Abstract
Meiosis is a specialized type of cell division that occurs physiologically only in germ cells. We previously demonstrated that MYC-associated factor X (MAX) blocks the ectopic onset of meiosis in embryonic and germline stem cells in culture systems. Here, we investigated the Max gene’s role in mouse primordial germ cells. Although Max is generally ubiquitously expressed, we revealed that sexually undifferentiated male and female germ cells had abundant MAX protein because of their higher Max gene expression than somatic cells. Moreover, our data revealed that this high MAX protein level in female germ cells declined significantly around physiological meiotic onset. Max disruption in sexually undifferentiated germ cells led to ectopic and precocious expression of meiosis-related genes, including Meiosin, the gatekeeper of meiotic onset, in both male and female germ cells. However, Max-null male and female germ cells did not complete the entire meiotic process, but stalled during its early stages and were eventually eliminated by apoptosis. Additionally, our meta-analyses identified a regulatory region that supports the high Max expression in sexually undifferentiated male and female germ cells. These results indicate the strong connection between the Max gene and physiological onset of meiosis in vivo through dynamic alteration of its expression.
Similar content being viewed by others
Introduction
Germ cells are the only cell type that transmits genetic information to the next generation. Meiosis, a fundamental process of gametogenesis, is a specialized type of cell division that halves the ploidy of the genome1,2,3. While the molecular bases of most events that occur during meiosis are highly conserved among eukaryotes, the regulatory mechanisms that control its onset in mammalian germ cells are largely unknown because of their limited evolutionary conservation2,3,4,5,6. Dysregulation of meiotic genes in germ cells is closely linked to infertility and cancer6,7,8,9, underscoring the importance of elucidating the detailed molecular mechanisms that control meiotic onset in mammals.
During fetal development, primordial germ cells (PGCs) are generated from several epiblast cells located in the posterior/proximal corner at approximately embryonic day (E)6.25. They migrate to the genital ridge by around E10.5. Up to E11.5, germ cells remain sexually undifferentiated while testis- or ovary-specific gene expression can be observed in somatic cells10. Germ cells undergo sexual differentiation by E12.5, based on differentiation signals from the surrounding somatic cells. Female germ cells initiate meiosis around E13.5 with the expression of meiotic genes in response to retinoic acid (RA)11,12. In contrast, male germ cells do not initiate meiosis during the embryonic stage, mainly due to degradation of RA by CYP26B1 and translational repression of mRNAs from meiosis-related genes by NANOS2, one of the crucial male-specific factors that promote and suppress male- and female-specific pathways, respectively13,14. The Stra8 (stimulated by retinoic acid 8) gene is a crucial regulator of meiotic onset in male and female germ cells15,16,17,18. Although the molecular mechanisms by which STRA8 sustains the early stages of meiosis have long been enigmatic, identification of MEIOSIN as a binding partner of STRA8 revealed that STRA8 functions as a transcriptional activator that activates numerous meiosis-related genes by interacting with MEIOSIN, including the Stra8 and Meiosin genes themselves19. Meiosis-related genes may need to undergo certain epigenetic changes before being activated by the STRA8/MEIOSIN transcriptional complex so that the complex can perform its function effectively. However, any such epigenetic regulation remains elusive.
Although MYC-associated factor X (MAX) is widely known as an obligate partner of MYC, activating numerous genes involved in cell proliferation and metabolism20,21,22, it also heterodimerizes with MGA. This MGA-MAX dimer constitutes polycomb repressive complex (PRC)1.6, an atypical subtype of PRC123,24. Previously, we demonstrated that PRC1.6 suppresses meiotic onset in mouse embryonic stem cells (ESCs) and germline stem cells (GSCs), which are derived from preimplantation embryos and spermatogonial stem cells, respectively25. We previously showed that knockout of the Max gene in ESCs is accompanied by cytological changes reminiscent of germ cells undergoing meiotic prophase I, with global derepression of meiosis-related genes such as Sycp3 and Stra825,26,27,28,29,30,31,32,33,34,35. These cytological changes are independent of the expression of Blimp1, the gene encoding a master regulator of primordial germ cells36, but dependent on Stra8 expression, suggesting that the meiotic germ cell-like cytological changes observed in Max-null ESCs are not secondary consequences of ESC differentiation into germ cells, but stem from these cells’ intrinsic ability to initiate meiosis ectopically. Consequently, we hypothesized that MAX acts as a direct and crucial regulator for the initiation of meiosis. In this study, we addressed this hypothesis by investigating the role of the Max gene in mouse primordial germ cells.
Results
Meiotic onset is preceded by a prominent decline in Max expression in female PGCs
We previously demonstrated that MAX protein levels decrease notably prior to meiosis during spermatogenesis25, although Max is generally known to be expressed ubiquitously22,37,38. To further explore the correlation between Max expression and meiotic onset in germ cells in vivo, we examined MAX protein levels in male and female germ cells during the sexual differentiation process by immunohistochemistry. At E11.5, when PGCs were not sexually differentiated, OCT4-positive male and female germ cells had significantly stronger MAX protein staining signals than the surrounding OCT4-negative somatic cells (Fig. 1A). However, at E13.5, the MAX protein signal in female germ cells at the preleptotene stage—the transition stage shortly before entering meiotic prophase I39—became less intense than in E11.5 germ cells, and this less intense signal was prominently weakened further in the early leptotene stage (Fig. 1B). Quantification of the MAX protein levels during the mid-gestation stage revealed that the MAX protein signal was maintained mostly at a constant level in somatic cells in both sexes during the entire mid-gestation stage. However, female but not male germ cells showed a considerable decline in the MAX protein signal at E13.5, coinciding with the timing of their meiotic onset (Fig. 1C). Male germ cells also showed a significantly decreased MAX protein signal at E14.5, although the biological significance and cause of this reduction are unknown. Notably, while the lowest levels of the MAX protein staining signal in male germ cells at E14.5 were comparable to the levels in somatic cells, female germ cells at E13.5 had a lower level of the MAX staining signal than somatic cells. Analyses using publicly available transcriptome analysis data40 revealed that the most significant difference in Max expression levels between male and female PGCs was evident at E13.5 (Supplementary Fig. 1A), indicating that the difference in the Max mRNA levels between male and female germ cells during the mid-gestation stage is largely reflected by the MAX protein levels.
Spatiotemporal alterations of MAX protein levels in embryonic male and female germ cells. (A) Double immunofluorescence staining of MAX and OCT4 (POU5F1) in male and female gonads at E11.5. Arrowheads indicate each OCT4-positive germ cell. (B) Gonadal sections of female germ cells at E13.5 were immunostained for MAX and SYCP3, and counterstained with DAPI. PL, preleptotene; eL, early leptotene. (C) Plots of the intensity of individual MAX staining signals in male and female germ cells and somatic cells. The relative intensities of MAX staining in individual germ and somatic cells in male and female gonads at E11.5, E12.5, E13.5, and E14.5 were determined using ImageJ software. The mean MAX signal intensity in somatic cells was arbitrarily set to 100. (D) Scatter plots representing the intensities of the MAX and SYCP3 staining signals in individual male and female germ cells at E13.5 localized in the anterior (blue dot), middle (red dot), or posterior (green dot) of the gonads. The intensities of the MAX and SYCP3 staining signals in somatic cells were set to 100.
Meiosis occurs in female gonads along the anterior–posterior axis. It occurs restrictively in germ cells located at the anterior portion of the gonad at approximately E13.5, and then the region containing the germ cells undergoing meiosis expands and/or moves to the posterior region in a wave-like fashion11,12,41. Therefore, we used publicly available global gene expression data42 to examine whether expression levels of the Max gene, as well as pluripotency- and meiosis-related genes, were influenced by the location of the female germ cells in the gonad. Female germ cells located in the posterior part of the gonad at E13.5 had higher expression levels of Max and pluripotency genes (Oct4 and Nanog), but lower expression levels of meiosis-related genes (Dazl, Stra8, and Sycp3), than germ cells located in the anterior portion (Supplementary Fig. 1B). Next, we examined the MAX and SYCP3 protein levels in male and female germ cells by double immunohistochemistry in relation to the spatiotemporal regulation of meiotic entry. Female germ cells in the posterior portion of the gonad also showed delays in the reduction and increase in MAX and SYCP3 protein levels, respectively, compared with those in the anterior portion (Fig. 1D, upper panels), strengthening the notion that the decline in MAX protein levels and meiosis are closely linked events. However, this inverse correlation between them was no longer evident at E14.5. In contrast to female germ cells, male germ cells, which do not produce SYCP3 protein at appreciable levels during the embryonic stages, maintained a mostly constant level of MAX protein until E13.5, followed by significantly decreased levels between E13.5 and E14.5. However, unlike female germ cells, there were no spatiotemporal differences (Fig. 1D, lower panels). Immunocytochemical analyses of female gonadal cells dissociated to single cells revealed the lowest MAX protein levels in the early leptotene stage. However, these levels elevated during the subsequent leptotene and zygotene stages (Supplementary Fig. 2). These results indicate that the decline in MAX protein levels in female germ cells is a transient event that occurs around their physiological meiotic onset.
Max gene disruption is coupled to meiotic onset in sexually undifferentiated male and female PGCs
To further explore the relationship between Max expression and meiotic onset in germ cells, we conducted germ cell-specific knockout of the Max gene (Fig. 2A). To this end, we crossed Maxfl/fl female mice43 with Maxfl/fl male mice with the Oct4-CreERT2 transgene to produce MAX conditional-knockout (cKO) mice (Maxfl/fl;Oct4-CreERT2 female and male mice, abbreviated as FC and MC, respectively), as well as control female and male Maxfl/fl mice (abbreviated as F and M, respectively). Both FC and MC mice harbor a homozygous Max-floxed allele (Maxfl/fl mice), wherein exon V can be excised through Cre activity, and a hemizygous Oct4dPE-CreERT2 transgene expressed under the control of a distal enhancer of the Oct4 gene (Fig. 2A). To evaluate the efficiency of tamoxifen administration-mediated Max gene disruption, we injected tamoxifen into the abdomen at E8.5 and then performed immunostaining for MAX protein at appropriate embryonic stages. At E11.5, while Cre-negative control germ cells maintained a high MAX protein signal that was approximately twice as strong as that in somatic cells irrespective of tamoxifen treatment, most Cre-positive germ cells decreased the signal to a background level (Fig. 2B, see also germ cells indicated by a yellow arrowhead in Fig. 2D). We next examined alterations in the expression levels of meiosis-related genes (Sycp3, Meiosin, and Stra8) due to Max gene disruption in male and female germ cells by tamoxifen administration (Fig. 2C). Significant activation of the Sycp3 and Meiosin genes in male germ cells was evident between E11.5 and E14.5 after the induction of Max gene disruption by tamoxifen administration. This activation was most prominent at E11.5 and E12.5, and the extent of the activation became less significant at E13.5 and E14.5, with the difference of Meiosin expression at E14.5 not being statistically significant. Female germ cells also showed the activation of these genes following Max gene disruption. However, such activation was evident only before sexual differentiation (E11.5 and E12.5). Indeed, expression of these genes in female germ cells subjected to Cre-mediated Max gene disruption was comparable to the level in controls without CreERT2 cDNA at E13.5, and lower at E14.5 (Fig. 2C). As for Stra8 expression, our data revealed that the effect of Max gene disruption was distinct from that on Sycp3 and Meiosin expression in which Max expression ablation did not notably affect Stra8 expression levels in both male and female germ cells, except for at one particular time point, namely, E12.5, in female germ cells. We will discuss the regulation of Stra8 expression of germ cells in the Max-null background, including its activation in female germ cells at E12.5, in more detail below (see Discussion). To further confirm that Max gene disruption in sexually undifferentiated germ cells leads to de-repression of other meiosis-related genes, we also compared expression levels of Taf7l and Ddx4 between male and female Max-null germ cells at E11.5 and their respective control germ cells (Supplementary Fig. 3). These analyses revealed that, similar to Sycp3 and Meiosin genes, the expression levels of both Taf7l and Ddx4 were significantly elevated upon Max gene disruption, although the difference in Ddx4 expression level in female germ cells was not statistically significant between tamoxifen-administered germ cells and their untreated controls.
Max conditional knockout in sexually undifferentiated germ cells induces meiotic onset. (A) The mating scheme to generate Max conditional-knockout mice. Male and female Maxfl/fl embryos with CreERT2 cDNA were designated MC and FC, while male and female embryos without the cDNA were designated M and F, respectively. (B) Plots of the intensity of individual MAX staining signals in germ and somatic cells at E11.5 from tamoxifen-administered Maxfl/fl mouse embryos with and without the Cre transgene. MAX protein signal intensity was quantified using ImageJ software, as in Fig. 1C. (C) Examination of the effect of Max disruption on the expression of meiosis-related genes. Sycp3, Meiosin, and Stra8 mRNA levels in germ cells from M, MC, F, and FC embryos at the indicated days were quantified by quantitative PCR. Data were normalized to the β-actin expression levels. A short vertical line within the dots represents the mean. Student’s t-tests were conducted, *P < 0.05. (D) Double immunostaining of germ cells subjected to Max gene disruption and their control counterparts. Fetal female and male embryos with or without CreERT2 cDNA were treated with tamoxifen at E8.5 and then sectioned genital ridges from embryos at either E11.5 or E13.5 were subjected to immunostaining of MAX (magenta) and SYCP3 (green), and counterstained with DAPI (blue). Each germ cell is indicated by an arrowhead. Yellow arrowheads indicate germ cells that completely lost the MAX protein signal. PL, preleptotene; eL, early leptotene. (E) Scatter plot representing the intensity of the MAX and SYCP3 staining signals in individual male and female germ cells at E11.5 and E13.5. Proportions of germ cells with a lower MAX protein signal than somatic cells are shown. (F) Proportions of germ cells in meiotic prophase l in gonads from F (n = 3305 total cells) and FC (n = 4456 total cells) embryos at E14.5. Germ cells were classified into four stages (preleptotene, leptotene, zygotene, and pachytene) by the SYCP3 staining pattern. The differences in the proportions between F and FC mice at each stage were examined statistically by Student’s t-test, *P < 0.05.
Double immunohistochemical staining of MAX and SYCP3 confirmed the high level of MAX and little or no SYCP3 signal in the nuclei of sexually undifferentiated germ cells of control Maxfl/fl mice (F and M) at E11.5 (Fig. 2D,E). More importantly, a marked reduction in MAX signal by tamoxifen administration-mediated Max gene disruption in Maxfl/fl mice with the CreERT2 transgene was accompanied in most cases by the acquisition of a significant SYCP3 signal in male germ cells, and an even stronger SYPC3 signal in female germ cells, at E11.5 (Fig. 2D,E). Cytologically, these SYCP3-positive Max-null germ cells at E11.5 resembled germ cells at the preleptotene stage because of their clumped SYCP3 signal (Fig. 2D). However, this significant SYCP3 protein signal became hardly detectable in Max-ablated male PGCs at E13.5 (Fig. 2D,E). These results indicate that Max-ablated male germ cells did not maintain the meiosis-like cytological changes observed before sexual differentiation because of the reduction in the expression of meiosis-related genes during the meiotic process. Unlike male germ cells, female germ cells subjected to Max gene disruption retained the SYCP3 signal even at E13.5 (Fig. 2D,E). However, the immunostaining pattern of SYCP3 in germ cells at E14.5 revealed that germ cells at earlier (preleptotene and leptotene) and more advanced (zygotene and pachytene) stages were enriched and decreased, respectively, among germ cells subjected to Max gene disruption compared with the findings in control female germ cells that initiated meiosis physiologically (Fig. 2F). Taken together, these findings suggest that complete ablation of Max expression is sufficient to force the onset of meiosis in sexually undifferentiated male and female PGCs. However, such Max-null germ cells failed to undergo subsequent meiotic processes progressively and stalled immediately after the forced induction of meiotic onset. Next, we examined the effect of the ablation of Max expression on migrating PGCs. The SYCP3 signal intensity was extremely low in most MAX-null migrating germ cells at E10.5 (Supplementary Fig. S4A). However, some Max-ablated germ cells that had migrated to the vicinity of the genital ridge showed a clear SYCP3 protein signal (Supplementary Fig. 4A,B). These results indicate that the germ cells had acquired competence to respond to forced meiotic induction by Max ablation during their migration.
Next, we examined whether Max ablation in germ cells would affect the expression of genes related to sexual differentiation in the surrounding somatic cells as a secondary consequence of forced meiotic induction in germ cells. However, quantitative PCR analyses revealed that Sox9, which is crucially involved in male-specific characteristics44,45, did not have noticeably altered expression in either male or female gonadal cells (Supplementary Fig. 5A). Similarly, Foxl2, which plays indispensable roles in oocyte development and meiotic progression during the embryonic stage46,47,48, did not have appreciably altered expression (Supplementary Fig. 5B).
Global expression analyses of the effects of Max gene disruption on embryonic germ cells
To examine alterations in the gene expression profile caused by embryonic germ cell-specific Max gene disruption in an unbiased manner, we conducted DNA microarray analyses to compare global expression profiles between germ cells subjected to CRE recombinase-mediated Max gene disruption and their control counterparts lacking CreERT2 cDNA at E12.5 (Fig. 3A). The genes upregulated by Max gene disruption in female germ cells showed a statistically significant overlap with those in male germ cells, suggesting a shared role for MAX between male and female germ cells (Fig. 3B). Notably, a significant number of meiosis-related genes are included in all three gene sets; that is, genes commonly activated in male and female germ cells and those activated only in either male or female germ cells. Consistent with this, gene ontology (GO) analyses identified terms related to meiosis with all three gene sets, in which genes commonly activated in both sexes yielded those terms with higher statistical significance than in the two other gene sets (Fig. 3C). We also noted significant overlap between the genes upregulated by Max gene disruption and genes denoted as germline reprogramming-responsive (GRR) genes, whose transcriptional activation is closely linked to extensive epigenetic reprogramming in germ cells at E10.5–E11.5 via DNA demethylation of their high CpG promoters (Fig. 3D)49. This also supports the notion that control of the Max expression level is crucial for the initiation of meiosis because GRR genes play important roles in gamete generation and the initiation of meiosis50.
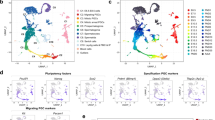
Alterations in the global expression profile of germ cells due to Max gene disruption. (A) Scatter plot of DNA microarray data from germ cells subjected to Cre-mediated Max gene knockout and their control counterparts without CreERT2 cDNA. The left and right panels show data from female and male germ cells at E12.5, respectively. Numerical values along the X- and Y-axes are the log2-transformed signal intensities of the microarray data. The upper and lower yellow diagonal lines in each panel indicate the boundaries of twofold up- and downregulation, respectively, due to Max gene disruption. Representative meiosis-related genes upregulated by Max disruption are individually marked by pink dots with gene symbols. (B) Venn diagram comparing genes activated more than twofold after Cre-mediated Max disruption in female and male germ cells at E12.5. The P-values for the significance of the overlap between two gene sets were calculated by hypergeometric tests. Meiosis-related genes included in each group are listed. (C) GO analyses of activated genes upregulated more than twofold after Cre-mediated Max disruption. Upregulated genes common to both male and female germ cells (middle panel) and those specific to either male (bottom panel) or female (upper panel) germ cells were subjected to GO analyses. Terms related to gametogenesis are indicated by red letters. (D) Venn diagram showing a significant overlap between genes activated by Max disruption in germ cells and genes designated as germline reprogramming-responsive genes50. P-values were calculated by hypergeometric tests, as in B.
Although MAX was originally cloned as an obligate heterodimerization partner of MYC proteins, which are predominantly associated with the transcriptional activation of genes involved in cell proliferation and metabolic processes20,21, most genes defined as cell-type-independent targets of the MYC/MAX transcription complex51 did not show significantly altered expression (Supplementary Fig. 6). These results indicate that the major role of MAX at these developmental stages is not the promotion of cell proliferation and cellular metabolism by interacting with MYC.
Germ cells with meiosis onset induced by Max ablation are eventually eliminated by apoptosis
Both male and female germ cells with forced meiosis onset induced by Max ablation stalled in the early stages of meiosis, and thus we determined the subsequent fate of these meiotically arrested germ cells. Although the total germ cell numbers in the genital ridges, assessed by the numbers of TRA98-positive cells, were unaffected by tamoxifen administration-mediated Max gene disruption at E12.5 in male and female embryos, a significant reduction in the germ cell number became evident in both sexes at E14.5 (Fig. 4A,B). To examine whether this reduction was due to apoptosis of Max-ablated germ cells, we performed immunostaining for cleaved PARP-1 as an indicator of the action of caspases52,53. In accordance with our hypothesis, there were significantly more cleaved-PARP-1-positive germ cells at E14.5 among the germ cells subjected to CRE recombination-mediated Max gene disruption than in controls without CreERT2 cDNA, for both males and females (Fig. 4C, Supplementary Fig. 7). Taken together, our data indicate that male and female germ cells that undergo artificial meiosis due to Max ablation are stalled at the early stages of meiosis, and are eventually eliminated by apoptotic cell death. Consistent with this idea, the skew of the distribution towards the earlier stages of meiosis at E14.5, observed in germ cells subjected to Max gene disruption (Fig. 2F), was no longer evident at E18.5 (Fig. 4D), indicating that the Max-null germ cells were mostly eliminated by apoptotic cell death before this stage, and that the vast majority of germ cells present at E18.5 are those that escaped CRE recombination-mediated Max gene disruption.
Apoptotic phenotype of primordial germ cells undergoing forced meiosis by Max ablation. (A, B) Total germ cell numbers assessed by TRA98 staining in gonads of M, MC, F, and FC mouse embryos at E12.5 (A) and E14.5 (B). (C) Percentages of cleaved-PARP-positive germ cells in gonads of M, MC, F, and FC mice at E14.5. (D) Proportions of germ cells in meiotic prophase l in E18.5 gonads of F and FC embryos. Germ cells were immunostained for SYCP3 after spreading onto a glass slide and their meiotic stages were classified as described in Fig. 2F.
Identification of the regulatory region that supports high Max expression in pluripotent embryonic stem cells and premeiotic germ cells
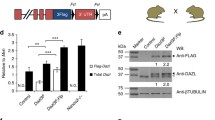
Several pluripotency marker genes that encode transcription factors, such as Oct4 and Nanog, are also highly expressed in premeiotic germ cells and ESCs, as well as Max. Therefore, we hypothesized that Max contains a regulatory element that has a common function in germ cells and ESCs of recruiting commonly expressed pluripotency factors. To test this hypothesis, we searched for genomic regions in the vicinity of Max that are bound by certain pluripotency factors in mouse ESCs using ChIP-Atlas, a database of ChIP-seq data for meta-analysis (https://chip-atlas.org). As a result, we identified a genomic region that met these criteria (Fig. 5A). This genomic region, which was named MUR (Max Upstream Region), was located approximately 3 kb upstream of the transcription start site of Max. Analyses using publicly available DNase-sequencing data54 confirmed that MUR has an open chromatin configuration in ESCs and germ cells, but not in somatic cells (Supplementary Fig. 8A). We also investigated whether MUR bears characteristic histone modifications that generally characterize enhancers in ESCs and germ cells. Since no publicly available histone modification data of germ cells in vivo were available, we inspected data of germ cells generated from ESCs in vitro55. We found that mono-methylated histone H3 lysine 4 (H3K4me1) was more or less present around the region of MUR at all stages during the conversion of ESCs to primordial germ cell-like cells (PGCLCs). Meanwhile, levels of acetylated histone H3 lysine 27 (H3K27ac) dynamically changed during the conversion, in which rather high levels of H3K27ac in ESCs declined substantially in epiblast-like cells (EpiLCs) and such low H3K27ac levels persisted in d2PGCLCs, which are known to be most similar to in vivo germ cells at E7.0. However, the level of this modification was remarkably elevated in d4c7PGCLCs, which are transcriptionally similar to germ cells at E11.5. Since active and poised enhancers are generally known to be H3K4me1/H3K27ac-double-positive and H3K4me1-positive, but H3K27ac-negative, respectively56, these findings further support the hypothesis that MUR is involved in the transcriptional activation of the Max gene in ESCs and premeiotic germ cells as an active enhancer. To directly evaluate the transcription-stimulating activity of MUR in ESCs, we conducted luciferase assays using a luciferase reporter carrying MUR. The reporter with the MUR sequence induced two- or threefold higher luciferase activity than the reporter without MUR in ESCs, whereas no statistically significant difference in luciferase activity with MUR was evident in mouse embryonic fibroblasts (Fig. 5B). Next, we generated mice lacking MUR in their genome (dMUR) to assess the function of MUR in supporting Max expression in premeiotic germ cells. Although this deletion did not affect the germ cell number (Supplementary Fig. 8C), immunohistochemical analyses of MAX revealed that, compared with wild-type male and female germ cells, sexually undifferentiated germ cells lacking MUR showed a significantly lower MAX protein signal, comparable to that of surrounding somatic cells (Fig. 5C). This indicates that MUR contributes to the high Max expression in ESCs and premeiotic germ cells, which was supported by the quantification of Max mRNA (Fig. 5D). However, unlike complete ablation of Max expression, the decline in Max expression following MUR deletion was not accompanied by the induction of meiotic gene expression (Fig. 5E). These findings indicate that the decline in Max expression level to a level comparable to that of somatic cells is not sufficient to induce the onset of meiosis in germ cells, and thus these levels of Max gene expression need to decrease below those of somatic cells for the initiation of meiosis.
Identification of the positive regulatory region of the Max gene common to ESCs and premeiotic germ cells. (A) Analysis of ChIP-Atlas metadata identified a genomic region (MUR) that recruits representative pluripotent cell-specific transcription factors upstream of the Max gene. (B) Examination of the transcription-stimulating activity of MUR. Two luciferase reporters with or without MUR were individually introduced into mouse ESCs and embryonic fibroblasts by transfection, together with an internal control plasmid carrying the Renilla reniformis luciferase gene. A dual-Glo luciferase assay was conducted 48 h post-transfection. Data represent the mean ± standard deviation of three independent experiments. Student’s t-tests were conducted. P-values for the differences in luciferase activity due to the presence of MUR were 0.011 (marked by an asterisk) and 0.217 for ESCs and mouse embryonic fibroblasts, respectively. (C) Immunohistochemical analyses of MAX protein in germ cells of dMUR mutant mice. Upper panel illustrates the genetic manipulation of dMUR mice. Middle panel shows representative images of immunostained MAX and OCT4 in sections of genital ridges from wild-type and dMUR mice. Arrowheads indicate each OCT4-positive germ cell. The lower panel shows a plot of MAX protein signal intensity quantified by ImageJ software, as in Fig. 1C. The mean of the MAX signal intensity in somatic cells was arbitrarily set to 100. (D) Effect of MUR deletion on Max mRNA levels in germ cells. Max mRNA levels in germ and somatic cells of wild-type and dMUR mutant mice were quantified by quantitative PCR using Rn18s RNA level as an internal control. Blue and red dots indicate data from female and male samples, respectively. The asterisk indicates P < 0.05. (E) Effect of MUR deletion on meiosis-related gene expression in germ cells. Expression levels of Dazl, Ddx4, and Sycp3 in germ cells of wild-type and dMUR mutant mice at E11.5 were quantified as in D. Expression levels of Max and meiosis-related genes in wild-type germ cells were arbitrarily set to 1. No statistically significant differences were evident in the expression levels between wild-type and dMUR mutant mice, except for in Max expression (P = 2.87 × 10−6; marked by an asterisk).
Discussion
In this study, we demonstrated that conditional Max gene knockout enforces meiosis in both sexually undifferentiated female and male PGCs. However, these Max-null germ cells failed to progressively undergo subsequent processes. Indeed, Max ablation in female germ cells did not coordinate with the physiological meiotic onset at approximately E13.5 to further drive meiotic progression, but instead meiosis stalled, mainly around the leptotene-to-zygotene transition. Most Max-null male germ cells were positive for SYCP3 before sexual differentiation, but the high percentage of SYCP3-positive cells among these cells subsequently declined substantially, and SYCP3-positive cells were hardly detectable at E13.5. Our data also showed that male and female germ cells subjected to forced meiotic induction were eventually eliminated by apoptosis.
Although our data demonstrated that Max gene disruption was accompanied with de-repression of many meiosis-related genes such as Meiosin and Sycp3 in sexually undifferentiated germ cells, that effect on the Stra8 gene was completely different from that on other meiosis-related genes, indicating that, like in ESCs35, Stra8 is not a direct target of PRC1.6. Our previous study also demonstrated that Meiosin expression in ESCs in a background of impaired PRC1.6 function sensitizes the Stra8 gene to activation by RA35. Given that germ cells bear the same regulatory mechanisms as ESCs, it may be that removal of the PRC1.6-mediated repression of the Meiosin gene renders germ cells highly sensitive to Stra8 activation by RA, which leads to the induction of meiotic onset. Therefore, we consider that the significant Stra8 gene activation observed in Max-null female germ cells at E12.5 represents a secondary consequence of Meiosin gene activation, making germ cells susceptible to RA secreted from the mesonephros. Although the Meiosin gene was also activated in Max-null male germ cells, this was not accompanied by the activation of Stra8, probably because of the extensive degradation of RA by CYP26B1 produced in the Sertoli and testicular interstitial cells18.
This study also importantly demonstrates the dynamic changes of Max expression in germ cells, especially because Max expression levels are generally comparable across numerous cell types and are not strongly influenced by environmental changes38,57. Indeed, our data reveal that male and female germ cells maintain approximately twofold higher expression levels of Max than the surrounding somatic cells in a mitotically active state, but Max expression declines prominently at or immediately prior to meiotic onset in female germ cells. Because this decline is assumed to be linked to attenuation of the transcription-repressing activity of PRC1.6, it may be an important prerequisite for germ cells to undergo meiosis. It is not presently known how female germ cells reduce Max expression to a level lower than that in somatic cells. However, it seems likely that MUR—herein identified as a regulatory region that sustains high Max expression in both ESCs and mitotically active germ cells—substantially contributes to the subsequent downregulation of Max expression in female germ cells because the expression of genes encoding OCT4 and other pluripotency factors, which probably confer transcription-enhancing activity on MUR, shows a profound decline in female germ cells during this period. Our data also reveal that Max regains high expression around the zygotene stage in female germ cells, which may lead to the functional recovery of MAX-dependent activities, namely, MYC and PRC1.6 activities. In conjunction with the finding that Max-null male and female germ cells did not progressively undergo meiotic processes but were stalled, it seems likely that the recovery of MYC and/or PRC1.6 from their inactivated state via upregulation of Max expression is also required for germ cells to complete meiosis. Therefore, as the next step towards obtaining a complete understanding of the events occurring during the early stage of meiosis in germ cells at the molecular level, we will conduct future studies to elucidate how Max regains high expression after its low-expression state during meiotic processes, and how upregulation of Max contributes to the progression of meiosis.
Methods
Ethical statement
Animal experiments were carried out in strict accordance with international and institutional guidelines. The protocol was approved by the Institutional Review Board on the Ethics of Animal Experiments of Saitama Medical University (Permission numbers: 3136, 3137, and 3138). The study is reported in accordance with ARRIVE guidelines (https://arriveguidelines.org).
Mice
Oct4dPE-CreERT2 transgenic mice were purchased from RIKEN BRC (BRC No. RBRC06509)58,59. Mice with a homozygous Max-floxed allele (Maxfl/fl mice)43 were bred with Oct4dPE-CreERT2 transgenic mice, and the resulting compound mice were bred with Maxfl/fl mice to obtain Maxfl/fl;Oct4dPE-CreERT2+ mice. For tamoxifen administration, tamoxifen (Sigma-Aldrich, MO) was first dissolved in corn oil at a concentration of 10 mg/ml, and 0.5 mg (50 µl of the prepared solution) was injected intraperitoneally into pregnant mice at E8.5 days.
To generate mice lacking MUR, we used the guide RNAs 5′-CCCACTGAAACATCGCCCCATGG-3′ and 5′-TCTTCACAGGCTAAGATCTCAGG-3′. Unfertilized oocytes were collected from female C57BL/6 J mice (> 10 weeks old) superovulated with gonadotropin and chorionic gonadotropin and subjected to in vitro fertilization with sperm from C57BL/6 J mice. Five hours post-treatment, the resultant zygotes were subjected to guide RNA (5 ng/µl) introduction and GeneArt Platinum Cas9 Nuclease (100 ng/µl) (Thermo Fisher Scientific, MA) by electroporation using a NEPA 21 electroplater (NEPA GENE, Chiba, Japan), as described by Sato et al.60. Two-cell-stage embryos developed by subsequent in vitro culture were transferred into oviducts of pseudopregnant ICR female mice for development and delivery. One of the originally obtained heterozygous dMUR mutant mice was used as a founder to maintain the colony by crossing with wild-type C57BL/6 J mice. Heterozygous mutant mice were backcrossed three times before being used to generate homozygous mutant mice by heterozygous intercrosses. PCR using the following nucleotides was used to distinguish between wild-type and deletion mice:
Wild-type allele with the MUR sequence: forward, 5′-TTTTGGCTGGACTGGCTAAC-3′; reverse, 5ʹ-TTTTGGCTGGACTGGCTAAC-3ʹ.
Mutant allele without the MUR sequence: forward, 5ʹ-TTTTGGCTGGACTGGCTAAC-3ʹ; reverse, 5ʹ-TTTTGGCTGGACTGGCTAAC-3ʹ.
Plasmid construction and luciferase assays
The 5ʹ flanking region of Max was amplified by PCR from genomic DNA of wild-type and mutant mice lacking MUR, using the following oligonucleotides: 5ʹ-GGTACCCCAGGAAGGATAGGCTGTGACG-3ʹ and 5ʹ-GTCGACGTAGTCCTCGAGCGTCGGAT-3ʹ. These PCR products were individually subcloned into the pGL4.10 vector carrying luciferase cDNA (Promega, WI). Plasmid transfection and subsequent luciferase assays were conducted as described previously61.
Isolation of germ cells by MACS
To isolate germ cells by magnetic-activated cell sorting (MACS), gonads at E10.5–14.5 were dissected from embryos and dissociated to single cells using 0.05% trypsin–EDTA in PBS after removal of the mesonephros. After treatment with MACS buffer (2 mM EDTA and 0.5% BSA in PBS), germ cells were recovered using an anti-SSEA-1 antibody conjugated to MicroBeads (Miltenyi Biotec, 130-094-530), in accordance with the manufacturer’s instructions.
Immunostaining
For immunohistochemical analyses, genital ridges isolated from embryos (E10.5–14.5) were fixed in 4% paraformaldehyde, embedded in OCT compound, and then frozen. Sections prepared from the frozen tissues were mounted on glass slides and subjected to immunostaining as described previously25. The primary antibodies used were as follows: anti-GENA (TRA98; BioAcademia, 73–003), anti-MAX (Santa Cruz Biotechnology, sc-197), anti-OCT4 (Santa Cruz Biotechnology, sc5279), and anti-cPARP (BD Pharmingen, 558,710). The anti-GENA antibody was used at 1:400 dilution and the other antibodies were used at 1:100 dilution. Immunostained tissues/cells were observed under a confocal laser scanning microscope (TCS SP8; Leica Microsystems, Wetzlar, Germany). The average staining intensities of MAX and SYCP3 in somatic cells were arbitrarily set to 100.
Immunofluorescence staining for stage classification of germ cells in meiotic prophase
Preparation of germ cell nuclear spreads was conducted in accordance with the method described by Hwang et al.62 with some modifications. Briefly, after washing with PBS, gonads recovered from embryos were immersed in a low-tension buffer (17 mM trisodium citrate dihydrate citrate, 50 mM sucrose, 30 mM Tris–HCl pH 8.2, 1 mM EDTA, 0.5 mM DTT, and 0.1 mM PMSF) for 7 min. Each pair of gonads was suspended in a drop of 100 mM sucrose on a glass slide and gently dispersed by pipetting. The solution containing gonadal cells was then dropped onto slides soaked in a fixative solution (1% PFA and 0.15% Triton X-100) and pipetted to spread the cells. After repeated cell spreading, the glass slide was incubated overnight at room temperature in a humidified box and then dried at room temperature. The cells were then used in immunocytochemical analyses using antibodies against SYCP3 and MAX.
Quantitative reverse-transcription PCR analysis
An RNeasy Micro Kit (QIAGEN) was used for total RNA preparation from germ cells isolated using MACS. The recovered RNAs were reverse-transcribed to cDNA using an FSQ-301 kit (Toyobo). TaqMan-based quantitative PCR was performed using the StepOnePlus Real-Time PCR System (Applied Biosystems, CA). All samples were normalized to the expression level of Actb (β-actin) or Rn18s (18S ribosomal RNA). The TaqMan probes used were as follows: Max, Mm00484802_g1; Sycp3, Mm00488519_m1; Meiosin, Mm01305445_m1; Stra8, Mm00486473_m1; Dazl, Mm03053726_s1; Ddx4, Mm00802445_m1; Foxl2, Mm00843544_s1; Sox9, Mm02619580_g1; Actb, Mm02619580_g1; and Rn18s, Mm03928990_g1.
DNA microarray and GO analyses
Biotin-labeled cDNA was synthesized and then hybridized to Affymetrix GeneChip Mouse Genome 430 2.0 arrays (Affymetrix, CA), in accordance with the manufacturer’s instructions. Microarray expression data were background subtracted and normalized by the robust multiarray analysis method using R statistical software. Gene ontology (GO) analyses were conducted using DAVID web tools (http://david.abcc.ncifcrf.gov). The selected GO terms were subjected to further analyses using the AmiGO1 (http://amigo1.geneontology.org/cgi-bin/amigo/go.cgi) and REVIGO (http://revigo.irb.hr) websites to eliminate synonymous terms. The significance of the overlap between gene sets was assessed using the hypergeometric test.
Data availability
Data from the DNA microarray analyses have been deposited in the NCBI Gene Expression Omnibus database under accession number GSE233556. Other data that support the findings of this study are available from the corresponding author upon request.
References
Handel, M. A. & Schimenti, J. C. Genetics of mammalian meiosis: Regulation, dynamics and impact on fertility. Nat. Rev. Genet. 11, 124–136 (2010).
Kimble, J. Molecular regulation of the mitosis/meiosis decision in multicellular organisms. Cold Spring Harb. Perspect. Biol. 3, a002683 (2011).
Spiller, C. & Bowles, J. Instructing mouse germ cells to adopt a female fate. Sex. Dev., 1–13 (2022).
Marston, A. L. & Amon, A. Meiosis: Cell-cycle controls shuffle and deal. Nat. Rev. Mol. Cell Biol. 5, 983–997 (2004).
Lesch, B. J. & Page, D. C. Genetics of germ cell development. Nat. Rev. Genet. 13, 781–794 (2012).
Farini, D. & De Felici, M. The beginning of meiosis in mammalian female germ cells: A never-ending story of intrinsic and extrinsic factors. Int. J. Mol. Sci. 23, 12571 (2022).
Feichtinger, J. & McFarlane, R. J. Meiotic gene activation in somatic and germ cell tumours. Andrology 7, 415–427 (2019).
Nicholls, P. K. & Page, D. C. Germ cell determination and the developmental origin of germ cell tumors. Development 148, 198 (2021).
Lingg, L., Rottenberg, S. & Francica, P. Meiotic genes and DNA double strand break repair in cancer. Front. Genet. 13, 831620 (2022).
Saga, Y. How germ cells determine their own sexual fate in mice. Sex. Dev. 16, 329–341 (2022).
Menke, D. B., Koubova, J. & Page, D. C. Sexual differentiation of germ cells in XX mouse gonads occurs in an anterior-to-posterior wave. Dev. Biol. 262, 303–312 (2003).
Bullejos, M. & Koopman, P. Germ cells enter meiosis in a rostro-caudal wave during development of the mouse ovary. Mol. Reprod. Dev. 68, 422–428 (2004).
Suzuki, A. & Saga, Y. Nanos2 suppresses meiosis and promotes male germ cell differentiation. Genes Dev. 22, 430–435 (2008).
Kato, Y., Katsuki, T., Kokubo, H., Masuda, A. & Saga, Y. Dazl is a target RNA suppressed by mammalian NANOS2 in sexually differentiating male germ cells. Nat. Commun. 7, 11272 (2016).
Baltus, A. E. et al. In germ cells of mouse embryonic ovaries, the decision to enter meiosis precedes premeiotic DNA replication. Nat. Genet. 38, 1430–1434 (2006).
Bowles, J. et al. Retinoid signaling determines germ cell fate in mice. Science 312, 596–600 (2006).
Koubova, J. et al. Retinoic acid regulates sex-specific timing of meiotic initiation in mice. Proc. Natl. Acad. Sci. USA 103, 2474–2479 (2006).
Anderson, E. L. et al. Stra8 and its inducer, retinoic acid, regulate meiotic initiation in both spermatogenesis and oogenesis in mice. Proc. Natl. Acad. Sci. USA 105, 14976–14980 (2008).
Ishiguro, K. I. et al. MEIOSIN directs the switch from mitosis to meiosis in mammalian germ cells. Dev. Cell 52, 429-445 e410 (2020).
Carroll, P. A., Freie, B. W., Mathsyaraja, H. & Eisenman, R. N. The MYC transcription factor network: Balancing metabolism, proliferation and oncogenesis. Front. Med. 12, 412–425 (2018).
Eilers, M. & Eisenman, R. N. Myc’s broad reach. Genes Dev. 22, 2755–2766 (2008).
Illi, B. & Nasi, S. Myc beyond cancer: Regulation of mammalian tissue regeneration. Pathophysiology 30, 346–365 (2023).
Ogawa, H., Ishiguro, K., Gaubatz, S., Livingston, D. M. & Nakatani, Y. A complex with chromatin modifiers that occupies E2F- and Myc-responsive genes in G0 cells. Science 296, 1132–1136 (2002).
Gao, Z. et al. PCGF homologs, CBX proteins, and RYBP define functionally distinct PRC1 family complexes. Mol. Cell 45, 344–356 (2012).
Suzuki, A. et al. Loss of MAX results in meiotic entry in mouse embryonic and germline stem cells. Nat. Commun. 7, 11056 (2016).
Kehoe, S. M. et al. A conserved E2F6-binding element in murine meiosis-specific gene promoters. Biol. Reprod. 79, 921–930 (2008).
Hisada, K. et al. RYBP represses endogenous retroviruses and preimplantation- and germ line-specific genes in mouse embryonic stem cells. Mol. Cell. Biol. 32, 1139–1149 (2012).
Qin, J. et al. The polycomb group protein L3mbtl2 assembles an atypical PRC1-family complex that is essential in pluripotent stem cells and early development. Cell Stem Cell 11, 319–332 (2012).
Leseva, M., Santostefano, K. E., Rosenbluth, A. L., Hamazaki, T. & Terada, N. E2f6-mediated repression of the meiotic Stag3 and Smc1beta genes during early embryonic development requires Ezh2 and not the de novo methyltransferase Dnmt3b. Epigenetics 8, 873–884 (2013).
Maeda, I. et al. Max is a repressor of germ cell-related gene expression in mouse embryonic stem cells. Nat. Commun. 4, 1754 (2013).
Yang, C. S., Chang, K. Y., Dang, J. & Rana, T. M. Polycomb group protein Pcgf6 acts as a master regulator to maintain embryonic stem cell identity. Sci. Rep. 6, 26899 (2016).
Endoh, M. et al. PCGF6-PRC1 suppresses premature differentiation of mouse embryonic stem cells by regulating germ cell-related genes. Elife 6, e21064 (2017).
Huang, Y. et al. Combinatorial control of recruitment of a variant PRC1.6 complex in embryonic stem cells. Cell Rep. 22, 3032–3043 (2018).
Stielow, B., Finkernagel, F., Stiewe, T., Nist, A. & Suske, G. MGA, L3MBTL2 and E2F6 determine genomic binding of the non-canonical polycomb repressive complex PRC16. PLoS Genet. 14, e1007193 (2018).
Uranishi, K. et al. Two DNA binding domains of MGA act in combination to suppress ectopic activation of meiosis-related genes in mouse embryonic stem cells. Stem Cells 39, 1435–1446 (2021).
Ohinata, Y. et al. Blimp1 is a critical determinant of the germ cell lineage in mice. Nature 436, 207–213 (2005).
Cascon, A. & Robledo, M. MAX and MYC: A heritable breakup. Cancer Res. 72, 3119–3124 (2012).
Grandori, C., Cowley, S. M., James, L. P. & Eisenman, R. N. The Myc/Max/Mad network and the transcriptional control of cell behavior. Annu. Rev. Cell Dev. Biol. 16, 653–699 (2000).
Shimada, R. & Ishiguro, K. I. Cell cycle regulation for meiosis in mammalian germ cells. J. Reprod. Dev. 69, 139–146 (2023).
Sabour, D. et al. Identification of genes specific to mouse primordial germ cells through dynamic global gene expression. Hum. Mol. Genet. 20, 115–125 (2011).
Yao, H. H., DiNapoli, L. & Capel, B. Meiotic germ cells antagonize mesonephric cell migration and testis cord formation in mouse gonads. Development 130, 5895–5902 (2003).
Soh, Y. Q. et al. A gene regulatory program for meiotic prophase in the fetal ovary. PLoS Genet. 11, e1005531 (2015).
Mathsyaraja, H. et al. Max deletion destabilizes MYC protein and abrogates Emicro-Myc lymphomagenesis. Genes Dev. 33, 1252–1264 (2019).
Kent, J., Wheatley, S. C., Andrews, J. E., Sinclair, A. H. & Koopman, P. A male-specific role for SOX9 in vertebrate sex determination. Development 122, 2813–2822 (1996).
Sekido, R. & Lovell-Badge, R. Sex determination involves synergistic action of SRY and SF1 on a specific Sox9 enhancer. Nature 453, 930–934 (2008).
Ottolenghi, C. et al. Foxl2 is required for commitment to ovary differentiation. Hum. Mol. Genet. 14, 2053–2062 (2005).
Uhlenhaut, N. H. et al. Somatic sex reprogramming of adult ovaries to testes by FOXL2 ablation. Cell 139, 1130–1142 (2009).
Tucker, E. J. The genetics and biology of FOXL2. Sex. Dev. 16, 184–193 (2022).
Hill, P. W. S. et al. Epigenetic reprogramming enables the transition from primordial germ cell to gonocyte. Nature 555, 392–396 (2018).
Mochizuki, K. et al. Repression of germline genes by PRC1.6 and SETDB1 in the early embryo precedes DNA methylation-mediated silencing. Nat. Commun. 12, 7020 (2021).
Ji, H. et al. Cell-type independent MYC target genes reveal a primordial signature involved in biomass accumulation. PLoS One 6, e26057 (2011).
Anzar, M., He, L., Buhr, M. M., Kroetsch, T. G. & Pauls, K. P. Sperm apoptosis in fresh and cryopreserved bull semen detected by flow cytometry and its relationship with fertility. Biol. Reprod. 66, 354–360 (2002).
Nguyen, D. H. et al. Apoptosis in the fetal testis eliminates developmentally defective germ cell clones. Nat. Cell Biol. 22, 1423–1435 (2020).
Li, J. et al. Accurate annotation of accessible chromatin in mouse and human primordial germ cells. Cell Res. 28, 1077–1089 (2018).
Nagano, M. et al. Nucleome programming is required for the foundation of totipotency in mammalian germline development. EMBO J. 41, e110600 (2022).
Buecker, C. & Wysocka, J. Enhancers as information integration hubs in development: lessons from genomics. Trends Genet. 28, 276–284 (2012).
Henriksson, M. & Luscher, B. Proteins of the Myc network: Essential regulators of cell growth and differentiation. Adv. Cancer Res. 68, 109–182 (1996).
Yeom, Y. I. et al. Germline regulatory element of Oct-4 specific for the totipotent cycle of embryonal cells. Development 122, 881–894 (1996).
Yoshimizu, T. et al. Germline-specific expression of the Oct-4/green fluorescent protein (GFP) transgene in mice. Dev. Growth Differ. 41, 675–684 (1999).
Sato, Y. et al. A mutation in transcription factor MAFB causes focal segmental glomerulosclerosis with duane retraction syndrome. Kidney Int. 94, 396–407 (2018).
Tomioka, M. et al. Identification of Sox-2 regulatory region which is under the control of Oct-3/4-Sox-2 complex. Nucleic Acids Res. 30, 3202–3213 (2002).
Hwang, G. H., Hopkins, J. L. & Jordan, P. W. Chromatin spread preparations for the analysis of mouse oocyte progression from prophase to metaphase II. J. Vis. Exp. 132, e56736 (2018).
Acknowledgements
We are indebted to Ms. Pei Feng Cheng (Fred Hutchinson Cancer Research Center) and Dr. Atsushi Suzuki (Yokohama National University) for mice with targeted Max deletion and Oct4dPE-CreERT2 mice, respectively. We are also grateful to Kazumi Ubukata for his technical assistance. We thank Catherine Perfect, MA (Cantab), from Edanz (https://jp.edanz.com/ac), for editing a draft of this manuscript. This study was supported in part by grants from the Japan Society for the Promotion of Science (JSPS) KAKENHI (Grant Numbers 21K06843, 23K06391, and 23H02678 to A.S., K.U., and A.O., respectively), and a grant from the National Institute of Health (Grant Number NIH/NCI R35 CA231989) to R.N.E. This study was also supported in part by grants from the Takeda Science Foundation to A.S. and K.U. and by internal grants from Saitama Medical University (Grant Numbers 20-G-1-01 and 21-B-1-16 to A.S. and K.U., respectively).
Author information
Authors and Affiliations
Contributions
A.S. and A.O. conceived and designed the study. A.S., K.U., M.N., Y.M., and S.M. performed experiments. A.S. and K.U. analyzed and interpreted the data. S.M., S.T., and R.N.E. provided resources. A.S. drafted the manuscript. S.T., R.N.E., and A.O. reviewed and edited the manuscript. A.S., K.U., R.N.E., and A.O. acquired funding for this study. A.O. administered the project. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Suzuki, A., Uranishi, K., Nishimoto, M. et al. MAX controls meiotic entry in sexually undifferentiated germ cells. Sci Rep 14, 5236 (2024). https://doi.org/10.1038/s41598-024-55506-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-55506-7
- Springer Nature Limited