Abstract
Defects in ATP synthase functioning due to the substitutions in its two mitochondrially encoded subunits a and 8 lead to untreatable mitochondrial diseases. Defining the character of variants in genes encoding these subunits is challenging due to their low frequency, heteroplasmy of mitochondrial DNA in patients’ cells and polymorphisms of mitochondrial genome. We successfully used yeast S. cerevisiae as a model to study the effects of variants in MT-ATP6 gene and our research led to understand how eight amino acid residues substitutions impact the proton translocation through the channel formed by subunit a and c-ring of ATP synthase at the molecular level. Here we applied this approach to study the effects of the m.8403T>C variant in MT-ATP8 gene. The biochemical data from yeast mitochondria indicate that equivalent mutation is not detrimental for the yeast enzyme functioning. The structural analysis of substitutions in subunit 8 introduced by m.8403T>C and five other variants in MT-ATP8 provides indications about the role of subunit 8 in the membrane domain of ATP synthase and potential structural consequences of substitutions in this subunit.
Similar content being viewed by others
Introduction
Mitochondrial diseases are a wide group of metabolic and neuromuscular diseases, often with mental disorders, most of which are caused by defects in the functioning of oxidative phosphorylation system (OXPHOS) and a deficit in the energy-rich ATP molecule1,2. OXPHOS is formed by five multi-subunit complexes built from around 90 subunits of dual genetic origin, nuclear and mitochondrial3. In humans, mitochondrial genome (mtDNA) encodes only 13 OXPHOS subunits, but mutations in these genes account for 7–15% of patients with respiratory chain deficiency (depending on the source4,5). This is due to the fact that mtDNA is replicated more often than the nuclear genome, in the organelle, which is also the source of harmful reactive oxygen species6. The mtDNA occurs in the mammalian cell in thousands of copies, which on the one hand is advantageous because the negative effects of accumulated variants may be dominated by the wild-type mtDNA molecules. However, mtDNA is inherited randomly from the mother and pathogenic variants may be at high heteroplasmy in progeny7,8. When the level of heteroplasmy overcomes the critical threshold, different for each variant and depending on tissue, the disease phenotype appears9. Studies on cell lines and tissues from patients with heteroplasmic variants allowed to define the threshold for different variants10and references therein. Patient cell lines are often used for production of homoplasmic cybrid cell lines, a reliable model for the evaluation of pathogenic effect of mtDNA variants. However the nuclear background of ρ0 cell line generated for create cybrid, often being the tumor cell line, and fact that to deplete the cell line from mtDNA the ethidium bromide is used, contribute to conflicting finding11. Recently developed methods of mtDNA editing in cell lines will allow the creation of cellular models carrying particular mtDNA variants, but they are limited to specific changes12,13,14. For these reasons, different model organisms are used for research and unicellular organism Saccharomyces cerevisiae is an ideal model due to possibility to introduce mutations into its always homoplasmic mtDNA in defined nuclear background and ability to survive on fermentative medium when mutations disrupt the functioning of the OXPHOS15. The effect of mtDNA variants is studied in this model at the level of the whole organism, not the particular cell line or tissue type.
In our previous studies, we used yeast to determine the nature of MT-ATP6 gene variants and define their mechanisms of pathogenesis16,17,18,19,20,21,22,23,24,25,26. This gene codes for the inner mitochondrial membrane located subunit a/ATP6 of the fifth OXPHOS complex—ATP synthase—providing the cell in ATP. Subunit a is directly involved in proton transport through the enzyme FO domain (the membrane embedded part, while the catalytic matrix oriented part is called F1)27,28,29. In this work, we focused on MT-ATP8 gene, which encodes the subunit 8/ATP8 not directly involved in the proton transport, adjacent to the a subunit. The MT-ATP8 and MT-ATP6 genes show a 46 nucleotide overlap10. We limited our analysis to variants in the MT-ATP8 gene fragment specific to subunit 8 only, excluding variants in a region common to both genes. To date, nine variants in this region have been described in the literature in patients with mitochondrial diseases (listed in Table 1), but due to the increasing use of NGS sequencing in diagnosis, the number of reported cases is increasing30,31,32,33. In the MITOMAP and ClinVar databases, these variants have the status “reported” and mostly uncertain significance. The subunit 8 primary sequence is not highly conserved, even between higher organisms. Only the beginning of yeast 8 subunit sequence shows great similarity to the human homologue. Its role in enzyme functioning is not well understood and there is no experimental work that investigated the activity and stability of ATP synthase with substitutions in this subunit. Basing on the structure of the FO domain it was proposed that subunit 8 serves to stabilize the positioning of subunit a34. Thanks to the complete structures of ATP synthases from different organisms now available28,35, we were able to compare the structures of mammalian and yeast subunits 8 in the context of the whole holoenzyme. And although the primary sequence is different, the structure of the membrane part of subunits 8 is preserved which allows for modeling substitutions in this region. We successfully introduced the mutation equivalent to the m.8403T>C into the yeast ATP8 gene and studied its effects in vivo and in vitro. We analyzed in silico the remaining amino acid substitutions in subunit 8 using the structure of the “humanized” bovine-derived FO domain, within which the sequence of subunit 8 was replaced by the Homo sapiens one and their consequences for the functioning of the enzyme were proposed. So far it is the first work aiming to define the consequences of MT-ATP8 gene variant in yeast model.
Results
Structural consequences of subunit 8 substitutions in the ATP synthase FO domain
Nine variants in the MT-ATP8 gene fragment specific for subunit 8 were described in patients suffering from mitochondrial diseases in the literature. Their positions in mtDNA, status in the MITOMAP and ClinVar databases, substitutions in subunit 8, associated diseases, pathogenicity score, and references are given in Table 1 (see also Fig. 1a,c). ATP synthase is a motor enzyme located in the inner mitochondrial membrane, built from the membrane-embedded FO and matrix exposed F1 domains, connected by the central and external stalks (Fig. 1b). During catalysis the ring of subunits c rotates together with the central stalk, while the catalytic hexamer of α/β subunits (F1 domain) and the stator does not rotate (Fig. 1b). The γ subunit of the central stalk protrudes into the F1 domain, causing conformational changes favoring substrates’ binding, ATP synthesis, and release36. Rotation of the c-ring is coupled to the proton transport through the channel formed between the c-ring and tightly adjusted subunit a. Subunit 8 is located in the membrane part of the ATP synthase stator and is tightly adjusted to subunit a and i/j, forms an α-helix, spanning the membrane and protrudes into the matrix (Fig. 1b–d). Subunit 8 is not involved in the catalytic proton transfer because it is remote from the c-ring. Due to the significant differences in the sequence of yeast and human subunit 8, the application of yeast model to study the effects of mutations equivalent to human MT-ATP8 variants on enzyme function is limited. For this reason, we first used the available bovine ATP synthase structure to analyze what structural consequences the particular substitution in subunit 8 described in patients has.
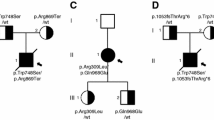
Conservation of subunits 8 and localization of the residues substituted due to variants in patients mtDNA. (a) Sequence alignment of subunit 8 from vertebrates. The abbreviations mean H.s.—Homo sapiens, B.t.—Bos taurus, S.s.—Sus scrofa, O.c.—Oryctolagus cuniculus, O.a.—Ovis aries, C.l.—Canis lupus, R.n.—Rattus norvegicus, M.m.—Mus musculus. The H.s. amino acids numbering is above the alignment. The mtDNA variants and subunit 8 substitutions are indicated. (b) Overall view of the mammalian ATP synthase monomer. Each subunit is marked by different color. The membrane is indicated by dotted line and the pathway along which protons move from the intermembrane space to the mitochondrial matrix is indicated by the arrow. (c) View from membrane of the mammalian FO domain subunits c, a, 8 and j. The amino acid residues cE58 directly involved in the proton translocation are shown as an orange belts. The positions of the substitutions in subunit 8 are indicated by deep violet belts and sticks. (d) View of the a (red), 8 (magenta) and j (green) subunits interface. Five side chains substituted in patients (8T6, 8I13, 8M16, 8L18 and 8L20) are drawn as deep violet sticks. The hydrophobic interactions are indicated as green arrows, the residues jM31 and jV35 are indicated by cyan sticks. 8W9 is colored in blue. The internal hydrogen bond of 8T6 side-chain oxygen with the backbone carbonyl group of 8L4 is indicated by yellow dotted line. The panels b, c and d were prepared with the Yasara Structure package (version 21.8.27, licensed to J.P.; http://www.yasara.org/)65, combining 6cp5 and 6zbb PDB records.
In bovine and porcine ATP synthase solely the structure of the first 29 residues of subunits 8 was solved and the remaining C-terminal region was missing28,37. Bearing in mind the identity of the two mammalian structures of the N-terminal fragment of subunit 8, we decided to analyze in silico “humanized” bovine FO domain, in which that of Homo sapiens replaced the sequence of bovine subunit 8. We used two classes of descriptors to quantify the thermodynamic effects of a particular residue replacement at a given position: total free energy change of the stability of the subunit 8 peptide and the stabilization/destabilization effect on the pairwise inter-subunits interactions.
The first six residues of subunit 8 helix are bent ninety degrees towards subunit a helix 4 (aH4, where a indicates the subunit, H means helix). The conserved threonine in position 6 causes the 8-helix to fold towards the a subunit. The internal hydrogen bond of its side-chain oxygen with the backbone carbonyl group of leucine in position 4 (8L4) stabilizes subunit 8 backbone bending (Fig. 1d, yellow dotted line). The neighboring tryptophan 8W9 interacts with aL98 and aS99 in the helix 4 of subunit a, stabilizing its positioning. The variants m.8381A>G and m.8382C>T led to substitution of threonine in position 6 to alanine or isoleucine (8T6A/I), respectively. Replacement of threonine by alanine moderately destabilizes the subunit 8 bending (ΔΔGfold = 0.6 kcal/mol), while isoleucine in this position has the opposite effect (ΔΔGfold = − 0.7 kcal/mol) and moreover it interacts with the 8W9 and makes this fragment more compact (Fig. 1d, green arrow). In both cases, the positioning of subunit a may be changed, which may minutely affect the proton channel functioning.
The m.8403T>C introduces threonine in the place of isoleucine in position 13 of subunit 8 (8I13T). 8I13 interacts with subunit j residues jV35 and jM31, forming the hydrophobic cluster (Fig. 1d, green arrows). Its side-chain is also involved in Van der Waals interactions with subunit a residues aG103 and aS99, which stabilize the positioning of subunit a relative to subunit 8. Threonine in this position breaks these interactions pattern (ΔΔGfold = 3.2 kcal/mol), which may affect the subunit a placement.
The m.8411A>G introduces valine in the place of methionine in position 16 of subunit 8 (8M16V). The 8M16 side-chain forms the hydrophobic cluster with residues aM100, aG103, aM104, 8M12, and jM31. Substitution with valine in this position substantially modifies these interactions (ΔΔGfold = 3.7 kcal/mol), which may destabilize the interface between these three subunits, especially with subunit a, and in consequence its functioning.
The m.8418T>C and m.8424T>C led to replacement of both 8L18 or 8L20 by proline, which destabilizes the whole FO domain (ΔΔGfold = 4.0 or 10 kcal/mol, respectively). 8L18 interacts with aT21 and aL25, while 8L20 interacts with aL75, aF78, aS74, aM71, and aM104. Substitution with proline in one of these two positions introduces substantial steric clashes, which must be further partially compensated by a global conformational change (not modeled in FoldX). That will strongly affect the positioning of subunit a and, in consequence, the functioning of the channel.
The two remaining substitutions: 8P39L, and 8M42T (Table 1, Fig. 1a,c), are located in not defined subunit 8 fragment in two mammalian ATP synthase structures28, so the results must be regarded roughly qualitative. The thermodynamic effect of 8P39L substitution (ΔΔGfold < 1 kcal/mol) remained almost negligible.
Consequences of the ATP8-I13T equivalent substitution in yeast subunit 8 (8L13T)
Yeast as a model can be used to study the effects of mutations in humans when these mutations affect conserved amino acid residues. Comparing the sequences of the human and yeast subunits 8, a similarity is seen within the first 18 amino acid residues, where the first 4 residues and leucine in position 18 are conserved while similar residues are present in positions 9 and 13: W9 and I13 in mammals is replaced by F9 and L13 in yeast, respectively (Fig. 2a). Basing on the sequence alignment we aimed to introduce the threonine in place of leucine in position 13 and valine in place of leucine in position 18 of yeast subunit 8, succeeding with the first substitution. With the publication of the subunit 8 structures in vertebrates and yeast, it was possible to compare them. As can be seen in Fig. 2b, the first 20 amino acid residues coincide quite closely in the structure of subunit 8 of vertebrates and yeast, validating our choice. Because subunit 8 was proposed to stabilize the subunit a34 we constructed two variants of 8L13T substitution: in the wild type mtDNA (strain CPY1-WT, Table 2) and in the mtDNA in which subunit a was fused to HisHA tags at the C-terminus (CPY1) in the goal to pulldown the whole enzyme complex and assess its stability (see “Methods” for description of strains construction38).
Mammalian and yeast subunits 8 overlap. (a) Sequence alignment of subunit 8 from vertebrates (the first 30 amino acids) and yeasts. The abbreviations mean: S.p.—Schizosaccharomyces pombe, Y.l.—Yarrowia lipolytica, C.g.—Candida glabrata and Saccharomyces cerevisiae (S.c.) and remaining are explained in the legend to Fig. 1. The human variants and subunit 8 substitutions are indicated above or below the alignment, respectively. (b) Super positioning of the subunits 8 from bovine (magenta) and yeast (cyan) (pdb codes 6zbb and 6cp5, respectively) done using the Pymol™, (version 2.3.2, licensed to R.K.; https://pymol.org/2/). The first 6 residues of the yeast and second part of the bovine (starting from 29th residue) subunits were not resolved in the published structures therefore are not visible. Positions found to be substituted in patients are indicated by black sticks while the corresponding in yeast subunit are shown in deep blue.
Respiratory growth
The growth of 8L13T cells was checked on solid and liquid fermentative (glucose) and respiratory (glycerol) media at both temperatures routinely used for yeast cells: physiological 28 °C and elevated 36 °C (conditions of the mild heat stress). While the 8L13T substitution in wild type enzyme does not slow down the growth of yeast cells at both temperatures and on both types of media liquid and solid, it leads to very poor respiratory growth when coexists with subunit aHisHA on solid medium and slows down the growth in liquid respiratory medium at elevated temperature (Fig. 3, Fig. S1). Interestingly these cells reached lower plateau also in liquid fermentative medium, especially when grown at elevated temperature, what additionally indicates on the lower capacity to use non-fermentative carbon sources (here the ethanol produced during logarithmic growth by fermentation of glucose, Fig. S1). The good respiratory growth does not imply that the mutation has no deleterious effects on ATP synthase, because decreasing ATP synthase activity by ~ 85% is required to affect significantly growth of yeast cells on non-fermentable substrates39. However, the ATP production deficits lower than 85% may be manifested by the growth phenotype on respiratory medium supplemented with sub-inhibitory concentration of oligomycin—a drug that inhibits rotation of the c-ring—because less of this drug is needed to reach the ATP synthase threshold activity19. Like 8L13T substitution alone does not impact the respiratory growth in the presence of oligomycin, coexistence of 8L13T with aHisHA does it at elevated temperature (Fig. 3).
Respiratory growth phenotypes of strains bearing the 8L13T in the background of aWT or aHisHA. Cells from the indicated strains grown in glucose pre-cultures were serially diluted and spotted on rich glucose or glycerol plates with or without oligomycin (0.5 μg/mL) and incubated at 28 or 36 °C. Plates without oligomycin were scanned after three days of incubation while those with the drug after four days of incubation.
Respiration and ATP synthase activities
To measure the efficiency of OXPHOS functioning, mitochondria were isolated from cells bearing 8L13T grown in rich galactose medium at two temperatures 28 and 36 °C. Galactose is a fermentative carbon source which does not repress the expression of mitochondrial genes40. To reflect the activities of respiratory chain we measured the rate of oxygen consumption by mitochondria using NADH as a substrate (state 4). To reflect the activity of respiratory chain and ATP synthase the ADP was added after NADH (oxygen consumption upon ATP synthase operation, state 3). Addition of the proton ionophore CCCP before NADH let to measure the uncoupled, maximal respiration. Ascorbate/TMPD in the presence of CCCP was used to measure the maximal complex IV activity. ATP synthesis was assayed in the conditions of state 3.
The activities reflect the respiratory growth deficits, described above. The 8L13T substitution in wild type enzyme does not slow down the oxygen consumption and ATP synthesis in mitochondria isolated from cells grown at both temperatures (Fig. 4a, Table S1). Consistently with slower respiratory growth, the oxygen consumption (state 4 and state 3) and ATP synthesis were reduced by ~ 45% and 35%, respectively, in mitochondria extracted from the 8L13T aHisHA cells, similarly at both temperatures (Fig. 4b, Table S2). Although the state 3 and the ATP synthesis were significantly reduced, the yield in ATP per electron transferred to oxygen in mitochondria was the same as the one measured in control mitochondria. The uncoupled, maximal respiration and activity of complex IV were reduced similarly, in accordance with the adaptation of complex IV biogenesis to the proton transport activity of FO (Table S241). Then we evaluated the F1-mediated ATP hydrolysis activity on non-osmotically protected mitochondria at pH 8.4, the conditions under which this activity is maximal. A 15% reduction in this activity was observed in mitochondria from 8L13T aHisHA cells grown at elevated temperature (Table S2).
State 3 and ATP synthesis in mitochondria from 8L13T aWT (a) and 8L13T aHisHA (b) cells. The oxygen consumption at state 3 is given in the nmol of O2 min−1 mg−1 and ATP synthesis values are represented in nmol of ATP min−1 mg−1. The histograms show the data from three biological repetitions. Statistical significance is shown: *p < 0.05. The figure is relative to Tables S1 and S2.
To further evaluate the influence of the 8L13T substitution in subunit 8 on oxidative phosphorylation we monitored changes in the mitochondrial transmembrane potential (Δψ) using a cationic dye Rhodamine123, the uptake rate of which is a highly reproducible and sensitive parameter for estimation of Δψ. Mitochondria are energized by ethanol, then ADP is added, what induce Δψ due to proton reentry through the F1FO-ATP synthase, which is normally rebuild during one minute in wild type mitochondria. Then the respiratory chain is blocked by addition of potassium cyanide (KCN) what induce the immediate drop in Δψ which is not total because the ATP synthesized during the previous step of the experiment become hydrolyzed by ATP synthase, which pumps protons to the IMS and sustains the potential. Addition of oligomycin inhibits ATP synthase what is manifested by fast drop in Δψ. The CCCP is added at the end of the experiment to normalize it, as it dissipates Δψ totally. The Δψ variations in mitochondria from 8L13T cells in wild type subunit a were similar to those in the control mitochondria, independently on the growth temperature of cells. When the 8L13T coexists with subunit aHisHA the changes are similar in mitochondria from cells grown at physiological temperature for yeast, but not in those from cells grown at elevated temperature. In these conditions a significantly longer time is needed to reestablish the Δψ after ADP addition, in proportion to the reduced capacity of those mitochondria to respire and synthesize ATP (Fig. 5).
Variations in mitochondrial inner membrane potential in mitochondria from 8L13T aHisHA cells grown at elevated temperature. The tracings show how the mitochondria responded to externally added ADP. The additions were 75 μM ADP, 0.5 μg/mL Rhodamine 123, 75 μg/mL mitochondrial proteins (Mito), 10 μL ethanol (EtOH), 2 mM KCN, 4 μg/mL oligomycin (oligo), and 4 μM CCCP. The shown tracings are representative of three biological repetitions.
ATP synthase stability
The stability and amount of fully assembled ATP synthase complexes were analyzed by BN- and SDS-PAGE electrophoresis followed by Western blotting. The complexes were liberated from the inner membrane of previously isolated mitochondria by the mild detergent—digitonin, the conditions under which the dimers and monomers of the enzyme are well preserved. The 8L13T bearing ATP synthase complexes, independently on the subunit a variant, were stable and accumulated in the amount comparable to the control mitochondria (Fig. 6a). The accumulation of ATP synthase subunits in total protein extracts from cells was not decreased, indicating on no impact of the 8L13T substitution on stability of the enzyme (Fig. 6b). Then we performed the pull-down of the ATP synthase complexes from the mitochondria using the Ni-NTA agarose to further study the enzyme stability. As shown on Fig. S2 the amount of ATP synthase subunits purified from mitochondria containing wild type subunit 8 and 8L13T variant were similar.
Assembly and stability of ATP synthase complexes in 8L13T cells. (a) The ATP synthase complexes were liberated from the mitochondrial membrane by digitonin (1.5 g/g protein) and 200 μg of proteins were separated by BN-PAGE in gels containing a 3–10% polyacrylamide gradient. The proteins were transferred to a PVDF membrane and probed with antibodies against, Atp1 (subunit α). Dimeric, monomeric and free F1 subdomains of ATP synthase are indicated. (b) Total cellular protein extracts were separated by SDS-PAGE and then transferred to a nitrocellulose membrane and probed with antibodies against the indicated proteins. The intensity of bands was calculated using ImageJ and normalized to the porin used as a loading control. The standard errors and p-values were calculated from results of at least three independent experiments. Samples from the indicated strains were analyzed together on one gel transferred to the membrane and the original blots are presented in SupplementaryRowImages pages 2–5. The representative gels fragments are shown.
Discussion
In the mitochondrial MT-ATP8 gene, nine variants introducing the substitutions into subunit 8 of ATP synthase have been described to date in patients with mitochondrial diseases (Table 1). The pathogenicity of any of these variants has not been confirmed (MITOMAP/ClinVar). All but one was described in single case/family. Biochemical data from patients' cells are missing, except one report in which m.8382C>T, m.8403T>C and m.8424T>C variants were studied permitting to conclude about pathogenicity of m.8424T>C33. Therefore, we aimed to use S. cerevisiae, which is an ideal model organism to study the effects of variants in the MT-ATP6 gene, to model variants in the MT-ATP8. The low sequence conservation of subunit 8 limits the use of this organism as a model to study the effects of substitutions in subunit 8 (Fig. 2a) however, the structures of the membrane fragments of human and yeast subunits 8 are highly conserved in evolution. Therefore, we believe that amino acid residues introduced at the corresponding positions in the yeast and human subunit 8 will result in similar changes in the structure and functioning of the FO domain. The substitution of 8L13 of yeast subunit 8, equivalent to 8I13 in the human subunit, has no detrimental consequences for the functioning of the yeast ATP synthase, arguing that this variant is not pathogenic. However, the significant (of about 40%) decrease of oxygen consumption and ATP production was found in mitochondria when this substitution coexisted with the modification of the C-terminus of subunit a by its fusion with the six histidine residues and HA epitope sequence of nine amino acid residues (YPYDVPDYA)38. As individually, both these modifications do not affect the activity of ATP synthase, when combined, they reduce the efficiency of proton transport by 40% (but not the stability of the enzyme). The explanation is that each of these modifications individually can be compensated within these subunits, but when they occurred together, such compensation is not possible. The C-terminal HA tag and 6 histidine residues are at the end of subunit aH6, which is directly adjacent to the c-ring. The compensation of changes within the subunit a introduced by HisHA tag may be located within subunit a helices 6, 5 and even 4, because the 8L13T substitution which disturbs the subunit aH4, prevents it. It is therefore possible that the 8L13T variant may be harmful in certain genetic background(s). With an energy change of ΔΔGfold = 3.2 kcal/mol, stability and interactions of subunit 8 with the subunit a neighboring residues (aG103 and aS99) may be disturbed, which may be of great importance for ATP synthase functioning. Interestingly the pathogenicity score of m.8403T>C was high42 and patient suffered from progressive neuropathy. The measurements performed on the patients’ primary fibroblasts showed decreased complex IV but normal ATP synthesis activity33,43. Further experimental studies are needed to clearly define the nature of m.8403T>C variant.
The structures of FO domain of ATP synthase from many organisms allowed to understand the enzyme mechanism of action. The in silico analysis of amino acid substitutions in subunits of ATP synthase provides much information about the potential mechanisms by which they affect the enzyme functioning. In the structures of mammalian ATP synthases, recently published, the ATP8 protein was solved within its first 29 amino acid residues permitting to analyze only six substitutions28,37. The in silico analysis of 8T6A, 8T6I, 8M16V, 8L18P and 8L20P substitutions provided important information’s about the role of these residues in subunit 8: The 8T6 is important for the shape of subunit 8, which stabilize the placement of the subunit aH4, while the 8M16, 8L18 and 8L20 are crucial for the hydrophobic interactions of subunit 8 with subunit a and j. This finding is in accordance with previously proposed role for subunit 8 in stabilization of subunit a in the FO34. The substitutions 8T6A and 8T6I minutely change the ΔΔG energy and may destroy the interactions of subunit 8 with the neighboring residues of subunit a. They may have slight indirect effects on the functioning of the proton channel, but experimental research is needed to define their consequences. In accordance to these finding substitutions of 8T6 were associated with milder diseases phenotypes: cardiomyopathy or diabetes with deafness. The subunit 8 energy change due to 8M16V was significant and this variant was associated to more severe mitochondrial disease leading to death in childhood33,44,45,46. This can be explained by the fact that 8M16 is located very close to the central part of the a subunit, where amino acid residues directly involved in proton transport through FO are located. The located close 8L18P and 8L20P changes are very costly energetically, proline is excluded from both these positions of subunit 8, moreover significant respiratory chain deficiencies were found in m.8424T>C cybrids. The huge structural destabilization assigned in silico to the 8L20P variant (ΔΔGfold = 10 kcal/mol) may directly affect the biogenesis of subunit 8 and cause its improper assembly to ATP synthase FO domain, as suggested in Ref.33. These findings correlate with severe diseases in patients and speak for pathogenic character of those two variants33,47. The super-positioning of subunits 8 from yeast and bovine indicates that both variants may be modelled in yeast model organism for experimental verification of this conclusion.
Conclusion
In silico analysis suggests that two variants m.8418T>C and m.8424T>C may be the cause of the mitochondrial disease in the reported cases. Mutation equivalent to the m.8403T>C variant in MT-ATP8 gene does not affect the ATP synthase functioning in S. cerevisiae, which indicates its non-pathogenic character. The obtained results on its two yeast models suggest that structural compensations of variants within the subunits of ATP synthase are possible. This fact, as well as our previous work based on the analysis of genetic suppressors of pathogenic variants in the a subunit, are an argument for designing small molecules/peptides, potential drugs, that would induce structural changes and restore the enzyme function.
Methods
Growth media
The media used for growing yeast were: fermentative: YPDA (1% Bacto yeast extract, 1% Bacto Peptone, 2% glucose for standard growth or 10% glucose for selection of the cytoductants, 40 mg/L adenine), YPGalA (1% Bacto yeast extract, 1% Bacto Peptone, 2% galactose, 40 mg/L adenine), respiratory: YPGlyA (1% Bacto yeast extract, 1% Bacto Peptone, 2% glycerol, 40 mg/L adenine). BIOL-Leu (5% glucose, 182.5 g/L sorbitol, 1.7 g/L Yeast Nitrogen base without amino acids, 5 g/L ammonium sulfate, 0.8 g/L CSM-Leu drop out mix, and 40 mg/L adenine) is a fermentative medium for selection of mitochondrial transformants. Media were solidified by the addition of 2% (w/v) Bacto agar. Yeast cells were grown at 28 or 36 °C, cultures in liquid media were shaken at 180 rpm. Oligomycin was added to the media at the concentration of 0.5 µg/mL. Growth curves were established with the Bioscreen CTM system.
ATP8 gene mutagenesis and construction of the atp8-L13T mutant strains
The equivalent mutation to m.8403T>C was introduced into ATP8 gene cloned into pMOS vector48 with the Q5® Site-directed Mutagenesis Kit of NEBiolabs. The sequences of primers used for mutagenesis reaction are: 5′ TATTATTAATTTTATTCTCACAATTCTTTTTACCTATG 3′ (the changed bases are indicated in bold) and 5′ GAATCATTAATAAGAAACCATATGTTGTTTGATTCATAAAATAAAATGGAAC 3′. The DNA fragment containing the atp8-L13T was liberated by XbaI-NdeI digestion from pMOS-ATP8-based plasmids resulting from mutagenesis reaction and ligated at the same sites with pJM2 giving the pCP1 plasmid49. The pJM2 contains the yeast mitochondrial COX2 gene, which serves for identification of mitochondrial transformants.
The mutations into mtDNA are introduced into the MR6 strain50. It is a derivative of W303-1B strain in which the nuclear ARG8 gene was replaced with HIS3, contains mitochondrial genome of S288C strain that has been entirely sequenced, and has a wild-type CAN1 gene, encoding a basic amino acid permease. The RKY194 strain was obtained by the change of the mating type of MR6-atp8Δ MATα strain (atp8Δ-ATP6-WT, provided by A. Tzagoloff) using a plasmid encoding the HO endonuclease51. The isogenic wild type control strain for RKY194 was constructed by crossing RKY194 with the RKY68, the ρ− synthetic bearing in the mtDNA only wild type ATP8 and COX2 genes, and selection the respiring progeny bearing the nucleus of RKY194. To receive the atp8-L13T mutant the pCP1 plasmid was introduced by co-transformation with the LEU2 gene containing plasmid Yep351 into the ρ0 strain DFS160 by microprojectile bombardment using a biolistic PDS-1000/He particle delivery system (Bio-Rad) as described15. To learn more about the technique developed by us see52. Mitochondrial transformants were identified among the Leu+ nuclear transformants by their ability to produce respiring clones when mated to the non-respiring NB40-3C strain bearing a deletion in the mitochondrial COX2 gene. The resulting clone ρ− synthetic CPY1synth (Table 2) was crossed to strains MR6 atp8Δ-ATP6-HisHA (provided by prof. A. Tzagoloff) or RKY194 (atp8Δ-ATP6-WT). Because the ρ− synthetic strain bears the mutation kar1-1 which blocks the fusion of the nuclei, during the crosses the cytoplasm and the mitochondrial network fuse permitting for homologous recombination between the two mtDNA molecules and the replacement of ARG8m gene present in the ATP8 locus by atp8 sequence bearing atp8-L13T mutation. During the two passages of cells resulting from the crosses in the rich 10% glucose medium the mtDNAs separate till the homoplasmic state. The resulting CPY1 (in background of ATP6-HisHA) and CPY1-WT (in background of ATP6-WT) strains bearing in the complete (ρ+) mtDNA the atp8-L13T gene variant was identified by ability to grow on non-fermentable carbon source glycerol and inability to grow in the absence of arginine. The presence of mutation was verified by DNA sequencing of the PCR amplified ATP8 locus with oligonucleotides oATP8-3 5′-TGTCAGTTATTTTATATTAATGTTTAATC-3′ and oATP8-4 5′-ATATATATATATAAATATATAGTCCGTAAGG-3′.
Measurement of mitochondrial respiration, ATP synthesis/hydrolysis and membrane potential
The mitochondria were prepared from yeast cells grown in rich galactose medium (YPGalA) at 28 °C or 36 °C by the enzymatic method described in Ref.53. YPGalA is a fermentative medium in which the mitochondrial genes are not repressed. This growth condition reduces the probability for selection of the spontaneous suppressors in the respiratory deficient mutants and is used in our routine analysis of mitochondrial mutants. For respiration and ATP synthesis assays, mitochondria were diluted to 0.075 mg/mL in respiration buffer (10 mM Tris-maleate (pH 6.8), 0.65 M mannitol, 0.36 mM EGTA, and 5 mM Tris–phosphate). Oxygen consumption rates were measured using a Clarke electrode after consecutively adding 4 mM NADH (state 4 respiration), 150 µM ADP (state 3) or 4 µM carbonyl cyanide m-chlorophenylhydrazone (CCCP) (uncoupled respiration), as previously described54. The rates of ATP synthesis were determined under the same experimental conditions in the presence of 750 µM ADP; aliquots were withdrawn from the oxygraph cuvette every 15 s and the reactions were stopped with 3.5% (w/v) perchloric acid, 12.5 mM EDTA. The ATP in samples was quantified using the Kinase-Glo Max Luminescence Kinase Assay (Promega) and a Beckman Coulter's Paradigm Plate Reader. Participation of F1FO-ATP synthase to ATP production was assessed by measuring the sensitivity of ATP synthesis to oligomycin (3 μg/mL). Variations in transmembrane potential (ΔΨ) were evaluated in the respiration buffer containing 0.150 mg/mL of mitochondria and the Rhodamine 123 (0.5 μg/mL), with λexc of 485 nm and λem of 533 nm under constant stirring using a Cary Eclipse Fluorescence Spectrophotometer (Agilent Technologies, Santa Clara, CA, USA)55. The specific ATPase activity at pH 8.4 of non-osmotically protected mitochondria was measured as described in Ref.56.
BN- and SDS PAGE analyses
Blue native-PAGE experiments were carried out as described57. Briefly 200 µg of mitochondrial proteins was suspended in 100 µL of extraction buffer (30 mM HEPES pH = 6,8, 150 mM potassium acetate, 12% glycerol, 2 mM 6-aminocaproic acid, 1 mM EGTA, 1.5% digitonin (Sigma)), supplemented with protease inhibitors cocktail tablet (Roche, one tablet per 10 mL of the buffer). After 26 min incubation on ice, the extracts were cleared by centrifugation (21,950×g, 4 °C, 30 min), supplemented with 4.5 µL of loading dye (5% Serva Blue G-250, 750 mM 6-aminocaproic acid) and run on NativePAGE™ 3–12% Bis–Tris Gels (Invitrogen). For SDS-PAGE analysis 10 OD of cells was centrifuged, pellet was suspended in 500 µL of 0.2 M NaOH and incubated for 10 min on ice. Then 50 µL of 50% TCA (trichloroacetic acid) was added, vortexed and after 10 min incubation on ice, centrifuged at 21,950×g for 10 min at 4 °C. The protein pellet was washed with 1 mL of 1 M Tris-base and suspended in 50 µL of 5% SDS for measurement of protein concentration by method of Lowry58. 50 µg of protein were loaded per lane of 10% SDS-PAGE gel59. After transfer onto a PVDF or nitrocellulose membrane, ATP synthase complexes were detected using polyclonal antibodies raised against Atp1, Atp2, Atp4, Atp7, Atp17, Atp18, OSCP subunits of yeast ATP synthase at 1:10,000 dilution (a kind gift from Marie-France Giraud, Bordeaux, France)60 or against Por1 (gift from prof. Teresa Zoladek, IBB PAS). The membranes were incubated successively with the indicated antibodies—membranes were stripped before incubation with anti-Por1 antibody (Restore™ PLUS Western Blot Stripping Buffer, Thermo Scientific, ref. 46430). Immunoreactivity was studied using secondary antibody conjugated to horseradish peroxidase (DAKO). The blots were stained with Immobilon Western Chemiluminescent HRP Substrate from Millipore (ref. WBKLS0500) used as a substrate. For image acquisition the Uvitec Cambridge Q4 Alliance System (Uvitec Cambridge, UK) was used, images were collected in the serial accumulation option. Images were processed with ImageJ and Adobe Photoshop CC 2019.
Purification of the ATP synthase holoenzyme from the mitochondria
The protocol described in the Ref.61 was applied. 5 mg of mitochondria were centrifuged in 2 mL tubes and suspended in 1 mL of sonication buffer (250 mM saccharose, 50 mM NaH2PO4, 5 mM 6-aminocaproic acid, 1 mM EDTA, pH 7.5, protease inhibitors cocktail tablet (Roche), 1 µM PMSF) and sonicated 6 times 10 s, with 10 s intervals on ice. After centrifugation of the extract at 5000×g for 10 min at 4 °C supernatant was ultracentrifuged at 268,526×g during 1 h (Thermo Scientific™ Sorvall™ WX ultracentrifuge, TFT80.2 rotor). The pellet was washed twice with the sonication buffer without EDTA (without suspending it) and then suspended with the use of the potter in 500 µL of MP extraction buffer (150 mM potassium acetate, 10% glycerol, 2 mM 6-aminocaproic acid, 30 mM HEPES, pH 7.4, 1% N-dodecyl-β-maltoside, 2 mM PMSF and protease inhibitors cocktail tablet) by 20 min incubation on ice. The membranes were centrifuged 30 min at 21,950×g at 4 °C and the extract was incubated with 200 µL of the Ni-NTA agarose washed previously by Binding buffer (50 mM NaCl, 10% glycerol, 10 mM imidazole, 20 mM NaH2PO4, pH = 7.9, 0,1% n-dodecyl-β-maltoside, 2 mM PMSF, protease inhibitors cocktail tablet) for the night. Next day the beads were washed twice with the Binding buffer, then suspended in 400 µL of Binding buffer, dosed and after addition of 100 µL of 5 × Laemmli sample buffer, boiled 5 min. The 50 µg of the extract and 2 µg of the bead’s eluate were loaded on the 15% gel.
In silico analysis of subunit 8 substitutions in the ATP synthase structures
Multiple sequences of ATP synthase 8-subunits of various origins were aligned and drawn using Clustal Omega62 and Espript 3.063. Molecular views of ATP synthase monomer and the FO subunits 8, a, j and c-ring were obtained from the monomer of bovine ATP synthase (pdb_id: 6zbb28). The pdb records: 6cp5 and 6zbb were used as the starting points for the in silico analysis of yeast and mammalian ATP synthase FO domains, respectively35 using Yasara Structure package (http://www.yasara.org/). The side-chain conformations were iteratively optimized for both structures with ten cycles of repairPDB procedure implemented in FoldX5 (http://foldxsuite.crg.eu/). The original sequence of subunit 8 of bovine ATP synthase was then replaced with a Homo sapiens one (A0A075X6N5_HUMAN), and the side-chain packing in the resulting structure was again re-optimized with FoldX5. The effect of single-residue replacement was analyzed in silico using FoldX564. The thermodynamic effect of residue replacement at a given position (ΔΔG, expressed in kcal/mol) was then assessed for the whole FO domain with FoldX5 using PositionScan, and further decomposed to the contribution of particular intra- and inter-domain interaction using AnalyseComplex. The stability (ΔG) of a protein is defined by the free energy. The lower it is, the more stable a protein is. The ΔΔG is a difference in free energy between a wild-type and substituted variant. A substitution that brings energy (ΔΔG > 0 kcal/mol) will destabilize the protein structure, while a substitution that remove energy (ΔΔG < 0 kcal/mol) will stabilize the protein structure. A common threshold ΔΔG > 1 kcal/mol is considered as a significant effect. The structure figures were drawn using PyMOL™ 2.3.2 (https://pymol.org/2/) or Yasara Strucure package (http://www.yasara.org/)65.
Statistical analysis
Three biological and three technical replicates were performed for all experiments. The unpaired two-tailed t test was used for all data sets. Significance and confidence level was set at 0.05.
Statement of ethics
The permission number for work with genetically modified microorganisms (GMM I) for RK is 01.2-28/201.
Data availability
Supplementary data and the materials (yeast strains, sequence of the ATP8 locus amplified by PCR using the mtDNA from the model strains) are available upon request from the corresponding author.
References
Gorman, G. S. et al. Mitochondrial diseases. Nat. Rev. Dis. Primers 2, 16080. https://doi.org/10.1038/nrdp.2016.80 (2016).
Chinnery, P. F. Mitochondrial disease in adults: What’s old and what’s new?. EMBO Mol. Med. https://doi.org/10.15252/emmm.201505079 (2015).
Saraste, M. Oxidative phosphorylation at the fin de siecle. Science 283, 1488–1493. https://doi.org/10.1126/science.283.5407.1488 (1999).
Wallace, D. C. Mitochondrial DNA mutations in disease and aging. Environ. Mol. Mutagen. 51, 440–450. https://doi.org/10.1002/em.20586 (2010).
Bannwarth, S. et al. Prevalence of rare mitochondrial DNA mutations in mitochondrial disorders. J. Med. Genet. 50, 704–714. https://doi.org/10.1136/jmedgenet-2013-101604 (2013).
Melvin, R. G. & Ballard, J. W. O. Cellular and population level processes influence the rate, accumulation and observed frequency of inherited and somatic mtDNA mutations. Mutagenesis 32, 323–334. https://doi.org/10.1093/mutage/gex004 (2017).
Edwards, D. M. et al. Avoiding organelle mutational meltdown across eukaryotes with or without a germline bottleneck. PLoS Biol. 19, e3001153. https://doi.org/10.1371/journal.pbio.3001153 (2021).
Stewart, J. B. & Chinnery, P. F. The dynamics of mitochondrial DNA heteroplasmy: Implications for human health and disease. Nat. Rev. Genet. 16, 530–542. https://doi.org/10.1038/nrg3966 (2015).
Wallace, D. C. & Chalkia, D. Mitochondrial DNA genetics and the heteroplasmy conundrum in evolution and disease. Cold Spring Harb. Perspect. Biol. 5, a021220. https://doi.org/10.1101/cshperspect.a021220 (2013).
Dautant, A. et al. ATP synthase diseases of mitochondrial genetic origin. Front. Physiol. 9, 329. https://doi.org/10.3389/fphys.2018.00329 (2018).
Wilkins, H. M., Carl, S. M. & Swerdlow, R. H. Cytoplasmic hybrid (cybrid) cell lines as a practical model for mitochondriopathies. Redox Biol. 2, 619–631. https://doi.org/10.1016/j.redox.2014.03.006 (2014).
Mok, B. Y. et al. CRISPR-free base editors with enhanced activity and expanded targeting scope in mitochondrial and nuclear DNA. Nat. Biotechnol. 40, 1378–1387. https://doi.org/10.1038/s41587-022-01256-8 (2022).
Lei, Z. et al. Mitochondrial base editor induces substantial nuclear off-target mutations. Nature 606, 804–811. https://doi.org/10.1038/s41586-022-04836-5 (2022).
Cho, S. I. et al. Targeted A-to-G base editing in human mitochondrial DNA with programmable deaminases. Cell 185, 1764-1776 e1712. https://doi.org/10.1016/j.cell.2022.03.039 (2022).
Bonnefoy, N. & Fox, T. D. Genetic transformation of Saccharomyces cerevisiae mitochondria. Methods Cell Biol. 65, 381–396. https://doi.org/10.1016/s0091-679x(01)65022-2 (2001).
Kabala, A. M., Lasserre, J. P., Ackerman, S. H., di Rago, J. P. & Kucharczyk, R. Defining the impact on yeast ATP synthase of two pathogenic human mitochondrial DNA mutations, T9185C and T9191C. Biochimie 100, 200–206. https://doi.org/10.1016/j.biochi.2013.11.024 (2014).
Kucharczyk, R., Dautant, A., Godard, F., Tribouillard-Tanvier, D. & di Rago, J. P. Functional investigation of an universally conserved leucine residue in subunit a of ATP synthase targeted by the pathogenic m.9176T>G mutation. Biochim. Biophys. Acta Bioenerg. 1860, 52–59. https://doi.org/10.1016/j.bbabio.2018.11.005 (2019).
Kucharczyk, R. et al. The pathogenic MT-ATP6 m.8851T>C mutation prevents proton movements within the n-side hydrophilic cleft of the membrane domain of ATP synthase. Biochim. Biophys. Acta Bioenerg. 1860, 562–572. https://doi.org/10.1016/j.bbabio.2019.06.002 (2019).
Kucharczyk, R. et al. Consequences of the pathogenic T9176C mutation of human mitochondrial DNA on yeast mitochondrial ATP synthase. Biochem. Biophys. Acta 1797, 1105–1112. https://doi.org/10.1016/j.bbabio.2009.12.022 (2010).
Kucharczyk, R. et al. Defining the pathogenesis of human mtDNA mutations using a yeast model: The case of T8851C. Int. J. Biochem. Cell Biol. 45, 130–140. https://doi.org/10.1016/j.biocel.2012.07.001 (2013).
Kucharczyk, R., Rak, M. & di Rago, J. P. Biochemical consequences in yeast of the human mitochondrial DNA 8993T>C mutation in the ATPase6 gene found in NARP/MILS patients. Biochem. Biophys. Acta 1793, 817–824. https://doi.org/10.1016/j.bbamcr.2009.02.011 (2009).
Su, X. et al. Molecular basis of the pathogenic mechanism induced by the m.9191T>C mutation in mitochondrial ATP6 gene. Int. J. Mol. Sci. 21, 5083. https://doi.org/10.3390/ijms21145083 (2020).
Su, X. et al. The pathogenic m.8993 T > G mutation in mitochondrial ATP6 gene prevents proton release from the subunit c-ring rotor of ATP synthase. Hum. Mol. Genet. 30, 381–392. https://doi.org/10.1093/hmg/ddab043 (2021).
Skoczen, N. et al. Molecular basis of diseases caused by the mtDNA mutation m.8969G>A in the subunit a of ATP synthase. Biochim. Biophys. Acta Bioenerg. 1859, 602–611. https://doi.org/10.1016/j.bbabio.2018.05.009 (2018).
Baranowska, E. et al. Molecular basis of diseases induced by the mitochondrial DNA mutation m.9032 T>C. Hum. Mol. Genet. https://doi.org/10.1093/hmg/ddac292 (2022).
Baranowska, E. et al. Probing the pathogenicity of patient-derived variants of MT-ATP6 in yeast. Dis. Model Mech. 16, dmm049783. https://doi.org/10.1242/dmm.049783 (2023).
Xu, T., Pagadala, V. & Mueller, D. M. Understanding structure, function, and mutations in the mitochondrial ATP synthase. Microb. Cell 2, 105–125. https://doi.org/10.15698/mic2015.04.197 (2015).
Spikes, T. E., Montgomery, M. G. & Walker, J. E. Structure of the dimeric ATP synthase from bovine mitochondria. Proc. Natl. Acad. Sci. U.S.A. 117, 23519–23526. https://doi.org/10.1073/pnas.2013998117 (2020).
Srivastava, A. P. et al. High-resolution cryo-EM analysis of the yeast ATP synthase in a lipid membrane. Science 360, eaas9699. https://doi.org/10.1126/science.aas9699 (2018).
Al-Kafaji, G. et al. Next-generation sequencing of the whole mitochondrial genome identifies functionally deleterious mutations in patients with multiple sclerosis. PLoS One 17, e0263606. https://doi.org/10.1371/journal.pone.0263606 (2022).
Jiang, W. et al. Mitochondrial DNA mutations associated with type 2 diabetes mellitus in Chinese Uyghur population. Sci. Rep. 7, 16989. https://doi.org/10.1038/s41598-017-17086-7 (2017).
Zhu, Y., Gu, X. & Xu, C. Mitochondrial DNA 7908–8816 region mutations in maternally inherited essential hypertensive subjects in China. BMC Med. Genom. 11, 89. https://doi.org/10.1186/s12920-018-0408-0 (2018).
Rucheton, B. et al. Homoplasmic deleterious MT-ATP6/8 mutations in adult patients. Mitochondrion 55, 64–77. https://doi.org/10.1016/j.mito.2020.08.004 (2020).
Hahn, A. et al. Structure of a complete ATP synthase dimer reveals the molecular basis of inner mitochondrial membrane morphology. Mol. Cell 63, 445–456. https://doi.org/10.1016/j.molcel.2016.05.037 (2016).
Guo, H., Bueler, S. A. & Rubinstein, J. L. Atomic model for the dimeric FO region of mitochondrial ATP synthase. Science 358, 936–940. https://doi.org/10.1126/science.aao4815 (2017).
Boyer, P. D. The ATP synthase—A splendid molecular machine. Annu. Rev. Biochem. 66, 717–749. https://doi.org/10.1146/annurev.biochem.66.1.717 (1997).
Gu, J. et al. Cryo-EM structure of the mammalian ATP synthase tetramer bound with inhibitory protein IF1. Science 364, 1068–1075. https://doi.org/10.1126/science.aaw4852 (2019).
Rak, M. & Tzagoloff, A. F1-dependent translation of mitochondrially encoded Atp6p and Atp8p subunits of yeast ATP synthase. Proc. Natl. Acad. Sci. U.S.A. 106, 18509–18514. https://doi.org/10.1073/pnas.0910351106 (2009).
Mukhopadhyay, A., Uh, M. & Mueller, D. M. Level of ATP synthase activity required for yeast Saccharomyces cerevisiae to grow on glycerol media. FEBS Lett. 343, 160–164. https://doi.org/10.1016/0014-5793(94)80310-2 (1994).
Ephrussi, B., Slonimski, P. P. & Perrodin, G. The adaptative synthesis of cytochromes in baker’s yeast. Biochem. Biophys. Acta 6, 256–267. https://doi.org/10.1016/0006-3002(50)90098-9 (1950).
Soto, I. C. et al. Synthesis of cytochrome c oxidase subunit 1 is translationally downregulated in the absence of functional F1F0-ATP synthase. Biochem. Biophys. Acta 1793, 1776–1786. https://doi.org/10.1016/j.bbamcr.2009.09.002 (2009).
Pereira, L., Soares, P., Radivojac, P., Li, B. & Samuels, D. C. Comparing phylogeny and the predicted pathogenicity of protein variations reveals equal purifying selection across the global human mtDNA diversity. Am. J. Hum. Genet. 88, 433–439. https://doi.org/10.1016/j.ajhg.2011.03.006 (2011).
Aure, K. et al. Episodic weakness due to mitochondrial DNA MT-ATP6/8 mutations. Neurology 81, 1810–1818. https://doi.org/10.1212/01.wnl.0000436067.43384.0b (2013).
Mkaouar-Rebai, E. et al. A de novo mutation in the adenosine triphosphatase (ATPase) 8 gene in a patient with mitochondrial disorder. J. Child Neurol. 25, 770–775. https://doi.org/10.1177/0883073809344351 (2010).
Perucca-Lostanlen, D. et al. Mitochondrial DNA variations in patients with maternally inherited diabetes and deafness syndrome. Biochem. Biophys. Res. Commun. 277, 771–775. https://doi.org/10.1006/bbrc.2000.3751 (2000).
Finsterer, J., Stollberger, C. & Schubert, B. Acquired left ventricular hypertrabeculation/noncompaction in mitochondriopathy. Cardiology 102, 228–230. https://doi.org/10.1159/000081015 (2004).
Bacalhau, M. et al. In silico analysis for predicting pathogenicity of five unclassified mitochondrial DNA mutations associated with mitochondrial cytopathies’ phenotypes. Eur. J. Med. Genet. 60, 172–177. https://doi.org/10.1016/j.ejmg.2016.12.009 (2017).
Godard, F., Tetaud, E., Duvezin-Caubet, S. & di Rago, J. P. A genetic screen targeted on the FO component of mitochondrial ATP synthase in Saccharomyces cerevisiae. J. Biol. Chem. 286, 18181–18189. https://doi.org/10.1074/jbc.M110.214825 (2011).
Steele, D. F., Butler, C. A. & Fox, T. D. Expression of a recoded nuclear gene inserted into yeast mitochondrial DNA is limited by mRNA-specific translational activation. Proc. Natl. Acad. Sci. U.S.A. 93, 5253–5257. https://doi.org/10.1073/pnas.93.11.5253 (1996).
Rak, M. et al. Yeast cells lacking the mitochondrial gene encoding the ATP synthase subunit 6 exhibit a selective loss of complex IV and unusual mitochondrial morphology. J. Biol. Chem. 282, 10853–10864. https://doi.org/10.1074/jbc.M608692200 (2007).
Herskowitz, I. & Jensen, R. E. Putting the HO gene to work: Practical uses for mating-type switching. Methods Enzymol. 194, 132–146. https://doi.org/10.1016/0076-6879(91)94011-z (1991).
Tribouillard-Tanvier, D. et al. Creation of yeast models for evaluating the pathogenicity of mutations in the human mitochondrial gene MT-ATP6 and discovering therapeutic molecules. Methods Mol. Biol. 2497, 221–242. https://doi.org/10.1007/978-1-0716-2309-1_14 (2022).
Guerin, B., Labbe, P. & Somlo, M. Preparation of yeast mitochondria (Saccharomyces cerevisiae) with good P/O and respiratory control ratios. Methods Enzymol. 55, 149–159. https://doi.org/10.1016/0076-6879(79)55021-6 (1979).
Rigoulet, M. & Guerin, B. Phosphate transport and ATP synthesis in yeast mitochondria: Effect of a new inhibitor: The tribenzylphosphate. FEBS Lett. 102, 18–22. https://doi.org/10.1016/0014-5793(79)80919-9 (1979).
Emaus, R. K., Grunwald, R. & Lemasters, J. J. Rhodamine 123 as a probe of transmembrane potential in isolated rat-liver mitochondria: Spectral and metabolic properties. Biochem. Biophys. Acta 850, 436–448. https://doi.org/10.1016/0005-2728(86)90112-x (1986).
Somlo, M. Induction and repression of mitochondrial ATPase in yeast. Eur. J. Biochem. 5, 276–284. https://doi.org/10.1111/j.1432-1033.1968.tb00368.x (1968).
Schagger, H. & von Jagow, G. Blue native electrophoresis for isolation of membrane protein complexes in enzymatically active form. Anal. Biochem. 199, 223–231. https://doi.org/10.1016/0003-2697(91)90094-a (1991).
Lowry, O. H., Rosebrough, N. J., Farr, A. L. & Randall, R. J. Protein measurement with the Folin phenol reagent. J. Biol. Chem. 193, 265–275 (1951).
Laemmli, U. K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227, 680–685. https://doi.org/10.1038/227680a0 (1970).
Grandier-Vazeille, X., Ouhabi, R. & Guerin, M. Antibodies against subunits of F0 sector of ATP synthase from Saccharomyces cerevisiae. Stimulation of ATP synthase by subunit-8-reactive antibodies and inhibition by subunit-9-reactive antibodies. Eur. J. Biochem. 223, 521–528. https://doi.org/10.1111/j.1432-1033.1994.tb19021.x (1994).
Velours, J. et al. Organisation of the yeast ATP synthase F(0): A study based on cysteine mutants, thiol modification and cross-linking reagents. Biochem. Biophys. Acta 1458, 443–456. https://doi.org/10.1016/s0005-2728(00)00093-1 (2000).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539. https://doi.org/10.1038/msb.2011.75 (2011).
Robert, X. & Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 42, W320–W324. https://doi.org/10.1093/nar/gku316 (2014).
Delgado, J., Radusky, L. G., Cianferoni, D. & Serrano, L. FoldX 5.0: Working with RNA, small molecules and a new graphical interface. Bioinformatics 35, 4168–4169. https://doi.org/10.1093/bioinformatics/btz184 (2019).
Land, H. & Humble, M. S. YASARA: A tool to obtain structural guidance in biocatalytic investigations. Methods Mol. Biol. 1685, 43–67. https://doi.org/10.1007/978-1-4939-7366-8_4 (2018).
Tansel, T., Pacal, F. & Ustek, D. A novel ATP8 gene mutation in an infant with tetralogy of Fallot. Cardiol. Young 24, 531–533. https://doi.org/10.1017/S1047951113000668 (2014).
Elango, S., Venugopal, S., Thangaraj, K. & Viswanadha, V. P. Novel mutations in ATPase 8, ND1 and ND5 genes associated with peripheral neuropathy of diabetes. Diabetes Res. Clin. Pract. 103, e49-52. https://doi.org/10.1016/j.diabres.2013.12.015 (2014).
Jarviaho, T. et al. Novel non-neutral mitochondrial DNA mutations found in childhood acute lymphoblastic leukemia. Clin. Genet. 93, 275–285. https://doi.org/10.1111/cge.13100 (2018).
Acknowledgements
Authors thank to Prof. A. Tzagoloff for the atp8Δ deletion strains and to Marie-France Giraud for the anti-ATP synthase subunits antibodies.
Funding
This work was supported by a grant from the National Science Center of Poland (2016/23/B/NZ3/02098) to RK.
Author information
Authors and Affiliations
Contributions
C.P. and E.B.: data curation—plasmids and strains construction, isolation of mitochondria, biochemical assays and protein electrophoresis; K.N.: ATP synthase purification; J.P.: structural modeling and analysis. R.K.: conceptualization, supervision, validation, writing of the manuscript—review and editing, funding acquisition. All authors analyzed the data and contributed to the writing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Panja, C., Niedzwiecka, K., Baranowska, E. et al. Analysis of MT-ATP8 gene variants reported in patients by modeling in silico and in yeast model organism. Sci Rep 13, 9972 (2023). https://doi.org/10.1038/s41598-023-36637-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-36637-9
- Springer Nature Limited