Abstract
Methicillin-resistant Staphylococcus aureus (MRSA) is responsible for hard-to-treat infections. The presence of 19 virulence genes in 120 MRSA isolates obtained from hospitalized patients and genetic relationships of these isolates were investigated. The eno (100%) and ebps (93.3%) genes encoding laminin- and elastin binding proteins, respectively, were ubiquitous. Other adhesion genes: fib (77.5%), fnbB (41.6%), bbp (40.8%), cna (30.8%) encoding proteins binding fibrinogen, fibronectin, bone sialoprotein and collagen, respectively, and map/eap (62.5%), encoding Eap, were also frequent. The etB and etD genes, encoding exfoliative toxins, were present in 15.6% and 12.5% isolates, respectively. The splA, splE and sspA, encoding serine protease were detected in 100%, 70.8% and 94.2% isolates, respectively. The tst gene, encoding toxic shock syndrome toxin-1 was found in 75% isolates. The cna, map/eap and tst genes were the most common in wound isolates and much less common in blood isolates. We identified 45 different spa types, t003 (21.7%) and t008 (18.8%) being the most common. The t003 was the most frequent among isolates from the respiratory tract (35.5%), while t008 in blood isolates (40%). Identification of virulence factors of MRSA is important for evaluation of pathogen transmission rate and disease development.
Similar content being viewed by others
Introduction
Methicillin-resistant Staphylococcus aureus (MRSA) cause hard-to-treat infections in various patient populations, and thus is a serious health problem. Treatment of MRSA infections is a significant financial burden for medical institutions1. The gene mecA in MRSA genome, encodes an enzyme responsible for crosslinking the peptidoglycans in the bacterial cell wall called penicillin-binding protein 2a (PBP2a). Low affinity PBP2a to β-lactams results in resistance to β-lactam antibiotics including penicillins, cephalosporins (except for ceftaroline and ceftobiprole), and carbapenems2. Multidrug-resistant (MDR) MRSA isolates are often resistant to commonly used antibiotic groups, such as aminoglycosides, fluoroquinolones, macrolides, tetracycline and chloramphenicol3,4.
MRSA is responsible for skin and surgical wound infections but also may infect different parts of the body including lower respiratory tract, cause bloodstream infection (BSI) and toxin-mediated syndromes as well as life-threatening diseases5,6,7. The studies show that the proportion of MRSA among all S. aureus isolates is between 13 and 89%8,9. MRSA remains a prominent pathogen with persistently high mortality10. The mortality rate of S. aureus bacteremia is around 20–30%11. A European report from a Finnish Hospital Infection Program showed that S. aureus ranked among the top three organisms causing bloodstream infections12. MRSA are widely spread in various countries, in hospital environments, in community and livestock13,14. MRSA infections are routinely detected in hospitalized patients (HAIs, health care-associated infections), also in high-income countries6,11. Infections caused by MRSA isolates lead to prolonged hospital stay, increased mortality, especially in patients with underlying diseases such as malignancy or chronic pulmonary diseases15. The virulence factors produced by MRSA can be divided into different groups, including degradative enzymes, adhesins, and superantigenic toxins. The infective process requires participation of numerous virulence determinants. First step of staphylococcal infection is the attachment of bacterial cells to the host tissues. S. aureus produces the surface-exposed proteins (MSCRAMMs—microbial surface components recognizing adhesive matrix molecules)16. Using these MSCRAMMs S. aureus can bind to one or more host extracellular matrix factors including laminin, elastin, fibrinogen, fibronectin and collagen17. Other protein expressed by S. aureus is extracellular adherence protein (Eap) belonging to SERAMs (secretable expanded repertoire adhesive molecules) can bind to various glycoproteins of extracellular matrix (ECM), such as fibronectin, fibrinogen, sialoprotein and some collagens18. According to Hussain et al.19, this protein is also involved in adhesion of S. aureus to fibroblasts as well as in bacteria internalization. S. aureus produce also proteases being important virulence factors that can cleave host proteins and allow the transition of MRSA cells from an adhesive to an invasive phenotype. Bacterial skin invasion is facilitated by exfoliative toxins, whereas enterotoxins and the toxic shock syndrome toxin-1, as superantigens suppress the host immune response and promote the persistence of S. aureus in host organism20. Evaluation of the virulence potential seems to be a reliable method of predicting the behaviour of these bacteria in host organisms, which is very important because it allows to predict the development and course of an infection. The role of virulence factors in the pathogenesis of MRSA may vary with the type of infection and the patient's health status. In Poland, only few works included the virulence potential of MRSA isolated from humans and these studies concerned the population of MRSA isolates from one specific clinical materials21,22 or from residents and personnel of nursing home23. In this study, our aim was to examine the presence of 19 important virulence genes encoding adhesins, proteases and superantigenic toxins in 120 of MRSA isolates from different clinical materials and to determine the genetic virulence profiles specific for MRSA isolates causing different infections in hospitalized patients during 2015–2017 in hospitals of Masovian district in Poland. We also determined clonal diversity of investigated MRSA isolates by spa-typing.
Results
The frequency of 19 genes encoding virulence factors investigated by PCR in 120 MRSA isolates is shown in Table 1. The results are presented for three groups created according to related genes: the adhesin genes, the protease genes and the superantigenic toxin genes encoding the enterotoxins and the toxic shock syndrome toxin-1. The eno gene encoding laminin binding protein was identified in all MRSA isolates. Among the remaining genes encoding adhesins, the ebps (93.3%) and fib (77.5%) encoding elastin and fibrinogen binding proteins, respectively, were the most frequently identified in the investigated MRSA isolates. The map/eap gene encoding Eap (62.5%) was also frequently identified. The fnbB and bbp genes encoding fibronectin and bone sialoprotein binding proteins, were found in more than 40% of isolates. The cna gene encoding collagen binding protein was one of the less frequently identified because it was found in only in over 30% of the isolates (Table 1). In a group of three investigated serine protease-encoding genes, splA gene was identified in all MRSA isolates. The sspA and splE genes were present in 94.2 and 70.8% investigated MRSA isolates, respectively. The etB and etD genes encoding exfoliative toxins were present in the genomes of 15 and 12.5% isolates, respectively. Among the genes encoding classical enterotoxins, the sea gene was the most frequently identified (25%). The tst gene, encoding the toxic shock syndrome toxin-1, was harboured by 75% isolates (Table 1). Cluster analysis of investigated genes showed that adhesin encoding genes (eno, ebps) and serine protease-encoding genes (sspA, splA) belonged to one group of genes that occurred most frequently in this population of MRSA isolates. While, the genes encoding exfoliative toxins (etB, etD) and enterotoxin-coding genes (seb, sec, sed, see) were rarely detected in MRSA isolates (Fig. 1). MRSA isolates differed in the presence of virulence genes. The most common virulotypes were eno, ebpS, fib, map/eap, splA, sspA, tst (6.7%), eno, ebpS, cna, fnbB, fib, bbp, map/eap, splA, splE, sspA, tst, sea (2.5%) and eno, ebpS, fnbB, fib, bbp, map/eap, splA, splE, sspA, tst, sea (2.5%) (Table 2). In case of isolates from respiratory tract and nose, the most frequent combination of adhesin genes was eno, ebps, fib, map/eap. In genome of isolates from anus and blood, the most frequent adhesin genes were eno, ebps, fib, while eno, ebps, cna, fnbB, fib, bbp, map/eap genes were frequently detected in isolates from wounds (Table S1). The simultaneous presence of genes encoding serine proteases (splA, splE, sspA) was the most frequently detected in the genomes of isolates from wound, anus and respiratory tract (Table S2). The tst and sea genes encoding toxins was the most frequent in the investigated MRSA isolates (19.2%) (Table S3). The presence of cna gene was the most common in MRSA isolates from wound (53.3%) (Table 3) and it was highly significantly more frequent than in isolates from respiratory tract (18.8%) (p = 0.0016) and in isolates from blood (9.1%) (p = 0.0119). The fib gene was present in all isolates from anus (Table 3). This gene was identified significantly less frequently in isolates from wounds (70%) (p = 0.019) than in isolates from anus. The frequency of fnbB gene in MRSA isolates obtained from different clinical materials did not differ significantly. The ebps gene was detected in all isolates from anus and blood, while the frequency of the bbp gene in various groups of isolates did not differ significantly. The map/eap gene was the most often identified among wound isolates (73.3%). Among the genes encoding exfoliative toxins, etB gene was the most frequently identified in isolates from respiratory tract (20.8%). There were no significant differences in the frequency of the etD gene. The splE gene was significantly more frequently detected in wound isolates (80%) than in blood isolates (45.5%) (p = 0.0336). Whereas, the sea gene was significantly less frequently detected in wound isolates (6.7%) than in respiratory tract isolates (29.2%) (p = 0.017). The presence of tst gene was shown in all nose isolates (Table 3). Moreover, this gene was significantly more frequent in wound isolates (90%) than in isolates from respiratory tract (68.8%) (p = 0.0310) and blood (54.5%) (p = 0.012). The spa typing of 106 MRSA isolates showed the presence of 45 different spa types (Table 4). The most common spa types were t003 (23 isolates, 21.7%), t008 (20 isolates, 18.8%), t4474 (5 isolates, 4.7%) and t5224 (5 isolates, 4.7%). t003 spa type was the most frequently detected among isolates from respiratory tract (35.5%), while t008 spa type was the most frequently present in isolates from blood (40%). The frequency of the most common spa types (t003 and t008) in isolates from patients hospitalized in Siedlce and Warsaw did not differ significantly.
The groups of genes encoding adhesins, proteases and superantigenic toxins in MRSA isolated from patients hospitalized in 2015–2017 created based on the cluster analysis. fnbB, bbp, cna, fib, ebpS, eno—genes encoding adhesins such as fibronectin B-, bone sialoprotein-, collagen-, fibrinogen-, elastin-, laminin-binding proteins, respectively; map—gene encoding Eap (extracellular adherence protein); etA, etB, etD—genes encoding exfoliative toxins; splA, sspA, splE—genes encoding serine proteases; sea, seb, sec, sed, see—genes encoding the enterotoxins A to E; tst—gene encoding the toxic shock syndrome toxin-1.
Discussion
We undertook studies to assess the virulence potential of MRSA isolates from different clinical materials from hospitalized patients because such an analysis is necessary to predict the development and course of infection in the host organism and is important to control it. We also assessed whether MRSA isolates causing infections localized at different sites in the human body differ in the set of pathogenicity-determining genes, and whether the genes encoding virulence factors were associated with the type of infection or with the spa type. According to our knowledge, the number of studies on the virulence factors of MRSA isolates from hospitalized patients in Poland is limited and concern small collections of MRSA isolates22,23 or only few virulence factors21. Among the genes encoding adhesins, the eno and ebps genes encoding the ability of adhesion to laminin and elastin, respectively, were ubiquitous in the investigated MRSA isolates. The eno gene encodes α-enolase, which is able to bind laminin and also functions as a plasminogen receptor. Laminin is a major component of the basal membrane of the vasculature and thus adherence to laminin may contribute to tissue invasion and dissemination of staphylococcal cells by blood to different sites in the host. Besides, α-enolase may cause laminin degradation by activation of plasminogen24. Similarly to our results, Kasela et al.23 also showed that eno gene was present in all MRSA isolates from the residents and personnel of a nursing home in Poland, while Haghi et al.25 reported high prevalence of this gene among different clinical isolates. Elastin is a major component of elastic fibres in the extracellular matrix that are present in the skin, lung and blood vessels. Elastin-binding protein of S. aureus may be significant mainly in colonisation of injured tissues abundant in elastin26. In our study the ebps gene was among the most frequently identified genes in the investigated population of MRSA. High frequency of ebps gene in MRSA isolates from hospitalised patients indicated that EbpS protein is an important virulence factor of MRSA that binds to elastin and this interaction may promote bacterial colonization and facilitate pathogenesis. In our earlier research27 we showed that the expression levels of the ebps gene were significantly higher in the first hours of growth under biofilm conditions compared to growth under planktonic conditions, which suggest that the product of this gene is important for the first phase of biofilm growth, in which bacterial cells interact with host extracellular ligands. In the investigated population of MRSA isolates, the fib gene encoding fibrinogen binding protein Fib was also frequently identified (77.5%) which is similar to the results (62.2%) obtained by Azmi et al.28. Fibrinogen is a glycoprotein present in the blood and it is one of main proteins deposited on implanted biomaterials. The Fib is an important adherence factor responsible for the ability of S. aureus to adhere to fibrinogen adsorbed on biomaterials and endothelial cells29. Among adhesin genes, the map/eap gene encoding a staphylococcal surface protein was present in 62.5% of MRSA isolates. Eap, besides the ability to bind to many different glycoproteins (fibronectin, fibrinogen, sialoprotein and some collagens)18 can also form oligomers, and by rebinding to the surface of staphylococcal cells, mediates bacterial agglutination19. Damaged tissues in wounds provide a matrix of proteins, including fibrinogen, fibronectin, collagen and albumin to which pathogens may adhere. In our study, the map/eap gene was present in many MRSA isolates, but was the most often identified in wound isolates, which indicate that Eap belongs to the group of adhesins important for wound bed colonisation and biofilm formation in wounds. In this study, similarly as the map/eap gene, the cna gene was also the most frequently identified in wound isolates. Collagen is present in tissues as the structural support but in a wounded tissue becomes available to adhering bacterial cells. The presence of cna gene mainly in wound isolates indicate that the product of this gene is essential for the occupation of wound environment and development of infection. Other authors also confirmed that adhesion of S. aureus to collagen promoted infection and the initiation of biofilm formation30. The ability of adhesion to fibronectin is important in the disease development process, because fibronectin is a ubiquitous host protein present in soluble form in the blood and in fibrillar form in cellular matrices31. In our study, we found no relationship between the presence of the fnbB gene and the source of MRSA isolation. Contrary to the results obtained by Szczuka et al.32 who did not show the presence of bbp gene in S. aureus from hospitalised patients in Poland, we identified this gene in a high percentage of isolates (40.8%). In our study, the frequency of bbp gene encoding the bone sialoprotein binding protein did not differ significantly among the groups of isolates, although the studies by other authors suggested that S. aureus with bbp were significantly associated with haematogenous osteomyelitis or arthritis33,34. In our study, none of the isolates was obtained from osteomyelitis or arthritis but we showed that other clinical materials could be an important source of isolates with bbp gene that may be probably also responsible for development of other diseases. Vazquez et al.35 (2011) showed that human fibrinogen is also a ligand for Bbp which indicated that other host extracellular matrix factors can be a Bbp-mediated S. aureus binding site. Campoccia et al.36 showed that co-existence of bbp and cna genes in S. aureus which caused orthopaedic implant infections indicate that adhesins encoded by these genes may act together during the colonization of an orthopaedic implant. In our study, we obtained MRSA isolates, except for blood isolates, in which both the cna and bbp genes were present simultaneously. In our research, we showed that the splE was significantly more frequently detected in wound isolates than in blood isolates isolates. The Spl proteases are a group of six serine proteases that are encoded on the pathogenicity island and are unique to S. aureus. Paharic et al.37 showed that SplA may promote invasion and spreading of S. aureus by removing mucin 16 from epithelial cells. Since mucin 16 is found on ocular, airway, and female reproductive tract epithelial cells, this function could facilitate infection of multiple body sites. The etA, etB and etD genes encode exfoliative toxins (ETs), which are unique serine proteases and are related to human infections38. ETA and ETB selectively recognize and hydrolyze desmosomal proteins in the skin and are responsible for localized epidermal infections such as bullous impetigo and generalized diseases like staphylococcal scalded skin syndrome (SSSS), predominantly in children39. ETA is encoded by the chromosomally located etA gene, while ETB by etB gene located on a large plasmid40. In the present study, we did not obtain MRSA isolates with etA gene, which is consistent with the results shown by other Polish authors23. Whereas, etB gene was present in 15% of MRSA isolates and this frequency was higher than study conducted in Turkey by Demir et al.41 who showed that this gene was present in 9.2% of S. aureus isolates. ETD mediates intra-epidermal cleavage through the granular layer of the epidermis of newborn mice and induces epidermal blisters. The etD gene is encoded in a pathogenicity island on the chromosome42. It was shown that S. aureus strains that produced ETD extracellularly were isolated mainly from other sources of infections and not from patients with bullous impetigo or staphylococcal scalded-skin syndrome. This indicates that ETD might play a role in a broader spectrum of bacterial infections than previously considered42. This is in line with our results because MRSA isolates with the etD gene were isolated from different clinical materials and we did not show significant differences in the frequency of etD in isolates from different sources. The tst gene encodes the toxin of toxic shock syndrome which mediates acute life threatening infections by stimulating release of cytokines such as IL-1, IL-2, TNF-α and others43. The presence of MRSA able to produce a toxic-shock syndrome toxin may increase the risk of the occurrence of more severe symptoms during infection. El-baz et al.44 showed the presence of this gene in about 66% of MRSA isolates, while Eftekhar et al.45 in 51.4%. In our research, this gene was present in all isolates from nose, while in isolates from blood and respiratory tract was detected significantly less often than in wound isolates. Our results are in line with those obtained by other authors who reported high frequency of this gene in MRSA isolates from wound and blood samples45 or respiratory tract46. Among genes encoding staphylococcal enterotoxins, we found the presence of sea gene in 25% of MRSA isolates, while remaining investigated enterotoxin genes were detected in a few isolates. The study of Aggarwal et al.47 concerning S. aureus isolated from a variety of nosocomial infections, showed that in this population 30.2% of strains were positive for sea gene.
Epidemiological monitoring of MRSA infection is essential to control the occurrence and spread of epidemic clones within and between hospitals. In our study, we used spa typing method to distinguish MRSA isolates. This method is based on the sequencing of short repetitive sequences of the polymorphic X region from the gene encoding protein A48,49,50. Main advantage of spa typing is the portability of sequence data which simplifies the sharing of information between laboratories and facilitates the creation of the large-scale database for the study of global and local epidemiology51. In addition, this method can be used to study the molecular evolution and genetic variability of S. aureus and enables the correct assignment of isolates to phylogenetic lines48. Therefore, this method can also be used to differentiate MRSA responsible for infection outbreaks in hospitals, and the results obtained can be compared between laboratories. Many studies showed that the spa types are area specific52,53. In our study, spa typing of MRSA isolates showed high diversity of isolates which indicates that these isolates have emerged from outside the hospitals. Nevertheless, t003 (21.7%), t008 (18.8%), t4474 and t5224 (4.7%) dominated among 45 different spa types. Spa type t003 corresponds to the epidemic clone isolated in the USA and in many European countries51. Several authors showed that spa type t003 is the most predominant among MRSA isolates in hospitals of Southern Poland21,22 and of Northern Poland54. Our results showed that spa type t003 is also the most predominant among MRSA isolates in hospitals of Central Poland. The results also revealed that among MRSA isolates spa type t008 belonged to the most frequent which was in line with the results of Wiśniewska et al.54, who investigated clinical MRSA in Northern Poland.
Conclusion
MRSA isolates responsible for variously localized infections in patients hospitalized in Central Poland showed high variation in terms of the presence of genes encoding important virulence factors. Irrespective of the source of isolation, the genes encoding laminin-, elastin- and fibrinogen binding proteins, serine proteases and toxic shock syndrome toxin-1 were the most commonly identified virulence factors. The results also showed that some genes: cna gene encoding collagen binding protein and map/eap encoding Eap were more common in wound isolates compared to the other sources. The obtained results showed genetic diversity of the analyzed group of MRSA isolates, which indicates that they originated from an outside hospital environment and did not show a relationship. The identification of pathogenic factors of MRSA isolates is important for evaluation of pathogen transmission rate, disease development rate and its severity. Therefore, analysis of potential virulence of MRSA provides information important for choice of appropriate methods of prevention and treatment of MRSA infections. Further studies should focus on development of factors preventing adhesion and biofilm formation by MRSA which would considerably increase treatment effectiveness.
Material and methods
MRSA isolates
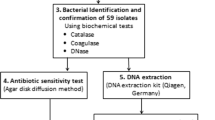
A total of 120 MRSA isolates from human clinical materials such as swabs from wound (30), anus (15), nose (8), blood samples (11), respiratory tract (48), and other samples: swabs from tracheostomy tube, endotracheal tube and catheter and urine (8) were used in this study. The MRSA isolates were obtained from hospitals in Siedlce (83) and Warsaw (37) (Poland) in 2015–2017 (Table 5). The MRSA isolates were collected as part of routine diagnostic microbiology and came from different patients. The numbers of MRSA isolates obtained in 2015, 2016 and 2017 were 18, 24 and 78, respectively. Identification of isolates as S. aureus was confirmed by Gram's staining, positive test for catalase activity and tube test for coagulase. The PCR analysis to amplify the part of the nuc gene, encoding a thermostable nuclease specific for S. aureus was used to confirm that the isolates belonged to the S. aureus species55. The presence of the mecA gene responsible for resistance against β-lactam antibiotics of these isolates was identified by PCR3.
DNA isolation
Genomic DNA from bacterial cells was isolated by using the NucleoSpin Microbial DNA (Macherey–Nagel GmbH&Co.KG, Düren, Germany) according to the manufacturer’s protocol. 2.5 µl of the total extracted material from each test sample was used as a template DNA for PCR application.
Primers and PCR conditions
The primer sequences specific for the genes of adhesins, proteases, gene of toxic shock syndrome toxin-1 and for A, B, C, D and E enterotoxin genes, synthesized at DNA-Gdańsk (Gdańsk, Poland), are listed in Table 6.
The monoplex PCR for each gene was performed in a 25 µL volume containing 2.5 µL of DNA template, 1 × PCR buffer, 0.2 mM each dATP, dCTP, dGTP and dTTP (Fermentas, Vilnius, Lithuania), the specific primers at 200 nM, and 1 U of RedTag Genomic DNA polymerase (Sigma Aldrich, Steinheim, Germany). The PCR method described by Tristan et al.33 was used for detection of genes: cna, fib, fnbB, ebps, bbp, eno encoding adhesins such as collagen-, fibrinogen-, fibronectin B-, elastin-, bone sialoprotein-, laminin-binding protein, respectively. The map/eap gene was amplified by PCR according to Rohde et al.56.
The thermal cycling conditions for detection of genes encoding protease (etA and etB) included predenaturation at 95 °C for 4 min, 35 cycles of denaturation at 95 °C for 0.5 min, primer annealing at 56 °C for 0.5 min and extension at 72 °C for 1 min. The primer annealing at 59 °C was used for the detection of genes coding the remaining proteases (etD, splA, splE, sspA). A 5-min extension at 70 °C was performed at the end of the final cycle. PCR for enterotoxin genes was carried out according to the protocol described earlier by Piechota et al.60. The tst gene coding the toxic shock syndrome toxin-1 was detected in accordance with the method described by Ote et al.57. Amplifications were carried out in the Eppendorf Mastercycler Nexus Gradient (Hamburg, Germany). A negative control was composed of all components of the PCR mixture, except for a template DNA. The positive controls included the genomic DNA from the MRSA isolates in which the presence of the searched genes was previously found in the genome. These controls were included in each test run. The PCR products were analysed by electrophoresis in 1.5% agarose gels stained with ethidium bromide. Molecular size markers (Sigma-Aldrich) were also run for verification of product size. The gel was electrophoresed in a 2 × Tris–borate buffer at 70 V for 1.5 h. The PCR amplicons were visualized using UV light (Syngen Imagine, Syngen Biotech, Wrocław, Poland).
Spa-typing
The highly polymorphic X region of Staphylococcus protein A (spa) gene was amplified by PCR according to Szweda et al.61. Amplified products were purified, and both standards were sequenced using an ABI PRISM® 3100 Genetic Analyzer (Applied Biosystems Hitachi, Waltham, Massachusetts, USA). The nucleotide sequences were analysed to assign the isolates to various types using the spa typing website Ridom SpaServer (http://spaserver.ridom.de).
Data analyses
Chi-squared statistics in Statistix 11.0 (Analytical Software, Tallahassee, FL, USA) was used for testing the significance of differences in the frequency of presence of each gene between pairs of proportions (percentages) of isolates from various isolation sources.
Ethical approval
According to Polish law and the regulations of the Scientific Research Ethics Committee established by the Rector's Order No 128/2020, Siedlce University of Natural Sciences and Humanities, no ethical approval was required for this study and all the data were anonymous.
References
Rocha, L. E. C. et al. Dynamic contact networks of patients and MRSA spread in hospitals. Sci. Rep. 10, 9336. https://doi.org/10.1038/s41598-020-66270-9 (2020).
Turner, N. A. et al. Methicillin-resistant Staphylococcus aureus: An overview of basic and clinical research. Nat. Rev. Microbiol. 17, 203–218. https://doi.org/10.1038/s41579-018-0147-4 (2019).
Kot, B., Wierzchowska, K., Piechota, M. & Grużewska, A. Antimicrobial resistance patterns in methicillin-resistant Staphylococcus aureus from patients hospitalized during 2015–2017 in hospitals in Poland. Med. Princ. Pract. 29, 61–68. https://doi.org/10.1159/000501788 (2020).
Nataraj, B. H. & Mallappa, R. H. Antibiotic resistance crisis: an update on antagonistic interactions between probiotics and methicillin-resistant Staphylococcus aureus (MRSA). Curr. Microbiol. 78, 2194–2211. https://doi.org/10.1007/s00284-021-02442-8 (2021).
Chen, X. et al. Molecular and virulence characteristics of methicillin-resistant Staphylococcus aureus in burn patients. Front Lab Med. 1, 43–47. https://doi.org/10.1016/j.flm.2017.02.010 (2017).
Wang, X. et al. Molecular characteristics of community-associated Staphylococcus aureus isolates from paediatric patients with bloodstream infections between 2012 and 2017 in Shanghai China. Front. Microbiol. 9, 1211. https://doi.org/10.3389/fmicb.2018.01211 (2018).
Ahmadishoar, S. et al. Genotypic and phenotypic characterisation of clinical isolates of methicillin-resistant Staphylococcus aureus in two different geographical locations of Iran. Indian J. Med. Microbiol. 38, 162–168. https://doi.org/10.4103/ijmm.IJMM_20_153 (2020).
Hassoun, A., Linden, P. K. & Friedman, B. Incidence, prevalence, and management of MRSA bacteraemia across patient populations—a review of recent developments in MRSA management and treatment. Crit. Care. 21, 211. https://doi.org/10.1186/s13054-017-1801-3 (2017).
Lim, W. W. et al. Determinants of MRSA prevalence in the Asia Pacific Region: A systematic review and meta-analysis. J. Glob. Antimicrob. Resist. 16, 17–27. https://doi.org/10.1016/j.jgar.2018.08.014 (2018).
Pinto, R. M. et al. Impact of nanosystems in Staphylococcus aureus biofilms treatment. FEMS Microbiol. Rev. 43, 622–641. https://doi.org/10.1093/femsre/fuz021 (2019).
Ansari, S. et al. Recent advances in Staphylococcus aureus infection: Focus on vaccine development. Infect. Drug Resist. 12, 1243–1255. https://doi.org/10.2147/IDR.S175014 (2019).
Huttunen, R. et al. Nosocomial blood stream infections in a Finnish tertiary care hospital: a retrospective cohort study of 2175 episodes during the years 1999–2001 and 2005–2010. Infect. Dis. 47, 20–26. https://doi.org/10.3109/00365548.2014.956791 (2015).
Tadesse, S. et al. Antimicrobial resistance profile of Staphylococcus aureus isolated from patients with infection at Tikur Anbessa Specialized Hospital, Addis Ababa Ethiopia. BMC Pharmacol. Toxicol. 19, 24. https://doi.org/10.1186/s40360-018-0210-9 (2018).
Kot, B. et al. Phenotypic and genotypic antimicrobial resistance of staphylococci from bovine milk. Pol. J. Vet. Sci. 15, 677–683. https://doi.org/10.2478/v10181-012-0105-4 (2012).
Zhen, X. et al. Clinical and economic impact of methicillin-resistant Staphylococcus aureus: A multicentre study in China. Sci. Rep. 10, 3900. https://doi.org/10.1038/s41598-020-60825-6 (2020).
Liu, Y., Zhang, J. & Ji, Y. Environmental factors modulate biofilm formation by Staphlococcus aureus. Sci. Prog. 103. https://doi.org/10.1177/0036850419898659 (2020).
Moormeier, D. E. & Bayles, K. W. Staphylococcus aureus biofilm: a complex developmental organism. Mol. Microbiol. 104, 365–376. https://doi.org/10.1111/mmi.13634 (2017).
Harraghy, N. et al. The adhesive and immunomodulating properties of the multifunctional Staphylococcus aureus protein Eap. Microbiology 149, 2701–2707. https://doi.org/10.1099/mic.0.26465-0 (2003).
Hussain, M. et al. Insertional inactivation of eap in Staphylococcus aureus strain newman confers reduced staphylococcal binding to fibroblasts. Infect. Immun. 70, 2933–2940. https://doi.org/10.1128/IAI.70.6.2933-2940.2002 (2002).
Dunyach-Remy, C., Essebe, C. N., Sotto, A. & Lavigne, J. P. Staphylococcus aureus toxins and diabetic foot ulcers: Role in pathogenesis and interest in diagnosis. Toxins. 8, 209. https://doi.org/10.3390/toxins8070209 (2016).
Gajda, M. et al. Virulence and drug-resistance of Staphylococcus aureus strains isolated from venous ulcers in polish patients. Int. J. Environ. Res. Public Health. 18, 4662. https://doi.org/10.3390/ijerph18094662 (2021).
Pomorska-Wesołowska, M. et al. Virulence and antimicrobial resistance of Staphylococcus aureus isolated from bloodstream infections and pneumonia in Southern Poland. J. Glob. Antimicrob. Resist. 11, 100–104. https://doi.org/10.1016/j.jgar.2017.07.009 (2017).
Kasela, M., Grzegorczyk, A., Nowakowicz-Dębek, B. & Malm, A. The prevalence of virulence determinants and antibiotic resistance patterns in methicillin-resistant Staphylococcus aureus in a nursing home in Poland. Pathogens. 10, 427. https://doi.org/10.3390/pathogens10040427 (2021).
Carneiro, C. R. et al. Identification of enolase as a laminin-binding protein on the surface of Staphylococcus aureus. Microbes. Infect. 6, 604–608. https://doi.org/10.1016/j.micinf.2004.02.003 (2004).
Haghi Ghahremanloi Olia, A., Ghahremani, M., Ahmadi, A. & Sharifi, Y. Comparison of biofilm production and virulence gene distribution among community- and hospital-acquired Staphylococcus aureus isolates from northwestern Iran. Infect. Genet. Evol. 81, 104262. https://doi.org/10.1016/j.meegid.2020.104262 (2020).
Downer, R. et al. The elastin-binding protein of Staphylococcus aureus (EbpS) is expressed at the cell surface as an integral membrane protein and not as a cell wall-associated protein. J. Biol. Chem. 277, 243–250. https://doi.org/10.1074/jbc.M107621200 (2002).
Kot, B., Sytykiewicz, H. & Sprawka, I. Expression of the biofilm-associated genes in methicillin-resistant Staphylococcus aureus in biofilm and planktonic conditions. Int. J. Mol. Sci. 19, 3487. https://doi.org/10.3390/ijms19113487 (2018).
Azmi, K., Qrei, W. & Abdeen, Z. Screening of genes encoding adhesion factors and biofilm production in methicillin resistant strains of Staphylococcus aureus isolated from Palestinian patients. BMC Genom. 20, 578. https://doi.org/10.1186/s12864-019-5929-1 (2019).
Hartford, O. M., Wann, E. R., Höök, M. & Foster, T. J. Identification of residues in the Staphylococcus aureus fibrinogen-binding MSCRAMM clumping factor A (ClfA) that are important for ligand binding. J. Biol. Chem. 276, 2466–2473. https://doi.org/10.1074/jbc.M007979200 (2001).
Foster, T. et al. Adhesion, invasion and evasion: the many functions of the surface proteins of Staphylococcus aureus. Nat. Rev. Microbiol. 12, 49–62 (2014).
Patti, J. M., Allen, B. L., McGavin, M. J. & Hook, M. MSCRAMM-mediated adherence of microorganisms to host tissues. Annu. Rev. Microbiol. 48, 585–617. https://doi.org/10.1146/annurev.mi.48.100194.003101 (1994).
Szczuka, E., Urbanska, K., Pietryka, M. & Kaznowski, A. Biofilm density and detection of biofilm-producing genes in methicillin-resistant Staphylococcus aureus strains. Folia Microbiol. (Praha) 58, 47–52. https://doi.org/10.1007/s12223-012-0175-9 (2013).
Tristan, A., Ying, M. B., Etienne, J., Vandenesch, F. & Lina, G. Use of multiplex PCR to identify Staphylococcus aureus adhesins involved in human hematogenous infections. J. Clin. Microbiol. 41, 4465–4467. https://doi.org/10.1128/JCM.41.9.4465-4467.2003 (2003).
Wiśniewska, K., Piórkowska, A., Kasprzyk, J., Bronk, M. & Świeć, K. Clonal distribution of bone sialoprotein-binding protein gene among Staphylococcus aureus isolates associated with bloodstream infections. Folia Microbiol. (Praha) 59, 465–471. https://doi.org/10.1007/s12223-014-0321-7 (2014).
Vazquez, V. et al. Fibrinogen is a ligand for the Staphylococcus aureus microbial surface components recognizing adhesive matrix molecules (MSCRAMM) bone sialoprotein binding protein (Bbp). J. Biol. Chem. 286, 29797–29805. https://doi.org/10.1074/jbc.M110.214981 (2011).
Campoccia, D. et al. The presence of both bone sialoprotein-binding protein gene and collagen adhesin gene as a typical virulence trait of the major epidemic cluster in isolates from orthopedic implant infections. Biomaterials 30, 6621–6628. https://doi.org/10.1016/j.biomaterials.2009.08.032 (2009).
Paharik, A.E. et al. The Spl serine proteases modulate Staphylococcus aureus protein production and virulence in a rabbit model of pneumonia. mSphere.1, e00208–16. https://doi.org/10.1128/mSphere.00208-16 (2016).
Mariutti, R. B. et al. Crystal structure of Staphylococcus aureus exfoliative toxin D-like protein: Structural basis for the high specificity of exfoliative toxins. Biochem. Biophys. Res. Commun. 467, 171–177. https://doi.org/10.1016/j.bbrc.2015.08.083 (2015).
Bukowski, M., Wladyka, B. & Dubin, G. Exfoliative toxins of Staphylococcus aureus. Toxins (Basel). 2, 1148–1165. https://doi.org/10.3390/toxins2051148 (2010).
Yamaguchi, T. et al. Phage conversion of exfoliative toxin A production in Staphylococcus aureus. Mol. Microbiol. 38, 694–705. https://doi.org/10.1046/j.1365-2958.2000.02169.x (2000).
Demir, C. et al. Investigation of toxin genes in Staphylococcus aureus strains isolated in Mustafa Kemal University Hospital. Turk. J. Med. Sci. 41, 343–352. https://doi.org/10.3906/sag-1003-657 (2011).
Yamaguchi, T. et al. Identification of the Staphylococcus aureus etd pathogenicity island which encodes a novel exfoliative toxin, ETD, and EDIN-B. Infect. Immun. 70, 5835–5845. https://doi.org/10.1128/IAI.70.10.5835-5845.2002 (2002).
Wilson, G. J. et al. A novel core genome-encoded superantigen contributes to lethality of community associated MRSA necrotizing pneumonia. PLoS Pathog. 7, e1002271. https://doi.org/10.1371/journal.ppat.1002271 (2011).
El-Baz, R., Rizk, D. E., Barwa, R. & Hassan, R. Virulence characteristics and molecular relatedness of methicillin resistant Staphylococcus aureus harbouring different staphylococcal cassette chromosome mec. Microb. Pathog. 113, 385–395. https://doi.org/10.1016/j.micpath.2017.11.021 (2017).
Eftekhar, F. et al. Distribution of adhesion and toxin genes in Staphylococcus aureus strains recovered from hospitalized patients admitted to the ICU. Arch. Pediatr. Infect. Dis. 5, e39349. https://doi.org/10.5812/pedinfect.39349 (2017).
Lim, K.T., Hanifah, Y.A., Mohd Yusof, M.Y. & Thong, K.L. Investigation of toxin genes among methicillin-resistant Staphylococcus aureus strains isolated from a tertiary hospital in Malaysia. Trop. Biomed. 29, 212–9 (2012).
Aggarwal, S. et al. Antibiotic susceptibility, virulence pattern, and typing of Staphylococcus aureus strains isolated from variety of infections in India. Front. Microbiol. 10, 2763. https://doi.org/10.3389/fmicb.2019.02763 (2019).
Koreen, L. et al. spa typing method for discriminating among Staphylococcus aureus isolates: implications for use of a single marker to detect genetic micro- and macrovariation. J. Clin. Microbiol. 42, 792–799. https://doi.org/10.1128/JCM.42.2.792-799.2004 (2004).
Sabat, A. J. et al. Overview of molecular typing methods for outbreak detection and epidemiological surveillance. Euro. Surveill. 18, 20380. https://doi.org/10.2807/ese.18.04.20380-en (2013).
Mazi, W., Sangal, V., Sandstrom, G., Saeed, A. & Yu, J. Evaluation of spa-typing of methicillin-resistant Staphylococcus aureus using high-resolution melting analysis. Int. J. Infect. Dis. 38, 125–128. https://doi.org/10.1016/j.ijid.2015.05.002 (2015).
Strommenger, B. et al. Spa typing of Staphylococcus aureus as a frontline tool in epidemiological typing. J. Clin. Microbiol. 46, 574–581. https://doi.org/10.1128/JCM.01599-07 (2008).
Hetem, D. J. et al. Molecular epidemiology of MRSA in 13 ICUs from eight European countries. J. Antimicrob. Chemother. 71, 45–52. https://doi.org/10.1093/jac/dkv298 (2016).
Fasihi, Y., Kiaei, S. & Kalantar-Neyestanaki, D. Characterization of SCCmec and spa types of methicillin-resistant Staphylococcus aureus isolates from health-care and community-acquired infections in Kerman Iran. J. Epidemiol. Glob. Health. 7, 263–267. https://doi.org/10.1016/j.jegh.2017.08.004 (2017).
Wiśniewska, K. et al. The use of spa and phage typing for characterization of clinical isolates of methicillin-resistant Staphylococcus aureus in the University Clinical Center in Gdańsk Poland. Folia Microbiol. (Praha) 57, 243–249. https://doi.org/10.1007/s12223-012-0148-z (2012).
Brakstad, O. G., Aasbakk, K. & Maeland, J. A. Detection of Staphylococcus aureus by polymerase chain reaction amplification of the nuc gene. J. Clin. Microbiol. 30, 1654–1660 (1992).
Rohde, H. et al. Polysaccharide intercellular adhesin or protein factors in biofilm accumulation of Staphylococcus epidermidis and Staphylococcus aureus isolated from prosthetic hip and knee joint infections. Biomaterials 28, 1711–1720. https://doi.org/10.1016/j.biomaterials.2006.11.046 (2007).
Ote, I., Taminiau, B., Duprez, J. N., Dizier, I. & Mainil, J. G. Genotypic characterization by polymerase chain reaction of Staphylococcus aureus isolates associated with bovine mastitis. Vet. Microbiol. 153, 285–292. https://doi.org/10.1016/j.vetmic.2011.05.042 (2011).
Park, J. Y. et al. Detection of classical and newly described staphylococcal superantigen genes in coagulase-negative staphylococci isolated from bovine intramammary infections. Vet. Microbiol. 147, 149–154. https://doi.org/10.1016/j.vetmic.2010.06.021 (2011).
Becker, K., Roth, R. & Peters, G. Rapid and specific detection of toxigenic Staphylococcus aureus: use of two multiplex PCR enzyme immunoassays for amplification and hybridization of staphylococcal enterotoxin genes, exfoliative toxin genes, and toxic shock syndrome toxin 1 gene. J. Clin. Microbiol. 36, 2548–2553. https://doi.org/10.1128/JCM.36.9.2548-2553 (1998).
Piechota, M. et al. Distribution of classical enterotoxin genes in staphylococci from milk of cows with– and without mastitis and the cowshed environment. Pol. J. Vet. Sci. 17, 407–411. https://doi.org/10.2478/pjvs-2014-0058 (2014).
Szweda, P., Schielmann, M., Milewski, S., Frankowska, A. & Jakubczak, A. Biofilm production and presence of ica and bap genes in Staphylococcus aureus strains isolated from cows with mastitis in the eastern Poland. Pol. J. Microbiol. 61, 65–69 (2012).
Funding
This research was carried out with the financial support of Siedlce University of Natural Science and Humanities (Scientific Research Project No. 74/20/B).
Author information
Authors and Affiliations
Contributions
B.K. conceptualization, methodology, formal analysis and investigation, original draft preparation; M.P. investigation; A.J. methodology, investigation; M.G. methodology, investigation; M.W. investigation; A.G. investigation; K.B. investigation; P.D. investigation. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kot, B., Piechota, M., Jakubczak, A. et al. The prevalence of virulence determinants in methicillin-resistant Staphylococcus aureus isolated from different infections in hospitalized patients in Poland. Sci Rep 12, 5477 (2022). https://doi.org/10.1038/s41598-022-09517-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-09517-x
- Springer Nature Limited