphatic index values of the Laccase proteins ranged from 74.44 to 90.4 with negative values of GRAVY index were attributed to the thermophilic and hydrophilic nature of Laccase during the comparative modelling and molecular docking analysis of white, brown and soft rot fungal Laccases using lignin model compounds for understanding the structural and functional properties of Laccases.
The findings of this study suggest that Laccase enzymes from TlFLU1 and TpFLU12 have the potential to degrade fluoranthene and anthracene, two commonly encountered PAHs of environmental concern. This observation is consistent with previous research highlighting the ability of fungi Laccase (50 U) purified from Leucoagaricus gongylophorus in degrading 10 ppm of anthracene (72 ± 1%), fluorene (40 ± 3%) and phenanthrene (25 ± 3%), respectively at pH 6 and 30 °C under 24 h incubation period28. Similarly, a previous study demonstrated the degradation of chrysene (75.8%) and Benzo[a]pyrene (35.6%), using a Laccase from Trametes versicolor29.
Conclusion
The study demonstrated the utilization of anthracene as carbon and energy sources by TlFLU1 and TpFLU12, resulted in the production of Laccase enzymes with broad catalytic potentials. The enzymes with molecular mass of 44 kDa and 68.7 kDa respectively, exhibit different optimum pH and temperature ranges with an implication potential application across various conditions. Despite the differences, both enzymes showed an enhanced activity under diverse metal ions and organic solvents. Structural predictions suggest that the differing in the copper atoms in the active sites contribute to their varied optimal pH. Additionally, the functional analysis indicates their involvement in key biological processes such as oxidation–reduction, lignin metabolism and aromatic compounds metabolism. This study provides valuable insights into the potential of Laccase-based systems for PAH removal and highlighting the potential of these enzymes as eco-friendly alternative for xenobiotic degradation.
Data availability
Data is provided within the manuscript or supplementary information files.
References
Mishra, S. et al. Recent advanced technologies for the characterization of xenobiotic-degrading microorganisms and microbial communities. Front. Bioeng. Biotechnol. 9, 1–26. https://doi.org/10.3389/fbioe.2021.632059 (2021).
Zafra, G. & Cortés-Espinosa, D. V. Biodegradation of polycyclic aromatic hydrocarbons by Trichoderma species: A mini review. Environ. Sci. Pollut. Res. 22, 19426–19433. https://doi.org/10.1007/s11356-015-5602-4 (2015).
Frisvad, J. C. Taxonomy, chemodiversity, and chemoconsistency of Aspergillus, Penicillium, and Talaromyces species. Front. Microbiol. 5, 1–7. https://doi.org/10.3389/fmicb.2014.00773 (2015).
Mate, D. M. & Alcalde, M. Laccase: A multi-purpose biocatalyst at the forefront of biotechnology. Microb. Biotechnol. 10, 1457–1467. https://doi.org/10.1111/1751-7915.12422 (2017).
Zhuo, R. & Fan, F. A comprehensive insight into the application of white rot fungi and their lignocellulolytic enzymes in the removal of organic pollutants. Sci. Total Environ. 778, 146132. https://doi.org/10.1016/j.scitotenv.2021.146132 (2021).
Chen, L., Zhang, X., Zhang, M., Zhu, Y. & Zhuo, R. Removal of heavy-metal pollutants by white rot fungi: Mechanisms, achievements, and perspectives. J. Clean. Prod. 354, 131681. https://doi.org/10.1016/j.jclepro.2022.131681 (2022).
Kumar, A. & Chandra, R. Ligninolytic enzymes and its mechanisms for degradation of lignocellulosic waste in environment. Heliyon 6, e03170. https://doi.org/10.1016/j.heliyon.2020.e03170 (2020).
Xu, P. et al. Degradation of several polycyclic aromatic hydrocarbons by Laccase in reverse micelle system. Sci. Total Environ. 708, 134970–134970. https://doi.org/10.1016/j.scitotenv.2019.134970 (2020).
Chen, S., Zhu, M., Guo, X., Yang, B. & Zhuo, R. Coupling of Fenton reaction and white rot fungi for the degradation of organic pollutants. Ecotoxicol. Environ. Saf. 254, 114697. https://doi.org/10.1016/j.ecoenv.2023.114697 (2023).
Gao, X. et al. Copper removal from aqueous solutions by white rot fungus Pleurotus ostreatus GEMB-PO1 and its potential in co-remediation of copper and organic pollutants. Bioresour. Technol. 395, 130337. https://doi.org/10.1016/j.biortech.2024.130337 (2024).
Falade, A., Jaouani, A., Mabinya, L., Okoh, A. & Nwodo, U. Exoproduction and molecular characterization of peroxidase from Ensifer adhaerens. Appl. Sci. 9, 1–15. https://doi.org/10.3390/app9153121 (2019).
Sondhi, S., Kaur, R. & Madan, J. Purification and characterization of a novel white highly thermo stable Laccase from a novel Bacillus sp. MSK-01 having potential to be used as anticancer agent. Int. J. Biol. Macromol. 170, 232–238. https://doi.org/10.1016/j.ijbiomac.2020.12.082 (2021).
Zhu, Y. et al. Laccase directed lignification is one of the major processes associated with the defense response against pythium ultimum infection in apple roots. Front. Plant Sci. 12, 1862–1862. https://doi.org/10.3389/FPLS.2021.629776/BIBTEX (2021).
Yang, C. H. et al. Involvement of Laccase2 in cuticle sclerotization of the whitefly, Bemisia tabaci middle East-Asia minor 1. Insects 13, 471. https://doi.org/10.3390/insects13050471 (2022).
Sakamoto, Y. Influences of environmental factors on fruiting body induction, development and maturation in mushroom-forming fungi. Fungal Biol. Rev. 32, 236–248. https://doi.org/10.1016/J.FBR.2018.02.003 (2018).
Yan, L., Xu, R., Bian, Y., Li, H. & Zhou, Y. Expression profile of Laccase gene family in white-rot basidiomycete Lentinula edodes under different environmental stresses. Genes 10, 1045–1045. https://doi.org/10.3390/GENES10121045 (2019).
Zhu, X. & Williamson, P. R. Role of Laccase in the biology and virulence of Cryptococcus neoformans. FEMS Yeast Res. 5, 1–10. https://doi.org/10.1016/J.FEMSYR.2004.04.004 (2004).
Williamson, P. R. Role of Laccase in the virulence of Talaromyces marneffei: A common link between AIDS-related fungal pathogens?. Virulence 7, 627–629. https://doi.org/10.1080/21505594.2016.1198867 (2016).
Santo, M., Weitsman, R. & Sivan, A. The role of the copper-binding enzyme–Laccase–in the biodegradation of polyethylene by the actinomycete Rhodococcus ruber. Int. Biodeteriorat. Biodegrad. 84, 204–210. https://doi.org/10.1016/j.ibiod.2012.03.001 (2013).
Chauhan, P. S., Goradia, B. & Saxena, A. Bacterial laccase: Recent update on production, properties and industrial applications. 3 Biotech 7, 323–323. https://doi.org/10.1007/s13205-017-0955-7 (2017).
Arregui, L. et al. Laccases: Structure, function, and potential application in water bioremediation. Microb. Cell Fact. https://doi.org/10.1186/s12934-019-1248-0 (2019).
Ali, W. B. et al. Enzyme properties of a laccase obtained from the transcriptome of the marine-derived fungus Stemphylium lucomagnoense. Int. J. Mol. Sci. 21, 1–16. https://doi.org/10.3390/ijms21218402 (2020).
Radveikienė, I., Vidžiūnaitė, R., Meškienė, R., Meškys, R. & Časaitė, V. Characterization of a yellow Laccase from Botrytis cinerea 241. J. Fungi 7, 1–16. https://doi.org/10.3390/jof7020143 (2021).
Agrawal, K. & Verma, P. Multicopper oxidase laccases with distinguished spectral properties: A new outlook. Heliyon 6, e03972–e03972. https://doi.org/10.1016/j.heliyon.2020.e03972 (2020).
Mot, A. C. et al. “Yellow” Laccase from Sclerotinia sclerotiorum is a blue laccase that enhances its substrate affinity by forming a reversible tyrosyl-product adduct. Plos One 15, e0225530. https://doi.org/10.1371/journal.pone.0225530 (2020).
Sekretaryova, A., Jones, S. M. & Solomon, E. I. O2 Reduction to water by high potential multicopper Oxidases: Contributions of the T1 copper site potential and the local environment of the trinuclear copper cluster. J. Am. Chem. Soc. 141, 11304–11314. https://doi.org/10.1021/jacs.9b05230 (2019).
Li, X. et al. Optimization of laccase-mediated benzo[a]pyrene oxidation and the bioremedial application in aged polycyclic aromatic hydrocarbons-contaminated soil. J. Health Sci. 56, 534–540. https://doi.org/10.1248/jhs.56.534 (2010).
Vipotnik, Z., Michelin, M. & Tavares, T. Development of a packed bed reactor for the removal of aromatic hydrocarbons from soil using Laccase/mediator feeding system. Microbiol. Res. 245, 126687–126687. https://doi.org/10.1016/j.micres.2020.126687 (2021).
Ike, P. T. L., Birolli, W. G., dos Santos, D. M., Porto, A. L. M. & Souza, D. H. F. Biodegradation of anthracene and different PAHs by a yellow laccase from Leucoagaricus gongylophorus. Environ. Sci. Pollut. Res. 26, 8675–8684. https://doi.org/10.1007/s11356-019-04197-z (2019).
Egbewale, S. O., Kumar, A., Mokoena, M. P. & Olaniran, A. O. Metabolic biodegradation pathway of fluoranthene by indigenous Trichoderma lixii and Talaromyces pinophilus spp. Catalysts 13, 791. https://doi.org/10.3390/catal13050791 (2023).
Lowry, O. H., Rosebrough, N. J., Farr, A. L. & Randall, R. J. Protein measurement with the Folin phenol reagent. J. Gen. Microbiol. 31, 3017–3027. https://doi.org/10.1016/S0021-9258(19)52451-6 (1951).
Zungu, N. S., Egbewale, S. O., Olaniran, A. O., Pérez-Fernández, M. & Magadlela, A. Soil nutrition, microbial composition and associated soil enzyme activities in KwaZulu-Natal grasslands and savannah ecosystems soils. Appl. Soil Ecol. 155, 103663–103663. https://doi.org/10.1016/j.apsoil.2020.103663 (2020).
Wu, Y. R., Luo, Z. H., Kwok-Kei Chow, R. & Vrijmoed, L. L. P. Purification and characterization of an extracellular Laccase from the anthracene-degrading fungus Fusarium solani MAS2. Bioresour. Technol. 101, 9772–9777. https://doi.org/10.1016/j.biortech.2010.07.091 (2010).
Ning, Y.-J. et al. An extracellular yellow Laccase with potent dye decolorizing ability from the fungus Leucoagaricus naucinus LAC-04. Int. J. Biol. Macromol. 93, 837–842. https://doi.org/10.1016/j.ijbiomac.2016.09.046 (2016).
Setlhare, B., Kumar, A., Mokoena, M. P., Pillay, B. & Olaniran, A. O. Phenol hydroxylase from Pseudomonas sp. KZNSA: Purification, characterization and prediction of three-dimensional structure. Int. J. Biol. Macromol. 146, 1000–1008. https://doi.org/10.1016/j.ijbiomac.2019.09.224 (2020).
Wei, Z. et al. Folding non-homology proteins by coupling deep-learning contact maps with I-TASSER assembly simulations. Cell Rep. Methods 1, 100014. https://doi.org/10.1016/j.crmeth.2021.100014 (2021).
Yang, J. & Zhang, Y. I-TASSER server: New development for protein structure and function predictions. Nucleic Acids Res. 43, W174–W181. https://doi.org/10.1093/nar/gkv342 (2015).
Zhang, C., Freddolino, P. L. & Zhang, Y. COFACTOR: Improved protein function prediction by combining structure, sequence and protein–protein interaction information. Nucleic Acids Res. 45, W291–W299. https://doi.org/10.1093/nar/gkx366 (2017).
Waterhouse, A. M., Procter, J. B., Martin, D. M. A., Clamp, M. & Barton, G. J. Jalview Version 2—A multiple sequence alignment editor and analysis workbench. Bioinformatics 25, 1189–1191. https://doi.org/10.1093/bioinformatics/btp033 (2009).
Tamura, K., Stecher, G. & Kumar, S. MEGA11: Molecular evolutionary genetics analysis version 11. Mol. Biol. Evol. 38, 3022–3027. https://doi.org/10.1093/molbev/msab120 (2021).
Zelinkova, Z. & Wenzl, T. The occurrence of 16 EPA PAHs in food—A review. Polycycl. Aromat. Compd. 35, 248–284. https://doi.org/10.1080/10406638.2014.918550 (2015).
Cambria, M. T., Minniti, Z., Librando, V. & Cambria, A. Degradation of polycyclic aromatic hydrocarbons by Rigidoporus lignosus and its laccase in the presence of redox mediators. Appl. Biochem. Biotechnol. 149, 1–8. https://doi.org/10.1007/s12010-007-8100-4 (2008).
Ezike, T. C., Ezugwu, A. L., Udeh, J. O., Eze, S. O. O. & Chilaka, F. C. Purification and characterisation of new laccase from Trametes polyzona WRF03. Biotechnol. Rep. 28, e00566. https://doi.org/10.1016/j.btre.2020.e00566 (2020).
Mukhopadhyay, M. & Banerjee, R. Purification and biochemical characterization of a newly produced yellow Laccase from Lentinus squarrosulus MR13. 3 Biotech 5, 227–236. https://doi.org/10.1007/s13205-014-0219-8 (2015).
Agrawal, K., Bhardwaj, N., Kumar, B., Chaturvedi, V. & Verma, P. Process optimization, purification and characterization of alkaline stable white laccase from Myrothecium verrucaria ITCC-8447 and its application in delignification of agroresidues. Int. J. Biol. Macromol. 125, 1042–1055. https://doi.org/10.1016/j.ijbiomac.2018.12.108 (2019).
Schückel, J., Matura, A. & van Pée, K. H. One-copper Laccase-related enzyme from Marasmius sp.: Purification, characterization and bleaching of textile dyes. Enz. Microb. Technol. 48, 278–284. https://doi.org/10.1016/j.enzmictec.2010.12.002 (2011).
Jeon, S. J. & Lim, S. J. Purification and characterization of the laccase involved in dye decolorization by the white-rot fungus Marasmius scorodonius. J. Microbiol. Biotechnol. 27, 1120–1127. https://doi.org/10.4014/jmb.1701.01004 (2017).
Mtibaà, R. et al. Purification and characterization of a fungal laccase from the ascomycete Thielavia sp. and its role in the decolorization of a recalcitrant dye. Int. Biodeteriorat. Biodegrad. 120, 1744–1751. https://doi.org/10.1016/j.ijbiomac.2018.09.175 (2018).
Zhao, D., Zhang, X., Cui, D. & Zhao, M. Characterisation of a novel white laccase from the deuteromycete fungus Myrothecium verrucaria NF-05 and its decolourisation of dyes. Plos One 7, e38817. https://doi.org/10.1371/journal.pone.0038817 (2012).
Othman, A. M., Elsayed, M. A., Elshafei, A. M. & Hassan, M. M. Purification and biochemical characterization of two isolated Laccase isoforms from Agaricus bisporus CU13 and their potency in dye decolorization. Int. J. Biol. Macromol. 113, 1142–1148. https://doi.org/10.1016/j.ijbiomac.2018.03.043 (2018).
Navada, K. K. & Kulal, A. Kinetic characterization of purified Laccase from Trametes hirsuta: A study on Laccase catalyzed biotransformation of 1,4-dioxane. Biotechnol. Lett. 43, 613–626. https://doi.org/10.1007/s10529-020-03038-1 (2021).
Sadeghian-Abadi, S., Rezaei, S., Yousefi-Mokri, M. & Faramarzi, M. A. Enhanced production, one-step affinity purification, and characterization of Laccase from solid-state culture of Lentinus tigrinus and delignification of pistachio shell by free and immobilized enzyme. J. Environ. Manag. 244, 235–246. https://doi.org/10.1016/j.jenvman.2019.05.058 (2019).
Bagewadi, Z. K., Mulla, S. I. & Ninnekar, H. Z. Purification and immobilization of laccase from Trichoderma harzianum strain HZN10 and its application in dye decolorization. J. Genet. Eng. Biotechnol. 15, 139–150. https://doi.org/10.1016/j.jgeb.2017.01.007 (2017).
Liu, X., Zain Ul Arifeen, M., Xue, Y. & Liu, C. Genome-wide characterization of laccase gene family in Schizophyllum commune 20R–7-F01, isolated from deep sediment 2 km below the seafloor. Front. Microbiol. 13, 923451. https://doi.org/10.3389/fmicb.2022.923451 (2022).
Xu, X., Zhang, Y., Liang, M., Kong, W. & Liu, J. The citrus Laccase gene CsLAC18 contributes to cold tolerance. Int. J. Mol. Sci. 23, 14509 (2022).
Sharma, V., Upadhyay, L. S. B. & Vasanth, D. Extracellular thermostable Laccase-like enzymes from Bacillus licheniformis strains: Production, purification and characterization. Appl. Biochem. Microbiol. 56, 420–432. https://doi.org/10.1134/S0003683820040146 (2020).
Gutiérrez-Antón, M. et al. Improvement of Laccase production by Thielavia terrestris Co3Bag1. Enhancing the bio-catalytic performance of the native thermophilic TtLacA via immobilization in copper alginate gel beads. J. Fungi 9, 308 (2023).
Chatterjee, R., Johansson, K., Järnström, L. & Jönsson, L. J. Evaluation of the potential of fungal and plant Laccases for active-packaging applications. J. Agric. Food Chem. 59, 5390–5395. https://doi.org/10.1021/jf103811g (2011).
Quynh, D. T., Hoang, N. H., Lan, N. N., Hoang, L. V. & Nghi, D. H. Cloning, experession, and characterization of a Laccase from the white rot fungi Pleurotus pulmonarius MPN18. VNU J. Sci. Nat. Sci. Technol. 39, 59–67. https://doi.org/10.25073/2588-1140/vnunst.5312 (2023).
Lin, Y. et al. Purification and characterization of a novel Laccase from Coprinus cinereus and decolorization of different chemically dyes. Mol. Biol. Rep. 40, 1487–1494. https://doi.org/10.1007/s11033-012-2191-x (2013).
Murugesan, K., Kim, Y. M., Jeon, J. R. & Chang, Y. S. Effect of metal ions on reactive dye decolorization by laccase from Ganoderma lucidum. J. Hazard. Mater. 168, 523–529. https://doi.org/10.1016/j.jhazmat.2009.02.075 (2009).
Nagai, M. et al. Purification and characterization of an extracellular laccase from the edible mushroom Lentinula edodes, and decolorization of chemically different dyes. Appl. Microbiol. Biotechnol. 60, 327–335. https://doi.org/10.1007/s00253-002-1109-2 (2002).
Zhang, G. Q., Wang, Y. F., Zhang, X. Q., Ng, T. B. & Wang, H. X. Purification and characterization of a novel Laccase from the edible mushroom Clitocybe maxima. Process Biochem. 45, 627–633. https://doi.org/10.1016/j.procbio.2009.12.010 (2010).
Rezaei, S., Shahverdi, A. R. & Faramarzi, M. A. Isolation, one-step affinity purification, and characterization of a polyextremotolerant Laccase from the halophilic bacterium Aquisalibacillus elongatus and its application in the delignification of sugar beet pulp. Bioresour. Technol. 230, 67–75. https://doi.org/10.1016/j.biortech.2017.01.036 (2017).
Sadhasivam, S., Savitha, S., Swaminathan, K. & Lin, F.-H. Production, purification and characterization of mid-redox potential Laccase from a newly isolated Trichoderma harzianum WL1. Process Biochem. 43, 736–742. https://doi.org/10.1016/j.procbio.2008.02.017 (2008).
Lueangjaroenkit, P. et al. Two Manganese peroxidases and a Laccase of Trametes polyzona KU-RNW027 with novel properties for dye and pharmaceutical product degradation in redox mediator-free system. Mycobiology 47, 217–229. https://doi.org/10.1080/12298093.2019.1589900 (2019).
Giardina, P. et al. Protein and gene structure of a blue laccase from Pleurotus ostreatus. Biochem. J. 341, 655–663. https://doi.org/10.1042/BJ3410655 (1999).
Cázares-García, S. V., Vázquez-Garcidueñas, M. S. & Vázquez-Marrufo, G. Structural and phylogenetic analysis of Laccases from Trichoderma: A bioinformatic approach. Plos One 8, e55295. https://doi.org/10.1371/journal.pone.0055295 (2013).
Nadar, S. S. & Rathod, V. K. Amino acid induced hyper activation of laccase and its application in dye degradation. Biocatal. Agric. Biotechnol. 18, 101064. https://doi.org/10.1016/j.bcab.2019.101064 (2019).
Gomez-Fernandez, B. J., Risso, V. A., Sanchez-Ruiz, J. M. & Alcalde, M. Consensus design of an evolved high-redox potential Laccase. Front. Bioeng. Biotechnol. 8, 354–354. https://doi.org/10.3389/FBIOE.2020.00354/BIBTEX (2020).
Mehra, R., Muschiol, J., Meyer, A. S. & Kepp, K. P. A structural-chemical explanation of fungal Laccase activity. Sci. Rep. 8, 1–16. https://doi.org/10.1038/s41598-018-35633-8 (2018).
Kallio, J. P. et al. Structure-function studies of a Melanocarpus albomyces Laccase suggest a pathway for oxidation of phenolic compounds. J. Mol. Biol. 392, 895–909. https://doi.org/10.1016/j.jmb.2009.06.053 (2009).
Fernández-Fueyo, E., Ruiz-Dueñas, F. J., MikiY, M. M. J., Hammel, K. E. & Martínez, A. T. Lignin-degrading peroxidases from genome of selective ligninolytic fungus Ceriporiopsis subvermispora. J. Biol. Chem. 287, 16309–16916. https://doi.org/10.1074/jbc.M112.356378 (2012).
Fathi-Roudsari, M., Behmanesh, M., Salmanian, A.-H., Sadeghizadeh, M. & Khajeh, K. Iranian biomedical functional surface display of laccase in a phenol-inducible bacterial circuit for bioremediation purposes. Iran Biomed. J. 22, 202–209. https://doi.org/10.22034/ibj.22.3.202 (2018).
Hu, Q. et al. Laccase GhLac1 modulates broad-spectrum biotic stress tolerance via manipulating phenylpropanoid pathway and jasmonic acid synthesis. Plant Physiol. 176, 1808–1823. https://doi.org/10.1104/pp.17.01628 (2018).
Bankole, P. O., Semple, K. T., Jeon, B. H. & Govindwar, S. P. Biodegradation of fluorene by the newly isolated marine-derived fungus, Mucor irregularis strain bpo1 using response surface methodology. Ecotoxicol. Environ. Saf. 208, 111619–111619. https://doi.org/10.1016/j.ecoenv.2020.111619 (2021).
Huang, W., Zhang, W., Gan, Y., Yang, J. & Zhang, S. Laccase immobilization with metal-organic frameworks: Current status, remaining challenges and future perspectives. Crit. Rev. Environ. Sci. Technol. 52, 1282–1324. https://doi.org/10.1080/10643389.2020.1854565 (2022).
Oliva-Taravilla, A., Tomás-Pejó, E., Demuez, M., González-Fernández, C. & Ballesteros, M. Inhibition of cellulose enzymatic hydrolysis by Laccase-derived compounds from phenols. Biotechnol. Prog. 31, 700–706. https://doi.org/10.1002/BTPR.2068 (2015).
Di Franco, A., Poujol, R., Baurain, D. & Philippe, H. Evaluating the usefulness of alignment filtering methods to reduce the impact of errors on evolutionary inferences. BMV Evol. Biol. 19, 21. https://doi.org/10.1186/s12862-019-1350-2 (2019).
Malhis, N., Jones, S. J. M. & Gsponer, J. Improved measures for evolutionary conservation that exploit taxonomy distances. Nat. Commun. 10, 1556. https://doi.org/10.1038/s41467-019-09583-2 (2019).
Singh, A. K., Katari, S. K., Umamaheswari, A. & Raj, A. In silico exploration of lignin peroxidase for unraveling the degradation mechanism employing lignin model compounds. RCS Adv. 11, 14632–14653. https://doi.org/10.1039/D0RA10840E (2021).
Vanajothi, R. et al. In silico and in vitro analysis of Nigella sativa bioactives against Chorismate synthase of Listeria monocytogenes: A target protein for biofilm inhibition. Appl. Biochem. Biotechnol. 195, 519–533. https://doi.org/10.1007/s12010-022-04157-3 (2023).
Nelapati, A. K., Meena, S., Singh, A. K., Bhakta, N. & PonnanEttiyappan, J. In silico structural and functional analysis of Bacillus uricases. Curr. Prot. 18, 124–142. https://doi.org/10.2174/1570164617999200512081127 (2021).
Guo, X. et al. Characterization of a highly thermostable and organic solvent-tolerant copper-containing polyphenol Oxidase with dye-decolorizing ability from Kurthia huakuii LAM0618T. Plos One 11, e0164810. https://doi.org/10.1371/JOURNAL.PONE.0164810 (2016).
Endo, K. et al. Enzymological characterization of EpoA, a laccase-like phenol oxidase produced by Streptomyces griseus. J. Biochem. 133, 671–677. https://doi.org/10.1093/jb/mvg086 (2003).
Kumar, A., Kameshwar, S., Barber, R. & Qin, W. Comparative modeling and molecular docking analysis of white, brown and soft rot fungal Laccases using lignin model compounds for understanding the structural and functional properties of Laccases. J. Mol. Graph Model 79, 15–26. https://doi.org/10.1016/j.jmgm.2017.10.019 (2018).
Acknowledgements
The authors appreciate Dr Mare Vlok of the central analytical facility, Stellenbosch, South Africa for the Liquid-Chromatography Mass-Spectrometry proteomics analysis of the protein.
Funding
National Research Foundation, South Africa (Grant No: 94036 and 92803).
Author information
Authors and Affiliations
Contributions
Conceptualization, S.O.E. and A.O.O.; methodology, S.O.E. and A.K.; vali-dation, S.O.E., A.K., and A.O.O.; formal analysis, S.O.E. and A.K.; investigation, S.O.E. and A.O.O.; resources, M.P.M. and A.O.O.; data curation, S.O.E. and A.K.; writing-S.O.E.; writing-A.K. and A.O.O.; visualization, S.O.E.; supervision, M.P.M. and A.O.O.; project administration, A.O.O.; funding acquisition, A.O.O. and M.P.M. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Egbewale, S.O., Kumar, A., Mokoena, M.P. et al. Purification, characterization and three-dimensional structure prediction of multicopper oxidase Laccases from Trichoderma lixii FLU1 and Talaromyces pinophilus FLU12. Sci Rep 14, 13371 (2024). https://doi.org/10.1038/s41598-024-63959-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-63959-z
- Springer Nature Limited