Abstract
[NiFe]-hydrogenases activate dihydrogen. Like all [NiFe]-hydrogenases, hydrogenase 2 of Escherichia coli has a bimetallic NiFe(CN)2CO cofactor in its catalytic subunit. Biosynthesis of the Fe(CN)2CO group of the [NiFe]-cofactor occurs on a distinct scaffold complex comprising the HybG and HypD accessory proteins. HybG is a member of the HypC-family of chaperones that confers specificity towards immature hydrogenase catalytic subunits during transfer of the Fe(CN)2CO group. Using native mass spectrometry of an anaerobically isolated HybG–HypD complex we show that HybG carries the Fe(CN)2CO group. Our results also reveal that only HybG, but not HypD, interacts with the apo-form of the catalytic subunit. Finally, HybG was shown to have two distinct, and apparently CO2-related, covalent modifications that depended on the presence of the N-terminal cysteine residue on the protein, possibly representing intermediates during Fe(CN)2CO group biosynthesis. Together, these findings suggest that the HybG chaperone is involved in both biosynthesis and delivery of the Fe(CN)2CO group to its target protein. HybG is thus suggested to shuttle between the assembly complex and the apo-catalytic subunit. This study provides new insights into our understanding of how organometallic cofactor components are assembled on a scaffold complex and transferred to their client proteins.
Similar content being viewed by others
Introduction
[NiFe]-hydrogenases activate dihydrogen to release electrons and protons, or they reduce protons to help balance the redox state of bacterial and archaeal cells1. These enzymes harbour a bimetallic NiFe(CN)2CO cofactor in their catalytic subunit2. The Fe2+ ion of this cofactor is unusual because it carries a carbonyl and two cyano (CN−) ligands3, which aid catalysis4,5. It is still unclear how the Fe(CN)2CO group, which is synthesised on a distinct scaffold complex, is ultimately transferred into the apo-form of the immature catalytic subunit. Here, we use the hydrogen-oxidising hydrogenase 2 of the bacterium Escherichia coli6 as a model system to address how the Fe(CN)2CO group of this[NiFe]-cofactor might be transferred into the immature (apo-form; called pre-HybC) of the hydrogenase 2 catalytic subunit.
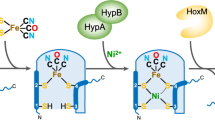
During hydrogenase 2 assembly, two highly conserved accessory proteins, called HybG and HypD, form a scaffold complex upon which the Fe(CN)2CO group is synthesised before it is introduced into pre-HybC7,8 (Fig. 1). While HypD is an iron-sulfur (FeS) cluster-containing enzyme9, HybG belongs to the HypC-family of small chaperone-like proteins7,10. HybG and HypD each has a highly conserved cysteinyl residue, and, together, these two residues are proposed to coordinate the Fe(CN)2CO group on the scaffold complex. These residues are Cys41 on HypD (E. coli numbering) and the N-terminal Cys2 on HypC (and HybG)8,11. Conveniently, the HybG–HypD scaffold complex carrying the Fe(CN)2CO group can be isolated under anoxic conditions, which has allowed its spectroscopic characterization12,13.
Schematic overview of iron group transfer into the apo-catalytic subunit of hydrogenase 2. The 4Fe-4S cluster in HypD is depicted as a distorted cube in HypD, while the red ‘squiggle’ on pre-HybC signifies a C-terminal peptide that is removed by endoproteolytic cleavage after Ni2+ insertion has been completed8.
Two pieces of circumstantial evidence suggest that HybG might be involved in the ultimate transfer of the Fe(CN)2CO group from the scaffold complex to pre-HybC. The first is based on the fact that many bacteria and archaea that synthesise more than one[NiFe]-hydrogenase often have HybG paralogues7,14. For example, E. coli has the HybG paralogue HypC, whereby these proteins share 57% amino acid sequence identity; HypC is specifically required for maturation of the catalytic subunit of the H2-producing hydrogenase 37,15. The second piece of evidence in support of a transfer function of HypC/HybG was provided when hydrogenase apo-catalytic subunits were shown to co-purify with HypC-family proteins16,17. Notably, however, all attempts so far to isolate HybG, or other members of the HypC-family of proteins, with an attached Fe(CN)2CO group have proved unsuccessful. Surprisingly, it has, however, been possible to isolate over-produced HypD carrying the Fe(CN)2CO group13. This is possibly due to the larger HypD protein providing a more stable coordination sphere for the chemically sensitive group. Nevertheless, HybG is still essential for synthesis of the Fe(CN)2CO group attached to over-produced HypD7,13. Hence, there is still uncertainty as to which scaffold protein carries and delivers the Fe(CN)2CO group to the hydrogenase large subunit precursor (Fig. 1). Therefore, the aim of this study is to understand better the pathway of cofactor assembly into pre-HybC, the catalytic subunit of hydrogenase 2.
So far it has not been possible to demonstrate that HybG (or paralogues thereof) carries the Fe(CN)2CO group and this is likely due to the lability of this putative intermediate on the ~ 10 kDa HybG protein. Therefore, we decided to adopt a mass spectrometry approach to address this issue. Native mass spectrometry (MS) has proven to be a powerful technique that can yield unequivocal and detailed molecular information on protein modifications, and protein–protein interactions within larger protein complexes. Moreover, it can provide snapshots of biosynthetic processes, in particular allowing detection of transient intermediates18,19,20,21. Therefore, dynamic assemblies of proteins that are not amenable to high-resolution structural or spectroscopic techniques can be investigated by native MS22. In the current study, we resolve the protein interaction network between HypD, HybG and pre-HybC, the hydrogenase 2 large-subunit precursor of E. coli, and demonstrate that HybG indeed carries the Fe(CN)2CO group. The results obtained herein serve as a blueprint for the analysis of cofactor biosynthesis, and maturation of metalloproteins in general.
Results
Stoichiometric HybG–HypD complexes identified by native MS
The HybG–HypD complex used in this study was affinity-purified under anoxic conditions by taking advantage of the C-terminal StrepII-tag attached to HybG and will be referred to as HybG–HypD throughout this study; unless specifically stated otherwise, HypD was untagged. Analysis by native MS revealed a 1:1 stoichiometry for the HybG–HypD complex (Fig. 2a; see Tables S1 and S2 for measured masses). As well as minor signals arising from free HybG and HypD, which arose due to the comparatively high collision energy used in the MS experiment, minor amounts of HybG–HypD complexes with stoichiometries of 2:1 and 2:2 were also measured (Fig. 2a).
Native mass spectra of StrepII-HybG–HypD complex. Native mass spectrum of (a) the StrepII-HybG–HypD complex at a collision energy of 40 V. Stoichiometry of the main complex species is StrepII-HybG:HypD 1:1, with minor species showing 2:1 and 2:2 ratios (indicated by green circles for HybG and red circles for HypD). (b) Mass spectrum of the dissociation of the 13+-charged ion species of the heterodimer into HybG (charge states + 4 to + 6, red circles) and HypD (charge states + 7 to + 9, green circles) at a collision energy of 80 V. The corresponding measured masses of the StrepII-HybG and HypD proteins are shown in Tables S1 and S2.
Signals for monomeric HybG and HypD were generated by dissociating the 1:1 complex using tandem MS/MS experiments (collision-induced dissociation, CID-MS/MS) of the isolated + 13 charged species of the observed complex signal (Fig. 2b). This dissociation yielded signals for highly charged HybG species (red circles, charge states + 4 to + 6, Fig. 2b) and signals corresponding to HypD (green circles, charge states + 7 to + 9, Fig. 2b) identical to those observed in the full mass spectrum (Fig. 2a).
StrepII-tagged HybG, when dissociated from the complex, revealed additional signals accounting for potential covalent modifications. To identify these modifications, high-resolution MS data for StrepII-tagged HybG and for StrepII-tagged HypD, purified individually, were acquired. While StrepII-tagged HypD purified in the absence of HybG was not observable in the high-resolution mass spectra, due its high susceptibility to aggregation and/or poor ionization behavior, StrepII-tagged HybG yielded the same pattern of modified ion species in comparison to the dissociation experiments of the HybG–HypD complex preparation (Fig. 3a).
High-resolution mass spectrum of StrepII-HybG modifications. (a) The isotope pattern shows high-resolution StrepII-tagged HybG (5+ charge state, red circle) relevant adduct species. The asterisk (*) indicates that the modification was not clearly identifiable. (b) High-resolution mass spectrum of StrepII-tagged HybG with the indicated species that was isolated to generate the top-down CID fragment ion spectrum. The signature fragment ion b2+ unambiguously shows that the additional mass of 26 Da is mostly located on the N-terminal cysteinyl residue (see also Table S2).
High-resolution MS measurements for StrepII-tagged HybG revealed a signal accounting for the non-modified protein at m/z 1923.98 (+ 5 charge state, deconvoluted mass: 9615 Da), as well as the signal for a highly abundant species at m/z 1929.17 (+ 5 charge state, deconvoluted mass: 9641 Da; Table S2) with an additional mass of 26 Da. The latter could indicate an addition of CO2 (+ 44 Da) followed by a loss of a water molecule (− 18 Da). A further species at m/z 1937.97, corresponding to StrepII-tagged HybG with an additional mass of 70 Da, was also identified. This could correspond to the addition of another CO2 molecule (44 Da) and the 26 Da modification. Notably, however, in all experiments, the + 26 Da modification on HybG was present, and almost always exhibited the peak with the highest relative intensity for all HybG species. Additional weak signals were identified in the range of m/z 1933.57 ± 3, but were not clearly resolved.
As is the case for all HypC-family members, the N-terminal cysteinyl residue (formally amino acid position 2) of HybG is essential for the biological function of the protein7,15,17. To determine whether the observed modifications were dependent on this cysteinyl residue, we analyzed a mutant StrepII-tagged HybG variant in which this cysteinyl residue was substituted by an alanyl residue (StrepII-HybGC2A; C2A variant). By comparing high-resolution mass spectra of the native HybG with the HybGC2A variant, it was observed that all of the additional modifications identified for the native HybG protein were no longer present on the HybGC2A variant (Fig. S1).
Finally, to prove that the modification accounting for the mass addition of 26 Da was localized on the N-terminal cysteinyl residue of native StrepII-tagged HybG, we performed top-down tandem mass spectrometric experiments (CID-MS/MS) of the ion species at m/z 1929.17 (5+ charged, deconvoluted mass: 9641 Da, Fig. 3a). The resulting fragment ion spectrum contained a b-type ion series (b9 to b2), which clearly demonstrated that the initial cysteinyl residue was modified, and that it carried the additional mass of 26 Da (Fig. 3b).
HybG interacts specifically with the large subunit precursor pre-HybC
The apo-form of the hydrogenase 2 large subunit, pre-HybC, is a target for the delivery of the Fe(CN)2CO group of the cofactor after its synthesis has been completed on the HybG–HypD scaffold complex6,8,23,24. Pre-HybC is subsequently converted to its mature species25,26, which we refer to here as HybC. It is unresolved whether transfer of Fe(CN)2CO to pre-HybC is mediated by the HybG–HypD scaffold, or HybG, or HypD. Therefore, to address this, we first tested whether the interaction between HybG with pre-HybC could be resolved using native MS. During analysis of a purified, N-terminally His-tagged pre-HybC preparation from E. coli, we detected an ion species corresponding to the expected mass of pre-HybC (Fig. 4a, dark blue squares; see also Table S1). Strikingly, we also detected minor amounts of processed HybC (Fig. 4a, light blue circles; see also Table S1), the origin of which remains unresolved. Purified N-terminally His-tagged pre-HybC was then supplemented with equimolar amounts of HybG to identify any complex formation between the proteins. Upon addition of HybG, signals were immediately observed that indicated the formation of a 1:1 HybG/pre-HybC heterodimeric complex (Fig. 4a, red circle + dark blue square). Notably, small amounts of HybG dimers were also observed, but these were not detected in complexes with pre-HybC and may result from the tendency of HybG to aggregate when not in complex with one of its partner proteins (see Fig. S2;27). The identity of the HybG/pre-HybC complex was again confirmed by tandem MS experiments (CID-MS/MS), whereby the + 16 charged ion species of the complex was dissociated into monomeric HybG (red circles) and monomeric pre-HybC (dark blue squares, Fig. 4b) species. Interestingly, we were never able to detect a complex between HybG and mature HybC, which is in agreement with previous observations15,17,25,26.
Native mass spectra of the StrepII-HybG-pre-HybC complex. (a) Native mass spectrum of the HybG (red circles)-pre-HybC (blue squares) heterodimeric complex. HybG has a C-terminal StrepII-tag and pre-HybC carries a N-terminal His-tag. C-terminally truncated HybC (light blue circles) and pre-HybC dimers, as well as minor amounts of HybG dimers were also observed. (b) The 16+ charged complex species was isolated and dissociated by collision-induced dissociation (CID) resulting in HybG monomer (+ 4 and + 5; red circles) and pre-HybC monomer (+ 10 to + 12, blue squares). The corresponding measured masses of pre-HybC and mature HybC are shown in Table S1.
Using pull-down interaction studies, it was also shown that when StrepII-tagged HybG was incubated with His-tagged pre-HybC, and the mixture was applied to a StrepTactin column, pre-HybC co-eluted with HybG (Fig. S2a). When this experiment was performed with the mutant HybGC2A variant, no retention of pre-HybC on the column was observed (Fig. S2b). This result indicates that the N-terminal cysteinyl residue of HybG is essential for the interaction with pre-HybC to occur.
Pre-HybC titrates HybG from the HybG–HypD complex
The data obtained so far clearly indicate that HybG interacts specifically with pre-HybC. To analyse whether pre-HybC also can interact with the HybG–HypD scaffold complex, increasing concentrations of His-tagged pre-HybC were added to the HybG–HypD complex (Fig. 5; molar ratios of pre-HybC to HypD–HybG complex ranged from 1:20 up to 1:3). The formation of a HybG-pre-HybC complex was observed directly upon addition of a low concentration of His-tagged pre-HybC (Fig. 5a), whereby StrepII-HybG could only have originated from the HybG–HypD complex. Increasing the concentration of His-tagged pre-HybC caused a concomitant increase in the signal intensity of the HybG-pre-HybC complex, while the intensity of the HybG–HypD complex decreased correspondingly (Fig. 5b,c). Pre-HybC was thus able to titrate HybG from the HybG–HypD complex, even at a molar ratio of ~ 1:20 in favour of HybG–HypD (Fig. 5a). The removal of HybG from the HybG–HypD complex appeared to have a negative impact on the stability of HypD, and probably led to its aggregation. Therefore, non-complexed HypD was not detected in the native mass spectra at higher concentrations of pre-HybC, which correlates with our earlier findings (see Fig. 2). Importantly, HypD was never identified to interact with pre-HybC, or to enter into a ternary complex with pre-HybC and HybG (Fig. 5). This indicates that HybG probably delivers the Fe(CN)2CO group to pre-HybC.
Native mass spectra showing pre-HybC titrating HybG from the HybG–HypD complex. Native mass spectrum of the StrepII-tagged HybG (red circles)-HypD (green circles) complex upon addition of increasing amounts of N-terminally His-tagged pre-HybC (blue squares). Ratios of pre-HybC to HybG–HypD complex were (a) 1:20, (b) 1:6, (c) 1:3.
StrepII-tagged HybG carries the Fe(CN)2CO group
In an attempt to identify a HybG species carrying the Fe(CN)2CO group, while simultaneously minimising detrimental exposure of the complex to oxygen, we performed native MS experiments on anaerobically isolated HybG–HypD complex after rapid buffer-exchange28. We also isolated a StrepII-HybG–HypDC41A complex in which native StrepII-tagged HybG was in complex with a mutant HypDC41A protein29. The exchange of C41 for an alanyl residue inactivates HypD and prevents synthesis of the Fe(CN)2CO group9. Both complexes (HybG–HypD and HybG–HypDC41A) were analysed by native MS, whereby we focused on the m/z region in the spectra harbouring the signals for dissociated HybG. Surprisingly, both spectra differed substantially from one another, despite the fact that both measured the same dissociated, native StrepII-tagged HybG protein (Fig. 6a). The mass spectrum of HybG dissociated from the native HybG–HypD complex revealed two main ion species. One ion species had a m/z 1929.2 (+ 5 charge state, deconvoluted mass: 9641 Da; Table S2), and one had an equally intense species at m/z 1924 (+ 5 charge state, deconvoluted mass: 9615 Da), which indicated the presence of a population of StrepII-HybG without any modification. The unique feature of this spectrum, however, was the presence of a signal at m/z 1956.6 (+ 5 charge state, deconvoluted mass: 9783 Da; Table S2), which showed StrepII-HybG modified with 26 Da plus an additional 136 Da, which is the mass of the Fe(CN)2CO group (Fig. 6a, upper panel). In contrast, StrepII-HybG that dissociated from the complex with HypDC41A showed a main ion species at m/z 1929.2 indicating HybG with the + 26 Da modification, but no unmodified HybG species. Importantly, however, no signal at m/z 1956.6 was observed (Fig. 6a lower panel), which is consistent with the inability of HypDC41A to synthesise the Fe(CN)2CO group. Furthermore, we observed minor species showing successive mass increases of ~ 16 Da, suggesting successive oxidation events that may have occurred either in solution or during the ESI process (Fig. 6a, lower panel). An overlay of both spectra highlights that only the StrepII-HybG protein dissociated from the native HypD scaffold complex carries the Fe(CN)2CO group (Fig. 6b).
Native mass spectrum of HybG carrying the Fe(CN)2CO group after dissociation from the HybG–HypD complex. (a) StrepII-HybG–HypD (upper panel) or StrepII-HybG–HypDC41A (lower panel) complexes were purified anaerobically after overproduction in E. coli DHP-D. Both StrepII-HybG spectra were generated by dissociation in MS mode without isolating a precursor ion. Collision energy was set to 80 V. (b) Overlay of both HybG spectra shown in part (a). Note that while both spectra appear to include a mixture of minor oxidation products, some minor modifications were only observed on the HybG from the native complex. These minor species were not identified.
Discussion
The findings presented in this study show that HybG, a member of the large and conserved HypC family of [NiFe]-hydrogenase accessory maturation proteins, shuttles between the HypD scaffold protein and its cognate hydrogenase 2 large subunit precursor, pre-HybC. HybG is thus not only involved in the biosynthesis of the Fe(CN)2CO group, but also is proposed to transfer it to pre-HybC. Although we could not demonstrate the direct transfer of Fe(CN)2CO from HybG to pre-HybC in the current study, we show unambiguously that HybG enters a 1:1 complex with pre-HybC, and that it carries the Fe(CN)2CO group. HypD, in contrast, does not enter into a ternary complex with HybG and pre-HybC. This indicates that HypD acts solely as a component of the central scaffold, where it serves to synthesise the Fe(CN)2CO group12,13.
There is strong evidence indicating that the cysteinyl residue at position 41 in HypD coordinates the Fe(CN)2CO group13,30. When we dissociated native HybG from an anaerobically isolated scaffold complex in which C41 on HypD had been exchanged for an alanyl residue, HybG lacked bound Fe(CN)2CO. This experimental evidence strongly supports the contention that the iron group is coordinated via C41 on HypD. Moreover, exchange of HybG’s N-terminal cysteinyl residue for an alanyl residue is known to prevent synthesis of active hydrogenase 2 in vivo30. This also supports the contention that C2 of HybG is the other main coordination site for the Fe(CN)2CO group on the scaffold complex8. Finally, we show here that the N-terminal cysteine on HybG is also required for interaction with pre-HybC (see model in Fig. 7), which indirectly supports a role for this residue in the delivery of the Fe(CN)2CO group to the active site cavity of pre-HybC15. The demonstration of direct transfer of the Fe(CN)2CO group from HybG to pre-HybC, presumably via a thiol-exchange reaction8, could not be achieved in the current study and will be elucidated further in future work.
Model depicting the interaction network of HybG in [NiFe]-hydrogenase 2 large subunit maturation (see text for details). The small ball-and-stick representation within HypD indicates a 4Fe-4S cluster. The C-terminally located thiocyanate on HypE is represented by –S–CN31; the penta-coordinated Fe ion attached to the HybG–HypD complex depicts the Fe(CN)2CO group. Pre-HybC, the apo-form of the catalytic subunit of hydrogenase 2.
It was notable that HybG with the bound Fe(CN)2CO group also had a further 26 Da modification directly associated with the N-terminal cysteinyl residue. The same 26 Da modification was also observed on HypC and on HybG isolated from a mutant strain lacking HypD (data not shown), which indicates that its provenance is independent of the hydrogenase accessory protein machinery. One explanation for this 26 Da mass increase is that it might arise through initial addition of CO2 (44 Da), resulting from an anaerobic decarboxylase reaction, followed by the loss of water (18 Da). Cyanide transfer via a HypE-dependent thiol-transferase side-reaction was also considered as an alternative source of this modification. The only way cyanide might yield a + 26 Da addition is, however, if one invokes generation of a transient radical species, to which a proton is then added. Consequently, this possibility is currently considered to be unlikely. Future analyses will be, however, required to define precisely the (bio)chemical origin of the modification and whether it is on the biosynthetic pathway of the Fe(CN)2CO group in vivo.
In addition to the 26 Da modification, the other minor modification to HybG identified was a further mass shift of 44 Da, attributable to binding of CO2. Modification by addition of CO2 has been identified previously using infrared spectroscopy and proposed as a potential source of the diatomic CO ligand27. Notably, this 44 Da modification also depends on the N-terminal cysteinyl residue, because conversion to an alanyl residue in the HybGC2A variant resulted in loss of all modifications of HybG. Moreover, the 44 Da modification was not observed on the species of HybG carrying the Fe(CN)2CO group. This suggests that at least two populations of HybG exist: one species that has the 44 Da (+ CO2) modification, while the other carries the Fe(CN)2CO group. Notably, however, both species also have the 26 Da (+ CO2 –H2O) modification on the cysteine at the N-terminus.
As well as interacting with the apo-form of the hydrogenase catalytic subunit, HybG (and other HypC-family proteins) also has an essential role in biosynthesis of the Fe(CN)2CO metal center. Data from a recent study suggest that HybG might even function in reductive synthesis of the CO and CN− ligands on the iron29. It might also be involved in recruiting the iron and CO2 molecules for further modification by HypD24,27 (see model in Fig. 7). These features of HybG are reminiscent of the role of the IscA/SufA carrier protein family in iron-sulfur cluster assembly32,33,34. In the case of hydrogenase maturation, the HybG–HypD complex is supplied with CN− ligands, originating from carbamoylphosphate35, and these ligands are generated and provided by two further accessory proteins HypE and HypF31,36. This modular form of cluster biosynthesis possibly represents an archetype for many complex metal-based cofactors that use a separate scaffold complex for cofactor synthesis, including, for example, molybdenum-iron cofactor biosynthesis for nitrogenase37. In the case of hydrogenase maturation, the controlled delivery of components to, and transfer from, the central HybG–HypD scaffold (Fig. 7) allows introduction of both temporal and regulatory checkpoints that help ensure correct and efficient cofactor synthesis, and ultimately enzyme assembly.
Based on the findings of the current study, the use of native MS to examine [NiFe]-cofactor biogenesis adds a new analytical dimension to dissect this maturation process. The results obtained in this study suggest that this approach can be used to investigate the temporal order of complex formation from the initiation of cofactor assembly to its ultimate insertion into the target apo-catalytic subunit. Based on the evidence presented here, this approach also allows identification of transient, unstable intermediates. Native MS also provides information on the stoichiometry of the protein complexes involved, and can even be used to determine changes in the redox states of metal clusters18. Future native MS studies on the hydrogenase maturation system will focus on optimising preparative steps so that transfer of Fe(CN)2CO between HybG and pre-HybC can be examined in detail.
Materials and methods
Bacterial strains, plasmids and growth conditions
The E. coli strains used included MC4100 (F−, araD139, ∆(argF-lac)U169, λ−, rpsL150, relA1, deoC1, flhD5301, ∆(fruK-yeiR)725(fruA25), rbsR22, ∆(fimB-fimE)38, its isogenic mutant derivatives BEF314 (like MC4100, but ΔhypB-E)39, and DHP-D (ΔhypD)39 and BL21(DE3) (F− ompT gal dcm lon hsdSB(rB− mB−) λ(DE3 [lacI lacUV5-T7 gene 1 ind1 sam7 nin5]) (Novagen, USA). The plasmids used included pASK-hybG, pASKhybGC2A30, pT-hybG-hypDEF40, pT-hybG-hypDC41AEF28, and pCAN-hybCstop41. pCAN-hybCproc was created by introducing an ochre stop at codon position 553 in the hybC gene on plasmid pCAN-hybCstop using site-directed mutagenesis (QuikChange II mutagenesis kit, Agilent) employing the oligonucleotides HybCfwd (5′-GCCTGTGCGGTACACTAAGTGGATGCTGACGGC-3′) and HybCrev (5′-GCCGTCAGCATCCACTTAGTGTACCGCACAGGC-3′).
E. coli strain BL21(DE3) was used as host for general overproduction of native StrepII-HybG–HypD complexes, while BEF314 (ΔhypB-E) or DHP-D (ΔhypD) was used to overproduce native StrepII-HybG–HypD and StrepII-HybG–HypDC41A protein complexes for analysis of HybG modifications. E. coli strains were transformed with the indicated plasmid using standard procedures42 and cultivated in modified TB medium43. Depending on the plasmid used, the medium contained either 100 μg ml−1 of ampicillin or 15 μg ml−1 of chloramphenicol to maintain plasmid selection. Cultivation was at 30 °C on a rotary shaker and incubation was continued until an optical density at 600 nm of 0.4 was reached. Expression of plasmid-borne hybG, or hybG plus the hyp genes, and which were under the control of the araBAD promoter, was induced by the addition of 0.2 μg ml−1 anhydrotetracycline (AHT) followed by continuing incubation at 30 °C for a further 3–5 h. Expression of the hybC gene variants on plasmids was induced by the addition of 0.1 mM isopropyl β-D-1-thiogalactopyranoside (IPTG) followed by further incubation of the culture at 30 °C for 3–5 h. Cells were harvested (OD600nm of between 1.0 and 1.2) by centrifugation of the culture for 15 min at 50,000 g at 4 °C and washed cell pellets were either used immediately or stored at − 20 °C until use.
Protein purification
All steps in cell disruption and protein purification were carried out under anoxic conditions in an anaerobic chamber (Coy Laboratories, Grass Lake, USA). Cells were suspended in 2 ml of 50 mM Tris–HCl, pH 8 containing 150 mM NaCl g−1 wet weight of cell paste. PMSF (phenylmethylsulfonyl fluoride) was added to a final concentration 0.8 mM and DNAse I was added to 10 μg ml−1 to the cell suspension and cells were disrupted by sonication (Sonotrode, 20–40 W with 0.5 s pulses for 5 min) on ice. Unbroken cells and cell debris were removed by centrifugation of the suspension at 45,000×g for 20 min at 4 °C. The resulting crude extract was used directly for protein purification.
The StrepII-tagged HybG derivatives or the HybG–HypD complexes were purified using StrepTactin Sepharose® (IBA, Göttingen) exactly as described27. N-terminally His-tagged pre-HybC and HybCproc were purified using cobalt-charged TALON® Superflow™ agarose (Cytiva) exactly as described41. When required, eluted, pooled protein fractions from the affinity chromatography steps were immediately buffer-exchanged into anaerobic 50 mM Tris–HCl, pH 8, using 5 ml PD-10 columns containing G-25 Sephadex matrix (Cytiva) and then chromatographed on a 1 ml Q-Sepharose® fast-flow column (Cytiva) equilibrated with the same buffer. Bound proteins were eluted stepwise using equilibration buffer containing 50, 150, 300 and 500 mM NaCl. Subsequently, samples were again buffer-exchanged into anaerobic 50 mM Tris–HCl, pH 8, containing 150 mM NaCl (buffer A). Protein samples were concentrated using Amicon centrifugal concentration filters (cut-off of 5, 10, 30, or 50 kDa, depending on the protein sample) and samples were stored at − 80 °C.
Protein concentration was determined as described44.
Pull-down assays
All experiments performed in this study used StrepII-tagged HybG (C-terminal tag), a N-terminally His-tagged pre-HybC or HybCproc, and HypD that was untagged. To examine the interaction between StrepII-tagged HybG or HypD–HybG with His-tagged pre-HybC or HybCproc, 150 μg (15 nmole) of each protein, or protein complex, were mixed and incubated for 30 min at 30 °C. After incubation, the mixture was loaded onto either a 0.5 ml StrepTactin Sepharose or a 0.5 ml nickel-NTA (Qiagen) column for the purification of StrepII- or His-tagged complexes, respectively. Each column was equilibrated with buffer A (see above). After sample-loading, columns were washed with 10 column volumes of buffer A and bound proteins were eluted from the StrepTactin column with buffer A containing 50 mM biotin, or from the Ni–NTA column with buffer A containing 300 mM imidazole. Elution fractions (E) of 0.5 ml were collected and analyzed immediately by electrophoresis on 12.5% (w/v) acrylamide SDS-PAGE45. Gels were subsequently stained with silver (Pierce™ Silver-staining Kit, Thermo Fischer), or were transferred to nitrocellulose membranes and challenged with antiserum (diluted 1:1000) raised against purified HybG.
Mass spectrometry analyses
For native MS measurements, as well as high resolution MS measurements, the buffer was exchanged to 500 mM ammonium acetate, pH 6.8, by using Amicon Ultra centrifugal filter units with an appropriate molecular weight cut-off (Merck Millipore, Darmstadt), unless stated otherwise.
The concentration of protein solutions after buffer-exchange was approximately 10 μM. Native MS was carried out on a High-Mass Q-TOF II instrument (Waters Micromass/MS Vision) equipped with a nano-electrospray ionization (ESI) source. The applied capillary voltage ranged from 1.0 to 1.4 kV, while the sample cone voltage varied from 100 to 160 V. The source pressure was adjusted to 9.2–10 mbar and the pressure in the collision cell was adjusted to 10−2 to 210−2 mbar. MS measurements were carried out using MS profile mode for the quadrupole to guide ions within the m/z region of interest. The acceleration voltage in the collision cell was set to 30–60 V for MS measurements. Dissociation experiments were carried out by collision-induced dissociation (CID) for the selected ion species. To achieve dissociation of protein complexes the acceleration voltage of the collision cell was increased in dependency of the measured complex (ranging between 60 and 90 V). Data were recalibrated by using cesium iodide (CsI).
High-resolution ESI–MS and MS/MS data were recorded on an Orbitrap Fusion Tribrid mass spectrometer (Thermo Fisher Scientific) equipped with a nano-ESI source. The resolving power was set to 120,000 at m/z 200. The fragment ion mass spectra (CID-MS/MS) were obtained in the linear ion trap.
Rapid online buffer exchange
Online buffer exchange (OBE) was carried out on an Ultimate 3000 RSLC nano-HPLC system (Thermo Fischer Scientific) using a size-exclusion column with a length of 12 cm, an inner diameter of 0.6 mm using Bio-Gel P6-Resin (Biorad) that was prepared in-house, as described in28. Buffer-exchange was not performed anaerobically. After equilibration of the column with target buffer solution (250 mM ammonium acetate, pH 6.8), 5 µl of 20 µM protein solution was injected onto the column. Buffer exchange was achieved by isocratic elution with a flow rate of 10 µl/min. The nano-HPLC system was directly coupled to a nano-ESI source (Waters) equipped with a stainless-steel emitter. The transfer flow of eluted protein from the SEC-column to the nano-ESI source was set to 300 nL min−1. The chromatographic step was tracked by UV-absorption at a wavelength of 280 nm. The method was adapted from28. The applied capillary voltage was adjusted to 2.0–2.3 kV for the OBE measurements, while all other settings were as described before.
References
Vignais, P. M. & Billoud, B. Occurrence, classification, and biological function of hydrogenases: An overview. Chem. Rev. 107, 4206–4272 (2007).
Volbeda, A. et al. Crystal structure of the nickel-iron hydrogenase from Desulfovibrio gigas. Nature 373, 580–587 (1995).
Pierik, A. J., Roseboom, W., Happe, R. P., Bagley, K. A. & Albracht, S. P. Carbon monoxide and cyanide as intrinsic ligands to iron in the active site of [NiFe]-hydrogenases. NiFe(CN)2CO, Biology’s way to activate H2. J. Biol. Chem. 274, 3331–3337 (1999).
Lubitz, W., Ogata, H., Rudiger, O. & Reijerse, E. Hydrogenases. Chem. Rev. 114, 4081–4148 (2014).
Ogata, H., Nishikawa, K. & Lubitz, W. Hydrogens detected by subatomic resolution protein crystallography in a [NiFe] hydrogenase. Nature 520, 571–574 (2015).
Sargent, F. The model [NiFe]-hydrogenases of Escherichia coli. Adv. Microb. Physiol. 68, 433–507 (2016).
Blokesch, M., Magalon, A. & Böck, A. Interplay between the specific chaperone-like proteins HybG and HypC in maturation of hydrogenases 1, 2, and 3 from Escherichia coli. J. Bacteriol. 183, 2817–2822. https://doi.org/10.1128/JB.183.9.2817-2822.2001 (2001) (PMID: 11292801).
Böck, A., King, P. W., Blokesch, M. & Posewitz, M. C. Maturation of hydrogenases. Adv. Microb. Physiol. 51, 1–71 (2006).
Blokesch, M. & Böck, A. Properties of the [NiFe]-hydrogenase maturation protein HypD. FEBS Lett. 580, 4065–4068 (2006).
Blokesch, M. & Böck, A. Maturation of [NiFe]-hydrogenases in Escherichia coli: The HypC cycle. J. Mol. Biol. 324, 287–294 (2002).
Miki, K., Atomi, H. & Watanabe, S. Structural insight into [NiFe] hydrogenase maturation by transient complexes between Hyp proteins. Acc. Chem. Res. 53, 875–886 (2020).
Bürstel, I. et al. A universal scaffold for synthesis of the Fe(CN)2(CO) group of [NiFe] hydrogenase. J. Biol Chem. 287, 38845–38853 (2012).
Stripp, S. T. et al. HypD is the scaffold protein for Fe-(CN)2CO cofactor assembly in [NiFe]-hydrogenase maturation. Biochemistry 52, 3289–3296 (2013).
Wolf, I., Buhrke, T., Dernedde, J., Pohlmann, A. & Friedrich, B. Duplication of hyp genes involved in maturation of [NiFe] hydrogenases in Alcaligenes eutrophus H16. Arch. Microbiol. 170, 451–459 (1998).
Magalon, A. & Bock, A. Analysis of the HypC-HycE complex, a key intermediate in the assembly of the metal center of the Escherichia coli hydrogenase 3. J. Biol. Chem. 275, 21114–21120 (2000).
Drapal, N. & Bock, A. Interaction of the hydrogenase accessory protein HypC with HycE, the large subunit of Escherichia coli hydrogenase 3 during enzyme maturation. Biochemistry 37, 2941–2948 (1998).
Jones, A. K., Lenz, O., Strack, A., Buhrke, T. & Friedrich, B. NiFe hydrogenase active site biosynthesis: Identification of Hyp protein complexes in Ralstonia eutropha. Biochemistry 43, 13467–13477 (2004).
Crack, J. C., Gray, E. & Le Brun, N. E. Sensing mechanisms of iron-sulfur cluster regulatory proteins elucidated using native mass spectrometry. Dalton Trans. 50, 7887 (2021).
Schmidt, C., Beilsten-Edmands, V. & Robinson, C. V. The joining of the Hsp90 and Hsp70 chaperone cycles yields transient interactions and stable intermediates: Insights from mass spectrometry. Oncotarget 6, 18276–18281 (2015).
Sharon, M. & Horovitz, A. Probing allosteric mechanisms using native mass spectrometry. Curr. Opin. Struct. Biol. 34, 7–16 (2015).
Snijder, J. et al. Structures of the cyanobacterial circadian oscillator frozen in a fully assembled state. Science 355, 1181–1184 (2017).
Liko, I., Allison, T. M., Hopper, J. T. & Robinson, C. V. Mass spectrometry guided structural biology. Curr. Opin. Struct. Biol. 40, 136–144 (2016).
Lacasse, M. J. & Zamble, D. B. [NiFe]-hydrogenase maturation. Biochemistry 55, 1689–1701 (2016).
Pinske, C. & Sawers, R. G. Anaerobic formate and hydrogen metabolism. EcoSal Plus https://doi.org/10.1128/ecosalplus.ESP-0011-2016 (2016).
Thomas, C., Muhr, E. & Sawers, R. G. Coordination of synthesis and assembly of a modular membrane-associated [NiFe]-hydrogenase is determined by cleavage of the C-terminal peptide. J. Bacteriol. 197, 2989–2998 (2015).
Senger, M., Stripp, S. T. & Soboh, B. Proteolytic cleavage orchestrates cofactor insertion and protein assembly in [NiFe]-hydrogenase biosynthesis. J. Biol. Chem. 292, 11670–11681 (2017).
Soboh, B. et al. The [NiFe]-hydrogenase accessory chaperones HypC and HybG of Escherichia coli are iron- and carbon dioxide-binding proteins. FEBS Lett. 587, 2512–2516 (2013).
VanAernum, Z. L. et al. Rapid online buffer exchange for screening of proteins, protein complexes and cell lysates by native mass spectrometry. Nat. Prot. 15, 1132–1157 (2020).
Nutschan, K., Golbik, R. P. & Sawers, R. G. The iron-sulfur-containing HypC-HypD scaffold complex of the [NiFe]-hydrogenase maturation machinery is an ATPase. FEBS Open Bio 9, 2072–2079 (2019).
Blokesch, M. et al. The complex between hydrogenase-maturation proteins HypC and HypD is an intermediate in the supply of cyanide to the active site iron of [NiFe]-hydrogenases. J. Mol. Biol. 344, 155–167 (2004).
Reissmann, S. et al. Taming of a poison: Biosynthesis of the NiFe-hydrogenase cyanide ligands. Science 299, 1067–1070 (2003).
Baussier, C. et al. Making iron-sulfur cluster: Structure, regulation and evolution of the bacterial ISC system. Adv. Microb. Physiol. 76, 1–39 (2020).
Braymer, J. J., Freibert, S. A., Rakwalska-Bange, M. & Lill, R. Mechanistic concepts of iron-sulfur protein biogenesis in biology. Biochim. Biophys. Acta Mol. Cell Res. 1868, 118 (2021).
Peters, J. W. & Broderick, J. B. Emerging paradigms for complex iron-sulfur cofactor assembly and insertion. Ann. Rev. Biochem. 81, 429–450 (2012).
Paschos, A., Bauer, A., Zimmermann, A., Zehelein, E. & Bock, A. HypF, a carbamoyl phosphate-converting enzyme involved in [NiFe] hydrogenase maturation. J. Biol. Chem. 277, 49945–49951 (2002).
Watanabe, S., Matsumi, R., Atomi, H., Imanaka, T. & Miki, K. Crystal structures of the HypCD complex and the HypCDE ternary complex: Transient intermediate complexes during [NiFe] hydrogenase maturation. Structure 20, 2124–2137 (2012).
Sickerman, N. S., Rettberg, L. A., Lee, C. C., Hu, Y. & Ribbe, M. W. Cluster assembly in nitrogenase. Essays Biochem. 61, 271–279 (2017).
Casadaban, M. J. Transposition and fusion of the lac genes to selected promoters in Escherichia coli using bacteriophage lambda and Mu. J. Mol. Biol. 104, 541–555 (1976).
Jacobi, A., Rossmann, R. & Böck, A. The hyp operon gene products are required for the maturation of catalytically active hydrogenase isoenzymes in Escherichia coli. Arch. Microbiol. 158, 444–451 (1992).
Soboh, B. et al. [NiFe]-hydrogenase maturation in vitro: Analysis of the roles of the HybG and HypD accessory proteins. Biochem. J. 464, 169–177 (2014).
Soboh, B. et al. Evidence for an oxygen-sensitive iron-sulfur cluster in an immature large subunit species of Escherichia coli [NiFe]-hydrogenase 2. Biochem. Biophys. Res. Comm. 424, 158–163 (2012).
Sambrook, J., Fritsch, E. F. & Maniatis, T. Molecular cloning: A laboratory manual (Cold Spring Harbor Laboratory Press, 1989).
Soboh, B. et al. [NiFe]-hydrogenase maturation: Isolation of a HypC-HypD complex carrying diatomic CO and CN-ligands. FEBS Lett. 586, 3882–3887 (2012).
Lowry, O. H., Rosebrough, N. J., Farr, A. L. & Randall, R. J. Protein measurement with the Folin phenol reagent. J. Biol. Chem. 193, 265–275 (1951).
Laemmli, U. K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227, 680–685 (1970).
Acknowledgements
We thank Claudia Granich for help with the construction of the plasmid expressing hybG and Mandy Waclawek for help in purifying N-terminally His-tagged pre-HybC. Constanze Pinske and Elisabeth Schwarz are thanked for discussion. This work was supported by the Deutsche Forschungsgemeinschaft (DFG) via RTG 2467, project number 391498659 “Intrinsically Disordered Proteins—Molecular Principles, Cellular Functions, and Diseases” to AS, and through the SPP1927 “Iron-sulfur for life” initiative to RGS.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
C.A., C.I., A.S. and R.G.S. designed the experiments and analysed the data. C.A., K.N., C.I., A.H. and D.T. carried out all the experiments. C.A., A.S. and R.G.S. drafted the manuscript and conceived the study. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Arlt, C., Nutschan, K., Haase, A. et al. Native mass spectrometry identifies the HybG chaperone as carrier of the Fe(CN)2CO group during maturation of E. coli [NiFe]-hydrogenase 2. Sci Rep 11, 24362 (2021). https://doi.org/10.1038/s41598-021-03900-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-03900-w
- Springer Nature Limited
This article is cited by
-
Evidence the Isc iron–sulfur cluster biogenesis machinery is the source of iron for [NiFe]-cofactor biosynthesis in Escherichia coli
Scientific Reports (2024)
-
Stepwise assembly of the active site of [NiFe]-hydrogenase
Nature Chemical Biology (2023)