Abstract
Primary cancer cells exert unique capacity to disseminate and nestle in distant organs. Once seeded in secondary sites, cancer cells may enter a dormant state, becoming resistant to current treatment approaches, and they remain silent until they reactivate and cause overt metastases. To illuminate the complex mechanisms of cancer dormancy, 10 transcriptomic datasets from the literature enabling 21 dormancy–cancer comparisons were mapped on protein–protein interaction networks and gene-regulatory networks to extract subnetworks that are enriched in significantly deregulated genes. The genes appearing in the subnetworks and significantly upregulated in dormancy with respect to proliferative state were scored and filtered across all comparisons, leading to a dormancy–interaction network for the first time in the literature, which includes 139 genes and 1974 interactions. The dormancy interaction network will contribute to the elucidation of cellular mechanisms orchestrating cancer dormancy, paving the way for improvements in the diagnosis and treatment of metastatic cancer.
Similar content being viewed by others
Introduction
Cancer is one of the most fatal and refractory diseases. Although advances in medicine and technological developments now allow treatment of many types of cancer, especially if patients were diagnosed at earlier stages; diagnosis of advanced disease, metastatic relapse and treatment at these late stages still remains as a difficult challenge. There is compelling evidence that dormant cancer cells are responsible for cancer recurrence1. Some of the disseminated tumor cells might escape immune surveillance and develop resistance against cancer treatment modalities. These single cells or micrometastases formed by disseminated tumor cells are beyond the detection limit of current cancer diagnosis techniques, and untraceable through CT scans, MRI approaches and PET-scans. Thus, residual disseminated tumor cells remain in the body, enter a dormant state (hereafter called dormancy) and cause recurrence of the disease months or even years after "the cure" (5 years tumor-free survival).
Cancer cell dormancy is a complex cellular phenomenon and remains to be explored. However, recent studies suggested that there are two major mechanisms of cancer dormancy: Tumor mass dormancy and cellular dormancy2. Tumor mass dormancy refers to the stagnation in tumor mass growth, when cell division and cell death take place at the same rate, keeping the total number of cells constant. Cell numbers in tumors, hence the tumor mass, depend on vascular supply of blood and anti-tumor immune responses. In cellular dormancy however, cell-intrinsic mechanisms that control proliferation and cell cycle entry are dominant. Rather than cell division versus death rates of cancer cells, in cellular dormancy, intrinsic mechanisms leading to a quiescence-like stage and cell cycle arrest are in play. Experimental models indicate that cellular dormancy can be triggered by drugs, growth factor/hormone deprivation, hypoxia, tumor microenvironment components such as extracellular matrix signals and stromal cell interactions1. An important characteristic of dormant cells is their ability to exit the dormancy state and continue to proliferate. Little is known about reasons and mechanisms of entry to dormant state and factors that trigger cell reactivation. Recent studies employed transcriptome-based analyses of dormant cells and revealed potential target genes that may play a role in dormancy in different cancer types3,4.
Network-based computational methods are frequently used in the systems biology field to elucidate disease-associated molecular mechanisms from omics data. Integration of omics data with biological networks is a promising approach for better understanding the disease etiology and to unravel efficient biomarkers and drug targets5. In the literature, numerous techniques were proposed for integrating omics data with biological networks. One of these techniques is active-module detection through network projection of omics data6, known also as subnetwork discovery. KeyPathwayMiner7 and BioNet8 are two powerful subnetwork discovery tools that are used for data integration9. KeyPathwayMiner has been applied to data from several diseases, including ovarian cancer10, breast cancer11 and multiple sclerosis12. Similarly, BioNet has been applied to data from prostate cancer13, breast cancer14, and hepatocellular carcinoma15, among others.
Although in the last couple of years, there is a sharp increase in the number of studies generating transcriptomic data from dormant cancer cells, a meta-analysis of those datasets is still missing. Moreover, available studies mostly lack a network-based advanced analysis of the gene expression signatures of dormant states. Here, by mapping 21 comparisons from 10 transcriptome studies on human or mouse protein–protein interaction (PPI) networks and gene-regulatory (GR) networks, we provide a comprehensive transcriptome-based catalogue of dormancy-related genes and interactions.
Results and discussion
Discovery of PPI and GR subnetworks from dormancy transcriptome datasets
The 10 dormancy-associated transcriptome datasets were obtained from GEO database (Supplementary Table S1). 7 datasets were collected from human cell lines and the other three were collected from mouse cell lines. Datasets were obtained using either microarray, RNAseq or single-cell RNAseq methods. The 10 datasets included 7 different cancer types (bladder cancer, acute lymphoblastic leukemia, colorectal cancer, acute myeloid leukemia, breast cancer, prostate cancer, and myeloma cells).
KeyPathwayMiner (KPM) and BioNet were applied separately to each dormancy–cancer comparison to extract subnetworks that are enriched in significantly changed genes. As a result, we obtained 34 subnetworks for PPI network-based analyses, and 35 subnetworks for GR and TF-free-GR network-based analyses of 21 comparisons. Sizes of the discovered subnetworks are shown in Supplementary Table S2. No subnetworks were found by BioNet in PPI and GR subnetwork discovery analyses for 8 and 7 of comparisons respectively. However, when both BioNet and KPM discover subnetworks, BioNet usually reports a bigger subnetwork. Besides, majority of the genes in the dormancy-interaction network have appeared in subnetworks identified either by both tools or only by BioNet (Supplementary Fig. S1).
Thereafter, we performed enrichment analysis for each subnetwork to show whether they were enriched with genes associated with dormancy. Even though the subnetworks were based on data from different cancer types and different cell lines, genes in the subnetworks were mainly enriched in terms that are known to be associated with dormancy-related mechanisms such as extracellular matrix organization, response to stress and cell cycle regardless of the network type they were derived from (results not shown). Indeed, during cellular dormancy, tumor cells repress cell cycle driving pathways and activate cell cycle inhibitory pathways in order to halt proliferation1. For example, dormant cancer cells directly interact with extracellular matrix, and cell–matrix contact regulates tumor cell growth, migration, differentiation and survival2. Stress conditions such as hypoxia, nutrient deprivation, and chemotherapy induce dormancy, hence activation of stress response pathways during dormancy contribute to the resistance of these cells to these unfavorable conditions16. Convergence of enrichment analysis results to above mentioned pathways justified our computational approach for the discovery of key genes in dormancy mechanisms using genome-wide networks.
Integrated PPI and GR subnetworks pinpoint genes playing roles in dormancy associated molecular mechanisms
In the next step, KPM and BioNet results of each dormancy–cancer comparison were combined to eliminate algorithm-specific effects on the results. Due to limited number of datasets in literature that compares cancer and dormancy data, datasets utilized in this study were obtained from different experimental conditions, cancer types and/or organisms. By consolidating the results of two different algorithms and scoring genes based on the number of appearances in all comparisons, we minimize the effect of these differences. We integrated all subnetworks to score genes based on their appearance in subnetworks. We aimed to extract the most important genes across subnetworks. Score tables were generated for PPI network and GR network analyses separately.
The score table of PPI subnetworks contains 4459 genes that appeared at least in one subnetwork. The genes were filtered based on their appearance in multiple dormancy–cancer comparisons or datasets and based on their significant upregulation (see “Methods” section for details). The filtered score table includes 74 genes based on significance score filter (Supplementary Table S3). The score tables of GR and TF-free GR subnetworks contain 2196 and 2969 genes respectively and filtered score tables include 70 and 111 genes (Supplementary Table S3). The genes in filtered score tables also have interactions with each other in the associated genome-wide networks. 47 out of 74 genes interact via 67 edges in PPI network, 15 out of 70 genes interact via 16 edges in GR network and 110 out of 111 genes interact with 1892 edges in TF-free GR network (Supplementary Fig. S2).
The top 10 genes in filtered score tables in both PPI and GR subnetworks were mainly associated with cell cycle and extracellular matrix (Table 1). Details on these genes in terms of the number of comparisons they were identified in and the corresponding tumor type (liquid or solid) are given in Supplementary Table S4. In concordance with the literature, our results demonstrated that ECM organization and cell cycle control are key pathways in dormancy mechanisms in different cancer types2,3. Based on this, genes that are important according to our study were examined in detail to discover other unknown pathways having role in dormancy mechanisms. Clusterin (CLU), H2B clustered histone 21 (HIST2H2BE) and F-box protein 32 (FBXO32) were found among top 10 genes of all PPI, GR and TF-free GR filtered lists. CLU reduces the sensitivity of prostate cancer cells to chemotherapy and it is highly expressed in drug-resistant cancer cells17 In addition, the CLU protein has been shown to induce epithelial–mesenchymal transition (EMT) in lung cancer18. Nayak et al. showed that HIST1H2BE is overexpressed in treatment resistant breast cancer cells19. FBXO32 is ubiquitinated to stabilize CtBP1, which induces transcription of genes for creating suitable microenvironment for EMT progression20. CDKN2B/p15INK4B was reported to attenuate the growth capacity of the tumors and sustain dormancy phenotype. CDKN2B binds to CDK4/CDK6, and controls transition of proliferative state and it also inhibits ENO1, a glycolytic enzyme, whose activation required for aerobic glycolysis21.
Many potential top hits from Table 1 have not been studied in cancer dormancy context before, yet several studies revealed their function in metastatic outgrowth, escape from immune surveillance, resistance to cell death and stemness. OPTN was identified as a key regulator of cell cycle arrest and stemness in cancer cells22. Ectopic expression of NEU1 suppressed migration and invasion in in vitro cancer models. Also, administration of NEU1 expressing cells into mice prevented metastasis significantly. On the other hand, loss of NEU1 was associated with increased mobility and invasiveness23,24. Also, targeting of NEU1 was reported to attenuate drug resistance25. In mice, loss of the cholesterol transporter ABCG1 resulted in excessive lipid accumulation in macrophages with an elevated proinflammatory cytokines secretion, leading to altered tumor-killing capacity of the macrophages26. Increased expression of ABCG1 was documented in 3D tumoroids and significantly correlated with low Ki67 levels27.
In addition to transcriptome data, one of the effective factors in this study is interactome data. Interactome databases store experimentally validated interactions or interactions predicted by methods such as text-mining. Only experimentally validated interactions were used in this study. However, the information contained in these databases does not yet include all protein–protein interactions due to the technical difficulties in detecting the interactions. Therefore, there is no interaction information for a set of genes in the analyzed transcriptome datasets, and these genes were automatically excluded from our subnetwork-based analysis approach. In order to dissect their possible roles in dormancy, a separate analysis was performed for the genes in the transcriptome datasets with no associated interactome data. These genes were scored based on the number of transcriptome datasets in which they were significantly changed (Supplementary Table S5). The highest scoring genes in the analysis are not associated with autophagy, ECM, cell cycle, or dormancy. This shows that the absence of these genes in our subnetwork analysis does not cause a loss of information. On the other hand, their significance in multiple dormancy-related datasets implies that, although experimentally not reported, they may have potential roles in dormancy mechanisms.
Network based meta-analysis of transcriptome datasets leads to a dormancy–cancer interaction network
Finally, we created a “consensus dormancy-interaction network” composed of the union of the genes in the three filtered lists and their interactions. Since down-regulated genes were mainly associated with cell cycle activation, the constructed dormancy–interaction network focused mainly on up-regulated genes during dormancy. These up-regulated genes have a high potential to be associated with molecular mechanisms of dormancy activation. Indeed, many of the genes in dormancy–interaction network were associated with cancer or dormancy pathways. Figure 1 summarizes our computational approach to construct the dormancy interaction network from transcriptome data and molecular interaction networks.
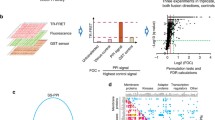
Summary of computational approach to construct the dormancy interaction network. Firstly, cancer–dormancy transcriptome data were downloaded from GEO database and Genome-wide interaction networks were obtained. Then, KPM and BioNet, two subnetwork discovery tools, were performed and subnetworks that are enriched in significantly changed genes were identified. Finally, the upregulated genes were scored and filtered based on the number of appearances in the subnetworks and according to this filtering, the dormancy-interaction network was constructed (see also “Methods”).
The dormancy-interaction network included 139 genes and 1974 interactions (Fig. 2). Enrichment analysis of these 139 genes led to terms that pinpoint cancer cell dormancy mechanisms, such as “negative regulation of programmed cell death”, “cell communication” and “response to stress” (Supplementary Table S6). The network included 36 genes that appeared in the score lists of all three network types. Enrichment analysis of the intersection-genes revealed that they were mainly associated with “cell-death” and “cell-cycle arrest”, similar to union of networks (Supplementary Table S6). Most of these genes were reported to be related with several types of cancer in literature such as EPAS128, SMAD329, NOTCH330. ENO2 has a role in adaptation to serum starvation, hypoxia and chemotherapy in glioblastoma31. BTG2 and CDKN2B cause cell cycle arrest in different cancers and BTG2 is upregulated in quiescent cells32 TIMP-2 also causes cell cycle arrest and inhibits angiogenesis by de-phosphorylation of VEGF33. HO-1 and VEGF-A induce angiogenesis. VEGF-A is found to be upregulated in our study. Although its inhibition has been associated with dormancy34, VEGF-A plays a role as not only angiogenesis induced factor but also tumor-induced immunosuppressor, which can explain its function in dormant cells35. On the other hand, VEGF-A has the highest degree (96 interactions) in the dormancy–interaction network, and the 3 cancer-related transcription factors (EPAS1, KLF4 and SMAD3) that control VEGF-A also appear in the dormancy interaction network. The fact that VEGF-A has a high number of interactions in the predicted dormancy network and the increase in the expression levels of transcription factors that are effective in VEGF-A regulation supports that VEGF-A plays an important role in the dormancy mechanism. THBS1 is another angiogenesis regulator, and its upregulation was associated with dormancy4,36. Another high degree gene in the network is the cyclin dependent kinase inhibitor CDKN1A (93 interactions), which was reported to arrest cells in the G0/G1 phase. Also, increased expression of CDKN1A was previously associated with dormancy37. All interactions of CDKN1A in the dormancy-interaction networks are TF-free regulatory interactions, which further supports our choice of including the TF-free network in our approach. There are many interactions in the dormancy interaction network originating from the TF-free network. This suggests that the genes associated with dormancy may not physically interact, but rather regulated by similar transcription factors. Most of the genes that were commonly found in all three network types were not directly linked with dormancy previously, but the genes regulate critical dormancy pathways including hypoxia, extracellular matrix organization and angiogenesis2. 15 out of 139 dormancy interaction network genes were previously associated with dormancy (Table 2).
Dormancy-interaction network. The network was constructed by integrative analysis of all subnetworks. Dormancy-interaction network includes genes found in different subnetwork analyses (shown in different clusters). Gene regulatory interactions between genes are shown by blue arrows, the direction of the arrow represents the direction of the interaction. Protein–protein interactions are shown by green lines and TF-free gene-regulatory interactions by orange lines.
It has been known that dormant cancer cells have molecular similarities with quiescent stem cells48. Accordingly, stem cell-related mechanisms may also play role in cancer cell dormancy. Indeed, six (CITED2, EPAS1, FOLR1, KDM3A, KLF4 and, SOX9) of the genes in the dormancy interaction network are associated with “stem-cell” term in GO database, and two (EPAS1 and KLF4) are associated with “transcriptional regulation of pluripotent stem-cell” term in REACTOME database.
To provide an independent validation for our findings, we analyzed two additional transcriptomic datasets49,50 that were made publicly available after we completed our analyses (Supplementary Tables S1, S2). The analysis of the datasets with the same pipeline led to a combined list of 40 genes (24 genes in one dataset, 21 genes in the other dataset) from our consensus dormancy-interaction network (Supplementary Table S7). Seven of those overlapping genes (ABCG1, CDKN2B, CLU, FBXO32, PLK2, VEGFA, THBS1) are among the top 10 genes reported in Table 1. As another independent validation, we scanned deletion/overexpression phenotypes of the yeast homologs of the genes in the dormancy-interaction network. Homologs of CCNG2, RASAL2, SLC3A2 and ABCG1 genes in the yeast S. cerevisiae exhibit haploproficiency, meaning that deletion of one of the alleles provided growth advantage to yeast cells. This is in line with the fact that these genes are all overexpressed in dormancy, implying their possible role in suppressing cell division. Similarly, the overexpression of the yeast homolog of FMNL2 gene from the network leads to lethality, which can show implication of the gene in driving the tumor cells to dormancy state. Therefore, (i) several literature findings for a number of genes in our dormancy-interaction network in terms of their role in dormancy and metastasis, (ii) analysis of two independent transcriptome datasets, (iii) comparison with yeast loss-of-function/overexpression studies provide validations and support the hypothesis that our dormancy–interaction network is a suitable model to explain cancer cell dormancy mechanisms.
Mapping transcriptome data from 21 dormancy–cancer comparisons on three different molecular interaction networks led us to construct a consensus dormancy-interaction network. The dormancy–interaction network introduced in this study provides a molecular framework for dormancy mechanisms by combining information repetitively reported in different cancer–dormancy datasets from literature. A number of the genes captured by the network was previously reported to be linked to dormancy or metastasis, providing a validation for the constructed network. However, majority of the nodes in the network are genes that have not been associated with cancer cell dormancy before. Hence, this network is an important model that will shed light on further studies on extended mechanisms for dormancy. As dormancy increases the resistance to therapy and plays a role in metastasis, targeting the genes featured in this study may increase impact of treatments and prevent recurrence.
Methods
Datasets
Transcriptome data of proliferative, dormant or post-dormant cancer cells of human or mouse origin were obtained from the GEO database51. The following criteria were taken into consideration in selection of transcriptome data to be included; it should include samples of both dormant and proliferative cancer states, at least two samples should be available from each condition, it should consist of human or mouse samples. A total of 10 datasets were found to meet all the criteria, which led to analysis of 21 dormancy–cancer comparisons (17 for solid tumors, 4 for liquid tumors) since a dataset can include data from multiple cancer cell lines (Supplementary Table S1). The datasets were first checked for outlier samples by creating PCA graphs for each of the 21 comparisons. Based on PCA results, one control sample from GSE57695 dataset (GSM1386901), one control and one dormant sample from HT55 cell line of GSE114012 dataset (GSM3130646 and GSM3130647) were identified as outliers and removed from the datasets prior to further analysis. Additionally, two datasets published after we completed the analyses in this study were selected as validation datasets and the same procedure was applied to these datasets. To obtain gene expression counts, RNAseq FASTQ files of both datasets were downloaded from The European Nucleotide Archive (ENA) [Project IDs: PRJNA644590 (GSE153944) and PRJNA610898 (GSE146592)]. Adapter sequences were removed from RNAseq fastq files using Trimmomatic (Version 0.39)52. Then using STAR (Version 2.7.8a)53, the trimmed FASTQ files were mapped to genome. For PRJNA610898 human genome GRCh38 and for PRJNA644590 mouse genome GRCm39 were used54. The counts were quantified using featureCounts55.
Statistical analysis of transcriptome data
Differential expression analysis was applied to all 21 dormancy–cancer comparisons from 10 datasets to obtain p-values, adjusted p-values and fold changes of genes. The R package Limma56 was used to identify differentially expressed genes in microarray data. Limma analyses were performed using the GEO2R tool of GEO database, which also generates corresponding R scripts. The R package DESeq257 was used for the differential expression analysis of RNAseq data. For the differential expression analysis of single-cell RNAseq data, DESeq2 was combined with R package zinbwave38.
Cellular networks
Human protein–protein interaction (PPI) network was downloaded from BioGRID 3.5.166 (release date: November, 2018)39. Duplicate edges and self-loops of interactome were removed by using Cytoscape40. Entrez ID of each gene in the interactome was retrieved via bioDBnet41, and integrated into the interactome. The final human PPI network consisted of 17,241 edges and 292,471 interactions. Mouse PPI network was obtained from a previous study of our research group42, which combined and merged PPI data from different databases. Duplicate edges and self-loops of mouse PPI network were removed, and gene names were converted to Entrez IDs as described above. The final mouse PPI network consists of 7713 nodes and 24,830 interactions.
Gene-regulatory (GR) networks of both human and mouse were downloaded from TRRUST version243 and RegNetwork44 databases. The networks from the two databases were merged, and duplicate edges and nodes and self-loops were removed. miRNA interactions, if any, were removed. The final human GR network consists of 6261 nodes, 19,146 edges and 945 transcription factors (TF) while the final mouse GR network has 4010 nodes, 13,485 edges and 1070 transcription factors. A modified TF-free version of GR network, termed TF-free network, was also created and used in the analysis. Here, all genes that were affected by the same TF were represented to be in interaction with each other, and all TF interactions were removed from the network. In this way, the direct effect of the transcription factor is eliminated, but gene regulation information is conserved. Human and mouse TF-free networks consist of 6048 nodes and 2,530,468 edges, and 3767 nodes and 4,622,077 edges, respectively.
Subnetwork discovery
Subnetwork discovery analysis was performed for all 21 dormancy–cancer comparisons from 10 datasets and 2 validation datasets. Data were mapped on PPI networks, GR networks and TF-free GR networks separately by using KeyPathwayMiner (KPM)7 and BioNet8 tools.
For KPM analysis, a binarized version of Benjamini–Hochberg corrected p-values was created using the associated p-value cut-off value for each comparison. Binarization was done by assigning the value 1 to significant genes and assigning the value 0 to non-significant genes. The appropriate cut-off values were chosen such that a similar-size subnetwork can be obtained for all comparisons (Supplementary Table S2). The cut-off values of each comparison remained the same across analyses based on protein–protein interactome and gene-regulatory interactome. Thereafter, the stand-alone version of KeyPathwayMiner (KPM 4.0) was executed with the following parameters: Individual Node Exceptions (INEs) strategy, Ant Colony Optimization (ACO) algorithm, K = 2 for PPI and K = 8 for GR networks. K is a parameter to define the maximum number of exception (not differentially expressed) nodes allowed in a subnetwork.
BioNet version 1.42.0 was run in R for each comparison. Calculated p-values were used as input, and False Discovery Rate (FDR) cut-off values for each pairwise comparison were selected such that similar-size subnetworks will be produced (Supplementary Table S2). The FDR values of each comparison remained the same across the analyses based on protein–protein and gene-regulatory interactomes.
Interpretation of subnetworks
In order to interpret the discovered subnetworks, we used three different approaches. Firstly, enrichment analyses were performed by using g:Profiler online tool45. g:Profiler allows to perform multiple types of enrichment analyses in one run. It retrieves data from different sources including GeneOntology, KEGG, Reactome and TRANSFAC databases. Its g:GOST module was used to perform functional enrichment analysis on input gene lists with default parameters.
Secondly, genes in all subnetworks were scored based on number of appearances in different subnetworks. To combine mouse and human subnetworks, human orthologues of mouse genes were retrieved via Ensembl/BioMart46. In our scoring approach, a gene gets a score of 1 for each subnetwork in which they appeared, and, the final score of a gene is sum of its scores across all constructed subnetworks. Two different scoring approaches were used. In the first scoring approach, called “significance score”, the gene gets a score of 1 if it is identified with at least one of the BioNet or KPM tools in a dormancy–cancer comparison and significantly changed in the comparison. In our second scoring approach, genes are scored for the number of datasets in which they were found. In this scoring, called “dataset score”, a gene gets a score of 1 if it is identified in at least one of all comparisons in a single dataset to minimize bias due to datasets with a higher number of comparisons. A filtering was applied to the scored genes afterwards by using their significance and dataset scores together. Here, genes were filtered if their dataset scores were at least 2 and their significance scores were at least 3. Additionally, we specifically chose genes that are mostly upregulated in dormancy. For this, we required that if the significance score of a gene is 3, it must be upregulated in all of these 3 comparisons. If the score is higher than 3, it must be upregulated in more than half of the subnetworks in which the gene was scored significant. All calculations and rearrangements were performed in R.
Thirdly, Gene Ontology terms in AmiGO database were used to construct lists of mouse and human genes that are associated with autophagy, cell cycle, and extracellular matrix terms47. Additionally, a list of dormancy-associated genes in human was manually curated based on abstracts of about 200 articles. Mouse homologs of dormancy genes were identified via Ensembl/BioMart46. Then, the genes in the subnetwork score tables were annotated for their association to autophagy, cell cycle, extracellular matrix and dormancy (Supplementary Tables S3, S5). Workflow of computational approach to construct the dormancy-interaction network is shown in Fig. 1.
Data availability
The datasets analysed during the current study are available in the Gene Expression Omnibus (GEO) repository. All data are incorporated into the article and its online Supplementary material.
References
Pradhan, S., Sperduto, J. L., Farino, C. J. & Slater, J. H. Engineered in vitro models of tumor dormancy and reactivation. J. Biol. Eng. 12, 37. https://doi.org/10.1186/s13036-018-0120-9 (2018).
Páez, D. et al. Cancer dormancy: A model of early dissemination and late cancer recurrence. Clin. Cancer Res. 18, 645. https://doi.org/10.1158/1078-0432.CCR-11-2186 (2012).
Al-Asadi, M. G. et al. A molecular signature of dormancy in CD34(+)CD38(−) acute myeloid leukaemia cells. Oncotarget 8, 111405–111418. https://doi.org/10.18632/oncotarget.22808 (2017).
Kim, R. S. et al. Dormancy signatures and metastasis in estrogen receptor positive and negative breast cancer. PLoS One 7, e35569. https://doi.org/10.1371/journal.pone.0035569 (2012).
Alcaraz, N. et al. De novo pathway-based biomarker identification. Nucleic Acids Res. 45, e151–e151. https://doi.org/10.1093/nar/gkx642 (2017).
Ideker, T., Ozier, O., Schwikowski, B. & Siegel, A. F. Discovering regulatory and signalling circuits in molecular interaction networks. Bioinformatics 18(Suppl 1), S233-240. https://doi.org/10.1093/bioinformatics/18.suppl_1.s233 (2002).
Alcaraz, N. et al. Efficient key pathway mining: Combining networks and OMICS data. Integr. Biol. (Camb.) 4, 756–764. https://doi.org/10.1039/c2ib00133k (2012).
Beisser, D., Klau, G. W., Dandekar, T., Muller, T. & Dittrich, M. T. BioNet: An R-Package for the functional analysis of biological networks. Bioinformatics 26, 1129–1130. https://doi.org/10.1093/bioinformatics/btq089 (2010).
Batra, R. et al. On the performance of de novo pathway enrichment. NPJ Syst. Biol. Appl. 3, 6. https://doi.org/10.1038/s41540-017-0007-2 (2017).
Salgado-Albarrán, M. et al. The epigenetic factor BORIS (CTCFL) controls the androgen receptor regulatory network in ovarian cancer. Oncogenesis 8, 41. https://doi.org/10.1038/s41389-019-0150-2 (2019).
Pauling, J. K. et al. Elucidation of epithelial–mesenchymal transition-related pathways in a triple-negative breast cancer cell line model by multi-omics interactome analysis. Integr. Biol. 6, 1058–1068. https://doi.org/10.1039/c4ib00137k (2014).
Elkjaer, M. L. et al. Molecular signature of different lesion types in the brain white matter of patients with progressive multiple sclerosis. Acta Neuropathol. Commun. 7, 205. https://doi.org/10.1186/s40478-019-0855-7 (2019).
Yang, Y., Yang, W. & Jin, L. The role of long non-coding RNA prostate cancer-associated transcript 1 in prostate cancer. J. Comput. Biol. 26, 975–984. https://doi.org/10.1089/cmb.2018.0240 (2019).
Zheng, T., Wang, A., Hu, D. & Wang, Y. Molecular mechanisms of breast cancer metastasis by gene expression profile analysis. Mol. Med. Rep. 16, 4671–4677. https://doi.org/10.3892/mmr.2017.7157 (2017).
Yan, H. et al. Aberrant expression of cell cycle and material metabolism related genes contributes to hepatocellular carcinoma occurrence. Pathol. Res. Pract. 213, 316–321. https://doi.org/10.1016/j.prp.2017.01.019 (2017).
Gao, X. L., Zhang, M., Tang, Y. L. & Liang, X. H. Cancer cell dormancy: Mechanisms and implications of cancer recurrence and metastasis. Onco Targets Ther. 10, 5219–5228. https://doi.org/10.2147/OTT.S140854 (2017).
Wilson, M. R. & Zoubeidi, A. Clusterin as a therapeutic target. Expert. Opin. Ther. Targets 21, 201–213. https://doi.org/10.1080/14728222.2017.1267142 (2017).
Jin, R., Chen, X., Han, D., Luo, X. & Li, H. Clusterin modulates transdifferentiation of non-small-cell lung cancer. BMC Cancer 17, 661. https://doi.org/10.1186/s12885-017-3649-y (2017).
Nayak, S. R. et al. A role for histone H2B variants in endocrine-resistant breast cancer. Horm. Cancer 6, 214–224. https://doi.org/10.1007/s12672-015-0230-5 (2015).
Sahu, S. K. et al. FBXO32 promotes microenvironment underlying epithelial–mesenchymal transition via CtBP1 during tumour metastasis and brain development. Nat. Commun. 8, 1523. https://doi.org/10.1038/s41467-017-01366-x (2017).
Xia, Y. et al. Dominant role of CDKN2B/p15INK4B of 9p21.3 tumor suppressor hub in inhibition of cell-cycle and glycolysis. Nat. Commun. 12, 2047. https://doi.org/10.1038/s41467-021-22327-5 (2021).
Cheng, J. & Cashman, J. R. PAWI-2 overcomes tumor stemness and drug resistance via cell cycle arrest in integrin β3-KRAS-dependent pancreatic cancer stem cells. Sci. Rep. 10, 9162. https://doi.org/10.1038/s41598-020-65804-5 (2020).
Uemura, T. et al. Contribution of sialidase NEU1 to suppression of metastasis of human colon cancer cells through desialylation of integrin β4. Oncogene 28, 1218–1229. https://doi.org/10.1038/onc.2008.471 (2009).
Zhou, B., Zhu, W., Jiang, X. & Ren, C. RASAL2 plays inconsistent roles in different cancers. Front. Oncol. 9, 1235. https://doi.org/10.3389/fonc.2019.01235 (2019).
O’Shea, L. K., Abdulkhalek, S., Allison, S., Neufeld, R. J. & Szewczuk, M. R. Therapeutic targeting of Neu1 sialidase with oseltamivir phosphate (Tamiflu®) disables cancer cell survival in human pancreatic cancer with acquired chemoresistance. Onco Targets Ther. 7, 117–134. https://doi.org/10.2147/ott.S55344 (2014).
Sag, D., Cekic, C., Wu, R., Linden, J. & Hedrick, C. C. The cholesterol transporter ABCG1 links cholesterol homeostasis and tumour immunity. Nat. Commun. 6, 6354. https://doi.org/10.1038/ncomms7354 (2015).
Namba, Y. et al. Depletion of lipid efflux pump ABCG1 triggers the intracellular accumulation of extracellular vesicles and reduces aggregation and tumorigenesis of metastatic cancer cells. Front. Oncol. 8, 376 (2018).
Endo, H. et al. The induction of MIG6 under hypoxic conditions is critical for dormancy in primary cultured lung cancer cells with activating EGFR mutations. Oncogene 36, 2824–2834. https://doi.org/10.1038/onc.2016.431 (2017).
Brown, J. A. et al. TGF-β-induced quiescence mediates chemoresistance of tumor-propagating cells in squamous cell carcinoma. Cell Stem Cell 21, 650-664.e658. https://doi.org/10.1016/j.stem.2017.10.001 (2017).
Indraccolo, S. et al. Cross-talk between tumor and endothelial cells involving the Notch3-Dll4 interaction marks escape from tumor dormancy. Can. Res. 69, 1314. https://doi.org/10.1158/0008-5472.CAN-08-2791 (2009).
Yan, T. et al. Neuronal markers are expressed in human gliomas and NSE knockdown sensitizes glioblastoma cells to radiotherapy and temozolomide. BMC Cancer 11, 524. https://doi.org/10.1186/1471-2407-11-524 (2011).
Yuniati, L., Scheijen, B., van der Meer, L. T. & van Leeuwen, F. N. Tumor suppressors BTG1 and BTG2: Beyond growth control. J. Cell. Physiol. 234, 5379–5389. https://doi.org/10.1002/jcp.27407 (2019).
Brew, K. & Nagase, H. The tissue inhibitors of metalloproteinases (TIMPs): An ancient family with structural and functional diversity. Biochim. Biophys. Acta 55–71, 2010. https://doi.org/10.1016/j.bbamcr.2010.01.003 (1803).
Kienast, Y. et al. Real-time imaging reveals the single steps of brain metastasis formation. Nat. Med. 16, 116–122. https://doi.org/10.1038/nm.2072 (2010).
Ohm, J. E. et al. VEGF inhibits T-cell development and may contribute to tumor-induced immune suppression. Blood 101, 4878–4886 (2003).
Tiram, G. et al. Reverting the molecular fingerprint of tumor dormancy as a therapeutic strategy for glioblastoma. FASEB J. 32, 5835–5850. https://doi.org/10.1096/fj.201701568R (2018).
Overton, K. W., Spencer, S. L., Noderer, W. L., Meyer, T. & Wang, C. L. Basal p21 controls population heterogeneity in cycling and quiescent cell cycle states. Proc. Natl. Acad. Sci. U.S.A. 111, E4386-4393. https://doi.org/10.1073/pnas.1409797111 (2014).
Risso, D., Perraudeau, F., Gribkova, S., Dudoit, S. & Vert, J.-P. A general and flexible method for signal extraction from single-cell RNA-seq data. Nat. Commun. 9, 284. https://doi.org/10.1038/s41467-017-02554-5 (2018).
Chatr-Aryamontri, A. et al. The BioGRID interaction database: 2017 update. Nucleic Acids Res. 45, D369–D379. https://doi.org/10.1093/nar/gkw1102 (2017).
Shannon, P. et al. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Mudunuri, U., Che, A., Yi, M. & Stephens, R. M. bioDBnet: The biological database network. Bioinformatics 25, 555–556. https://doi.org/10.1093/bioinformatics/btn654 (2009).
Emanetçi, E., Çakır, T. Network-Based Analysis of Cognitive Impairment and Memory Deficits from Transcriptome Data. J. Mol. Neurosci. https://doi.org/10.1007/s12031-021-01807-9 (in press).
Han, H. et al. TRRUST v2: An expanded reference database of human and mouse transcriptional regulatory interactions. Nucleic Acids Res. 46, D380–D386. https://doi.org/10.1093/nar/gkx1013 (2018).
Liu, Z.-P., Wu, C., Miao, H. & Wu, H. RegNetwork: An integrated database of transcriptional and post-transcriptional regulatory networks in human and mouse. Database https://doi.org/10.1093/database/bav095 (2015).
Raudvere, U. et al. g:Profiler: A web server for functional enrichment analysis and conversions of gene lists (2019 update). Nucleic Acids Res. 47, W191–W198. https://doi.org/10.1093/nar/gkz369 (2019).
Kinsella, R. J. et al. Ensembl BioMarts: A hub for data retrieval across taxonomic space. Database (Oxford) 2011, bar030. https://doi.org/10.1093/database/bar030 (2011).
Carbon, S. et al. AmiGO: Online access to ontology and annotation data. Bioinformatics 25, 288–289. https://doi.org/10.1093/bioinformatics/btn615 (2009).
Talukdar, S. et al. Dormancy and cancer stem cells: An enigma for cancer therapeutic targeting. Adv. Cancer Res. 141, 43–84. https://doi.org/10.1016/bs.acr.2018.12.002 (2019).
Duy, C. et al. Chemotherapy induces senescence-like resilient cells capable of initiating AML recurrence. Cancer Discov. 11, 1542–1561. https://doi.org/10.1158/2159-8290.Cd-20-1375 (2021).
Perego, M. et al. Reactivation of dormant tumor cells by modified lipids derived from stress-activated neutrophils. Sci. Transl. Med. https://doi.org/10.1126/scitranslmed.abb5817 (2020).
Barrett, T. et al. NCBI GEO: Archive for functional genomics data sets—Update. Nucleic Acids Res. 41, D991–D995. https://doi.org/10.1093/nar/gks1193 (2012).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120. https://doi.org/10.1093/bioinformatics/btu170 (2014).
Dobin, A. et al. STAR: Ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21. https://doi.org/10.1093/bioinformatics/bts635 (2013).
Frankish, A. et al. GENCODE reference annotation for the human and mouse genomes. Nucleic Acids Res. 47, D766-d773. https://doi.org/10.1093/nar/gky955 (2019).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: An efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930. https://doi.org/10.1093/bioinformatics/btt656 (2014).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47–e47. https://doi.org/10.1093/nar/gkv007 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550–550. https://doi.org/10.1186/s13059-014-0550-8 (2014).
Liu, W., Guan, M., Hu, T., Gu, X. & Lu, Y. Re-expression of AKAP12 inhibits progression and metastasis potential of colorectal carcinoma in vivo and in vitro. PLoS One 6, e24015. https://doi.org/10.1371/journal.pone.0024015 (2011).
Chaterjee, M. & van Golen, K. L. Breast cancer stem cells survive periods of farnesyl-transferase inhibitor-induced dormancy by undergoing autophagy. Bone Marrow Res. 2011, 1–7 (2011).
Schrader, J. et al. Matrix stiffness modulates proliferation, chemotherapeutic response, and dormancy in hepatocellular carcinoma cells. Hepatology 53, 1192–1205. https://doi.org/10.1002/hep.24108 (2011).
Kim, R. S. et al. Dormancy signatures and metastasis in estrogen receptor positive and negative breast cancer. PLoS One 7, e35569–e35569. https://doi.org/10.1371/journal.pone.0035569 (2012).
Sazawa, A. et al. Adenovirus mediated gelsolin gene therapy for orthotopic human bladder cancer in nude mice. J. Urol. 168, 1182–1187. https://doi.org/10.1016/S0022-5347(05)64622-8 (2002).
McGrath, J., Panzica, L., Ransom, R., Withers, H. G. & Gelman, I. H. Identification of genes regulating breast cancer dormancy in 3D bone endosteal niche cultures. Mol. Cancer Res. MCR 17, 860–869. https://doi.org/10.1158/1541-7786.MCR-18-0956 (2019).
Yang, J., Ikezoe, T., Nishioka, C., Nobumoto, A. & Yokoyama, A. IL-1β inhibits self-renewal capacity of dormant CD34+/CD38− acute myelogenous leukemia cells in vitro and in vivo. Int. J. Cancer 133, 1967–1981. https://doi.org/10.1002/ijc.28198 (2013).
Kobayashi, A. et al. Bone morphogenetic protein 7 in dormancy and metastasis of prostate cancer stem-like cells in bone. J. Exp. Med. 208, 2641–2655. https://doi.org/10.1084/jem.20110840 (2011).
Aguirre-Ghiso, J. A., Liu, D., Mignatti, A., Kovalski, K. & Ossowski, L. Urokinase receptor and fibronectin regulate the ERK(MAPK) to p38(MAPK) activity ratios that determine carcinoma cell proliferation or dormancy in vivo. Mol. Biol. Cell 12, 863–879. https://doi.org/10.1091/mbc.12.4.863 (2001).
Touil, Y. et al. Melanoma dormancy in a mouse model is linked to GILZ/FOXO3A-dependent quiescence of disseminated stem-like cells. Sci. Rep. 6, 30405. https://doi.org/10.1038/srep30405 (2016).
Acknowledgements
We thank Cemal Yıldız for manually curated dormancy associated gene list; Çiğdem Hazal Timuçin for helping with RNAseq data analyses; and Merve Kutay for helping with statistical tests. This work was supported by the TUBITAK (Project Code: 216S489).
Author information
Authors and Affiliations
Contributions
T.Ç., D.G., and P.P. conceived and supervised the study. D.U. performed all computational analyses and wrote the manuscript. T.Ç., D.G., P.P., Y.A., and N.P. analyzed the results and contributed to the writing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Uzuner, D., Akkoç, Y., Peker, N. et al. Transcriptional landscape of cellular networks reveal interactions driving the dormancy mechanisms in cancer. Sci Rep 11, 15806 (2021). https://doi.org/10.1038/s41598-021-94005-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-94005-x
- Springer Nature Limited
This article is cited by
-
Proteomic insights into breast cancer response to brain cell-secreted factors
Scientific Reports (2024)
-
mTOR pathway occupies a central role in the emergence of latent cancer cells
Cell Death & Disease (2024)
-
The genomic regulation of metastatic dormancy
Cancer and Metastasis Reviews (2023)
-
Dormancy, stemness, and therapy resistance: interconnected players in cancer evolution
Cancer and Metastasis Reviews (2023)
-
Dormant cancer cells: programmed quiescence, senescence, or both?
Cancer and Metastasis Reviews (2023)