Abstract
Vascular endothelial cells exhibit substantial phenotypic and transcriptional heterogeneity which is established during early embryogenesis. However, the molecular mechanisms involved in establishing endothelial cell diversity are still not well understood. Zebrafish has emerged as an advantageous model to study vascular development. Despite its importance, the single-cell transcriptomic profile of vascular endothelial cells during zebrafish development is still missing. To address this, we applied single-cell RNA-sequencing (scRNA-seq) of vascular endothelial cells isolated from zebrafish embryos at the 24 hpf stage. Six distinct clusters or subclusters related to vascular endothelial cells were identified which include arterial, two venous, cranial, endocardial and endothelial progenitor cell subtypes. Furthermore, we validated our findings by characterizing novel markers for arterial, venous, and endocardial cells. We experimentally confirmed the presence of two transcriptionally different venous cell subtypes, demonstrating heterogeneity among venous endothelial cells at this early developmental stage. This dataset will be a valuable resource for future functional characterization of vascular endothelial cells and interrogation of molecular mechanisms involved in the establishment of their heterogeneity and cell-fate decisions.
Similar content being viewed by others
Introduction
Vascular endothelial cells in different vascular beds exhibit considerable heterogeneity. This endothelial cell diversity is established during early embryogenesis when vascular endothelial cells are specified from mesodermal progenitors and differentiate into arterial, venous, capillary, lymphatic, endocardial, hemogenic and other subtypes of endothelial cells1. Despite advances in our understanding of mechanisms that govern vascular development, the extent of vascular heterogeneity and molecular mechanisms involved in its establishment are still poorly understood.
Because it is technically challenging to study embryonic vascular development in mammalian embryos, the zebrafish has emerged as an advantageous model to study early vascular development. Transparent zebrafish embryos develop externally and allow researchers to study spatiotemporal events in vivo and in real time. Furthermore, the signaling pathways and transcriptional programs that regulate early vascular development are highly conserved between zebrafish and other vertebrates2.
The advent of large-scale single cell RNA-sequencing (scRNA-Seq) in recent years has enabled robust access to the transcriptomic profile of single cells. Detailed analysis of scRNA-Seq data has revealed similarities and differences between different cell types and allowed researchers to carry out analysis of complexity and heterogeneity existing in the biological systems3,4. Numerous studies involving single-cell studies in health and disease have identified new endothelial cell subtypes and functions5,6,7,8,9,10. Several scRNA-Seq studies using zebrafish embryos have revealed cellular heterogeneity and developmental trajectory during tissue and organ development in zebrafish11. In addition, a single cell transcriptomic atlas of day one to five zebrafish embryos has been generated recently12. However, the transcriptional profile of zebrafish vascular endothelial cells, which might provide insights into vascular endothelial cell heterogeneity and cell fate decision process during zebrafish development, is still missing.
Kinase insert domain receptor like (Kdrl), also termed as flk1, is strongly expressed in developing endothelial precursors and differentiated vascular endothelial cells in zebrafish embryos13. We have previously generated Tg(kdrl:mCherry) transgenic line which faithfully recapitulates endothelial kdrl expression in zebrafish embryos14. Etv2/Etsrp, an ETS transcription factor, is one of the earliest markers of vascular progenitor cells in multiple vertebrates including zebrafish15,16,17. We have previously generated etv2ci32Gt gene trap line which has an insertion of the Gal4 transcriptional activator within the etv2 genomic locus. Furthermore, etv2ci32Gt+/−; UAS:GFP embryos are phenotypically normal and faithfully recapitulate GFP expression in vascular endothelial, myeloid and red blood cells18.
Here, we performed scRNA-seq using a combination of Tg(kdrl:mCherry) and etv2ci32Gt;UAS:GFP reporter lines to generate transcriptomic profile of vascular endothelial cells isolated from zebrafish embryos at 24 hpf (hours post fertilization) stage, when blood circulation is first initiated. We identified transcriptional profile of 6 different subtypes of vascular endothelial cells, which include vascular progenitor, arterial, two different venous, endocardial, and mixed arteriovenous subtypes. We validated our findings by characterizing novel markers for arterial, venous, and endocardial cells. We experimentally confirmed the presence of two transcriptionally different venous cell subtypes, demonstrating unexpected heterogeneity among venous endothelial cells at this early developmental stage. This data provides a valuable resource for future functional characterization of vascular endothelial cells and interrogation of molecular mechanisms involved in the establishment of their heterogeneity.
Results
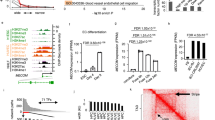
To obtain a single-cell transcriptional profile of vascular endothelial cells, we crossed the previously established Tg(kdrl:mCherry) and etv2ci32Gt; UAS:GFP lines14,18. GFP and mCherry positive embryos were collected at 24 hpf. Because the etv2ci32Gt; UAS:GFP line labels vascular endothelial and some other cell types, such as blood, we decided to collect two different cell populations to help with subsequent annotation and line characterization. Thus, GFP + mCherry + and GFP + mCherry- cell populations were isolated by FACS (fluorescent activated cell sorting) and subjected to single-cell RNA-seq analysis using the Chromium Controller (10 × Genomics) followed by the next-generation sequencing (Fig. 1A). After filtering, transcriptomes of 5275 and 5543 cells were obtained from GFP + mCherry + and GFP + mCherry- cell populations, respectively. Transcriptomes from both cell populations were pooled for further analysis. Unbiased graph-based clustering was performed using the Partek flow analysis package, and 17 distinct clusters were identified and visualized using Uniform Manifold Approximation and Projection (UMAP) approach (Fig. 1B,C). We then assigned cell identities to these clusters based on previously known marker genes which were significantly enriched in each cluster and showed specific gene expression patterns (Fig. 1B–F, Fig. S1 and Tables S1, S2). This revealed major cellular subtypes including five vascular endothelial, three red blood cell (RBC), sclerotome, two neural, macrophage, endothelial progenitor cell (EPC), epidermis, neutrophil, and myocytes. Vascular endothelial cells were mostly GFP + mCherry +, while RBC and macrophage clusters were largely GFP + mCherry− (Fig. 1E). This is consistent with known etv2 reporter expression in both endothelial and blood cells18, while kdrl:mCherry expression is largely restricted to vascular endothelial cells14.
The scRNA-seq analysis of etv2ci32Gt; UAS:GFP; kdrl:mCherry embryos at the 24 hpf stage. (A) Schematics of the experimental design for scRNA-seq. (B) UMAP plot of 10,818 cells with 17 distinct clusters. Classifications were based on previously known marker genes which were significantly enriched in each cluster. RBC, red blood cells; VE, vascular endothelial; EPC, endothelial progenitor cells. (C) UMAP plot showing the distribution of GFP + mCherry + and GFP + mCherry- cells. (D) A heatmap showing expression of top marker genes in different clusters. Enlarged heatmap and gene list is shown in Fig. S1. (E) Fraction of GFP + mCherry + and GFP + mCherry- cells within each cluster. (F) A dot plot showing the expression of selected marker genes in different clusters.
Five distinct clusters related to vascular endothelial cells including vascular progenitor, arterial, two different venous, and mixed arteriovenous subtypes were obtained, underscoring the heterogeneity of vascular endothelial cells. We discuss these clusters in greater detail below.
Identification of an arterial cluster
Top marker genes for cluster #1 included dll4, dlc, and flt1, which are known to be expressed in the arterial cells19,20,21 (Fig. 2A–E and Table S1). Many other known arterial markers, including vegfc, aqp1a.1 and hey2, were also enriched in cluster #120,22,23 (Fig. 2E and Table S1). In addition, cluster #1 also showed increased expression of unknown or previously uncharacterized genes. We hypothesized that many of these genes would be also expressed in arterial cells. Indeed, in situ hybridization (ISH) analysis of two selected previously uncharacterized genes, esm1 and notchl, demonstrated their expression in the dorsal aorta and intersegmental vessels (ISV) (Fig. 2G,H and Table S1). esm1 expression appeared enriched in the sprouting ISVs and was reduced in non-sprouting segments of the DA, suggesting further heterogeneity within the arterial population. In an attempt to gain insights into the biological processes represented by enriched genes in cluster #1, we performed Gene ontology (GO) and Pathway enrichment analysis. Top GO biological processes represented by gene set in cluster #1 included dorsal aorta morphogenesis, aorta morphogenesis, and dorsal aorta development (Fig. 2F and Table S3). Furthermore, pathway analysis revealed enrichment of Hedgehog signaling pathway, known to play an important role in arterial development24, including the regulation of size of the dorsal aorta25 (Table S4). Other enriched pathways included histamine H1 receptor signaling pathway, which is known to regulate arterial vasodilation26, as well as PI3 kinase, integrin signaling and EGF receptor signaling pathways (Table S4), which have been implicated in several steps of vascular development including angiogenesis27, vasculogenesis28 and arterial development29,30.
Arterial cluster #1. (A,B) UMAP feature plots showing expression of selected arterial top markers dll4 and flt1. (C,D) Violin plots showing expression of dll4 and flt1 in different cell populations. (E) List of top 20 marker genes differentially expressed in arterial cluster. (F) List of top 20 pathways enriched in arterial cluster. (G,H) In situ hybridization analysis at 24 hpf for selected marker genes esm1 and notchl. Note the expression of esm1 and notchl in the dorsal aorta (arrowhead) as well as the intersegmental vessels (arrow). Scale bars: 100 μm.
Identification of two subtypes of venous cells
Top marker genes for cluster #2 included lyve1b and sele, both known to be enriched in venous cells31,32 (Fig. 3A,C and Table S1). Many other known venous marker genes, such as flt4, stab1, mrc1a and stab2, were also enriched in cluster #233,34,35,36 (Fig. 3E and Table S1). In addition to known venous markers, cluster #2 included many genes with unknown or previously uncharacterized expression pattern, such as si:dkey-28n18.9 (Fig. 3B,D,E and Table S1). We hypothesized that many of these uncharacterized genes would also be expressed in venous cells. We selected two previously uncharacterized genes, bcl6b and si:dkey-28n18.9, and analyzed their expression by ISH. Consistent with our hypothesis, we observed expression of bcl6b and si:dkey-28n18.9 in the posterior cardinal vein (PCV) (Fig. 3G,H). Top GO biological processes included endothelial tube morphogenesis, blood vessel lumenization and morphogenesis of an endothelium as well as PI3 signaling and lymphangiogenesis (Fig. 3F and Table S3). Highly represented pathways included PI3 kinase, Insulin/IGF, Integrin and Ras signaling pathways (Table S4). PI3 kinase pathway has been previously implicated in arterial-venous specification30, while Insulin/IGF pathway has established roles in angiogenesis37. Other top pathways such as Integrin signalling have been shown to play a crucial role in vasculogenesis28, while Ras signaling pathway has also been implicated to regulate arterial-venous specification in zebrafish38. In addition, VEGF, FGF and Notch signaling pathways, which have established roles in vascular development and arteriovenous differentiation, were also among top pathways identified by the gene list in cluster #239,40 (Table S4).
Venous-1 cluster #2. (A,B) UMAP feature plots showing expression of selected top markers for cluster #2, lyve1b and si:dkey-28n18.9. (C,D) Violin plots showing expression of lyve1b and si:dkey-28n18.9 in different cell populations. (E) List of top 20 marker genes differentially expressed in Venous-1 cluster. (F) List of top 20 pathways enriched in Venous-1 cluster. (G,H) In situ hybridization analysis at 24 hpf for previously uncharacterized genes bcl6b and si:dkey-28n18.9. Note the expression of bcl6b and si:dkey-28n18.9 in the posterior cardinal vein (arrowheads). Scale bars: 100 μm.
Interestingly, we also identified a second cluster, cluster #7, which showed enriched expression of known venous marker genes. Top marker genes for cluster #7 included glula, known to be enriched in venous cells41 (Fig. 4A,C,E and Table S1). Other known venous markers, including mrc1a, stab2 and dab2, were also enriched in cluster #735,36,42 (Fig. 4E and Table S1). In addition to the known venous markers, top marker genes from this cluster also included many previously uncharacterized/unknown genes, such as cx30.3 and otc (Fig. 4B,D,E and Table S1). We performed ISH to confirm if these genes were expressed in venous cells in zebrafish. Indeed, both cx30.3 and otc demonstrated expression in the PCV and the caudal vein (Fig. 4G,H). cx30.3 expression was enriched in the caudal vein, and only weak expression in the PCV was apparent. In addition, cx30.3 showed strong expression in the cranial and trunk neural crest cells, while otc showed strong expression in the lens. Highly represented GO biological processes for cluster #7 included terms such as apical protein localization, endothelial tube morphogenesis, blood vessel lumenization, and morphogenesis of an endothelium (Fig. 4F and Table S3). Enriched pathways included Insulin/IGF, Hypoxia-induced factor (HIF), PI3 kinase, and Hedgehog signaling pathways (Table S4), which have been all implicated in several steps of vascular development, including angiogenesis37,43, arteriovenous specification30, and establishing vascular integrity44.
Venous-2 cluster #7. (A,B) UMAP feature plots showing expression of selected top markers for cluster #7, glula and otc. (C,D) Violin plots showing expression of glula and otc in different cell populations. (E) List of top 20 marker genes differentially expressed in Venous-2 cluster. (F) List of top 20 pathways enriched in Venous-2 cluster. (G) In situ hybridization analysis at 24 hpf for a previously uncharacterized marker gene cx30.3. Note the expression of cx30.3 in the caudal vein (black arrowheads) and weak expression in the PCV (green arrowheads). cx30.3 is also expressed in melanocytes (blue arrowheads). (H) In situ hybridization analysis at 24 hpf for a marker gene otc. Note the expression of otc in the CV, PCV (black arrowheads) and the lens (red arrowhead). Scale bars: 200 μm. (I–K) Two color fluorescent ISH analysis for the expression of lyve1b and glula at the 24 hpf stage. Note that many cells in the CV are positive for glula and not lyve1b (arrows), while cells with strong lyve1b expression in the PCV have only weak glula expression (arrowheads). Trunk and tail region is shown, anterior is to the left. PCV posterior cardinal vein, CV caudal vein. Scale bar in (K), 50 μm for (I–K).
We next asked if there was a significant biological difference between the two separate venous clusters and if they represented two distinct subsets of venous cells. To examine this, we performed fluorescent in situ hybridization for selected top markers of the two venous clusters, lyve1b (cluster #2) and glula (cluster #7), using Tg(kdrl:GFP) embryos at 24 hpf stage. There was a significant overlap between lyve1b and glula expression within the PCV in the trunk region. Intriguingly, the caudal vein (CV) displayed glula expression, while lyve1b expression was nearly absent (Fig. 4I–K). In contrast, a subset of endothelial cells within the PCV displayed strong lyve1b expression and showed very little glula expression (Fig. 4I–K), suggesting heterogeneity in the venous cell population. These results suggest that cluster #7 is enriched in the caudal vein cells. tfec, a previously characterized marker for caudal endothelial cells45, was preferentially enriched in the cluster #7 (Table S1), supporting its annotation as the caudal vein.
We have previously reported transcriptomic profiles of arterial and venous endothelial cells identified by scRNA-seq analaysis of zebrafish trunks at 30 hpf46. A substantial overlap was observed between 24 and 30 hpf datasets in both arterial and venous marker expression (Table S5). While there was only a single venous specific cluster identified at 30 hpf, its top marker genes were shared between 24 hpf Venous-1 and Venous-2 clusters, reflecting a substantial overlap between the two venous clusters and suggesting that many arterial and venous markers identified at 24 hpf continue their specific expression through later stages.
Identification of vascular endothelial cluster co-expressing both arterial and venous markers
Top marker genes for cluster #6 included cldn5b, known to be enriched in arterial vascular endothelial cells47 (Fig. 5A,C). In addition, known venous markers dab2 and lyve1b as well as pan-endothelial marker kdrl were also enriched in cluster #631,33 (Fig. 5B,D,E and Table S1). Other top markers for cluster #6 included rab11bb, cox4i2, fank1, and krt18 (krt18 had the lowest p value). Expression of these markers either has not been previously characterized or they are known to be expressed in many different cell types, including the vascular endothelium (Table S1). To identify which cells in a zebrafish embryo co-express arterial and venous markers, we performed fluorescent in situ for the arterial marker cldn5b and the venous marker dab2 in Tg(kdrl:GFP) embryos. Intriguingly, cldn5b and dab2 showed co-expression within a subset of cranial vasculature including venous primordial hindbrain channels (PHBCs) and mid cerebral vein (MCeV) at 24 and 30 hpf (Fig. 6A–F). In contrast, consistent with previous studies, expression of cldn5b and dab2 was restricted to the DA and PCV, respectively, in the trunk vasculature and did not overlap48 (Fig. 6G–I). Other cluster #6 markers cox4i2 and krt18 also showed expression in PHBCs and MCeV (Fig. 6J,K). These results suggest that cluster #6 corresponds to a subset of cranial vasculature, which shows a distinct molecular identity from other types of vascular endothelial cells.
Vascular endothelial (VE)-other cluster #6. (A,B) UMAP feature plots showing expression of selected markers for cluster #6, cldn5b and dab2. (C,D) Violin plots showing expression of cldn5b and dab2 in different cell populations. (E) List of top 20 genes differentially expressed in Vascular endothelial (VE)-other cluster. (F) List of top 20 pathways enriched in Vascular endothelial (VE)-other cluster.
Vascular endothelial (VE)-other cluster #6 corresponds to cells in cranial vasculature. (A–F) Two color fluorescent ISH analysis for the expression of cldn5b and dab2 at the 24 hpf and 30 hpf stages. Note that cldn5b and dab2 are co-expressed in the PHBC (white arrows) and MCeV (yellow arrows) at the 24 hpf and 30 hpf stages. (G–I) Two color fluorescent ISH analysis for the expression of arterial cldn5b and venous dab2 in the trunk region of 24 hpf embryos. Note that cldn5b and dab2 have distinct non-overlapping expression in the DA (arrows) and PCV (arrowheads) respectively. (J,K) Fluorescent ISH analysis for the expression of krt18 and cox4i2 in the head region of 30 hpf embryos. Both markers are expressed in the PHBC (white arrows) and MCeV (yellow arrows). PHBC primordial hindbrain channels, MCeV mid cerebral vein, DA dorsal aorta, PCV posterior cardinal vein. Scale bars: 100 μm.
Highly represented GO biological processes for cluster #6 included vascular endothelial growth factor receptor, Arp2/3 complex-mediated actin nucleation, regulation of vascular permeability and endothelium development (Fig. 5F and Table S3). Enriched pathways included histamine H2 receptor signaling pathway, implicated in vasodilation in brain capillaries49, cytoskeletal regulation by Rho GTPase, implicated in vascular integrity and tubulogenesis50,51, and angiotensin II-stimulated signaling through G proteins and beta-arrestin, known to regulate vasoconstriction and blood pressure52 (Table S4).
Identification of endothelial progenitor cells (EPC)
Top marker genes for cluster #14 included npas4l and lmo2, known to be enriched in endothelial progenitor cells33,53 (Fig. 7A,C and Table S1). Other known EPC markers, including tal1/scl and etv2, were also enriched in cluster #1415,54 (Fig. 7B,D,E and Table S1). Highly represented GO biological processes included actin filament-based transport, endothelial cell migration, vasculogenesis, non-canonical Wnt signaling pathway, endothelial cell differentiation, and artery development (Fig. 7F and Table S3). Interestingly, GO biological processes pertaining to lymph vessel development were also enriched. Enriched pathways included PDGF, integrin, TGF-beta and Wnt signaling pathways (Table S4). PDGF signaling pathway has been shown to induce proliferation, migration, and angiogenesis of EPCs55. Similarly, Integrin signaling has been shown to be a major regulator of EPC mobilization, homing, invasion and differentiation56. TGF-beta and Wnt signaling pathways have been implicated in differentiation and specification of EPCs57,58.
Endothelial progenitor cell (EPC) cluster. (A,B) UMAP plots showing expression of selected EPC top markers npas4l and etv2. (C,D) Violin plots showing expression of selected arterial marker genes npas4l and etv2 in different cell populations. (E) List of top 20 genes differentially expressed in the EPC cluster. (F) List of top 20 pathways enriched in the EPC cluster. (G) Developmental trajectory plot of vascular endothelial cells generated using Monocle 3.
To identify developmental relationship between all 5 vascular endothelial clusters, we performed developmental trajectory analysis using Monocle 359 (Fig. 7G). The trajectory analysis suggested that EPC cells transition into 3 distinct branches: an arterial and 2 venous. The two venous branches merged again, perhaps reflecting a significant overlap between the two cell populations. Venous-1 population transitioned into VE-other, which, as our results suggest, corresponds to a subset of cranial vasculature. Some VE-other cells were positioned in the vicinity of arterial population, reflecting mixed arteriovenous identity of VE-other cluster.
Identification of the endocardial cell cluster
While unsupervised clustering identified 5 different endothelial clusters, some other known endothelial cell subtypes, including endocardial cells, lymphatic progenitors, hemogenic endothelium, tip and stalk cells, were not identified using this approach. In an attempt to define additional vascular endothelial cell types, we performed subclustering using vascular endothelial cells only. However, such approach did not identify any additional meaningful clusters beyond what we have already demonstrated. Therefore, we focused on the endocardial cells and attempted to identify their transcriptional profile using the targeted clustering approach based on expression of known marker genes. Endocardial cells are known to share a large transcriptional similarity with other vascular endothelial cells33,60,61,62,63, but they also have their unique identity known by expression of marker genes such as nfatc1 and gata564,65. Recent studies have described endocardial transcriptome at 15 and 20-somite stages in zebrafish embryos48,66. However, endocardial transcriptome at 24 hpf stage has not been previously reported. Therefore, to identify the transcriptomic signature of endocardial cells, we subclustered endothelial cells based on the expression of nfatc1 and gata5 (Fig. 8A–C). This resulted in the identification of a single subcluster comprised of 21 cells. Top marker genes for this subpopulation, besides gata5, included has2 and fn1a, all known to have endocardial specific expression64,65,67,68,69 (Fig. 8D and Table S6). Known zebrafish endocardial markers klf2a and notch1b were also enriched in this cluster70,71 (Table S6). Heart-specific expression has been reported for many other genes in this cluster including id2b, tbx20, spock3, ece1 and smad6a, although it was not investigated whether this expression was endocardial or myocardial72,73,74,75. We previously reported the transcriptome of an endocardial cluster identified during scRNA-seq analysis of etv2ci32Gt; UAS:GFP embryos at the 20-somite stage (19 hpf)48. There was a significant overlap of marker genes, such as fn1a, gata5, id2b, wnt11r, bmp16 and bambib between the two endocardial clusters (Table S7). A recent study reported the zebrafish endocardial transcriptome at the 15-somite stage66. There was an extensive overlap between our endocardial dataset and the one reported by Capon et al. (Table S7), further validating endocardial identity of the nfatc1 + gata5 + cell cluster. As an additional confirmation, we analyzed expression of 3 selected markers wnt11r, bmp16 and bambib, endocardial expression of which has not been previously reported. In situ hybridization confirmed the expression of wnt11r, bambib and bmp16 in the heart at 24 hpf (Fig. 8E–G). To gain further insights into the signature of the endocardial cells relative to endothelial cells, we used Monocle 3 to analyze the position of endocardial cells in the trajectory plot. Majority of the endocardial gata5 + nfatc1 + cells clustered within or next to the arterial cluster. However, a few endocardial cells were also present within the venous cluster (Figure S2), suggesting a unique signature and heterogeneity between endocardial cells at 24 hpf. Altogether, we have identified the transcriptomic signature of zebrafish endocardial cells and validated new zebrafish endocardium markers.
Endocardial subcluster. (A) Endocardial cells were identified based on the expression of markers genes gata5 and nfatc1 (log2 > 0). (B,C) UMAP feature plots showing expression of gata5 and nfatc1. (D) List of top 30 genes differentially expressed in the endocardial subcluster. (E–G) In situ hybridization analysis at 24 hpf for selected endocardial enriched genes wnt11r, bambib and bmp16. Note the expression of wnt11r, bambib and bmp16 within the heart tube (arrowhead). Ventral view of deyolked and flat-mounted embryos, anterior is to the left. Scale bars: 100 μm.
Discussion
In this study we used scRNA-seq analysis to identify transcriptomic signatures of vascular endothelial cells in zebrafish embryos at 24 hpf, at the onset of blood circulation. Our profiling of 10,818 cells identified 6 different subtypes of vascular endothelial cells which included vascular progenitor, arterial, two different venous, endocardial, and cranial endothelial subtypes. Importantly, we have validated new markers for arterial, two different venous, and endocardial populations.
Intriguingly, two different venous populations were identified during scRNA-seq analysis. The PCV is known to harbor a heterogenous cell population including progenitors of lymphatic and subintestinal vasculature76,77,78. Many markers including lyve1b overlap between venous and lymphatic cells during development79 making it difficult to distinguish between the two lineages. Prox1a and prox1b are considered to be the definitive markers of lymphatic lineage80,81,82,83. Both markers were similarly enriched in the venous-1 (cluster #2, log2 values 2.85 and 3.96, respectively) and venous-2 (cluster #7, log2 values 2.97 and 3.46, respectively, Table S1) clusters. Therefore, it is unlikely that either of the clusters corresponds to lymphatic progenitors. Only a small number of cells were positive for either prox1a or prox1b, and they did not cluster together, preventing us from identifying lymphatic transcriptome at this developmental stage. While subintestinal progenitors also originate from the PCV starting at 28 hpf76,77,78, it is unclear if these cells have a unique transcriptomic signature at these early stages. Our data suggest that the venous-2 population is enriched in markers for the caudal vein. The caudal vein is known to have a unique transcriptional profile84,85 and also serves as the niche for hematopoietic stem cells86. Therefore, transcriptional characterization of caudal vein endothelium will help to understand pathways involved in its development.
An identity of cluster #6 (VE-other) is the most nebulous. Many top marker genes for this cluster are poorly characterized. Marker genes with known endothelial specific expression pattern include pan-endothelial kdrl, arterial cldn5b and venous specific lyve1b and dab2 (Table S1). Although the spatial and temporal origin of arteries and veins has been well established, the precise timing when arterial and venous-fated precursor cells differentiate into distinct arterial and venous vessels has been controversial87,88,89. We have previously shown co-expression of venous and arterial genes in many arterial progenitors at the 20-somite stage in the trunk region of zebrafish embryos. However, at later stages of development (24 hpf) the expression of venous and arterial markers becomes restricted to veins and arteries, respectively48, in the trunk vasculature. In contrast, we show here that arterial and venous genes are co-expressed in a subset of cranial vasculature including PHBC and MCeV at 24 hpf as well as 30 hpf. These results suggest mixed arteriovenous identity of cranial vasculature during embryonic development. Cells within cranial vasculature also show strong expression of other cluster #6 markers, such as cox4i2 and krt18, further supporting the argument that cluster #6 corresponds to a subset of cranial vasculature. However, we cannot rule out the possibility that cells with similar gene expression signature are also located elsewhere in the embryo. It is also possible that such expression pattern may represent a transitory phase of differentiation during embryonic development.
Endocardial cells share many similarities with vascular endothelial cells. However, they are also known to harbor a unique transcriptomic signature as evident by the expression of nfatc164,90. Differences in the transcriptomes between endocardial and other types of vascular endothelial cells have not been well characterized during embryonic development, and only a limited number of endocardial-specific markers are currently known in any vertebrate model system. We have previously reported a transcriptional signature of zebrafish endocardial progenitors identified during scRNA-seq analysis of etv2 reporter embryos at the 20-somite stage (19 hpf)48. In addition, a recent study performed RNA-seq using an endocardial specific reporter line to describe the endocardial transcriptome at the 15-somite stage66. Here we performed subclustering based on the established markers gata5 and nfatc1 to define the endocardial transcriptome at 24 hpf. A significant overlap between the endocardial marker genes at 15-somite, 20-somite and 24 hpf stages was observed which confirms the endocardial identity of gata5 + nfatc1 + cells. Multiple genes in this subcluster have been previously reported to show heart-specific expression, although most studies did not distinguish between the endocardial and myocardial expression. We further validated and demonstrated cardiac expression for 3 previously uncharacterized genes from this subcluster. Identification of the endocardial signature will be a useful resource for many investigators involved in cardiovascular research.
In summary, we identified transcriptional signatures of vascular endothelial cells during zebrafish development and described six different subtypes of endothelial-related cells at 24 hpf, when blood circulation is first initiated. This is one of the first reported scRNA-seq datasets of vascular endothelial cells during zebrafish development. We expect that this will be a useful resource for many researchers in cardiovascular development and will promote investigation of molecular mechanisms involved in the establishment of vascular heterogeneity.
Materials and methods
Zebrafish lines and maintenance
All studies were performed following standard guidelines and protocols approved by the University of South Florida and Cincinnati Children’s Hospital Medical Center Institutional Animal Care and Use Committee. Embryos were raised at 28.5 °C in embryo medium and staged by hours post fertilization (hpf)91. Zebrafish lines used in the study are as follows: wild-type AB, wild-type EKK, Tg(kdrl:GFP)s84392, Tg(kdrl:mCherry)14 and Tg(etv2ci32Gt; UAS:GFP)18.
Embryo dissociation and single-cell RNA-seq
Tg(kdrl:mCherry) fish were crossed to etv2ci32Gt; UAS:GFP line. Embryos double positive for both mCherry and GFP fluorescence were selected at 24 hpf. Embryos were dechorionated and immediately transferred to a 1.5 ml Eppendorf tube placed on ice for the generation of single-cell suspension. Whole embryos were then dissociated into a single-cell suspension by cold protease tissue dissociation protocol93. Cells expressing GFP only and both GFP and mCherry were sorted using a BD/FACSAria II flow cytometer cell sorter. A suspension of ~ 10,000 single cells was loaded onto the 10XGenomics Single Cell 3’ (V3 chemistry) chip at the CCHMC Gene Expression Core. 12 cDNA amplification cycles were used to generate cDNA. Samples were sequenced on an Illumina NovaSeq 6000 instrument (Illumina, San Diego, CA) running an S2 flow cell with parameters as follows: Read1, 28 cycles; Index Read1: 8 cycles; Read 2: 91 cycles at the CCHMC DNA Sequencing core. The raw Fastq files obtained from the sequencing core were then mapped to the Danio rerio genome (version zv11) to generate single-cell feature counts using Cell Ranger version 3.0.2. Counts were performed on fastq data from each of the populations individually. Filtered matrix files were then imported into Partek Flow analysis Suite. Cells were filtered on total read per cell (477–61,000), expressed genes per cell (96–5456) and mitochondrial counts (0–10%) which resulted in 10,818 cells. The gene count CPM (counts per million) values were normalized by adding 1.0 and calculating log2 values. GFP + and GFP + mCherry + samples were combined, and an unbiased graph-based clustering was performed. Finally, UMAP plots, violin plots, scatter plots, a dot plot, and a heatmap were obtained and exported from Partek. Endocardial cluster was identified based on the expression of marker genes gata5 and nfatc1 (log2 gata5 and nfatc1 > 0).
Gene ontology (GO) and pathway analysis
GO analysis was performed by applying gene list from respective clusters into GO site http://geneontology.org/ (GO Ontology database https://doi.org/10.5281/zenodo.6399963 Released 2022–03-22). To perform pathway analysis, PANTHER overrepresentation test (PANTHER version 17.0 Released 2022–02-22) was used. Statistical significance was calculated by Fisher’s exact test. A False discovery rate (FDR) corrected P-values less than 0.05 were considered statistically significant.
WISH and HCR
Whole mount in situ hybridization was performed as described before94.
To synthesize bcl6b, cx30.3, esm1, otc and si:dkey-28n18.9 probes, partial cDNA was first amplified from the cDNA of 24 hpf embryos using reverse oligo containing T7 promoter sequence. Resulting PCR product was then used as a template for RNA synthesis. Following primers were used for amplification of partial cDNA (Table S8). Anti-sense RNA was synthesized using T7 RNA polymerase (Promega) and DIG-labeling mix (Sigma-Aldrich). To synthesize bambib, bmp16, notchl and wnt11r probes, partial cDNA sequence was amplified using gene specific primers (Table S8). cDNA for each gene was subcloned into PCRII-TOPO vector using TOPO T/A cloning (ThermoFisher). Plasmids were linearized and antisense RNA was synthesized using SP6 (bmp16, wnt11r) or T7 (bambib, notchl) RNA polymerase (Promega) and DIG-labeling mix (Sigma-Aldrich). WISH embryos were imaged with a Nikon Eclipse Ni-E or Zeiss Axio imager compound microscope. HCR v3.0 probes for cldn5b, dab2, lyve1b, glula and krt18 were obtained from Molecular Technologies, Inc., Los Angeles, CA, USA. HCR in situ was performed as described previously95. Embryos were mounted in 0.6% low-melting agarose and imaged using Nikon Eclipse confocal microscope. At least 15 embryos processed for WISH and HCR were analyzed.
Ethical approval
The study is reported in accordance with ARRIVE guidelines96.
Data availability
The scRNA-seq datasets generated and/or analyzed during the current study are available in the NCBI GEO database under accession number GSE202912. All processed image datasets are included in this published article (and its Supplementary information files). Original image datasets generated during this study are available from the Lead contact upon reasonable request.
References
Marcelo, K. L., Goldie, L. C. & Hirschi, K. K. Regulation of endothelial cell differentiation and specification. Circ. Res. 112, 1272–1287. https://doi.org/10.1161/CIRCRESAHA.113.300506 (2013).
Matsuoka, R. L. & Stainier, D. Y. R. Recent insights into vascular development from studies in zebrafish. Curr. Opin. Hematol. 25, 204–211. https://doi.org/10.1097/Moh.0000000000000420 (2018).
Deng, Q., Ramskold, D., Reinius, B. & Sandberg, R. Single-cell RNA-seq reveals dynamic, random monoallelic gene expression in mammalian cells. Science 343, 193–196. https://doi.org/10.1126/science.1245316 (2014).
Jaitin, D. A. et al. Massively parallel single-cell RNA-seq for marker-free decomposition of tissues into cell types. Science 343, 776–779. https://doi.org/10.1126/science.1247651 (2014).
Kalucka, J. et al. Single-cell transcriptome atlas of murine endothelial cells. Cell 180, 764–779. https://doi.org/10.1016/j.cell.2020.01.015 (2020).
Jakab, M. & Augustin, H. G. Understanding angiodiversity: Insights from single cell biology. Development https://doi.org/10.1242/dev.146621 (2020).
Chavkin, N. W. & Hirschi, K. K. Single cell analysis in vascular biology. Front. Cardiovasc. Med. 7, 42. https://doi.org/10.3389/fcvm.2020.00042 (2020).
Paik, D. T. et al. Single-cell RNA sequencing unveils unique transcriptomic signatures of organ-specific endothelial cells. Circulation 142, 1848–1862. https://doi.org/10.1161/CIRCULATIONAHA.119.041433 (2020).
Augustin, H. G. & Koh, G. Y. Organotypic vasculature: From descriptive heterogeneity to functional pathophysiology. Science https://doi.org/10.1126/science.aal2379 (2017).
Koch, P. S., Lee, K. H., Goerdt, S. & Augustin, H. G. Angiodiversity and organotypic functions of sinusoidal endothelial cells. Angiogenesis 24, 289–310. https://doi.org/10.1007/s10456-021-09780-y (2021).
Jiang, M. M. et al. Characterization of the zebrafish cell landscape at single-cell resolution. Front. Cell Dev. Biol. https://doi.org/10.3389/fcell.2021.743421 (2021).
Farnsworth, D. R., Saunders, L. M. & Miller, A. C. A single-cell transcriptome atlas for zebrafish development. Dev. Biol. 459, 100–108. https://doi.org/10.1016/j.ydbio.2019.11.008 (2020).
Liao, W. et al. The zebrafish gene cloche acts upstream of a flk-1 homologue to regulate endothelial cell differentiation. Development 124, 381–389 (1997).
Proulx, K., Lu, A. & Sumanas, S. Cranial vasculature in zebrafish forms by angioblast cluster-derived angiogenesis. Dev. Biol. 348, 34–46. https://doi.org/10.1016/j.ydbio.2010.08.036 (2010).
Sumanas, S. & Lin, S. Ets1-related protein is a key regulator of vasculogenesis in zebrafish. PLoS Biol. 4, 60–69. https://doi.org/10.1371/journal.pbio.0040010 (2006).
Sumanas, S. et al. Interplay among Etsrp/ER71, Scl, and Alk8 signaling controls endothelial and myeloid cell formation. Blood 111, 4500–4510. https://doi.org/10.1182/blood-2007-09-110569 (2008).
Lee, D. et al. ER71 acts downstream of BMP, Notch, and Wnt signaling in blood and vessel progenitor specification. Cell Stem Cell 2, 497–507. https://doi.org/10.1016/j.stem.2008.03.008 (2008).
Chestnut, B. & Sumanas, S. Zebrafish etv2 knock-in line labels vascular endothelial and blood progenitor cells. Dev. Dyn. 249, 245–261. https://doi.org/10.1002/dvdy.130 (2020).
Leslie, J. D. et al. Endothelial signalling by the Notch ligand Delta-like 4 restricts angiogenesis. Development 134, 839–844. https://doi.org/10.1242/dev.003244 (2007).
Lawson, N. D. et al. Notch signaling is required for arterial-venous differentiation during embryonic vascular development. Development 128, 3675–3683 (2001).
Zygmunt, T. et al. Semaphorin-PlexinD1 signaling limits angiogenic potential via the VEGF decoy receptor sFlt1. Dev. Cell 21, 301–314. https://doi.org/10.1016/j.devcel.2011.06.033 (2011).
Covassin, L. D., Villefranc, J. A., Kacergis, M. C., Weinstein, B. M. & Lawson, N. D. Distinct genetic interactions between multiple Vegf receptors are required for development of different blood vessel types in zebrafish. Proc. Natl. Acad. Sci. USA 103, 6554–6559. https://doi.org/10.1073/pnas.0506886103 (2006).
Rehn, K., Wong, K. S., Balciunas, D. & Sumanas, S. Zebrafish enhancer trap line recapitulates embryonic aquaporin 1a expression pattern in vascular endothelial cells. Int. J. Dev. Biol. 55, 613–618. https://doi.org/10.1387/ijdb.103249kp (2011).
Lawson, N. D., Vogel, A. M. & Weinstein, B. M. sonic hedgehog and vascular endothelial growth factor act upstream of the notch pathway during arterial endothelial differentiation. Dev. Cell 3, 127–136. https://doi.org/10.1016/S1534-5807(02)00198-3 (2002).
Moran, C. M., Salanga, M. C. & Krieg, P. A. Hedgehog signaling regulates size of the dorsal aortae and density of the plexus during avian vascular development. Dev. Dyn. 240, 1354–1364. https://doi.org/10.1002/dvdy.22600 (2011).
Carrier, G. O., White, R. E. & Kirby, M. L. Histamine-induced relaxation of rat aorta: Importance of H-1 receptor and vascular endothelium. Blood Vessels 21, 180–183 (1984).
Kobialka, P. & Graupera, M. Revisiting PI3-kinase signalling in angiogenesis. Vasc. Biol. 1, H125–H134. https://doi.org/10.1530/VB-19-0025 (2019).
Somanath, P. R., Malinin, N. L. & Byzova, T. V. Cooperation between integrin alphavbeta3 and VEGFR2 in angiogenesis. Angiogenesis 12, 177–185. https://doi.org/10.1007/s10456-009-9141-9 (2009).
Schreier, B. et al. Endothelial epidermal growth factor receptor is of minor importance for vascular and renal function and obesity-induced dysfunction in mice. Sci. Rep. 11, 15. https://doi.org/10.1038/s41598-021-86587-3 (2021).
Hong, C. C., Peterson, Q. P., Hong, J. Y. & Peterson, R. T. Artery/vein specification is governed by opposing phosphatidylinositol-3 kinase and MAP kinase/ERK signaling. Curr. Biol. 16, 1366–1372. https://doi.org/10.1016/j.cub.2006.05.046 (2006).
Flores, M. V., Hall, C. J., Crosier, K. E. & Crosier, P. S. Visualization of embryonic lymphangiogenesis advances the use of the zebrafish model for research in cancer and lymphatic pathologies. Dev. Dyn. 239, 2128–2135. https://doi.org/10.1002/dvdy.22328 (2010).
Sun, G. J. et al. Identification and expression analysis of zebrafish (Danio rerio) E-selectin during embryonic development. Molecules 20, 18539–18550. https://doi.org/10.3390/molecules201018539 (2015).
Thompson, M. A. et al. The cloche and spadetail genes differentially affect hematopoiesis and vasculogenesis. Dev. Biol. 197, 248–269. https://doi.org/10.1006/dbio.1998.8887 (1998).
Swift, M. R. et al. SoxF factors and Notch regulate nr2f2 gene expression during venous differentiation in zebrafish. Dev. Biol. 390, 116–125. https://doi.org/10.1016/j.ydbio.2014.03.018 (2014).
Weber, G. J. et al. Mutant-specific gene programs in the zebrafish. Blood 106, 521–530. https://doi.org/10.1182/blood-2004-11-4541 (2005).
Wong, K. S., Proulx, K., Rost, M. S. & Sumanas, S. Identification of vasculature-specific genes by microarray analysis of Etsrp/Etv2 overexpressing zebrafish embryos. Dev. Dyn. 238, 1836–1850. https://doi.org/10.1002/dvdy.21990 (2009).
Hellstrom, A. et al. IGF-I is critical for normal vascularization of the human retina. J. Clin. Endocr. Metab. 87, 3413–3416. https://doi.org/10.1210/jc.87.7.3413 (2002).
Ren, C. G. et al. Activated N-Ras signaling regulates arterial-venous specification in zebrafish. J. Hematol. Oncol. https://doi.org/10.1186/1756-8722-6-34 (2013).
Akil, A. et al. Notch signaling in vascular endothelial cells, angiogenesis, and tumor progression: An update and prospective. Front. Cell Dev. Biol. https://doi.org/10.3389/fcell.2021.642352 (2021).
Oladipupo, S. S. et al. Endothelial cell FGF signaling is required for injury response but not for vascular homeostasis. Proc. Natl. Acad. Sci. USA 111, 13379–13384. https://doi.org/10.1073/pnas.1324235111 (2014).
Thisse, B. et al. Expression of the zebrafish genome during embryogenesis. ZFIN Direct Data Submission (2001).
Cermenati, S. et al. Sox18 and Sox7 play redundant roles in vascular development. Blood 111, 2657–2666. https://doi.org/10.1182/blood-2007-07-100412 (2008).
Manalo, D. J. et al. Transcriptional regulation of vascular endothelial cell responses to hypoxia by HIF-1. Blood 105, 659–669. https://doi.org/10.1182/blood-2004-07-2958 (2005).
Kallakuri, S. et al. Endothelial cilia are essential for developmental vascular integrity in zebrafish. J. Am. Soc. Nephrol. 26, 864–875. https://doi.org/10.1681/Asn.2013121314 (2015).
Mahony, C. B., Fish, R. J., Pasche, C. & Bertrand, J. Y. tfec controls the hematopoietic stem cell vascular niche during zebrafish embryogenesis. Blood 128, 1336–1345. https://doi.org/10.1182/blood-2016-04-710137 (2016).
Metikala, S., Chetty, S. C. & Sumanas, S. Single-cell transcriptome analysis of the zebrafish embryonic trunk. PLoS ONE 16, e254024. https://doi.org/10.1371/journal.pone.0254024 (2021).
Xie, J., Farage, E., Sugimoto, M. & Anand-Apte, B. A novel transgenic zebrafish model for blood-brain and blood-retinal barrier development. Bmc Dev. Biol. 10, 1–76. https://doi.org/10.1186/1471-213x-10-76 (2010).
Chestnut, B., Casie Chetty, S., Koenig, A. L. & Sumanas, S. Single-cell transcriptomic analysis identifies the conversion of zebrafish Etv2-deficient vascular progenitors into skeletal muscle. Nat. Commun https://doi.org/10.1038/s41467-020-16515-y (2020).
Marshall, I. Characterization and distribution of histamine H-1-receptor and H-2-receptor in precapillary vessels. J. Cardiovasc. Pharm. 6, S587–S597. https://doi.org/10.1097/00005344-198406004-00005 (1984).
Pillay, L. M. et al. In vivo dissection of Rhoa function in vascular development using zebrafish. Angiogenesis https://doi.org/10.1007/s10456-022-09834-9 (2022).
Bayless, K. J. & Davis, G. E. Microtubule depolymerization rapidly collapses capillary tube networks in vitro and angiogenic vessels in vivo through the small GTPase Rho. J. Biol. Chem. 279, 11686–11695. https://doi.org/10.1074/jbc.M308373200 (2004).
Mehta, P. K. & Griendling, K. K. Angiotensin II cell signaling: Physiological and pathological effects in the cardiovascular system. Am. J. Physiol. 292, C82–C97. https://doi.org/10.1152/ajpcell.00287.2006 (2007).
Reischauer, S. et al. Cloche is a bHLH-PAS transcription factor that drives haemato-vascular specification. Nature https://doi.org/10.1038/nature18614 (2016).
Liao, E. C. et al. SCL/Tal-1 transcription factor acts downstream of cloche to specify hematopoietic and vascular progenitors in zebrafish. Gene Dev. 12, 621–626. https://doi.org/10.1101/gad.12.5.621 (1998).
Wang, H. et al. Over-expression of PDGFR-beta promotes PDGF-induced proliferation, migration, and angiogenesis of EPCs through PI3K/Akt signaling pathway. PLoS ONE 7, e30503. https://doi.org/10.1371/journal.pone.0030503 (2012).
Caiado, F. & Dias, S. Endothelial progenitor cells and integrins: Adhesive needs. Fibrogenesis Tissue https://doi.org/10.1186/1755-1536-5-4 (2012).
Maeng, Y. S., Choi, Y. J. & Kim, E. K. TGFBIp regulates differentiation of EPC (CD133(+)c-kit(+)lin(-) cells) to EC through activation of the notch signaling pathway. Stem Cells 33, 2052–2062. https://doi.org/10.1002/stem.2003 (2015).
Hubner, K. et al. Wnt signaling positively regulates endothelial cell fate specification in the Flila-positive progenitor population via Lef1. Dev. Biol. 430, 142–155. https://doi.org/10.1016/j.ydbio.2017.08.004 (2017).
Cao, J. Y. et al. The single-cell transcriptional landscape of mammalian organogenesis. Nature https://doi.org/10.1038/s41586-019-0969-x (2019).
Brown, L. A. et al. Insights into early vasculogenesis revealed by expression of the ETS-domain transcription factor Fli-1 in wild-type and mutant zebrafish embryos. Mech. Dev. 90, 237–252. https://doi.org/10.1016/S0925-4773(99)00256-7 (2000).
Larson, J. D. et al. Expression of VE-cadherin in zebrafish embryos: A new tool to evaluate vascular development. Dev. Dyn. 231, 204–213. https://doi.org/10.1002/dvdy.20102 (2004).
Lyons, M. S., Bell, B., Stainier, D. & Peters, K. G. Isolation of the zebrafish homologues for the tie-1 and tie-2 endothelium-specific receptor tyrosine kinases. Dev. Dyn. 212, 133–140 (1998).
Harris, I. S. & Black, B. L. Development of the endocardium. Pediatr. Cardiol. 31, 391–399. https://doi.org/10.1007/s00246-010-9642-8 (2010).
Wong, K. S. et al. Hedgehog signaling is required for differentiation of endocardial progenitors in zebrafish. Dev. Biol. 361, 377–391. https://doi.org/10.1016/j.ydbio.2011.11.004 (2012).
Reiter, J. F. et al. Gata5 is required for the development of the heart and endoderm in zebrafish. Gene Dev. 13, 2983–2995. https://doi.org/10.1101/gad.13.22.2983 (1999).
Capon, S. J., Uribe, V., Dominado, N., Ehrlich, O. & Smith, K. A. Endocardial identity is established during early somitogenesis by Bmp signalling acting upstream of npas4l and etv2. Development https://doi.org/10.1242/dev.190421 (2022).
Bonetti, M. et al. Noonan and LEOPARD syndrome Shp2 variants induce heart displacement defects in zebrafish. Development 141, 1961–1970. https://doi.org/10.1242/dev.106310 (2014).
Trinh, L. A. & Stainier, D. Y. R. Fibronectin regulates epithelial organization during myocardial migration in zebrafish. Dev. Cell 6, 371–382 (2004).
Smith, K. A. et al. Rotation and asymmetric development of the zebrafish heart requires directed migration of cardiac progenitor cells. Dev. Cell 14, 287–297. https://doi.org/10.1016/j.devcel.2007.11.015 (2008).
Walsh, E. C. & Stainier, D. Y. R. UDP-glucose dehydrogenase required for cardiac valve formation in zebrafish. Science 293, 1670–1673. https://doi.org/10.1126/science.293.5535.1670 (2001).
Vermot, J. et al. Reversing blood flows act through klf2a to ensure normal valvulogenesis in the developing heart. PLoS Biol. 7, e1000246. https://doi.org/10.1371/journal.pbio.1000246 (2009).
Walton, R. Z. et al. Fog1 is required for cardiac looping in zebrafish. Dev. Biol. 289, 482–493. https://doi.org/10.1016/j.ydbio.2005.10.040 (2006).
Ikle, J. M. et al. Nkx2.5 regulates endothelin converting enzyme-1 during pharyngeal arch patterning. Genesis 55, doi: https://doi.org/10.1002/dvg.23021 (2017).
de Pater, E. et al. Bmp signaling exerts opposite effects on cardiac differentiation. Circ. Res. 110, 578-U179. https://doi.org/10.1161/Circresaha.111.261172 (2012).
Thisse, B. & Thisse, C. Fast Release Clones: A High Throughput Expression Analysis. ZFIN Direct Data Submission (2004).
Hen, G. et al. Venous-derived angioblasts generate organ-specific vessels during zebrafish embryonic development. Development 142, 4266–4278. https://doi.org/10.1242/dev.129247 (2015).
Koenig, A. L. et al. Vegfa signaling promotes zebrafish intestinal vasculature development through endothelial cell migration from the posterior cardinal vein. Dev. Biol. 411, 115–127. https://doi.org/10.1016/j.ydbio.2016.01.002 (2016).
Nicenboim, J. et al. Lymphatic vessels arise from specialized angioblasts within a venous niche. Nature 522, 56-U100. https://doi.org/10.1038/nature14425 (2015).
Okuda, K. S. et al. lyve1 expression reveals novel lymphatic vessels and new mechanisms for lymphatic vessel development in zebrafish. Development 139, 2381–2391. https://doi.org/10.1242/dev.077701 (2012).
Hogan, B. M. et al. ccbe1 is required for embryonic lymphangiogenesis and venous sprouting. Nat. Genet. 41, 396–398. https://doi.org/10.1038/ng.321 (2009).
Lim, A. H. et al. Motoneurons are essential for vascular pathfinding. Development 138, 3847–3857. https://doi.org/10.1242/dev.068403 (2011).
Yaniv, K. et al. Live imaging of lymphatic development in the zebrafish. Nat. Med. 12, 711–716. https://doi.org/10.1038/nm1427 (2006).
Del Giacco, L., Pistocchi, A. & Ghilardi, A. prox1b activity is essential in zebrafish lymphangiogenesis. PLoS ONE 5, E13170. https://doi.org/10.1371/journal.pone.0013170 (2010).
Kashiwada, T. et al. beta-catenin-dependent transcription is central to Bmp-mediated formation of venous vessels. Development 142, 497–509. https://doi.org/10.1242/dev.115576 (2015).
Wiley, D. M. et al. Distinct signalling pathways regulate sprouting angiogenesis from the dorsal aorta and the axial vein. Nat. Cell Biol. 13, 686-U138. https://doi.org/10.1038/ncb2232 (2011).
Tamplin, O. J. et al. Hematopoietic stem cell arrival triggers dynamic remodeling of the perivascular niche. Cell 160, 241–252. https://doi.org/10.1016/j.cell.2014.12.032 (2015).
Kohli, V., Schumacher, J. A., Desai, S. P., Rehn, K. & Sumanas, S. Arterial and venous progenitors of the major axial vessels originate at distinct locations. Dev. Cell 25, 196–206. https://doi.org/10.1016/j.devcel.2013.03.017 (2013).
Quillien, A. et al. Distinct Notch signaling outputs pattern the developing arterial system. Development 141, 1544–1552. https://doi.org/10.1242/dev.099986 (2014).
Zhong, T. P., Childs, S., Leu, J. P. & Fishman, M. C. Gridlock signalling pathway fashions the first embryonic artery. Nature 414, 216–220. https://doi.org/10.1038/35102599 (2001).
Castillo-Robles, J., Ramirez, L., Spaink, H. P. & Lomeli, H. smarce1 mutants have a defective endocardium and an increased expression of cardiac transcription factors in zebrafish. Sci. Rep. https://doi.org/10.1038/s41598-018-33746-8 (2018).
Kimmel, C. B., Ballard, W. W., Kimmel, S. R., Ullmann, B. & Schilling, T. F. Stages of embryonic-development of the Zebrafish. Dev. Dyn. 203, 253–310. https://doi.org/10.1002/aja.1002030302 (1995).
Jin, S. W., Beisl, D., Mitchell, T., Chen, J. N. & Stainier, D. Y. R. Cellular and molecular analyses of vascular tube and lumen formation in zebrafish. Development 132, 5199–5209. https://doi.org/10.1242/dev.02087 (2005).
Potter, A. S. & Potter, S. S. Dissociation of tissues for single-cell analysis. Methods Mol. Biol. 1926, 55–62. https://doi.org/10.1007/978-1-4939-9021-4_5 (2019).
Jowett, T. Analysis of protein and gene expression. Method Cell Biol. 59, 63–85 (1999).
Choi, H. M. T. et al. Third-generation in situ hybridization chain reaction: Multiplexed, quantitative, sensitive, versatile, robust. Development https://doi.org/10.1242/dev.165753 (2018).
du Sert, N. P. et al. Reporting animal research: Explanation and elaboration for the ARRIVE guidelines 2.0. PLoS Biol. 18, e3000411. https://doi.org/10.1371/journal.pbio.3000411 (2020).
Acknowledgements
This research was supported by postdoctoral fellowship from the American Heart Association to Suman Gurung (19POST34450021) and an award from the National Institutes of Health (R01HL153005) to Saulius Sumanas. We thank Shawn Smith and Kelly Rangel from CCHMC Gene expression core for assistance in processing samples for scRNA-seq. We would also like to thank FACS core at CCHMC for their assistance with the project.
Author information
Authors and Affiliations
Contributions
S.G. performed most experiments in the study and wrote the manuscript. N.K.R, B.C. and L.K. contributed to in situ experiments. S.S. conceived and supervised the project and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gurung, S., Restrepo, N.K., Chestnut, B. et al. Single-cell transcriptomic analysis of vascular endothelial cells in zebrafish embryos. Sci Rep 12, 13065 (2022). https://doi.org/10.1038/s41598-022-17127-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-17127-w
- Springer Nature Limited