Abstract
Familial Hypercholesterolemia (FH) is an inherited disorder of lipid metabolism, characterized by increased low density lipoprotein cholesterol (LDLc) levels. The main purpose of the current work was to explore alternative classification methods to traditional clinical criteria for FH diagnosis, based on several biochemical and biological indicators. Logistic regression (LR), decision tree (DT), random forest (RF) and naive Bayes (NB) algorithms were developed for this purpose, and thresholds were optimized by maximization of Youden index (YI). All models presented similar accuracy (Acc), specificity (Spec) and positive predictive values (PPV). Sensitivity (Sens) and G-mean values were significantly higher in LR and RF models, compared to the DT. When compared to Simon Broome (SB) biochemical criteria for FH diagnosis, all models presented significantly higher Acc, Spec and G-mean values (p < 0.01), and lower negative predictive value (NPV, p < 0.05). Moreover, LR and RF models presented comparable Sens values. Adjustment of the cut-off point by maximizing YI significantly increased Sens values, with no significant loss in Acc. The obtained results suggest such classification algorithms can be a viable alternative to be used as a widespread screening method. An online application has been developed to assess the performance of the LR model in a wider population.
Similar content being viewed by others
Introduction
Familial Hypercholesterolemia (FH) is an autosomal dominant disorder of lipid metabolism, characterized by increased low density lipoprotein cholesterol (LDLc) levels1. The high cholesterol levels from birth lead to its accumulation in arterial walls, promoting the early development of atherosclerosis, which represents a major risk factor for cardiovascular disease (CVD)1,2. This disorder can be divided into a heterozygous (HeFH), and a more severe homozygous form (HoFH). Due to the fact that it can go undetected for many years, HeFH is the focus of the present study, and unless stated otherwise, will be referred simply as FH. FH is caused by mutations in three identified genes that encode key proteins involved in the LDL receptor (LDLr) endocytic and recycling pathways, LDL receptor (LDLR), apolipoprotein B (APOB) and proprotein convertase subtilisin kexin type 9 (PCSK9), which account for around 90%, 5–10% and 1–5% of FH cases respectively3,4,5. A small percentage of FH-like phenotype cases is attributed to rare variants in other genes linked to dyslipidaemia, or to a form of polygenic hypercholesterolemia2,6. Recent studies suggest worldwide FH prevalence rates to be between 1:200 and 3007,8.
The early diagnosis of FH has been associated with a significant reduction in CVD risk, supporting the introduction of adequate and more aggressive therapeutic measures9. There are different clinical criteria available for the diagnosis of FH, although only genetic testing can positively confirm the diagnostic. The importance of molecular diagnosis to characterize FH is evidenced by large cohort studies, in which individuals with clinical criteria for FH that are confirmed to have a causative pathogenic variant, present a significant increase in the risk of coronary heart disease compared to clinical FH patients in whom a causative variant is not found10,11. However, because molecular diagnosis is costly and time-consuming, it cannot be used as a first line screening method, and previous selection of patients based on more accessible parameters should be performed. Simon Broome (SB) clinical criteria are among the most commonly used in the paediatric population, and are based on LDLc and total cholesterol (TC) levels, family history and presence of tendon xanthomas4,12. Since physical signs are a rare finding in individuals at paediatric age6, and information concerning family history is frequently absent13, the decision regarding a possible FH diagnosis is often made based solely on biochemical criteria. Due to the fact biochemical cut-off values used in SB criteria are very conservative, application of these criteria results in a high false positive rate when compared to molecular study results6,7. This issue constitutes a heavy burden in terms of healthcare costs, limiting the access of a larger universe of true FH cases to the genetic study.
There are several algorithms available to handle a classification problem with a binary response variable, in this case representing the positive or negative diagnosis for FH. Logistic regression (LR) is a special case of the generalized linear models methodology. The expected value of the dependent variable given by the logistic function represents the probability of the outcome variable to be FH14. The decision tree (DT) model is an algorithm derived from information theory. The classification rule in DT is created by repeatedly dividing the data into increasingly more homogeneous groups, with respect to the variable of interest, a method defined as recursive partitioning15,16. The RF algorithm aggregates the results of a multitude of individual DT to classify or predict an observation17. Two components of randomness are introduced into the construction of the individual trees. Firstly, each tree is built using a random bootstrapped sample of the training data, a method known as bagging18. Secondly, a random subset of predictor variables is tested in each of the trees, a procedure designated as the random subspace selection method19. Naïve Bayes (NB) is a probabilistic classifier based on Bayes’ theorem. This classification algorithm earned the term “naïve” since it relies on the strong, and often erroneous assumption, that predictor features are conditionally independent. Despite its relative simplicity, NB classifier has been shown to outperform even highly sophisticated classification algorithms20.
A common way to summarize the results of a classification model is through a confusion matrix, a contingency table where the observed outcome is cross-classified with the predicted outcome. Different operating characteristics (OC) can then be derived, and used to assess the performance of the model21. For this purpose, it is necessary to define the cut-off value that best discriminates successes from failures, to use as the classification rule. The default choice of a cut-off value of probability equal to 0.5, is not always the best decision. A particularly useful criterion in clinical diagnostic procedures where the two classes are imbalanced is to maximize Sens and Spec summation, also known as Youden index (YI). This method attributes equal importance to Sens and Spec values, ignoring the relative size of the populations22,23,24.
Previous studies have reported an overall good performance of LR25,26,27,28, RF28,29 and DT models30 in identifying FH cases. To the knowledge of the authors, there are no published studies assessing the performance of a NB classifier for FH diagnosis. The current study presents however several differences in relation to previous work. Most of the mentioned articles were developed using adult cohorts, while the current paper focuses on the paediatric population. The mentioned studies also have included a vast number of predictor variables routinely available in primary care, while in the current work, fewer parameters, including specific biochemical markers, were included. Another important difference is the fact that ROC curve analysis has been incorporated in the current study, in order to adjust for a cut-off value to optimize Sens and Spec values, hence addressing the class imbalance problem.
The main purpose of this work was to explore alternative classification procedures for FH diagnosis, based on different biological and biochemical indicators, with improved ability to screen for FH cases in comparison to traditional clinical criteria. The classification algorithms developed for this purpose were LR, NB RF and DT models. For the first three methods, the threshold was further adjusted by maximizing YI, in order to account for class imbalance issues.
Methods
Study sample
The sample used in this work was taken from the Portuguese FH study, an ongoing study started in 1999, with the purpose of identifying and characterizing FH in the Portuguese population31. 389 observations, corresponding to index patients at paediatric age (2 to 17 years), of both sexes, meeting clinical criteria for hypercholesterolemia (TC ≥ 170 mg/dL or LDLc ≥ 110 mg/dL)32, were initially retrieved from the Portuguese FH database. All subjects were white, of European ancestry. From this initial set of data, subjects under hypolipidemic medication at time of biochemical assessment were excluded, together with cases presenting a variant of unknown significance, a monogenic variant in a FH phenocopy gene or HoFH. The final dataset was comprised of 286 paediatric patients, of which 104 had a positive molecular diagnosis for FH. At time of assessment, participants were receiving standard healthcare and nutritional advice. All participants had an informed consent form signed by the legal guardian, and information was registered in a confidential database, approved by the National Data Protection Commission. The study complies with the Declaration of Helsinki and was approved by the ethics committee of the Instituto Nacional de Saúde Doutor Ricardo Jorge (INSA), in Lisbon.
Candidate predictor and outcome variables
Serum concentrations for a panel of several biochemical variables related to lipid metabolism were used as candidate predictor variables: TC, LDLc, high density lipoprotein cholesterol (HDLc), triglycerides (TG), apolipoproteins AI (ApoAI) and B (ApoB), and lipoprotein(a) (Lp(a)). Concentrations were determined in mg/dL, by enzymatic and colorimetric methods, using a Cobas Integra 400 Plus (Roche) analyser33. Additional information, regarding biological and anthropometric variables was also included, specifically age, sex and body mass index z-scores (zBMI), calculated according to the World Health Organization (WHO) standards34,35. Molecular diagnosis was performed by the study of the LDLR, APOB and PCSK9 genes, through fragment amplification by PCR, followed by direct sequencing using Sanger’s method. The promotor, coding and splicing regions of the 18 exons of LDLR gene, plus fragments of exons 26 and 29 in APOB gene, and five target exons (1, 2, 4, 6 and 9) and flanking regions in PCSK9 gene were studied, according to the protocol described by Medeiros et al.36. Large rearrangements in LDLR gene were also searched through MLPA technique. The found variants were classified according to the American College of Medical Genetics and Genomics (ACMG) guidelines37. Participants with a positive molecular diagnosis were classified as FH, and participants with a negative molecular diagnosis classified as non-FH.
Classification algorithms and model comparison
Exploratory and significance analysis of biological and biochemical variables was initially performed for FH and non-FH subjects. Missing values were imputed by means of k-nearest neighbours method using Gower’s distance38. Variable selection for the LR model was performed by purposeful selection methods, after assessing for collinearity. Continuous predictor variables were previously evaluated for linearity for the logit, and when non-linearity occurred, variables were categorized. For the NB model, variables for which significant differences between FH and non-FH patients were verified, and presenting low correlation values were retained. Between highly correlated variables, selection of the variable to retain was made through ROC curve analysis. Continuous variables were log-transformed, whenever a better adjustment to the Normal distribution was provided. The DT was built based on information gain measures, using the rpart package from R16. The final tree was pruned to avoid overfitting, adopting a complexity parameter (cp) of 0.01. Search for an optimal RF was conducted iteratively, using out-of-bag (OOB) Acc estimates for performance comparisons. In a first step, hyperparameter tuning was performed for ntree, mtry and node size, which respectively represent the total number of DT used, the number of predictor variables sampled for each DT, and the amount of pruning in each DT. RF with ntree between 10 and 2000, by increments of 10 trees, mtry ranging from 1 to 7 variables, and node size = 1, 5, 10, 15 and 20 were explored. In a second step, less informative variables were sequentially excluded from the initial set of candidate predictors. The RF model was developed and implemented using the randomForest package from R39.
A new cut-off value based on YI was calculated for LR, RF and NB models, in opposition to the default cut-off c = 0.5. For all classification algorithms, and respective cut-off values, a confusion matrix was generated, by comparison with molecular study results, and different OC were calculated: Acc, Sens, Spec, PPV, NPV (negative predictive value) and G-mean21,40.
Model comparison was performed by means of tenfold cross validation (CV). For each fold, mean values of different OC were calculated (an example of the source code used to test the models is provided on Supplementary Methods S1). Models were compared with SB biochemical criteria (TC > 260 mg/dL or LDLc > 155 mg/dL) regarding these OC, by means of Wilcoxon signed-rank (MWW) test. A diagram of overall study design is represented on Fig. 1. Only biochemical cutpoints in SB criteria were considered, since familial history of elevated cholesterol or CVD is not always available in standard clinical records, and physical signs are very rare to find in paediatric patients6. For each OC, different classification algorithms were compared among each other through Friedman test, and in cases where significant differences were observed, pairwise comparison was performed through MWW test. Bonferroni method was used to correct for multiple testing. Statistical analysis was performed using R and R Studio software (v3.5.2), adopting a significance level of α = 0.05.
Results
Exploratory analysis
Biological and biochemical characteristics for FH and non-FH patients are presented in Table 1. Significant differences were observed for all predictor variables (p < 0.05), except for Age (p = 0.19), zBMI (p = 0.1) and Lp(a) (p = 0.17). The 3 non-significant variables were also non-linear with the logit, and were therefore categorized. Age was divided into approximately equal range age groups (2–7, 8–12 and 13–17 years), overweight was defined as a zBMI > 1, following the WHO guidelines35, and high Lp(a) was defined as Lp(a) serum concentrations above 50 mg/dL, as this cut-off value has been previously established as a risk factor for CVD41.
Around 36% of the sample presented a positive molecular diagnosis for FH, the great majority with an identified pathogenic variant in the LDLR gene (94%), and remaining subjects in APOB gene. FH subjects presented higher values for TC, LDLc, and ApoB, and lower values for HDLc, TG, ApoAI and Lp(a) (p < 0.05). Missing values were detected for the variables zBMI, ApoAI, ApoB and Lp(a), with a percentage of missing cases ranging from 2 to 10%.
Trained classification algorithms
Final model fit for the LR model is presented in Table 2.
The final LR model included the continuous variables LDLc, TG and ApoAI, and the categorical variables overweight, sex and high Lp(a). Variable selection was the same as the one obtained for NB model, except the fact the log of continuous variables was adopted in the latter case, since better adjustment to Normal distribution has been shown.
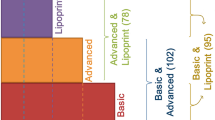
A representation of the pruned DT model, with respective cut-off values at each node, and showing the tree performance in the complete learning set, is shown in Fig. 2.
Decision tree model. At each node, it is represented the biochemical indicator used to divide the sample, the respective cut-off value, and the way the original sample is divided throughout the tree. FH familial hypercholesterolemia, LDLc low density lipoprotein cholesterol, in mg/dL, TG triglycerides, in mg/dL, ApoAI apolipoprotein AI, in mg/dL, pos positive cases.
Regardless of chosen mtry, RF performance seemed to stabilize from around 700 ntree. Since increasing the number of trees does not deteriorate RF predictive ability, and computational costs are not very high for the used sample size and number of predictors, a ntree = 1000 was defined. The model seemed to achieve better performance without inclusion of variables with very small information loss values (≈1), hence the final RF included only six candidate predictors: TC, LDLc, HDLc, TG, ApoAI and ApoB. For this RF model, optimal mtry = 3 and node size = 1 were defined.
Model comparison
The performance of the different classification models regarding the several OC, as obtained by tenfold CV, together with values that result from the application of SB criteria to the entire sample, are represented in Fig. 3. Corresponding mean values obtained for the different OC by each classification method can be consulted in Supplementary Table S1.
Results concerning OC comparison between pairs of classification algorithms for which significant differences were found, as well as between each classification algorithm and SB criteria, are presented in Table 3.
Finally, in Table 4, the mean OC values obtained by the chosen YI method are compared to the ones obtained using the default cut-off value c = 0.5, for each classification method.
Discussion
Four different classification algorithms were implemented and tested in the current study: LR, DT, RF and NB. The cut-off point for three of these models was further adjusted through ROC curve analysis, and defined as the value which maximized YI. This process was not applied to the DT algorithm, since very few candidate cut-points are provided by this model. Due to the fact YI maximization method attributes equal importance to Sens and Spec, it is considered particularly useful in classification problems in which the two classes are imbalanced24. This was the case in the current study, and also in most clinical diagnostic procedures, since the occurrence of a given disease is generally a relatively rare event. Models were posteriorly tested through tenfold CV.
When comparing the different models to each other (Table 3), significant differences were found in G-mean, Sens and NPV values, with the DT model presenting significantly lower values than the LR and the RF models regarding these OC (p < 0.05). Since the DT will return as splitting rule the cut-off point that decreases the total entropy of the system the most, it will implicitly favour the majority class, thus decreasing Sens and NPV values. Several techniques have been suggested to deal with this class imbalance problem, ranging from data sampling methods, to algorithmic modifications, to cost-sensitivity learning42. Further work is being prepared incorporating some of these features.
When compared to SB biochemical criteria, all classification algorithms presented significantly higher Acc, G-mean, Spec and PPV (p < 0.01). This means that overall classification error is smaller, and also that individuals with a positive screening following these methods are more likely to truly be FH. On the other hand, the DT and NB models revealed significantly lower Sens levels than SB criteria (p < 0.05). Elevated Sens in SB criteria is essentially due to very conservative cut-off values, which results in a high number of false positive cases7. Reducing the total number of potential candidates to be submitted to molecular diagnosis can have important repercussions in terms of the process cost-effectiveness, and should therefore be taken into account.
Previous studies applying these classification algorithms to FH diagnosis have reported an overall good performance in identifying FH cases25,26,27,28,29,30. Specifically, Niehaus et al.28 have studied the performance of a RF against a LR model for FH diagnosis in adults, finding an improved AUC (0.91 vs 0.82), Sens (0.61 vs 0.56) and Spec (0.96 vs 0.91) value for the RF model. One of the potential factors that can account for not observing such differences in the current study, is the fact relatively few predictor variables are used in this work, therefore decreasing the variability of the constituent trees of the RF. Additionally, although overall performance for LR and RF models was similar to the one observed in previous studies25,26,27,28,29, as assessed by the AUC, Sens levels were higher in the current work, and Spec values more reduced. Such differences are attributed to the fact that cut-off values in the current study have been defined by maximizing YI, which will make the cutpoint invariably lower, increasing Sens and decreasing Spec values. An important observation that arises from these results is that it is possible to keep overall Acc of a given classifier, while increasing Sens levels significantly (Table 4). This is particularly relevant in a clinical screening procedure, where besides obtaining a small classification error, is very important to retain as much disease-positive cases as possible.
Another interesting finding in the current work is the fact that, for all classification algorithms, the most relevant variables selected to classify the individual seem to be LDLc, followed by TG and ApoAI. While LDLc concentration was directly related to the probability of being FH positive, TG and ApoAI levels were inversely associated with the presence of this disorder. While the effect of FH on LDLc metabolism has been extensively studied1,2,3,4,5,6,7,9, fewer studies that have investigated the relation of this genetic disorder with TG43, and ApoAI or HDLc44 levels, support the lower levels of these biomarkers found FH individuals. A thorough discussion of the variables included in this study from a metabolic perspective is being addressed in a separate article.
In sum, any of the tested classification algorithms presented higher Acc, Spec and PPV when compared to SB biochemical criteria (p < 0.01). Additionally, LR and RF models presented similar Sens values than SB biochemical criteria, revealing the ability to detect the same rate of FH cases, with much less false positive retention. An improved AUC for LR model, in relation to RF, may favour the selection of this algorithm as a tool for FH classification in the paediatric population. In practice, this means that for a given laboratory annual budget p.e., we would obtain a significantly higher number of cases with a confirmed molecular diagnosis of FH, which would allow adjusting pharmacotherapy, and optimizing subsequent screening measures, by means of cascade screening. As for patients that are classified as non-FH, although it remains possible that a small percentage might carry a causative monogenic variant, as already happens with established clinical criteria, a large proportion of these individuals may have a form of polygenic hypercholesterolemia6, or dyslipidaemia triggered essentially by environmental factors, which as already stated, have a significantly lower associated CVD risk10,11.
A pilot internet application, implementing LR algorithm, with cut-off values defined by maximizing YI, can be consulted at https://fhidtool.shinyapps.io/dyslipid/. This application was developed using R Studio Shiny package, and named FH.ID.Tool. Paediatricians that collaborate with the Portuguese FH study will be encouraged to use and provide comments to improve the application utilities and interface to a final version, that will then be proposed for implementation at the national health system. Future directions of this work will comprise the development of classification models for the Portuguese adult population, validation of such models on national FH studies of different populations, and study of the relevance of additional biomarkers for FH classification.
Conclusions
Several conclusions can be taken from the current study, regarding the performance of the different classification algorithms. Higher Sens and G-mean values were achieved with the LR and RF models, while the DT model was the one that presented lowest values regarding these OC (p < 0.05). No significant differences were observed between methods regarding Acc, Spec and PPV values. In comparison with SB biochemical criteria, all classification models presented significantly higher Acc, Spec and PPV (p < 0.01). However, even after correction for multiple comparisons, the DT model still presented lower Sens values (p < 0.01). Low Sens values can be a problem in a screening procedure like the one presented in the current study, where retaining the highest possible percentage of FH cases is a major concern. Regarding this matter, cut-off values can be adjusted, based on ROC curve analysis, in order to increase Sens significantly, while maintaining Acc levels. Based on the results of the current study, LR may be considered the most parsimonious model, when accounting for all OC. The implementation of the proposed LR algorithm as a screening tool will allow retaining a similar number of FH paediatric cases to the clinical diagnosis criteria, while excluding a significant amount of false positive cases. The widespread implementation of such a tool may be beneficial in terms of cost-effectiveness.
References
Najam, O. & Ray, K. K. Familial hypercholesterolemia: A review of the natural history, diagnosis, and management. Cardiol. Ther. 4(1), 25–38 (2015).
Bouhairie, V. E. & Goldberg, A. C. Familial hypercholesterolemia. Cardiol. Clin. 33(2), 169–179 (2015).
Sharifi, M., Futema, M., Nair, D. & Humphries, S. E. Genetic architecture of familial hypercholesterolaemia. Curr. Cardiol. Rep. 19(5), 44 (2017).
Austin, M. A., Hutter, C. M., Zimmern, R. L. & Humphries, S. E. Genetic causes of monogenic heterozygous familial hypercholesterolemia: A HuGE prevalence review. Am. J. Epidemiol. 160, 407–420 (2004).
Henderson, R., O’Kane, M., McGilligan, V. & Watterson, S. The genetics and screening of familial hypercholesterolaemia. J. Biomed. Sci. 23(1), 39 (2016).
Defesche, J. C. et al. Familial hypercholesterolaemia. Nat. Rev. Dis. Primers 3(1), 1–20 (2017).
Nordestgaard, B. G. et al. Familial hypercholesterolaemia is underdiagnosed and undertreated in the general population: Guidance for clinicians to prevent coronary heart disease: Consensus statement of the European Atherosclerosis Society. Eur. Heart J. 34(45), 3478–3490 (2013).
Beheshti, S. O., Madsen, C. M., Varbo, A. & Nordestgaard, B. G. Worldwide prevalence of familial hypercholesterolemia: Meta-analyses of 11 million subjects. J. Am. Coll. Cardiol. 75(20), 2553–2566 (2020).
Martin, A. C., Gidding, S. S., Wiegman, A. & Watts, G. F. Known and unknowns in the care of paediatric familial hypercholesterolaemia. J. Lipid Res. 58, 1765–1776 (2017).
Abul-Husn, N. S. et al. Genetic identification of familial hypercholesterolemia within a single US health care system. Science 354(6319), 7000 (2016).
Patel, A. P. et al. Association of rare pathogenic DNA variants for familial hypercholesterolemia, hereditary breast and ovarian cancer syndrome, and lynch syndrome with disease risk in adults according to family history. JAMA Netw Open 3(4), e203959 (2020).
Scientific Steering Committee on behalf of the Simon Broome Register. Risk of fatal coronary heart disease in familial hypercholesterolaemia. Br. Med. J. 303, 893–896 (1991).
Dhiman, P., Kai, J., Horsfall, L., Walters, K. & Qureshi, N. Availability and quality of coronary heart disease family history in primary care medical records: Implications for cardiovascular risk assessment. PLoS ONE 9(1), 1–7 (2014).
Hosmer, D. W. Jr., Lemeshow, S. & Sturdivant, R. X. Introduction to the logistic regression model. In Applied Logistic Regression Vol. 398 (eds Balding, D. J. et al.) 1–20 (Wiley, 2013).
Breiman, L. Introduction to tree classification. In Classification and Regression Trees (ed. Breiman, L.) 20–42 (Routledge, 2013).
Therneau, T. M. & Atkinson, E. J. An introduction to recursive partitioning using the RPART routines. Mayo Foundation 5–17 (2019).
Breiman, L. Random forests. Mach. Learn. 45(1), 5–32 (2001).
Breiman, L. Bagging predictors. Mach. Learn. 24(2), 123–140 (1996).
Ho, T. K. The random subspace method for constructing decision forests. IEEE Trans. Pattern Anal. Mach. Intell. 20(8), 832–844 (1998).
Kononenko, I. Inductive and Bayesian learning in medical diagnosis. Appl. Artif. Intell. 7(4), 317–337 (1993).
Fawcett, T. An introduction to ROC analysis. Pattern Recognit. Lett. 27(8), 861–874 (2006).
Unal, I. Defining an optimal cut-point value in ROC analysis: An alternative approach. Comput. Math. Methods Med. 4, 1–14 (2017).
Habibzadeh, F., Habibzadeh, P. & Yadollahie, M. On determining the most appropriate test cut-off value: The case of tests with continuous results. Biochem. Med. 26(3), 297–307 (2016).
Hand, D. J. Assessing the performance of classification methods. Int. Stat. Rev. 80(3), 400–414 (2012).
Besseling, J. et al. Selection of individuals for genetic testing for familial hypercholesterolaemia: Development and external validation of a prediction model for the presence of a mutation causing familial hypercholesterolaemia. Eur. Heart J. 38(8), 565–573 (2017).
Weng, S. F., Kai, J., Neil, H. A., Humphries, S. E. & Qureshi, N. Improving identification of familial hypercholesterolaemia in primary care: Derivation and validation of the familial hypercholesterolaemia case ascertainment tool (FAMCAT). Atherosclerosis 238(2), 336–343 (2015).
Benlian, P. et al. Diagnosis scoring for clinical identification of children with heterozygous familial hypercholesterolemia. J. Pediatr. Gastroenterol. Nutr. 48(4), 456–463 (2009).
Niehaus, K. E., Banda, J. M., Knowles, J. W. & Shah, N. H. FIND FH—A phenotype model to identify patients with familial hypercholesterolemia. In Proc. Data Mining for Medical Informatics Workshop, 1–8 (2015).
Banda, J. M. et al. Finding missed cases of familial hypercholesterolemia in health systems using machine learning. NPJ Digit. Med. 2(1), 1–8 (2019).
Albuquerque, J., Alves, A. C., Medeiros, A. M., Bourbon, M. & Antunes, M. Modified classification trees applied to pediatric familial hypercholesterolemia: A comparative study including Simon Broome criteria. J. Stat. Health Dec. 1(1), 1–3 (2019).
Medeiros, A. M., Alves, A. C., Francisco, V. & Bourbon, M. Update of the Portuguese familial hypercholesterolaemia study. Atherosclerosis 212, 553–558 (2010).
Jolliffe, C. J. & Janssen, I. Distribution of lipoproteins by age and gender in adolescents. Circulation 114(10), 1056–1062 (2006).
Benito-Vicente, A. et al. The importance of an integrated analysis of clinical, molecular, and functional data for the genetic diagnosis of familial hypercholesterolemia. Genet. Med. 17(12), 980–988 (2015).
Onis, M. D. et al. Development of a WHO growth reference for school-aged children and adolescents. Bull. World Health Organ. 85, 60–667 (2007).
World Health Organization. WHO Child Growth Standards: Length/Height-for-Age, Weight-for-Age, Weight-for-Length, Weight-for-Height and Body Mass Index-for-Age: Methods and Development 301–305 (WHO, 2006).
Medeiros, A. M., Alves, A. C. & Bourbon, M. Mutational analysis of a cohort with clinical diagnosis of familial hypercholesterolemia: Considerations for genetic diagnosis improvement. Genet. Med. 18(4), 316–324 (2016).
Richards, S. et al. Standards and guidelines for the interpretation of sequence variants: A joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 17(5), 405–423 (2015).
Kowarik, A. & Templ, M. Imputation with the R Package VIM. J. Stat. Softw. 74(7), 1–16 (2016).
Liaw, A. & Wiener, M. randomForest: Breiman and Cutler’s random forests for classification and regression. R Package Version 4, 6–10 (2015).
Tharwat, A. Classification assessment methods. Appl. Comput. Inform. 17, 168–192 (2018).
Nordestgaard, B. G. & Langsted, A. Lipoprotein (a) as a cause of cardiovascular disease: Insights from epidemiology, genetics, and biology. J. Lipid Res. 57(11), 1953–1975 (2016).
López, V., Fernández, A., García, S., Palade, V. & Herrera, F. An insight into classification with imbalanced data: Empirical results and current trends on using data intrinsic characteristics. Inf. Sci. 250, 113–141 (2013).
Kolovou, G. D., Kostakou, P. M. & Anagnostopoulou, K. K. Familial hypercholesterolemia and triglyceride metabolism. Int. J. Cardiol. 147(3), 349–358 (2011).
Miltiadous, G., Cariolou, M. A. & Elisaf, M. HDL cholesterol levels in patients with molecularly defined familial hypercholesterolemia. Ann. Clin. Lab. Sci. 32(1), 50–54 (2002).
Acknowledgements
The current work was supported by the programme Norte2020 [Grant Number NORTE-08-5369-FSE-000018] and by Fundação para a Ciência e Tecnologia (FCT) [Grant Numbers UID/MAT/00006/2019 and PTDC/SAU-SER/29180/2017].
Author information
Authors and Affiliations
Contributions
M.B., J.A. and M.A. conceived and design the project. J.A. performed data processing and statistical analysis under M.A. supervision, literature revision and wrote the draft version. A.M.M. and A.C.A. performed molecular studies and dataset curation; A.M.M. managed the database with clinical and biochemical data. M.B. coordinated and supervised the whole study. M.A. supervised data processing and statistical analysis methods. M.B. and M.A. critically revised the manuscript. All authors contributed to the final version of the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Albuquerque, J., Medeiros, A.M., Alves, A.C. et al. Performance comparison of different classification algorithms applied to the diagnosis of familial hypercholesterolemia in paediatric subjects. Sci Rep 12, 1164 (2022). https://doi.org/10.1038/s41598-022-05063-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-05063-8
- Springer Nature Limited