Abstract
Perennial shrub willow are increasingly being promoted in short-rotation coppice systems as biomass feedstocks, for phytoremediation applications, and for the diverse ecosystem services that can accrue. This renewed interest has led to widespread willow cultivation, particularly of non-native varieties. However, Canadian willow species have not been widely adopted and their inherent diversity has not yet been thoroughly investigated. In this study, 324 genotypes of Salix famelica and Salix eriocephala collected from 33 sites of origin were analyzed using 26,016 single nucleotide polymorphisms to reveal patterns of population structure and genetic diversity. Analyses by Bayesian methods and principal component analysis detected five main clusters that appeared to be largely shaped by geoclimatic variables including mean annual precipitation and the number of frost-free days. The overall observed (HO) and expected (HE) heterozygosity were 0.126 and 0.179, respectively. An analysis of molecular variance revealed that the highest genetic variation occurred within genotypes (69%), while 8% of the variation existed among clusters and 23% between genotypes within clusters. These findings provide new insights into the extent of genetic variation that exists within native shrub willow species which could be leveraged in pan-Canadian willow breeding programs.
Similar content being viewed by others
Introduction
Shrub willow (Salix spp.) occur naturally in mesic areas adjacent to watercourses and wetlands across much of North America1. They were traditionally used in the production of woven baskets and as a source for analgesics2,3. Willow has been planted extensively as riparian buffers and shelterbelts in agricultural regions, and it has been promoted as a promising biomass feedstock for bioenergy applications4. The carbon dioxide that is released from burning willow wood is largely offset by its fixation during photosynthesis, making willow biomass a nearly carbon–neutral energy source5. In addition, shrub willow responds well to coppicing and is capable of reaching maximum annual growth rates at high planting densities in just a few growing seasons6. However, the non-native species that are routinely deployed can suffer from pests and disease, and may elicit concerns with regards to invasiveness7. As locally adapted genotypes offer unparalleled resistance to climatic and biotic stressors, the development of genetic resources for native willow species could facilitate the development of more resilient and productive commercial willow cultivars adapted for Canadian landscapes8.
Willow are dioecious and obligate outcrossers. There are approximately 450 recognized willow species in the world, falling into six sub-genera, of which over 100 occur in North America9. Canada alone has 76 native species10. Despite this enormous natural diversity, native willow taxa have received comparatively little research attention and few studies have sought to exploit this extensive variation. Three related shrub willow species occur across much of the temperate and boreal regions of North America: Salix eriocephala Michx. (2n = 2 × = 38; subgenus Vetrix; section Cordatae), Salix famelica (C.R. Ball) Argus (2n = 2 × = 38; subgenus Vetrix; section Cordatae), and Salix prolixa Andersson (2n = 2 × = 38; subgenus Vetrix; section Cordatae)11. Previously, these taxa have been circumscribed as subspecies: S. eriocephala ssp. eriocephala (mainly in Ontario, Quebec, New Brunswick, Nova Scotia, Prince Edward Island, and Newfoundland and Labrador), S. eriocephala ssp. famelica (Alberta, Saskatchewan, Manitoba, and Ontario), and S. eriocephala ssp. mackenzieana (Hooker) Dorn (Alaska, Yukon, British Columbia, and the Northwest Territories). However, the most recent taxonomic revision of North American Salix12 recognizes these as three separate species: S. eriocephala sensu stricto, S. famelica, and S. prolixa. Argus et al.12 note that although a small portion of the ranges of these taxa overlap, the distribution and morphology are sufficiently distinct to support separate species designations; however, genetic support for this has been lacking13.
The largest subgenus, Vetrix, are the most diverse and currently has an estimated 125 species. Only a small fraction of these is currently employed in short-rotation coppice (SRC) systems, whereby willow productivity is maximized by exploiting the propensity of shrub willow to regrow quickly after cutting the stems near the base. Over the past fifty years, willow breeding efforts in New Zealand, Sweden, the UK, and the USA have resulted in the development and deployment of both intra- and interspecific hybrids with important agronomic traits needed for SRC8,14,15,16. Yields of 4–6 oven dry tonnes (odt) ha−1 y−1 have been reported for hybrids when grown in commercial biomass production systems, while experimental plots have produced as much as 24–30 odt ha−1 y−1 in some studies17. On Prince Edward Island, the willow cultivars S. viminalis (5027) and S. dasyclados (SV1) can yield up to 18–20 odt ha−1 y−1 when planted as riparian buffer strips along watercourses18. Consequently, shrub willow are increasingly garnering attention in SRC systems for their uses in managing wetlands19, intercepting excess nutrient runoff20, remediating industrial soils and wastewater21,22, and improving the biodiversity of birds and insects23.
Despite the economic and ecological importance of willow, the development of genetic resources has lagged behind other woody species such as poplar and eucalyptus24. Nonetheless, molecular markers have been used to explore the genetic diversity in willow species, including those in a study of European S. viminalis that formed the foundations for the development of an association mapping population25. As in other tree species with well-developed genetic resources, single nucleotide polymorphisms (SNPs) have become the preferred marker as they are both easily distinguished and abundant across the genome, providing broad coverage of the species’ genetic architecture by capturing both adaptive and neutral diversity. For example, SNP markers were recently used to assess the genetic diversity of S. purpurea from Europe that has naturalized in the northeastern USA26. Genome-wide approaches are advantageous because they can simultaneously discern population structure and facilitate marker development for molecular breeding.
At Agriculture and Agri-Food Canada (AAFC), in-house willow resources are being managed to develop biomass feedstocks for bioenergy and environmental applications, which has led to the establishment of the Agriculture Canada Salix (AgCanSalix) germplasm collection. The priority has been to amass large genetically diverse base populations from which parents can be selected to develop locally adapted genotypes tailored for specific applications. Historically, progeny selection has been performed both under controlled and field environments for traits such as cold hardiness, salinity tolerance, and disease resistance. Moving forward, the objectives are to expand the breeding pool and select new cultivars with agronomic traits that maintain high biomass yields.
Willow breeding programs have already had some success in developing new cultivars optimized for SRC production. For example, the Swedish breeding pipeline has generated several high-yielding willow varieties with improved disease resistance and better frost tolerance by crossing genotypes from across Europe, most notably from central and eastern Russia27. Similar success has been achieved at Rothamsted in the UK and in upstate New York in the USA, where commercial cultivars have been established that perform well in those respective climates24,28. As such, there is every reason to believe that concerted efforts with Canadian shrub willow germplasm will help cultivate a productive short-rotation coppice industry in Canada.
Given that the AgCanSalix collection exhibits vast phenotypic diversity in terms of growth, phenology, physiology, and wood chemistry4,29,30, we anticipated that there would be considerable underlying genetic diversity to enable future breeding work. In this study, we investigated the geographic pattern of genetic diversity of S. famelica and S. eriocephala using genotyping-by-sequencing (GBS) to: (1) determine whether geoclimatic variables influenced the population structure, and (2) determine the level of genetic variation within and among populations. By uncovering the extent of diversity within the AgCanSalix collection and the relatedness of genotypes, this work will bolster future efforts to breed willow varieties with superior adaptive traits for a host of Canadian climatic regions. The ultimate aim is to develop multi-functional feedstocks that can simultaneously support a burgeoning bioenergy industry and also deliver environmental benefits.
Materials and methods
AgCanSalix collection
AAFC assembled a collection of Canada’s native willow germplasm from wild populations of S. amygdaloides, S. bebbiana, S. discolor, S. eriocephala, S. famelica, S. interior, and S. petiolaris during the summer of 2012. This collection, known as AgCanSalix, includes willow genotypes from Alberta, Saskatchewan, Manitoba, Ontario, Quebec, New Brunswick, Prince Edward Island, Nova Scotia, and Newfoundland and Labrador (Table S1; Fig. S1).
For S. famelica and S. eriocephala collectively, 34 sites were sampled with 15 genotypes per site of origin for a total of 510 genotypes representing the range across Canada. The geographic range of this collection spanned 10° in latitude (45–55 °N), 55° in longitude (57–113 °W), and 796 m in elevation (4–800 m). Sampling was done across a west–east transect to capture the geographic variation that these species occupy. The risk of clonal sampling was mitigated by selecting genotypes that were separated by a minimum of 1 km. The cuttings were collected during the dormancy period, bagged separately, and stored at − 4 °C. In the spring, dormant cuttings were dipped in rooting hormone powder (0.8% indole-3-butyric acid, Plant Products Co. Ltd., Brampton, Ontario, Canada) and induced to root in Spencer-Lemaire rootrainers (Beaver Plastics, Acheson, Alberta, Canada). Willow plants were grown in a greenhouse located at Indian Head, Saskatchewan (51 °N, 104 °W; elevation 605 m) with daytime and nighttime temperatures maintained at 23 and 18 °C, respectively, and a relative humidity of 40%. After two months of growth in the greenhouse, the willow plants were transferred to a shade house and allowed to undergo natural senescence. In late October, the frozen root plugs were individually bagged and stored at − 4 °C until the following spring when a common garden plot was established at Indian Head.
Geoclimatic data at sites of origin
Long-term climate normals (1981–2010) were obtained from nearby government-run weather stations operated by Environment Canada31 to provide environmental variables associated with the sites of origin, including frost-free days (FFDs, number of days), mean annual precipitation (MAP, mm), mean summer temperature (MST, °C), and growing degree days > 5 °C (GDD, °C) (Table S1). We define frost-free days as the number of days with a minimum temperature above 0 °C, a proxy for growing season length. The MAP range for the sites of origin varied markedly from 316 to 1709 mm. The MST ranged from 9.4 to 15.5 °C, values of FFD varied from 151 to 210 days, and the GDD fluctuated from 1023 to 1955 °C.
DNA extraction and genotyping-by-sequencing (GBS)
For this study on the genetic diversity of the AgCanSalix collection, 324 genotypes were selected (8–10 genotypes per site of origin, from 33 sites of origin). Young leaf tissues from the common garden were collected in Eppendorf tubes, immediately flash-frozen in liquid nitrogen, and stored at − 80 °C.
DNA was isolated from frozen leaf tissue using the CTAB method32 and quantified using a PicoGreen dsDNA kit (Molecular Probes, Life Technologies Inc., Burlington, Ontario, Canada). Library generation was based on the 96-plex GBS protocol described by Poland et al.33, employing a double-digest with PstI and MspI as the restriction enzymes. The resulting libraries were then sequenced using an Illumina HiSeq 2000 system (Illumina Inc., San Diego, California, USA).
Putative GBS markers were identified following the TASSEL 5.0 GBSv2 Discovery Pipeline34, using the Salix purpurea v1.0 genome35 as a mapping reference. This resulted in an initial dataset comprising 55,453 putative markers. Quality filtering and missing marker imputation were completed using the synbreed package36 in R version 3.5.137. First, markers were recoded to reflect the number of copies of a reference allele: aa = 0, Aa = 1, and AA = 2. Then, markers with a minor allele frequency (MAF) below 1% and > 50% missing data were removed. Missing markers (16.2%) were imputed using the random method in the codeGeno function, in which the missing values were sampled from the existing marginal allele distribution. This resulted in a final filtered set of 26,016 polymorphic markers for 324 genotypes. Among these markers, the highest number of genotypes with missing data was 162 out of 324 total genotypes. However, most of the markers had far fewer genotypes requiring imputation. The mean number of markers with imputed data was 52 out of 324 genotypes, with a standard deviation of 49.
Data analysis
To gain preliminary insights into population structure, a principal component analysis (PCA) was carried out using the glPca function in the R package adegenet38. Genotypes were broadly classified as either western or eastern, based on the reported ranges for S. famelica and S. eriocephala discerned by examining plant morphology39.
The population structure was assessed using a Bayesian approach for posterior inference implemented in fastSTRUCTURE40. The default convergence threshold was selected and runs were performed for a number of clusters ranging from K = 1 to K = 10 using all 324 genotypes. The output was then visualised using Distruct for many K’s as implemented in CLUMPAK41. The optimal value of K was determined using the chooseK.py algorithm, from which K = 4 was identified as the number of model components that maximized marginal likelihood, and K = 5 was the number of model components used to explain structure in data. The R package ggplot242 was then used to plot the average membership across fastSTRUCTURE runs at K = 5.
Pie charts representing admixture results for K = 5 were generated by the R package scatterpie43 and were plotted onto a geographical map of Canada imported from the R package rnaturalearth44 using R packages ggplot242 and ggspatial45. The species ranges proposed by Argus39 were added manually onto this map using Adobe Illustrator (Adobe Inc., San Jose, CA, USA).
For comparison, another evaluation of the population structure was carried out by performing a discriminant analysis of principal components (DAPC) using the dapc function in the R package adegenet38. Here, the optimal number of clusters, K, was determined for the 324 genotypes sourced from 33 sites of origin using the find.clusters function in the R package adegenet38. K = 5 was inferred by applying the Elbow method, a heuristic commonly used to manually evaluate the number of clusters as the point of inflection on a plot of the Bayesian information criterion (BIC) values against sequential numbers of clusters (K values).
To evaluate genetic diversity, the allelic frequency (p and q), MAF, expected heterozygosity (HE), observed heterozygosity (HO), Nei’s gene diversity (GD), polymorphism informative content (PIC), and the χ2 statistic for the Hardy–Weinberg equilibrium test and its corresponding P-value were calculated for each cluster using the R package snpReady46. To determine the level of population genetic differentiation, pairwise fixation indexes (FST) for each cluster were calculated based on the method of Weir and Cockerham47, using the R package StAMPP48.
The correlation between genetic distance (pairwise FST values) and geographic distance among 33 sites of origin was assessed by performing an isolation-by-distance (IBD) analysis using the R package adegenet49. The significance of the associations was tested based on a Mantel test with 10,000 permutations using the mantel.randtest function. Local density was plotted with two-dimensional kernel density estimations determined using the R package MASS50.
To calculate the extent of hierarchical population structure, an analysis of molecular variance (AMOVA) was performed based on the infinite alleles model (F-statistics) using the R package poppr51. The AMOVA partitioned variation among geographic clusters, between genotypes within clusters, and within genotypes.
To examine the influence of climate at the sites of origin on genetic structure, a redundancy analysis (RDA) was conducted using the R package vegan52 following the methodology described by Forester et al.53 and using the cluster assignments from the Bayesian analysis.
Plots were formatted using Adobe Illustrator (Adobe Inc., San Jose, CA, USA).
Results
Population structure
From the set of 324 S. famelica and S. eriocephala genotypes, 26,016 polymorphic markers were obtained that were distributed evenly across all 19 chromosomes. Using this dataset, we set out to characterize the AgCanSalix collection by examining the population structure with PCA (Fig. 1). Genotypes were assigned as either western or eastern based on the reported ranges determined previously from plant morphology39. The first two factors from this analysis explained 9.8% of the total variation: PC1, which explained 7.7% of the variation and corresponded to a longitudinal divide, and PC2, which explained 2.1% of the variation and reflected more complex contributions of biogeography. Many of the eastern genotypes formed a compact cluster, although some occurred separately, including those from Newfoundland and Labrador. The majority of western genotypes grouped loosely, while those from Alberta formed a compact cluster. Some of the genotypes from Ontario, which were initially assigned to the eastern group, clustered with western genotypes while others were more scattered, indicating that Ontario genotypes may represent a transition between the western and eastern groups.
Principal component analysis (PCA) showing 324 genotypes of S. famelica and S. eriocephala, plotted as western and eastern groups, separated along the first two principal component (PC) axes (explaining 7.7% and 2.1% of the total variation, respectively). AB, Alberta; SK, Saskatchewan; MB, Manitoba; ON, Ontario; QC, Quebec; NB, New Brunswick; NS, Nova Scotia; PEI, Prince Edward Island; NL, Newfoundland and Labrador.
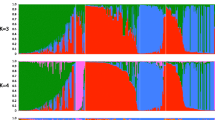
Next, we used fastSTRUCTURE to further assess the population structure and evaluate genetic variation within the collection. This analysis revealed the existence of five clusters, based on the optimal model complexity (Fig. 2A). The proportion of each individual genotype’s Bayesian assignment contributing to each cluster (K = 5) is depicted in Fig. 2B. As these largely corresponded to geographic areas, we assigned the following cluster names: Far West (orange), West (yellow), Central (pink), East (blue), and Far East (green). Far West included 68 genotypes from Alberta; West included 128 genotypes from sites in Saskatchewan, Manitoba, and northwestern Ontario; Central included 28 genotypes from central Ontario; East included 92 genotypes from sites in Quebec, New Brunswick, Nova Scotia, Prince Edward Island, and Newfoundland and Labrador; and the final cluster, Far East, had two genotypes from Ontario, one from Quebec, and five from Newfoundland and Labrador. For comparison, the results of fastSTRUCTURE runs for K = 2 to K = 10 are provided as Fig. S2. Overall, the genotypes from Ontario and Newfoundland and Labrador displayed the greatest admixture, while those from Alberta, New Brunswick, and Nova Scotia had the least admixture.
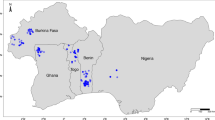
Inference of population structure based on 26,016 polymorphisms in 324 genotypes of S. famelica and S. eriocephala from 33 sites of origin using Bayesian simulations. (A) Pies show mean admixture results (K = 5) across genotypes for each site of origin. Shaded areas indicate the species distributions reported in Argus39. Map generated using R packages ggplot242, scatterpie43, rnaturalearth44, and ggspatial45. (B) Distribution of fastSTRUCTURE-defined clusters among 324 genotypes, sorted from west to east, for K = 5 where the y-axis indicates the estimated admixture coefficients for each genotype.
As a further confirmation of the population structure, a discriminant analysis of principal components (DAPC) was performed (Fig. 3A). This analysis was conducted by retaining the first 200 principal components with the first two linear discriminants accounting for 77.3% of the cumulative variance (Fig. 3B, C). In agreement with the Bayesian analysis, this method identified the optimal number of clusters to be K = 5 (Fig. S3). Again these clusters were reflective of geography such that the same five names were assigned: Far West, West, Central, East, and Far East. The membership of these groups was identical to those in the Bayesian analysis except that five genotypes from Alberta fell into the West cluster rather than Far West. As was observed with the PCA analysis, the western genotypes grouped together, apart from the eastern genotypes. Strikingly, this analysis showed that the Far East cluster was by far the most divergent of all the clusters, based on the minimum-spanning tree which represents the between-group differentiation.
(A) Discriminant analysis of principal components (DAPC) scatterplot showing positions of 324 S. famelica and S. eriocephala genotypes along the first two linear discriminants for K = 5. Each dot represents a genotype and a minimum spanning tree (black line) based on the squared distances between the inferred geographic clusters. (B) Principal component analysis (PCA) eigenvalues plot shows that 77.3% of the cumulative variance can be explained by the first 200 retained principal components (PCs). (C) Discriminant analysis eigenvalues plot shows the proportion of total variance captured by the first two linear discriminants.
Genetic diversity
Substantial levels of genetic polymorphism were observed among the collection of S. famelica and S. eriocephala genotypes (Table 1). For the five clusters, the expected heterozygosity ranged from 0.161 to 0.189, while the observed heterozygosity ranged from 0.125 to 0.162. Nei’s gene diversity (GD) and mean PIC were 0.179 and 0.150 for the collection, respectively, while the MAF ranged from 0.112 to 0.133 with an overall MAF of 0.121 across the entire collection. The conditions of Hardy–Weinberg equilibrium were not met for all five clusters.
Next, we performed pairwise FST comparisons between the five clusters, with the lowest FST observed between Far East and East (− 0.049, Table 2). The highest FST was observed between East and Central (0.097), followed closely by West and Central (0.090). This could again be an indication that the Central genotypes represent a transition zone from west to east. Alternatively, the genetic diversity across this region could be underestimated due to the relatively low number of genotypes and sites of origin included from Ontario and southern Quebec. Significant isolation-by-distance was evident across the range, and the correlation between geographic distance and FST was high (r = 0.887, p = 0.001, Fig. S4).
The hierarchical AMOVA revealed that 8.2% of the genetic variance existed among the five clusters identified by the Bayesian analysis (Table 3). Greater proportions of genetic polymorphisms were observed between the clusters (22.9%) and within genotypes (68.9%). As further support for these conclusions, the highest phi-statistic values resulted for the variance between clusters and within genotypes. However, as a caveat, only the analysis of variance within genotypes was associated with a significant p-value (< 0.01).
Influence of geoclimatic variables
RDA analysis was performed in an effort to tease apart the impact of geoclimatic variables on population structure. The first three constrained axes explained 86.4% of the total variance. RDA1 (x-axis, Fig. 4A, B) accounted for 61% of the variation and corresponded to a west–east divide of clusters. Genotypes from Central, East, and Far East were positively correlated to mean annual precipitation (MAP) and frost-free days (FFD). By contrast, genotypes from the West and Far West clusters were sourced from higher elevations (ELEV) and were characterized by higher mean summer temperatures (MST). RDA2 (y-axis, Fig. 4A) explained 16.8% of the variation and revealed a positive relationship for Central genotypes with growing degree days (GDD). RDA3 (y-axis, Fig. 4B) revealed considerable separation within each cluster, particularly for the East and the Far East clusters, that was largely correlated with GDD and MST. Overall, this analysis showed that MAP, FFD, and ELEV were strongly related to the population structure and genetic diversity observed among the S. famelica and S. eriocephala genotypes.
Redundancy analysis (RDA) ordination plots showing the relationship between population structure and environmental variables at the sites of origin along (A) RDA axes 1 and 2 (explaining 77.8% of the total variation), and (B) axes 1 and 3 (explaining 69.6% of the total variation). The dark grey cluster at the centre of each plot represents the 26,016 polymorphisms, while each dot represents a genotype (324 in total), coloured by geographic cluster as shown in the legend. Blue arrows represent five environmental variables: mean annual precipitation (MAP), frost-free days (FFD), growing degree days (GDD), mean summer temperature (MST), elevation (ELEV).
Discussion
Distribution of Canadian shrub willow
To our knowledge, this is the most comprehensive range-wide genetic study of S. famelica and S. eriocephala to date. This study examined 324 willow genotypes sourced from an expansive geographic range (~ 4000 km) spanning much of the temperate and boreal zones of Canada. Our results reveal that the AgCanSalix collection falls into a clear pattern of western and eastern genotypes and simultaneously provides genetic evidence to support separate designations for S. famelica and S. eriocephala.
The contemporary distribution of these taxa has been shaped by both ongoing and historical processes. As the majority of the present-day range of these species was covered by the Laurentide Ice Sheet during the Pleistocene epoch54, the Canadian genotypes likely arose following northward expansion from glacial refugia. Notably, Allard and Leonard55 described disjunct S. eriocephala populations as far south as Georgia and Alabama in the USA. Following the northward range expansion, there may have been physical and/or climatic barriers that contributed to genetic isolation and divergence. Furthermore, it is likely that willow habitats have been drastically altered by centuries of human colonization and extensive changes to the landscapes across North America56.
In this study, willow genotypes sourced from broad natural ranges were examined using Bayesian and principal component analyses to elucidate population structure. Both approaches captured five clusters corresponding to five geographic areas. At present, the AgCanSalix collection does not include genotypes from west or north of the Rocky Mountains. Further work in this region is needed to examine whether any additional western cluster(s) may exist or whether these genotypes would group with those from Alberta in the Far West cluster. There is reason to believe that such analyses could capture additional clusters as salicologists have previously described shrub willow west of the Rocky Mountains and in the northwestern boreal region as a separate species, S. prolixa (previously S. eriocephala ssp. mackenzieana)12. Moreover, in this scenario, the Alberta genotypes in the Far West cluster could perhaps represent a transition between S. prolixa and S. famelica.
After the last glacial retreat, range expansion and recolonization patterns may have contributed to the modern-day regional population structure in S. famelica and S. eriocephala. In addition, the barrier of the Great Lakes and the Canadian Shield along the longitudinal axis could plausibly limit gene flow between western and eastern populations. However, more genotypes from areas north of the Great Lakes throughout Ontario and southern Quebec are needed to validate this hypothesis. Evident genetic discontinuities along latitudinal and longitudinal transects have previously been observed for other North American tree and shrub species, including Pinus contorta, Salix melanopsis, Populus trichocarpa, and Populus balsamifera57,58,59,60.
With the evidence at hand, it appears that germplasm collected from Ontario may represent a zone of transition between western (S. famelica) and eastern (S. eriocephala) genotypes. Of particular note are the genotypes from Batchawana Bay in Ontario in which significant admixture was observed with genotypes from the West, Central, East, and even Far East clusters. The genotypes from Newfoundland and Labrador are also noteworthy as they appear to be the most divergent, an observation which conforms with decades of research on the genetic isolation of Newfoundland flora61.
Genetic variation within the AgCanSalix collection
High recombination rates among outcrossing and sexually propagated plant species generally result in high genetic diversity. Among the genotypes sampled in this study, the observed heterozygosity (HO) ranged from 0.125 to 0.162 for the five clusters that were identified by Bayesian analysis. Comparable levels of HO were reported among 45 genotypes of S. eriocephala collected from Ontario (mean HO = 0.136;62). Conversely, 58 genotypes collected from Quebec and New Brunswick exhibited higher heterozygosity (mean HO = 0.592;63). However, it is difficult to directly compare SNP-based studies on genetic diversity, such as ours, with earlier techniques.
Generally, levels of genetic differentiation are relatively lower among wind-pollinated taxa than for insect-pollinated species. Shrub willow are mainly, but not exclusively, insect-pollinated64. Within the AgCanSalix collection, we observed an overall FST = 0.024. An earlier microsatellites analysis by Lin et al.65 reported a slightly higher degree of differentiation (FST = 0.055) for 416 genotypes of S. eriocephala collected from three watersheds in upstate New York, USA. S. silicicola, a willow species endemic to the sand dunes of northern Saskatchewan, also displayed a higher FST = 0.156 in a study of 204 genotypes66.
The Far East cluster showed the highest levels of heterozygosity and also had the lowest FST values in comparison with all other clusters. This observation challenges the results of the population structure analyses which suggested that this cluster is the most divergent. It could be that the smaller sample size in this cluster is the source of the high level of heterozygosity. These contrasting observations bring into question the validity of the Far East cluster, particularly given the vast geographic distances between many of these genotypes. Of the 30 genotypes from Newfoundland, only five (from Stephenville) grouped in the Far East cluster, along with two genotypes from Ontario (Batchawana Bay and Kenora), and one from Quebec (Quebec City). Further analyses of the Far East genotypes and of additional germplasm from Newfoundland are needed to assess the extent of genetic differentiation in this region.
Analysis of isolation-by-distance revealed that genetic diversity was strongly correlated with the geographic distance between genotypes. However, the AMOVA revealed that the greatest variance occurred within genotypes (68.9%), while variance among the five clusters represented only 8.2%, indicating that S. famelica and S. eriocephala are highly outcrossing species. Previous molecular genetics studies on S. eriocephala have also reported that, although there exists a high level of allelic diversity, the majority of genetic variation occurs within populations and subpopulations rather than between populations62,65. Keller et al.60 analysed a collection of Populus balsamifera, a species with a comparably extensive range, and similarly found a low variance (4.4%) among groups. Such patterns of genetic diversity offer both challenges and opportunities for breeding efforts since highly heterozygous outcrossing populations can exhibit diverse phenotypes in crossings, but it can be difficult to tease apart adaptive alleles of interest from the abundant neutral genetic diversity.
Biogeography of Salix famelica and Salix eriocephala
A redundancy analysis that asked whether biogeographic factors can help explain some of the genetic diversity and population structure pointed to precipitation, frost tolerance, and summer temperatures. This is hardly surprising given the vastly different climates that occur across the west–east spatial separation of the clusters. A single willow plant produces thousands of small seeds which lack storage reserves and require suitably mesic sites for establishment. Since both S. famelica and S. eriocephala rely on wind for long-distance pollen and seed dispersal67, extensive gene flow could theoretically occur between distant populations. More locally, shrub willow are prone to extensive clonal reproduction since broken twigs and branches frequently root and take hold nearby68.
While western sites of origin in this study receive less annual precipitation compared to eastern sites, they are also exposed to more extreme summer and winter temperatures. Higher precipitation coupled with high soil moisture offer biologically suitable habitats for seed germination and seedling establishment. On average, the sites of origin from the Central, East, and Far East clusters received two-and-a-half times more precipitation annually (1,171 mm) compared to West and Far West sites of origin (468 mm). Labrecque and Teodorescu69 found that the growth of shrub willow in southern Quebec was evidently more limited by precipitation than by summer temperatures. On the other hand, western genotypes which occur in more continental climes may be better adapted to hotter, drier conditions in spring and summer and could be more resistant to frost damage in winter. Western sites of origin had MST values ranging from 11.2 to 15.5 °C and eastern sites ranged from 9.4 to 15.4 °C, while the average number of FFD was 189 for the eastern sites of origin compared to 166 for western sites. It is highly plausible that the west–east differences in climate contributed to the divergence of S. famelica and S. eriocephala. Of course, the willow genotypes examined in this study occur across gradients of climate and geography such that genotypes in the Central cluster may be adapted to intermediate conditions between the western and eastern regions.
Although geoclimatic variables clearly influenced the diversification of Canadian shrub willow, the enduring question is whether genetics trump environmental conditions with regards to performance in SRC and environmental applications. While the growth and yield of willow clones showed strong genetic-by-environmental interactions in common garden coppicing trials, S. eriocephala genotypes consistently performed the best on more productive sites70. However, in a study on the effectiveness of willow in phytoremediation of contaminated soils from industrial sites, it was observed that genetics accounted for the majority of performance differences71. Accordingly, it will be vital to account for the effects of local growing conditions when testing the performance of novel cultivars by conducting breeding trials in common garden experiments.
Conclusions
Opportunities abound to breed advanced willow cultivars by exploiting the expansive genetic diversity that exists across the ranges of S. famelica and S. eriocephala. One of the goals in establishing the AgCanSalix collection was to identify traits that influence climate adaptation, particularly with regards to drought, frost, and thermal stresses. Given that these are evidently key drivers for diversity in S. famelica and S. eriocephala, it should be eminently feasible to select for such traits in controlled crosses. Ultimately, the aim is to develop cultivars that are better suited to warmer and drier environments, particularly in the face of rapid global climate change.
In many ways, the use of Canadian shrub willow in SRC and environmental applications is still in its infancy. S. eriocephala has been promoted for bioenergy production as it is known to produce good yields of high-quality biomass4,72. Future breeding work with the AgCanSalix collection will seek to prioritize traits for high productivity in SRC. There have already been some successes in identifying genotypes of S. eriocephala for greater biomass yields73 and for phytoremediation applications as well74. However, the breeding work to date has been limited to a relatively small number of genotypes, just a fraction of those available in the AgCanSalix collection.
This study helps lay the groundwork for future controlled crossing experiments. Locally adapted genotypes that are more resilient to environmental and biotic stresses could be selected by exploiting the allelic diversity of the AgCanSalix collection. For example, genotypes with improved resistance to insect herbivores were readily identified among a collection of S. eriocephala hybrids75. However, the challenge of selecting and maintaining such adaptive traits from highly heterozygous outcrossing populations must not be underestimated76. The next steps will be to identify candidate genotypes for breeding in order to harness the inherent diversity of Canadian shrub willow and advance efforts to develop elite cultivars.
Data availability
The datasets generated during and/or analysed during the current study are available in the Dryad repository, https://doi.org/10.5061/dryad.905qfttp7.
References
Newsholme, C. Willows: The Genus Salix (Timber Press Inc., 1992).
Hubbard, W. F. The Basket Willow (United States Department of Agriculture, 1904).
Erichsen-Brown, C. Use of plants for the past 500 years (Breezy Creeks Press, 1979).
Murphy, E. K., Mottiar, Y., Soolanayakanahally, R. Y. & Mansfield, S. D. Variations in cell wall traits impact saccharification potential of Salix famelica and Salix eriocephala. Biomass Bioenergy 148, 106051 (2021).
Kuzovkina, Y. A. & Quigley, M. F. Willows beyond wetlands: uses of Salix L. species for environmental projects. Water Air Soil Pollut. 162, 183–204 (2005).
Willebrand, E. & Verwijst, T. Population dynamics of willow coppice systems and their implications for management of short-rotation forests. For. Chron. 69, 699–704 (1993).
Volk, T. A. et al. The development of short-rotation willow in the northeastern United States for bioenergy and bioproducts, agroforestry and phytoremediation. Biomass Bioenergy 30, 715–727 (2006).
Kopp, R. F. et al. The development of improved willow clones for eastern North America. For. Chron. 77, 287–291 (2001).
Argus, G. W. Infrageneric classification of Salix (Salicaceae) in the new world. Syst. Bot. Monogr. 52, 1–121 (1997).
Argus, G. W. Salix L. in Flora of North America North of Mexico, Volume 7. Magnoliophyta: Salicaceae to Brassicaceae (ed. Flora of North America Editorial Committee) 23–51 (Oxford University Press, 2010).
Dorn, R. D. A taxonomic study of Salix section Cordatae subsection Luteae (Salicaceae). Brittonia 47, 160–174 (1995).
Argus, G. W., Eckenwalder, J. E. & Kiger, R. W. Salicaceae: Flora of North America (Oxford University Press, 2010).
Lauron-Moreau, A., Pitre, F. E., Argus, G. W., Labrecque, M. & Brouillet, L. Phylogenetic relationships of American willows (Salix L., Salicaceae). PLoS ONE 10, e0121965 (2015).
Gullberg, U. Towards making willows pilot species for coppicing production. For. Chron. 69, 721–726 (1993).
Lindegaard, K. N. & Barker, J. H. A. Breeding willows for biomass. Asp. App. Biol. 49, 155–162 (1997).
Smart, L. B. & Cameron, K. D. Genetic improvement of willow (Salix spp.) as a dedicated bioenergy crop. In Genetic Improvement of Bioenergy Crops (ed. Vermerris, W.) (Springer, 2008).
Keoleian, G. A. & Volk, T. A. Renewable energy from willow biomass crops: life cycle energy, environmental and economic performance. Crit. Rev. Plant Sci. 24, 385–406 (2005).
Schroeder, W. R., Murray, B. J., Poppy, L., & Svendsen, E. Willow riparian buffers. AAFC Factsheet: AAFC No. 12433E. http://publications.gc.ca/collections/collection_2016/aac-aafc/A22-12433-2015-eng.pdf (2015).
Schroeder, W. R., Kort, J., Savoie, P. & Preto, F. Biomass harvest from natural willow rings around prairie wetlands. Bioenergy Res. 2, 99–105 (2009).
Da Ros, L. M., Soolanayakanahally, R. Y., Guy, R. D. & Mansfield, S. D. Phosphorus storage and resorption in riparian tree species: environmental applications of poplar and willow. Environ. Exp. Bot. 149, 1–8 (2018).
Mosseler, A. & Major, J. E. Phytoremediation efficacy of Salix discolor and S. eriocephela on adjacent acidic clay and shale overburden on a former mine site: growth, soil, and foliage traits. Forests 8, 475 (2017).
Bilek, M. A., Soolanayakanahally, R. Y., Guy, R. D. & Mansfield, S. D. Physiological response of Populus balsamifera and Salix eriocephala to salinity and hydraulic fracturing wastewater: potential for phytoremediation applications. Int. J. Environ. 17, 7641 (2020).
Mushanski, M. D. Habitat selection by birds in willow-ringed wetlands: management implications for harvesting willow biomass. MSc thesis, University of Regina (2015).
Karp, A. et al. Genetic improvement of willow for bioenergy and biofuels. J. Integr. Plant Biol. 53, 151–165 (2011).
Hanley, S. J. et al. A genetic linkage map of willow (Salix viminalis) based on AFLP and microsatellite markers. Theor. Appl. Genet. 105, 1087–1096 (2002).
Gouker, F. E., DiFazio, S. P., Bubner, B., Zander, M. & Smart, L. B. Genetic diversity and population structure of native, naturalized, and cultivated Salix purpurea. Tree Genet. Genomes 15, 47 (2019).
Larson, S. Genetic improvement of willow for short-rotation coppice. Biomass Bioenergy 15, 23–26 (1998).
Smart, L. B. et al. Genetic improvement of shrub willow (Salix spp.) crops for bioenergy and environmental applications in the United States. Unasylva 56, 51–55 (2005).
Shunmugam, A. S. K., Soolanayakanahally R. Y. S. & Guy R. D. Geo-climatic gradient shapes functional trait variations in Salix eriocephala Michx. bioRxiv. https://doi.org/10.1101/057836 (2016)
Hu, Y., Guy, R. D. & Soolanayakanahally, R. Y. Genotypic variation in C and N isotope discrimination suggests local adaptation of heart-leaved willow. Tree Physiol https://doi.org/10.1093/treephys/tpab010 (2021).
Environment Canada, Canadian climate normals. https://climate.weather.gc.ca/climate_normals/index_e.html (2021).
Doyle, J. J. & Doyle, J. L. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem. Bull. 19, 11–15 (1987).
Poland, J. A., Brown, P. J., Sorrells, M. E. & Jannink, J. L. Development of high-density genetic maps for barley and wheat using a novel two-enzyme genotyping-by-sequencing approach. PLoS ONE 7, 32253 (2012).
Glaubitz, J. C. et al. TASSEL-GBS: a high capacity genotyping by sequencing analysis pipeline. PLoS ONE 9, e90346 (2014).
DOE-JGI, Salix purpurea v1.0. http://phytozome.jgi.doe.gov/pz/portal.html#!info?alias=Org_Spurpurea (2015).
Wimmer, V., Albrecht, T., Auinger, H.-J. & Schön, C.-C. synbreed: a framework for the analysis of genomic prediction data using R. Bioinformatics 28, 2086–2087 (2012).
R Core Team. R: a language and environment for statistical computing. R Foundation for Statistical Computing (2019).
Jombart, T., Devillard, S. & Balloux, F. Discriminant analysis of principal components: a new method for the analysis of genetically structured populations. BMC Genet. 11, 94 (2010).
Argus, G. W. Salix (Salicaceae) distribution maps and a synopsis of their classification in North America, north of Mexico. Harv. Pap. Bot. 12, 335–368 (2007).
Raj, A., Stephens, M. & Pritchard, J. K. fastSTRUCTURE: variational inference of population structure in large SNP data sets. Genetics 197, 573–589 (2014).
Kopelman, N. M., Mayzel, J., Jakobsson, M., Rosenberg, N. A. & Mayrose, I. Clumpak: a program for identifying clustering modes and packaging population structure inferences across K. Mol. Ecol. Resour. 15, 1179–1191 (2015).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer, 2016).
Guangchuang, Y. scatterpie: Scatter Pie Plot (2021).
South, A. rnaturalearth: World Map Data from Natural Earth (2017).
Dunningtonn, D. ggspatial: Spatial Data Framework for ggplot2 (2018).
Granato, I. S. C. et al. snpReady: a tool to assist breeders in genomic analysis. Mol. Breed. 38, 102 (2018).
Weir, B. S. & Cockerham, C. C. Estimating F-statistics for the analysis of population structure. Evolution 38, 1358–1370 (1984).
Pembleton, L. W., Cogan, N. O. I. & Forster, J. W. StAMPP: an R package for calculation of genetic differentiation and structure of mixed-ploidy level populations. Mol. Ecol. Resour. 13, 946–952 (2013).
Jombart, T. adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics 24, 1403–1405 (2008).
Venables, W. N. & Ripley, B. D. Modern applied statistics with S 4th edn. (Springer, 2002).
Kamvar, Z. N., Tabima, J. F. & Grünwald, N. J. Poppr: an R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. PeerJ 2, e281 (2014).
Oksanen, J. et al. vegan: Community Ecology Package (2019).
Forester, B. R., Lasky, J. R., Wagner, H. H. & Urban, D. L. Comparing methods for detecting multilocus adaptation with multivariate genotype-environment associations. Mol. Ecol. 27, 2215–2233 (2018).
Dyke, A. S. & Prest, V. K. Late wisconsinan and holocene history of the laurentide ice sheet. Géogr. Phys. Quat. 41, 237–263 (1987).
Allard, H. A. & Leonard, E. C. The Canaan and the Stony river valleys of West Virginia, their former magnificent spruce forests, their vegetation and floristics today. Castanea 17, 1–60 (1952).
Argus, G. W. Classification of Salix in the new world. Bot. Electron. News 227, 1–6 (1999).
Wheeler, N. C. & Guries, R. P. Biogeography of lodgepole pine. Can. J. Bot. 60, 1805–1814 (1982).
Brunsfeld, S. J., Miller, T. R. & Carstens, B. C. Insights into the biogeography of the Pacific Northwest of North America: evidence from the Phylogeography of Salix melanopsis. Syst. Bot. 32, 129–139 (2007).
Xie, C.-Y., Ying, C. C., Yanchuk, A. D. & Holowachuk, D. L. Ecotypic mode of regional differentiation caused by restricted gene migration: a case in black cottonwood (Populus trichocarpa) along the Pacific Northwest coast. Can. J. For. Res. 39, 519–525 (2009).
Keller, S. R., Olson, M. S., Silim, S., Schroeder, W. R. & Tiffin, P. Genomic diversity, population structure, and migration following rapid range expansion in the Balsam poplar, Populus balsamifera. Mol. Ecol. 19, 1212–1226 (2010).
Fernald, M. L. Recent discoveries in the Newfoundland flora. Rhodora 35, 364–386 (1933).
Aravanopoulos, F. A., Kim, K. H. & Zsuffa, L. Genetic diversity of superior Salix clones selected for intensive forestry plantations. Biomass Bioenergy 16, 249–255 (1999).
Lauron-Moreau, A., Pitre, F. E., Brouillet, L. & Labrecque, M. Microsatellite markers of willow species and characterization of 11 polymorphic microsatellites for Salix eriocephala (Salicaceae), a potential native species for biomass production in Canada. Plants 2, 203–210 (2013).
Argus, G. W. An experimental study of hybridization and pollination in Salix (willow). Can. J. Bot. 52, 1613–1619 (1974).
Lin, J., Gibbs, J. P. & Smart, L. B. Population genetic structure of native versus naturalized sympatric shrub willows (Salix; Salicaceae). Am. J. Bot. 96, 771–785 (2009).
Purdy, B. G. & Bayer, R. J. Allozyme variation in the athabasca sand dune endemic, Salix silicicola, and the closely related widespread species, S. alaxensis. Syst. Bot. 20, 179–190 (1995).
Richardson, J., Isebrands, J. G. & Ball, J. B. Ecology and physiology of poplars and willows. In Poplars and Willows: Trees for Society and the Environment (eds Isebrands, J. G. & Richardson, J.) 92–123 (CAB International and Food and Agriculture Organization of the United Nations Rome, 2014).
Raven, J. A. The physiology of Salix, in Willow Symposium Proceedings of The Royal Society of Edinburgh, Vol. 98 (eds. Watling, R., Raven, J. A.) 49–62 (1992).
Labrecque, M. & Teodorescu, T. I. High biomass yield achieved by Salix clones in SRIC following two 3-year coppice rotations on abandoned farmland in southern Quebec, Canada. Biomass Bioenergy 25, 135–146 (2003).
Mosseler, A., Major, J. E. & Labrecque, M. Genetic by environment interactions of two North American Salix species assessed for coppice yield and components of growth on three sites of varying quality. Trees 28, 1401–1411 (2014).
Grenier, V., Pitre, F. E., Nissim, W. G. & Labrecque, M. Genotypic differences explain most of the response of willow cultivars to petroleum-contaminated soil. Trees 29, 871–881 (2015).
Mosseler, A., Zsuffa, L., Stoehr, M. U. & Kenney, W. A. Variation in biomass production, moisture content, and specific gravity in some North American willows (Salix L.). Can. J. For. Res. 18, 1535–1540 (1998).
Cameron, K. D. et al. Quantitative genetics of traits indicative of biomass production and heterosis in 34 full-sib F1 Salix eriocephala families. Bioenerg. Res. 1, 80–90 (2008).
Huang, X., Soolanayakanahally, R. Y., Guy, R. D., Shunmugam, A. S. K. & Mansfield, S. D. Differences in growth, and physiological and metabolic responses among Canadian native and hybrid willows (Salix spp.) under salinity stress. Tree Physiol. 40, 652–666 (2020).
Hochwender, C. G. & Fritz, R. S. Plant genetic differences influence herbivore community structure: evidence from a hybrid willow system. Oecologia 138, 547–557 (2004).
Hanley, S. J. & Karp, A. Genetic strategies for dissecting complex traits in biomass willows (Salix spp.). Tree Physiol. 34, 1167–1180 (2014).
Acknowledgements
This work was supported with funding from Agriculture and Agri-Food Canada to E.K.M. and R.Y.S, an NSERC Discovery Grant to S.D.M., and the National Agency for Scientific and Technological Promotion of Argentina (PICT 2016 1048) to E.P.C. The authors thank Lily Tang and Yogendra Khedikar at the Saskatoon Research and Development Centre for their technical support in sequencing. The authors would also like to thank Angela Karp, Steve Hanley, and Rachel Rossiter at Rothamsted Research for their contributions to earlier microsatellite marker work which helped inform the current study. We gratefully acknowledge Chris Stefner, Don Reynard, and Garth Inouye for their help in establishing the AgCanSalix collection, Lawrence Smart for the use of the unpublished Salix purpurea v1.0 genome sequence from Cornell University, and Quentin Cronk for his valuable insights into willow taxonomy. Finally, we thank Yaseen Mottiar for his invaluable guidance and support during the preparation of this manuscript.
Author information
Authors and Affiliations
Contributions
E.K.M., R.Y.S., and S.D.M designed the research; R.Y.S. and W.R.S. developed the willow germplasm collection; I.A.P.P. performed the GBS; E.K.M., E.P.C., and Y.A.E. analyzed the data. E.K.M. and R.Y.S. drafted the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Murphy, E.K., Cappa, E.P., Soolanayakanahally, R.Y. et al. Unweaving the population structure and genetic diversity of Canadian shrub willow. Sci Rep 12, 17254 (2022). https://doi.org/10.1038/s41598-022-20498-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-20498-9
- Springer Nature Limited